class: center, middle, inverse, title-slide # Analyzing high dimensional metabolic data in epidemiological studies ### Luke Johnston ### Aarhus University --- layout: true <div class="my-footer"> <span> <img src="../../common/au_logo_black.png" alt="Aarhus University", width="160"> </span> </div> --- # Aims: - Describe NetCoupler - Get feedback and/or comments on it -- - ... and to see if I explained it well --- class: center, middle # THE PROBLEM --- ## Modern epidemiology studies generate a lot of metabolic data .pull-left[ - -omics type data - Metabolic biomarkers ] .pull-right[ - High dimensionality - Complex networks ] ??? Like metabolomics, fatty acid panels. Classic epidemiology meaning having an exposure with an output --- ## "Traditional" analysis may use: Dimensionality reduction .center[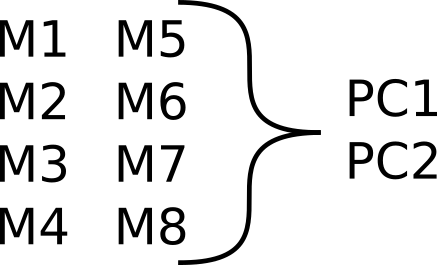] ??? This has the advantage of making things simpler while trying to maximize variance in the data. Afterward you can do modelling on each principal component. The disadvantage of this approach is that it loses a lot of information since the interdependence and connections between variables it not maintained. --- ## "Traditional" analysis may use: Many regression-type models ``` O1 = M1 + covariates O1 = M2 + covariates ... O1 = M7 + covariates O1 = M8 + covariates ``` ??? Some ways you might go about analyzing this data is by running many regression models, one for each metabolic variable for instance. This of course has problems since you're simply running a bunch of models and not taking account of the inherent interdependencies between variables. --- ## "Traditional" analysis may use: Network analysis <img src="data:image/png;base64,iVBORw0KGgoAAAANSUhEUgAAAfgAAAFoCAMAAACMkBkOAAADAFBMVEUAAAABAQECAgIDAwMEBAQFBQUGBgYHBwcICAgJCQkKCgoLCwsMDAwNDQ0ODg4PDw8QEBARERESEhITExMUFBQVFRUWFhYXFxcYGBgZGRkaGhobGxscHBwdHR0eHh4fHx8gICAhISEiIiIjIyMkJCQlJSUmJiYnJycoKCgpKSkqKiorKyssLCwtLS0uLi4vLy8wMDAxMTEyMjIzMzM0NDQ1NTU2NjY3Nzc4ODg5OTk6Ojo7Ozs8PDw9PT0+Pj4/Pz9AQEBBQUFCQkJDQ0NERERFRUVGRkZHR0dISEhJSUlKSkpLS0tMTExNTU1OTk5PT09QUFBRUVFSUlJTU1NUVFRVVVVWVlZXV1dYWFhZWVlaWlpbW1tcXFxdXV1eXl5fX19gYGBhYWFiYmJjY2NkZGRlZWVmZmZnZ2doaGhpaWlqampra2tsbGxtbW1ubm5vb29wcHBxcXFycnJzc3N0dHR1dXV2dnZ3d3d4eHh5eXl6enp7e3t8fHx9fX1+fn5/f3+AgICBgYGCgoKDg4OEhISFhYWGhoaHh4eIiIiJiYmKioqLi4uMjIyNjY2Ojo6Pj4+QkJCRkZGSkpKTk5OUlJSVlZWWlpaXl5eYmJiZmZmampqbm5ucnJydnZ2enp6fn5+goKChoaGioqKjo6OkpKSlpaWmpqanp6eoqKipqamqqqqrq6usrKytra2urq6vr6+wsLCxsbGysrKzs7O0tLS1tbW2tra3t7e4uLi5ubm6urq7u7u8vLy9vb2+vr6/v7/AwMDBwcHCwsLDw8PExMTFxcXGxsbHx8fIyMjJycnKysrLy8vMzMzNzc3Ozs7Pz8/Q0NDR0dHS0tLT09PU1NTV1dXW1tbX19fY2NjZ2dna2trb29vc3Nzd3d3e3t7f39/g4ODh4eHi4uLj4+Pk5OTl5eXm5ubn5+fo6Ojp6enq6urr6+vs7Ozt7e3u7u7v7+/w8PDx8fHy8vLz8/P09PT19fX29vb39/f4+Pj5+fn6+vr7+/v8/Pz9/f3+/v7////isF19AAAACXBIWXMAAAsSAAALEgHS3X78AAAgAElEQVR4nO2dCXwURb7HBxXFE90nq66u67quux5vdZ/vyRzJTO6EKxwh3PcRQG4BI4oERSDc4RCNIDcI4RQRRRCQILfKrRwhCVfkikBCyDXTr3uSmVT1VHdP9/S/etip735Wpqqr0/+p33R3nf+/iWOEJCajDWAYAxM+RGHChyhM+BCFCR+iMOFDFCZ8iMKED1GY8CEKEz5EYcKHKEz4EIUJH6Iw4UMUJnyIwoQPUZjwIQoTPkRhwocoTPgQhbrwrsPL3k9p3Sy5+9DZ2SW0L87wQlf4kqUtbb0+2nTifOHlnN0LhkfHjD9H9foMLzSFz+sfPuksllOc1bTJVooWMLzQE/5yzwZbCdln+sXspWYDwws14edavpU4cqZlnyJaVjA8UBL+RrthpdJH11oP0jGD4YWO8OfCvpI9nh/xJRU7GF6oCH/cfEihRHHifBqGMLzQEP6cOUexTHnSSgqWMLxQEP5GmNL9LnArLhvcEkYNFITv8IVv3q2HTM+JsgrrF8DbwvAAL/zcYYTMLJOP8NyOpuC2MLyAC3/VcouQm0gQnhvCXvP0ABe+1yZC5v47SMLfMLNZG2pAC3+mESHz4J9NJOG5j2cCW8PwAi18/63inIK18bVMZOFLLeXA5jA8AAtfEo6nd4Y9YqqGIDz3AaEDwAABWPilk/D0MpNJTvi8VrDmMLwAC98Sn39XEp6LIHUBGADACu8KE2Vc2e1miZTw72wFtYfhBVb4w73I+UekhN+UBmcMAwVW+M8/IudLCv9bMqQ5jBpghR+1mZwvKTxnB7SGgQArfM9fyfnSwjvgjGGgwArf9gI5nwlvOLDCJ14n50sLHy+zNI+hI7DCJ18i50sLH+EENIdRA6zwXU+T89mj3nBghR+2i5wvLXw4MZehO7DCz15IzpcUvrgBpDmMGmCFz36bnC8p/P6BkOYwaoAV/mYMOV9S+KnLIc1h1AA8OxdTTMyWFD7xN1BzGF6AhR+fRcyWEv5GJKw5DC/Awp9vRsyWEn7+DFhzGF6g19w1OUPKlRI++jKwOQwP0MJv6U/KlRB+S19gaxhewNfVx5BuebLwrpg8aGsYHsCF309aWkEWfvFwaGMYXuD3zvVe65tHFP5y/ZvgxjA8wAtfbM31ySMJ72qxHdwWhhcK26QPRvjcySTh09LhTWF4oeERY31CmXKhzE4ueEsYXqj4wJmfpLhPIjO5goYlDA90vF6tjLsie9yV1oPpThdKfu6yzTtkjl5uks6e85Sh5dnyUtMhEu4rD5csMTO/R9Sh58t2hfkT4grakc8MZ/13+lD0Xl0yw/KBePHljfnRfeN/pGcDwwNVf/Xl65LNzb+9WJ0q3p/RNHLGZS4/opKmEQw3tCNUjBuV1tLusMdERobFD1petex+ylTKRjDoCx97zf0PNqJTGZ5L2QoGbeGvJZBy97WkawWDuvDLphCz+xNm8BigUBa+4wli9nUzeTUuAwy6wjutEgeWvUXVDgZl4XcOljqSeICmHQzKwr8r4RqF407EstF6qtAVPlx6Yn7kAop2MOgKfzZJ+liJ+Xd6hjDoCp85V+bg18QV+AwgqArf7Lzs0Z9o2cGgK3xphOzhnGjWvqMHTeE3jpQ/ztp3FKEp/MDd8sdL6l+jYwiDrvBWJVdmqyXHdxh6Q1H4U50UizQ6SsEOhgBF4acuUyxyjDhrywCAovAJV5XLvLka3g6GAD3hi2P9KHTN7NlzU3kD0hgGPeHXjvOn1NzR1R92jgC0hUFR+JSD/pRyRuRXfdiWBmgLg6LwVv/G5XZ2rPp38weAtjDoCX8wxc+CHXa6/9k4Fs4WBkXhx63xs+DZCPej4avxgMYw6Akf43crfZR7yH7dZDhbGPSEv9bQ76Ilrwu/kdUZcMYw6Am/nLygHqXsm+oPS4We3AoWUhwUWsJ3kQhEhlAxrOEv7g+uKL5Lt+xjaJNCG0rCu2z+lDrRZIA7bNXu9hy3aDasRaEOJeH3+bmgbp1lgdCmb7eDmz8P0h4GJeHf3+BnwZtpcYc4Ls/inCMRzoahD5SEj/Tf28nJpgOucWEDM5cCmsOgJPzlxmpKr7NkXnhwEotOAwod4RerizxRkhae8NIqIFsYbugI3/6UyhNyEu+YBWIJoxoqwjvD1J+T6m9zkKEJKsLvGkTjKgw1UBF+5EYaV2GogYrwDkXn1fJcWj22c4yd/1+nMatYREJ9oCH8paaBnH18RFjTyd+cFtyhluZunNLM9s4vOtkV0tAQfkEADfQN8e024I4vKzd2iF3PtlcGCg3h2+RoPXN3xJALhOyLqY4fArCHwVERvlJDZ85Nce/ksxKHzrftIeEFneEfFITfMUTbeUdscoN366x+LddmSEBB+BGSrq5k2azg4faM42tNf5fhhoLwDj9iUPmyOv66QomiRp9r+cMMN/DCFzTXctbmeOW+f1njr7T8aYYAvPDzMjWcdNguut8Pvmn7y92Pvjb0MJpZFMH8JWkFXvhWGkJEi8OSnrabqqnVDnWHV2BjzlM0Ai58pV3DST1xN+ZrHjLV8De0Z/+1spcNBhFw4bOHqT9nTzssufMeQfDnWnS23Ct8+Hc5cqzzd4GZF7KAC//OFtWnuByYI8Tf/sir/ay7T3ihg6A8Gn34YpiSRyUGEXDh7eo7c+uGY8n+vNYvF1YnOvKJR9E/mbZSu22hDLTwBS3UnxNzEU3l320y1fHuwykRbn9U66sOraaFNtDCf/ap6lOOdsGSI3il+9YkB/JJbHdGCotXqAVo4Vvmqz7lLby99jKvNBLJ5tzKlSu3o8ez2bouLQALXxGu/hwb1lw7xev+T7niLqk4Nww5gIX/PlX1KeeTseR8/ElPoIPm6f5QBlj44VtVn/I5vvniLV54+RU8s+epvgYDWvjwcuUyIt7biiUb88LzffhjI+s/dfdj//Pmdt8TdrLQZRqAFf6ChtihrfEwFs/zwh++8UYtz4htxCHxCVcDWssZqsAKP2eO+nMi8IWUj/Fi//jfyFj9/T7ubh1azQtlYIVPOqf+HAeevJ/X+iX+/3f+o2XyP+4UlK8tXmgpH/GEQQRU+AqHhpNwGV1Vz/j7JrqDz14fcQefeOqW3BkMvwAVftvbGk5yYKkSt+4PewOQZgnJqXJnMPwCVPjU7zWc5MDe8VV3PDI435VPvig6Q8NVQh5Q4cPUd+b4dgE2ReN+x/8LSR8WfgjYcvvrjTRcJeSBFP5ssnIZX97JxpLCbBzqut5Zh8/A9s7v1bhuP7SBFP5TuYiikizCF2e+wuu8CM0Qunbz0YwFzCOeBiCFbyEbUVSK0+2xZBteZ2xHjTCU9wma0Y3tntUAoPDlGrtZeESDUbzO2DMgnM/AxnDY7JwWAIXfMly5DIn+2ADNBl7nHkja+QifsQ/J+NHfCAgMFEDhh2UrlyGxtw+aKn3AZKqHuEf8TujWozvmB+7QdpkQB1D4sAqNJ9qxXRLJvNI18WmcwpO+DXK4yMacJGgBTvizrbWeuXQMmtojDNR/4UkJ0/O1fkYOT5in9TKhDZzwn8zTemalFYtJ2VJQ/h33Xro84bOpM3Jwz2taBokYgMI3Izkx8Y9ve6GpC08LateJS+lmEaZoTK+gDpHbvJzGXGNoAUz4skDmzFpvQ1NH6yHT8aYwdEh3ZzNX1usZWhsToQyY8JsDCRFaWL8ATV5s6ZW97mh0G02hmS9XlhmWFcC1QhQw4YcE5JdqZ1wplj4y0vpMnYf+3mJOIZpb1rDqyXA1NZJ5wVIJmPC2gJ6/5dHRlYqFXB28USxy2nTQsA0/lIES/kwb5TLSFCYs+KR1qUKh8o5oaLo9cW9eCeSSoQaU8LMWBHDyadtWjvs8QV7IwkaiS2wKS2Ux5/0GSvimATgb3mVxz7dtN8sNxu42bxVnObPCMlkD30+AhC8NoDO3LOZy1YdLzftLubi5MTiR9MsqywzPYiO4fgEk/MaRmk8d267m5f6FZXIxocjNDLOU08vC1IhtEocYKEDCD9ql8cSK3gPQzbLlc61vHhAVOTzMMlvGzcaZri2Oarx4KAEkvEW5M0bkRhNxSFnX992svZYeqFpKf+vgsj62LlsU3N4cbtb1jLbLhxAwwouWT/nNuXBSCCLXoY/6xtmjbVH22DdmHvDH2dGu2JSLyqVCGhjhZyzWdNo+8xG9LOD7dkq+cEMbGOEba7rfvo4qUC7kL86s+ulKQ0ChDIjwJVFazspI8j8ArT+UZb6eyZzgSQEi/Ib31Z9TOWCA7ipdf8/OAp9JACJ8/33KZUQUN58OYAj3W7+E3RB/9/YHRHiL6nv3gmMdhCE8+SnxLIYJAQjhj3dWe8YRs/pnhN8cbZ3M/GL5ACH81GUqT9hsg51M3xWXwgJUioAQvuFV5TIo8xqAhxvYFJ7KQhpgAAhfHK2quCutK4UV0s7F5mmagiL9pwIg/BdjlMvUUNpeVXHtlM00z9c4g/CfCIDwvX9WLuPlSsxS/S2Q4Ga6hc3WewAQ3qyicnOsWtzkaKYwLVpb9MP/PPQX/nAP5TIedpl/VS6kK5dSGwH2HG8j9Bd+vFxEWJwVUZd0v7wieWxER0A34W95AkXE+t1vGttOOZokBEfYiI6OwhfFV/17Pd7PE8q7v2NYSys7ZvBlo64dJOj3qK+OHbxion/FfRdZUSXkR3T0E37oTvc/3fxb6Xg23OAg4K4sc7oxb5rgQD/h140T/usy+1V4j9n4lbDlmfVDeP+FfsIXuj2LLrdUpQ7KBn1dHRkUkyY3M0J3/4WO3Tm7MCAaNcr9eb9NzlP9xFYl+l02IG6k20J0b72Owg/Yz//nEXdrOdsh00Gv6Kv/IivtFPQNzTU6Ogq/cgrHHX9M+LQ1Qmaj641EQ5vzvuR0TDK+wUEdHYW/1Jzjegk+ztbHySxpPxsZfIG/j7ZuFXIjOnoO2dqc3NPZfPuuiegNjjadd1uO6XhF3dgdn6Ljmv7bAT2F732orC7HLW4m6h7vQnzaZkUbMDrvF5vsoTWio6fwS2fOfJ37uK14PU1KzXRYRutgac77IozoBK91uqOn8BdamadN6CgeE7ll83wq65Ia1L3m8szXQ2dER9dpWdtDo/v49NSWegbvr8YtEh8LNm6Ot64M6t+mfugqfJP7CC7qG1a7Nj1pJcSFDTqK0qNCY42OrsKnEDZLnkus+ndjWK6el4IjRNboAEeT5rixVUOimY1vnzZzXq82x422ARxw4a3CJvXKASm3VavpSOvkXFGWTyjj2xxo4Xf15f9zPXEm8GV054cYkTOVN9YbZAkQ0ML33stxp2zBN0qrzBrrWNRRw3UzyfHa7Quw8EInfuLTJ2EvAkTlXDParV8YiB/24ANY+GXjuQ6P3bZ+pXFHmQlBOcuglQCFPz57YII9OtwRHtV9wg7C3sfGp195oWro3pmzLr3j2MCuJsGV1SOSI8Pt9vDI5BFrVO7UVYDv1ntfU8fj5MZ2yrLHd4/iKyLanjBo9m3QKQhE+P0DLF0W/FgVEqbixBfD7C1Wizakno96tBv/300ZKVGvJ6dlHQFYgFEw2dF40rbqlVwFWyc2dkzRdVnXpdSGnh0Dw5dIFSpd1dzx1rqTVS+Goh/ndzYP/FGqbJCgWXjnKkevH0S3QP4YyxhsKr5T7ead7PZO49efBhoIPdKh4RLRzMrNxQ066fpQPt6qVVUjpcTyO7HA7x9axolcabqyUyLWBPXgr1bhs8PeI62yqVxinYnsRY7s+2UO4Nf/rVPLn0j5PyV11XX2d098vNuR+oZ+hIOVGZalpO3Xl0fYd+pphM5oE/5mz7ZSkaLLJ9ppNYLmW7dKHdpi1XdCaFxdty/15r7L846GT5IamzrXppe+nvv0RJPwR21y41g5joUyR3XjZpeBMi4uSvt313Vy3Tn4gea3uHyH+N6eH3Fa5qwVYUEb4lyL8N8rzLeU9tTurd5vrkYp7MrNita5gd+h7mBn+iw8893e8v5VcmwaIyuDo0H4DTGFSkVGvgHdsLloUwwivd2mswfr0+aHp4eh8TNdfUYpnXM1muSPOwhQL/zuSD9ier7/rgZbVHA93I+G0z4buRWunb0v1I1Ekm+PVj7lZkxw3vOqhc+x1rTmr5tE1ESi6TNHH/vIOBtvwtKuDf1e/WPtR1/qvhwbRNqYGPjAwY7UgSkpbZKbx8RYo2MsMS/cXfPWzuxLPGOhyVQHSV625gZsBABqhS+PRjYf/CgtfEU0saOlEx9MwpIb/+m14CmsNT8+cI9al/fvP56TV1jou7P2UATRTdupB3HhuSOOYPSzplb44XORxHJp4bl8G5z3ur1NsCbE+7VQG7oid7mr8X4wI8qsxPgn5a+bRMJzc4Dfe5pQKTw+Xj2G/4533IWAujb8aLwe9pFwRuaiyeluvR+L7Br3Z/enwcixfAfYNr0xnxCzU00+wnMJQdipUyl8E8xLVRf+O4pjRHlx2qHCwnyWjqby7uWteGK++we5/nn+cy10lGXMfCAjCsg/qe/uIAh/tCmQEQGgTvifumLJML6Wpcemvnxbk0WKVFqwbkU/vqIf8Mz4Fz7Jp5ogR4vNQN4shxG7aZf/ZCIIz3UKvuWb6oRPxt3SPW4yPS1d2BUOEw5oGd5iE6q6JsjBF3zqHvSHMRpm//s1O3Gkogl/KxCEP9YaxIhAUCX8lYZYsoj/irEyxWd9psUiRRpg066/C3dYzfx3sfCoRf3YXUTvf/3IzCTlzuAv/hZBeC4u6JxsqRJ+Oh6/+QD/FUnzVR6uxmmxSAnPQv1qTvBG1EZet8/yaayT3+QCBwBx9PJQHZPJcZUk/NyPIIwIBFXCJ+Br41fwX3GGXPmGeo+cCXw6F0sK9WyqmSqsEJp6mBflORDPnSuNCZklL5pMj5y9RhK+ykFQMKFG+LJIPD2O/4rfcqXbJr89YsZGkuuw9LWBmCZBm1w8/QhvxQpvah+fuhO7G3M6ABixiuTNrxd/7ZUcUXguPNgGcdQIv3sInu7Of8Wc2U9VjZvc2/6Ezwm7hgZmHJEwUXoof/FnPI+iivp8Kkn+BD14c49v3mr+0j04CeEH7wWwIhDUCD8Pf8hyDv4rPlczZHaXz4jNdYCGVXFDUcZvT/PXth12f85ryn+uK4pTmgCwGqLhDZ+ss38wmZ4vlhJ+NpU1CipQI/zbopnQJ8Ujtr3FZ4QHaB2BHweKc359XHi8N0qd9W4S37gyPSjelNtfTeAEP/H9Yk7+NqgtrLAkC7/9Hf2NCAg1wnc7hSVvuofIH3p7/9XSM8sbuZUXj2I6AjXPl6/H+WRdbIT8+MJ8VsSMAYg26fDJGc1f2/3iJwt/QoUXfyqoEb4VPgZ7SKjn/82tTn35gDCCJlqAGaH/eowswja8ZXW8ut/5ic8lp6/U3QhXpDhn510mU4z70mThC4JtCEeN8In4SJzQmHmlpsO2UXgAiOah4vV3E7zQZ6I/PxZ735jFaz3n6O+I45bYN/u1Z0ym/6oaMCALfz3RJ8tY1AjfGvcIVpSXl4duJOwgtPXwMxxazZJmpTgG7dm/CnOEXVYcuXl87cC7+c/1RO4KM/Tf4eyKEGW04a9b3XclC3++re5GBIYa4Xv4dthQ9gr3Wy6W5VBtjyIbRWsrSoQJuVc80+4nIvnU4/hWmg+/1d8KB56cy1+1T/VnsvC/puhvRECoEf69rbKHnQ/zXxmfswJo1R8SrXcSZuP/XDMSXvKqMFyOlegt6t7pAf7FTtxvMr3oWc1NFv67NP2NCAg1wi8mTkzUYOG/Mjaaf7W5BosUEL9ehfGjeUh6i9Cjw+ZtYwDiESRiS7ebmUz3eNclkIWfRS+8nn+oEf6n/vLHG/NfGVt3/j1E5xUfiCvgr3k3OprieoLPQae/XRAjd29jS2djxAMaVR1LtEhfyQUrBqFG+Eq7/PF4/ttis9+jvtFgkRJdsD3IO3xalFF8zudI+ihED/qrD9GUH8Lbgi26qarZuUS0I3/1wIED+dhh4fW6Fc2IgfAeshB7qGzkr/kydlwYzUHfSTMkNzcHwA1sxllZ+AKAl15gqBL+M7TOf+C/27PoaEnxXSbTnehE7HmQvuvVBDR1kLfiD9jxV/icdUg6WsZ3vnYaoV3bCV0Q2gnDSMIHtPsxHWrpn2ZUCX8dXUVb8aDoBl/Mp+ujxScuC8AwaVrkIokbtfmromuoLwpLcJAVYnnJIEYsmSJ1hNi4i/Cd1DEYdWvuuqKtJmHAJqZm7cutF/n0VOR4hQXGGfRXqWgqzoTPww7m039H0sNgwpyVWKVe2iTh93QHMSIQ1An/a0sksVt4kXlX0la051N/RGdAF/lOp+hDBLrQQlgGZJrgTS4VbnhklcSVcKDtmx9+LnGAJHyz4HOKo3JdfTt0AYIw+W1qXTUbtssmJNB2VKkZyonpqmFoKky4cETVfPzJZkLi2dKao2+uATLimlViTQ1B+B8gFgEFiErhz9qRJ9z5ekI11341uU+rZ9ztWKyf/wHctskm6DLas+79M7X+kpCS8Ff3boa6yNEj4mUb+pEpsS/PV/jKcLlYbAahdu/cZHQXy6F6WP/lXfSpeigGLsbY8Uh0X97xlzEr/ryr5lB5hPz0QiA4o8lDwb7Cj51KLGgsaoV3NkaX4VxoUVPhr2KjNcVWORchgfIp9rAvefshrxV1eqMNgDdhVvZXkR9OXNPlI/wuHTZr64/q/fGXLLlo8vjI2Cfvr/2n1wZvwhpRlUlfBmiYPJ3xOfbrczu+8Ohdf3g+eRbm7WpBN1Ajvkj2ZzjutCXoNlMIqPeI8YtV2ZWYK2WWYpmAKGvkh4eRrxrD7dR2M7O3cpfh83qHYY3QiAYfOHus+QolKrpB9eS8FMUq+D7iO3pxfvhsCYyx3ZXu+VxreoOgDNGgxevVUbP8IvHCJrO1GaOGW0lTZW8316RkCuHhM5vKbxbabT7GfScbYNkoNPm5u5CQIVPpE81btFqjhsohkTLet662eJ9Kk2qTmbC1woNrinuD5yHbr9JljEKbZ8vK0TEHJQ5d6/dyqsQhfXE2mGFeKPH7c82n8+PjudB0gNSj/EDUmKof32nrLokixqHVl+2p5p1J/j2uj7Ouc7WgEsBr1BSu5L2ILwjSu9Y60ig85j2staSTpmCOdWrhjVR8NTrootRrd1u+v2WTpfgsjGtH7/DFfGun0AyyMxlna1NB8kvvWMWOo/PHWt+l24GqWBQeNwv/Ad5c0jgZdVx+q02wReUJxF99weSouA/W/CK0ncvObJ7Z2jawen50V0Pw92uBufoFX7oyKWzAor2X+Ip3XdqzsH9Y0qpS+VMheK23tc3MzWeE4fuiY6s/iI2eIgpOXfnGgOAaxgkwQkXRN5NSGthj7DHt31+G3HlT0wL7s4o4GyCeLSv3fTqkWbTDEd1s6Kf7Dane7wfwT5plo9rzFWFvkDJ5I6kfmdGK4utHGaCYNK1gZsG9DAceIFJJK3+a7QuDqkMPJHwRrB/P9cG1L+WCf/4uttiURr4oAhWF6mA0oAuIXBv4mJwq0vz0/HHEHDyRasDCjy0eAPWXuVIHwN6YAKjw25feBftXoJaoAC7uXM8FymW00SPIvEus9MN7eTVFiWRHqPSBEx7svlwE9yzRRoJUfB4C5d3TgiM4FWCkyVwryJrin6OCzIHUqVaqimd0CQr7IUOMbmgF8OMutJzV/48GxFCVI9RZIO7/1AIaWzaN5A0uMJxNAHa7B0SZWe3Pe4stCKLtggoPoNKoCcpl6LJUvV/+o2a4+An+AhtNutCs8xqEr5ODo2mEEK/BLf8FB/DIpjLAYcR/0rcldtoK4wk9AE5ocmdV3Mxor8bAwnOLyZGatHEryEZuBN7apFyGQGVfg2froIXn+sxVLuMv3Zbr97d0osKqVb+MVjB7Sv0EXPjyGN3Gp2cNUy5Dm5XaA5ytjNI15LVKwIXnzloUI5L6x+54qbDNBtJYxaidmGxrjnIhKOCF53Y00sX/y0ULVFQrDUypdp95tkUgf+WURTFALhgUhOcmKYbe9YPKRsZVki8zq519jFonX06BKzGGtVpoCM+1XR/43xjyaeB/Qz82Vk3IOS0Bvn1K26YrFwKBivBFYaeUC8mzHHb/o1pyO7n/2RBw8FBX2kBjunVUhOdOhHsdn+1T8abe5232/hJuaN/HB2eVF+OWJxXK+cG0loZ8NTrCc+vaez5NWyFXDqePZ7zmhg1yt70W3M5sLyYoFfOHNYZ06ygJz73r8Tb+pYo5DU80GVey4UPbYpoJzmwnVDtA+i6wH8AeswF762gJX9m42vvrUZ/ANdJ4XKiOCTwMvN4MFSIXh1WtlFfaMqvIaas4jg48tITnrlTvq/IJ7iCNszoAyLctgm5KjstcxHHbBwmfilqnBtw8uxor5TwNDGrCe5dMKXhCRsjr6P4n3xpMGxGq2TqS4zof4j+ctOrhUO1WK9RBUrZkOf2gJzy3sCpwWKTf43hb0oT/lkYEoy+Rc225a4KD1/WOgHuqbpyDB9U8NxpR2PRJUfjqBdedcv0tP8ft+bcH2DLtQHA5uJmfca70dro56J6W5O3WZcK5CPRCU/hbDuHeTfvO3/LvCk2ez4JtMXU1du65outJeo67rYn0dOsuw4Q+x6ApPHfGwr+tF/jtIKftOcGrRFAsRvalZcaLB+vr6wFit9njjjFWpwlNGWgIf7PNuur3+ubmLm7HcH/Pi3ByhZbqRXs3Ph4iX5g27zzd1673YtlfLdXewafrHypPDJU7/kJ6/dSq1fDvT+DOt/H3NDvnbFy1TPfXVEtGcO2T5KbdmaK/A4bfHFXu+84FNNvrF5Qe9ZXrGrYSfF86X5vmE6xPiqJGXENhzU1ZFn8upHGaWAUSAKG4em8dfGADeu/446mW9ELu9AM/+duRP9B/0hNO7nx6/bQgWoEBTWWK22fYRHBnSVRb9QvCUw4se6SBn0ukVw96MH9T64SsIFxwBYgrre48I0oAAAoZSURBVCv/hXP9fh9qhWqrnuP2p4T9Xz0/47mPu7+7dYCBq9KM4uMmfJveDj1XS1l4jrs0rvZQ/0r+7bGFBvivCgImPXKC+/AL4ItQF16C20PiSsCwgVsmbveMAs6tu+uXTtIldakrw4UvWNo/zh4RFRMeGZ408rvgWmdTg+tQRhdHhD0mJjzC0WXaYYjZwt/XDI8N7znnsPDbWv/AsjCCy3U968pY4S9mOJrP+NnzDS5tHhGRtBbYx7wWfh5k7rUsp/pur8z5PMU8GChUbM6CAfFRAxYc+fnhBHF4Vp3rykjhc3rFLhJH98gdbZkeZLf9N3Hds0W3uGt7t3iwoYWb2ZNb2+LveRXL1L2ujBO+aGg8cd65NNOsYlkeOMfiBxE3y5wb2ABywdTFz9EQfwB1ZZjwW+tL7iUoGtLUyF1lKM4P40lOut0cixtLaWU0RF0ZJLxrdPOrMod3mGksQlHmUsNJMu0414TGIAGLxZcBqStjhK/oMka+YXwldiUlU+TItf4gXyDbCu/NBqiuDBG+IklxicmtJOMDb580H1MqctSsz8oraaDqygjhXd39WItRmaQYZgqYi1Y/Wm+nLAXKhQIArK6MEH70h55Pj5sIhFUduxkhFfaGDqURuEcH57oB//tknQeejZ+IzR/sjwRdIlRTV27yJjR6tm7tx/7VcznWhddQVwYIvy3R+86SE547YzHU01F/fJHnque8Bt7ZHg29MncwoBFIXfFc717ba8SfsMe7+rqiL3xR/Zo2qqzw3No+1I2rYRM+WD4MM7Eeutiug9+rR1WD1hV/u/8TM6Ib+ptQXVf0hR+KrDF48i4RQhzwrt7DSbupW+ehDA+kmuGu6kcdvdu+dJfw6T4k5OIVC9gME1pXXOlr7qjJr7TuF/+k25yRaFG1dUVd+JwGMgddkSbTP2pWHeXHwtsjwRTMvfgpQe1HV7hvsbPNhUp/Ghkr/WgakBF4XY0VrvuG+xHgmv0H/nNttEehtq6oC99Lbrhhksn0wFEk3XcbtDkSlNbHmmy9+Gp+0NvGd9/+yD7OcivQLY/VVanwXvQuNT4g/BT7o4VV1hVt4S/K/TAP3WMyLUUz8hOBzZFizgw0VXYfX8tITiM++TxyfJqOvvwQ8Lr6lr/oH2t+Yal88m/ocZV1RVv4qYulj5X+y2QSrTVLNMhHeRS28Xm7cMMjc2MHhFseec5eiwMxAq+rEcKDvia5U7ABm69TV1e0hbfL7DUbYjI9LhqVXq6/43N/OIk7qE3n69iBpCvv5TPQtnwyiMMOvK6a8tecXpO8LgiPXVZdXVEW/oLMToEttUymL0V5RTD3khLTsRcO15+v435oxit8BjpYtgjCJbGoriz8NZFQRmVCBwgL5aqurigLv3SG5KGSp00m359FlCGrMlrg47Ct+DrGWvmP8hnoGpwLELsrRHU1bcSIEYjQR4WuHT57o6quKAvfX3pp9Rj+i+T65KYqzI/BEI4nL+bl5aGVepyv9Fo3ZU7QBZm64vmYtyEGz1JVV5SFjxMvH/Jy+SHRiEQV8z6DNEeC3xW2KXfiK92C5TQCGF2WriuePf/F2yCK/6GqrigLL31nDOBbdoRvujMV0Bop9sjvzF3KN0ZM+FrIwfv0t0KirkoLju2Y01x4w4snCVTVFWXhHVIHTtU2maYS8k92hzNGkq9kXbItEWZKbHjeWPGaWB1wkLM98xv3fyg+oqqu6ArvjJI6wrefHie1TS6qC+qmD8tlGulX2gq1/uwFPHem/utDpeqqWviHfcMAqKorusJL+jrbW4t8w3PXjBi7WyS56KVs0sNCrT8p7rfP0d+TgVRdee74e3qLNxGrqivKj3qpvfENTKZ7iU7NzrUDtEaKVVKzLsv/6q7zOJ9w4FP18HkmQqKulsz56N2urwpm/F1khqq6Co53fA5/w3ckHjnWC84YSb71eX26OWF3y16X8Dz4QF9vOG4ccge/F36Cf8UX/6iqq+Bo1b/Ff4utxCPfkDWAhViDlROFgVrT3f1+IxzscVx/K+THBvKEiVn8J6iqrigL34Louq+sHt9cIq8hnm7ErpqyGN+8I68Lst/RkTwqHwWw449cV17G8Ob8H5ajqq4oCz+S+Ehcwn8HiV9rD8UVzhCE+/wKv71f0D1awsmmE2LkjlxXXr4XmvZYjqq6oiz85vdIuWH8rSQxp2gxxH9xb/Gi1U1389X8iOTI2I96hlX0gNfV9+np6dhSi0Lhl4gNGKqqK8rCl0QSMg/z30DC4zu8LxgiWZPxdL4wQPryGcnyEyH2/eB1NcMkWqxwWRAedYSorq5oz8cnEd6RA/lvsJBcPN2YjbNFosGTZN7C+jJO6e0gTviwulrPm/ACenSb0NJEM9TVFW3h1472zRNWrEvsR6kOBUCdDtgmmoN8b/N+mbUWR2X8lgQAVlcFvA13ok/2aXyt/Q9aXF1d0Ra+3Hct8kn+G7xCLr1pELQ9EmzHlqkLi+rlJkB6wYTEw+vq37wRyBxCxT/4NNoKUFlX1FfZTs/0yeG/gUTU2FifITJaxKLeEF7gLZSZ6z7rf9ANdWB1NVFYzu8doHf1FJ706GYulXVFXfgSs/h92MDku+Sqiq8gGsv+8V3Pms9OoUl/Tx0RNfXcdSuQEVhdFdfjrXjg46qHwME4k2heVm1d0d9JkyVyc3frXvHiMQ+lFgqRGqRovdP7Md9EwrszfkdbMCOwulorLAMw3fevpJ6RT7gtCEPeBKrryoBNk03xN+I3wmwXseBgiaY+Fc5ZvWtct8oKX2QNIKC0ElhdzbgTMyAB7cuprisDhL9UH/txDuK/A3E+8UsjJuZIl/9MVvhOABNzXvC62vZSzeWfmokO16ivKyP2x2fH+dPx+Dlct3Av2nhrgj+l0v2Ou6AJUV1tGmz+0313P/HvvquwyQENdWWIK5SVScpzGictxPc+RVwd5ikXmtsJeEwZqq6McX40P1FuBanATxYd4vUGSHnr6UpFMtqAO+IEqiuD3J2ti5Dvda4NN/p+F3C+8aasrmUD+1PwdAdTV0Y5ODxoXSt98Nbgzga/3z3MjpIZqc2JhNklKwakrgzzbFnUp6XUbNcGi5H9OJxDkWMknBuVjo46Qj6iOxB1ZaAT430xfUnPsM3x/eCjrvmP81PzTEInpGSGeQ4lh6YC+teVoW7LtyUmLsMHcHPTwwadM8gaKUo/tvXdiYlc+cMbtk8oh1bQu64MDlRwfmJsdOq8H3IuFZ77ZeOMHrY2Kwyah5Vn98Cw5mNXHjhdWHj6wMqxzcIG7zHACH3ryvAIFVzJD3Pe6pHcrG3Kh1nHgi9OvJffvs7o1ykpqXO/jG9Iy2zpoGNdGS88wxCY8CEKEz5EYcKHKEz4EIUJH6Iw4UMUJnyIwoQPUZjwIQoTPkRhwocoTPgQhQkfojDhQxQmfIjChA9RmPAhChM+RGHChyhM+BDl/wEYWpw0bvb5LAAAAABJRU5ErkJggg==" style="display: block; margin: auto;" /> ??? This approach is nice in that you can extract information about the connection between metabolic variables. But there is no way to incorporate the disease outcome with this approach and in order to construct the network properly most methods require you provide a prespecified base network, which you might not know. --- ## So, what if we... - want info about network structure? -- - don't know the network structure? -- - have an exposure, metabolites, and outcome? -- - are interested in causal links? --- ## What if we want to know something like this? <img src="data:image/png;base64,iVBORw0KGgoAAAANSUhEUgAABTIAAAI0CAYAAAAurtbHAAAABHNCSVQICAgIfAhkiAAAAAlwSFlzAAASdAAAEnQB3mYfeAAAABl0RVh0U29mdHdhcmUAd3d3Lmlua3NjYXBlLm9yZ5vuPBoAACAASURBVHic7N13vFx1nf/x100jPfQQepGugKCAgChLU7CgiIIIxJa1LbiWX3TXVdRV41rZxRJ3UYwiiuiCCkpREWFXpSmC9N4JLYT05N7fH5+5e+acmTtzy8x8p7yej8d55J4zM2fep9wk85lv6UOSJElSL1oP2BbYFJgNbFb6ebPS+ibAJGA6MBGYDEwBxgMzS/tYAvQDy4DVpWUZsAp4HHik9Odi4OHSz48C9wNrmnt4kiSp2/SlDiBJkiSpqbYAdgZ2Kv05+PO2RFEyhTXAPcBtpeX20vI3ougpSZJUwUKmJEmS1D2mA3sB+wAHAi8lWlh2kkeA60rLVcD/AMuTJpIkSW3BQqYkSZLUuWYBhwNHAAcAuwLjkiZqvLXAX4GrgUuA3xLd1yVJUo+xkClJkiR1jj6ixeUrSssBwIRR7msAeAB4iGzsyseIrt2DPy8HVgIrgHXAs6XXPkMUTAfHypxVWp9KjL05DZhDNtbmHGL8zU2ArYHNR5kZYvzNK4FfAb8EbhnDviRJUgexkClJkiS1v/2BE4FjiaLgSCwjxp68jcoxKVc0MONIzCDG6SyO27kbManQSNwL/Bj4PnBj4yJKkiRJkiRJGo5tgPlE4XFgBMtdwCLgNOAgYubxTjEB2B2YRxzDzcSs6MM99puB04HtW5xbkiRJkiRJ6inrAW8F/sDIindfIrqaz2p95KbbGHg1cCZwJ8M7J+uIsTTfyOi73kuSJEmSJEkq2JRoffkg9Yt0y4DLiBaX26QIm9j2RIvN84gxO+udr4eJVpobJsgqSZIkSZIkdYXnA2cRE+rUKsatAM4HjqGzuoo321TgzcBFwBpqn8OlwL8DOyRJKkmSJEmSJHWgrYGFwFpqd4++imh5uXGamB1lQ6Kl5lXUHldzDXHuxzJzuiRJkiRJktTVNiVaBa5i6ELbM8AXgW3TROwKuwLfJLrh1+qi/xm6c1xRSZIkSZIkaVSmAB+n9piOdxKtL2ckytiNNgL+CXiIoc/7E8AHgImJMkqSJEmSJElt4WDgdoYupN0AvA4YlypgD5gEnET967B3qoCSJEmSJElSKjOBM4ixLqsVzu4jxnQcnypgD5pInPOHGXr8zDOAaakCSpIkSZIkSa10NPAA1YtljwOn4uzjKU0DPgYsofo1ug04MFk6SZIkSZIkqcnGA5+l+qzZ64CvES011R42Bc5h6NaZH0gXTZIkSZIkSWqOjYBLGHoin0PSRVMdryS6+le7dj/EruaSJEmSJEnqEnsBdzP0mItT00XTME0DFlB9TNNbgF3TRZMkSZIkSZLG7pXAciqLXw8BByTMpdE5CniKyuv5FLBfwlySJEmSJEnSqB0NrKCy6HUVMCdhLo3N1sA1VF7X53CIAEmSJEmSJHWY44DVVBa7FuKM5N1gMvAdKq/vMuCwhLkkSZIkSZKkYTsRWEu+wNUPvDtlKDXFJ6ksZi4HjkgZSpIkSZIkSarnIGAV+cLWWuBtKUOpqT5MZTFzKfCClKEkSeo1fakDSJIkSR1kG+BPwKZl29YRRcxFSRKNzkbArCEeWw082KD32RiYOcRjq4gJkTrFu4Gvkf8MdS+wL7A4RSBJkiRJkiSpmmnADeRb5a0D3pQy1Ch9lcoWhuXdptdv0PtcX+N9/tCg92ilU6k8jl8DE1KGkiRJkiRJksr9kMoi1r8kTTR6tQqZA8A7G/Aez6/zHp1YyAT4OpXHckbSRJIkSZIkSVLJKVQWr35C5w7VVK+QeVUD3uMrdd6jUwuZE4HfUDnR0ytThpIkSZIkSZK2AJ4mX7j6M9HVvFPVK2QOALuMYf8TgEfr7L9TC5kAmwD3kD+e+xl6PFBJktQA41IHkCRJktrcAvJjRi4H3ggsSxOnaR4rrJ80hn29CphdY9+dbjFwAjFG6qCtgH9OE0eSJEmSJEm9bl+i23B5y7v3J03UGNVaZH6xsP4gMH6U+7+gsK8vVHm/Tm6ROah4XCuB7ZMmkiRJkiRJUk/6FZVdykdb3Gsn1QqZBxMtDcu3HT6KfW8KrC7sZ48q79cNhcwpwH3kj+uspIkkSZIkSZLUc15EZfHt0KSJGqdaIXN/4MzCtu+PYt8fKOzjamBSlffrhkImRBf88uNaDWydNJEkSZIkSZJ6yiLyBarfpY3TUEMVMl9c2Lac/Pigw/Hnwj7eSXcXMscDt5E/ts8lTSRJkiRJkqSeMYuYzKe8OPWKpIkaa6hCJsBfqCxEDtc+VC+EdnMhE+Dt5I/tEWBi0kSSJHUhZy2XJEmSKh0LTC1bvxe4NE2UlvteYf2UEby2+NyfAs+MLU5HOBdYUra+GXBYoiySJHUtC5mSJElSpdcU1hcRs5f3gu8Ba8rWDwR2GcbrJgEnFLZ9t1Gh2txy4EeFba9OEUSSpG5mIVOSJEnKm0hla7oLUgRJ5DHgksK2twzjda8BNi5bfxD4TaNCdYALC+uvTJJCkqQuZiFTkiRJynsBMK1s/WFiApteUmxJOZeY1KaWuYX1s4F1jYnTEX4DrCpb35boYi5JkhrEQqYkSZKUt29h/Q/EBC695GfAE2XrWwB/V+P5s4EjC9u+3+hQbW4lcENhW/FekiRJY2AhU5IkScrbqbB+fZIUaa0mJrApV2vSn5OBCWXrVwG3NTpUB7iusL5zkhSSJHUpC5mSJElS3naF9buTpEiv2L389cD6Qzz3pDqv7RX3FNa3TRFCkqRuZSFTkiRJytuksP5gkhTpXQfcWLY+BTiuyvP2JcYVHbQCOL+JudrZA4X12UlSSJLUpSxkSpIkSXnTC+vPJUnRHhYV1qt1L59bWD8feKYpadpf8V6ZVvVZkiRpVCxkSpIkSXkTC+trkqRoD98jf/wHAruUrU8Gji+8ple7lUN+1nKA9ZKkkCSpS1nIlCRJkvJWFtYnJ0nRHh4HflXY9payn48BNihbfxC4osmZ2tnUwvryJCkkSepSFjIlSZKkvKWF9aEmuOkVZxfW5wLjSz8Xu5p/B1jX5DztrHiv9PKwBJIkNZyFTEmSJCnvocL61klStI9fAIvL1rcA/q705+Fl2wfo7W7lUDlLeXHyH0mSNAYWMiVJkqS8ewrrOyVJ0T5WA+cWtp0CnEzWMhPg98BdrQrVpnYsrN+bIoQkSd3KQqYkSZKUd1Nhfd8kKdrL2YX11wPvKGzr9daYAPsV1ov3kiRJkiRJktQw2xPdpAeX5+i+2ae/Sv4YB4D967zmz1VeU36OZtR47aQqr/nD6OO3pdlAP9nxrQGmJU0kSVKXsUWmJEmSlHc3+XEypxFjQva6Wi0uf0LlJEm95migr2z9BmBZoiySJHUlC5mSJElSpV8U1t+QJEV7OYdoZVjN2S3M0a6OLaxflCSFJEmSJEmSesoryHeDXgpMT5qosUbTtRzggiqvu5f6DSS6vWv5VsBa8se3V9JEkiR1oQmpA0iSJElt6DKie/kWpfXpwFzgzFSB2sSHiZaZ5e4jxobsZe8hP4P7DcSYopIkSZIkSVLTfYp8C7v7iJaF3WC0LTJHq5tbZG4ALCF/bO9KmkiSpC7lGJmSJElSdV8Dlpetbw28O1EWta+PAjPL1h8HFiXKIklSV7OQKUmSJFX3GPBfhW2fADZOkEXt6XnAqYVtXyZfAJckSZIkSZKablPgGfLdhr+fNFFj2LV87PqAS8kf04PAtJShJEnqZrbIlCRJkob2OPDZwrYTgdclyKL28m7g8MK2jwDLEmSRJEmSJEmSmABcS77l3bPA7ilDjZEtMsfmJcBK8sfza6KVpiRJahL/oZUkSZLq2xP4E/lZy+8B9gWeSJJobPYCdixs+w3wZJPebxxwbGHbU0Txr9NsDlxT+nPQMmAP4O4kiSRJkiRJkqQy76OyVeGlRItN9YYpREG7/B7oB45PGUqSJEmSJEkq+k8qi5ln4djzvWA94OdUXv9PpwwlSZIkSZIkVTMRuILKYta52DKzm60H/IzK634BFrElSZIkSZLUpmYD91NZ1PoRUehUd5kGXE7l9f4rMCNhLkmSJEmSJKmuHalezLwQmJ4wlxprI+BKKq/zzcCchLkkSZIkSZKkYdsGuJPKItetwK4Jc6kxXkjMQm4RU5IkSZIkSR1vK+B2KotdzwKvT5hLY/MWYBmV1/U6YOOEuSRJkiRJkqRR2xz4M5VFr37gX3HczE4yFfgmlddygJjkaf1kySRJkiRJkqQGmAx8m+oFsBuBF6eLpmE6iBgWoNo1XIgFaUmSJEmSJHWRecAqKgtha4AziBmw1V5mEtdmHZXXbQUwN1kySZIkSZIkqYkOJsbIrNay7zbg8HTRVKYPOBZ4gOrX6k5gj2TpJEmSJEmSpCbaALiI6oWx8uUy4EWJMgoOAK6k+rXpJ7qSz0iWTpIkSZIkSWqiPYlWfOUFsQuA+xm6YHYesGOKsD1qd+KcD1VgvhM4JFk6SZIkSZIkqclOBJaRFcSWAseVHpsJfIMoXFYrnq0mJgqyG3PzvIQoYFYbB3NwDNPPEZM2SZIkSZIkSV1nArCAynEwd6/y3BcBlzN0a8AB4CqiADq+2cF7wDjg1cQ5rdfN3yKyJEmSJEmSutbmwNXki2IXArPqvO4I4DpqF9duA94HbNSM4F1uc2A+Q3fpH1x+R7TUlCRJkiRJkrrWS4GHyYpia4niWd8wX98HHA/cTu1i2yqiOPpGYErj4nedGcApwKXEtah1Tm8AjkoTU5IkSZIkSWqdecS4loOFscXAYaPc1zjgGOAKahffBoAlxFiaR+BYjgDTgNcC5wLLqX3u+onZ5A9j+MVmSZIkSZIkqSNNB35IvkB2LbBNg/a/F7AQWEH9ouZyYmzH+cBuDXr/TrA9UUj+ObCS+udpJbCI6mOWSpIkSZIkSV1nR+Cv5ItkC4FJTXivOcA/A3+jfqFucLkdOJOYPX3HJmRKYRxRpJ0LfAu4j+Gfj+uAfwQ2bHVoSZIkSZIkKZVXAU+TFclWAO9o0XvvDXyZ/Hicw1meILpSfwJ4BTEBTrvbmphl/NNEa9NnGNkx3wP8K7Brq4NLkqTWcHwYSZIkqbrxwL+UlnGlbQ8AxwLXJMhyKPAGojC51Sj28SzRcvM24Naynx8BHm9MzJrGAZsAWwA7F5adiDEvR+pO4GLgx2QzyEuSpC5lIVOSJEmqtBHwA2JinUG/JWYab0XRr57nA68kipoHMfYu7muJ41pMtP5cXFqWEBMbrSLG5VwLLC28dhZRpJxWyjEZmAlsWlrmEAXMTYiC7FisICZG+mVpuXOM+5MkSZIkSZI61guBu8nPeL2AsRfhmmU60SX7s8BviJaXI+mS3c7LU0TB8pPAkcCUBp0zSZIkSZIkqaOdTLQ8HCykLQFelzTRyI0nWmy+jZgk51qiFWXqouRwipb/C/wHcBLR3dweZJIk6f/4HwNJkiQJ1gP+HZhXtu1W4PXALUkSNd7g2JQ7lf7cBdiO6Po9s0UZ1gATSz+fBfyRbMzOduiyL0mS2piFTEmSJPW6LYHzgf3Ktp0LvBNYliRR600mxrDcnGxsy42pPv7lFKIY2Q+sI1qwriI/juazZGNuPgY8Wlr/PPD+0nu+FvhZ049MkiRJkiRJ6gIvJ4psg92b1wDzUwbqcvPIzvVHEmeRJEmSJEmS2l4fUbBcS1ZYeww4JGWoHvBSsvP93cRZJEmSJEmSpLY2g+hKXj7RzO+JsSLVXBuRnfM/Jc4iSZIkSZIkta2dgZvJFzEXEuM/qjUWE+d9KY7ZL0mSJEmSJFU4BlhCVsBcDsxNGahHXUl2DbZMnEWSJEmSJElqGxOABcQs24MFtDuAF6QM1cO+SXYdDk+cRZIkSZIkSWoLmwCXk+9K/gtgg5ShetxpZNfi1MRZJEmSJEmSpOReBNxHVjTrJ1pmjksZShxBdk2+kTiLJEmSJEmSlNQ8YBVZwewJ4MikiTRoK7LrckXaKJIkSZIkSVIak4GzyHclvx7YLmUo5fQBzxLX5vHEWSRJkiRJkqSW2xq4hnwRcxEwNWUoVVV+nTZOnEWSJEmSJElqmaOAp8iKYyuJSWXUnhaRXauDEmeRJEmSJEmSmq4PmA+sIyuMPQDsnzKU6voo2fV6Z+IskiRJkiRJUlPNBP6bfFfyK4DZCTNpeI4hu2ZfTpxFkiRJkiRJapo9gTvJimH9wBnAxJShNGw7kV27XybOIkmSJEmSJDXFicAyskLYs8BxSRNppCYQ45gOAPemjSJJkiRJkiQ11gRgAfmu5LcBu6cMpVG7iaw17fTEWSRJkiRJkqSG2AK4mnwR80JgVspQGpMfk13LfRJnkSRJkiRJksbspcAjZEWvtcRM5X0pQ2nMPkV2Td+SOIskSZIkSZI0JvOA1WQFr8XAYUkTqVFOILuun0mcRZIkSZIkSRqV6cAPyXclvxbYJmUoNdReZNf2p4mzSJIkSZIkSSO2I/BX8kXMhcCklKHUcJOJYQIGgFsSZ5EkSZIkSZJG5NXA02QFzBXA25MmUjPdRVzn1VioliRJkiRJUgcYD5wOrCMrYt4PvDhhJjXfL8iu966Js0iSJEmSJEk1bQRcQr4r+cXAhilDqSW+QHbNj02cRZIkSZIkSRrS3sA9ZMWsfmABMC5lKLXM28iu/ccSZ5EkSZIkSZKqOhlYTlbIWgK8LmkitdpLyK7/OYmzSJIkSZIkSTnrAd8i35X8b8AuKUMpifXJ7oHrE2eRJEmSJEmS/s+WwB/IFzF/AExLGUpJPULcB8twSAFJkiRJkiS1gZcDj5IVMNcA81MGUlv4Ndk9sW3aKJIkSZIkSeplfUTBci1Zweox4JCUodQ2ziS7L16ZOIskSZIkSZJ61AzgfPJdyX8PzEkZSm3lvWT3xgcTZ5EkSZIkSVIP2gW4mXwRcyEwMWUotZ2/I7s//itxFkmSJEmSJPWY44HnyApUy4G5KQOpbc0hu0+uTpxFkiRJkiRJPWICsADoJytO3QG8IGUotb2niHvl6dRBJEmSJEmS1P02AS4n35X858D6KUOpI/wv2T0zO3EWSZIkSZIkdbEXAfeRFaP6iZaZ41KGUsf4Ntm98/K0USRJkiRJktSt5gGryApRTwBHJk2kTvNhsvvn3YmzSJIkSZIkqctMBs4i35X8emC7lKHUkY4mu4f+PXEWSZIkSZIkdZGtgWvIFzEXAVNThlLH2p7sProscRZJkiRJkiR1iaPIZpkeAFYCpyVNpE43DlhG3E8PJc4iSZIkSZKkDtcHzAfWkRUxHwD2TxlKXePPZPfVrMRZJEmSJEmS1KFmAv9Nviv5FcDshJnUXc4lu7f2S5xFkiRJkiRJHWgv4C6yIlM/cAYwMWUodZ2Pk91jc9NGkSRJkiRJUqc5kWzswgHgWeANSROpWx1Hdp99PnEWSZIkSZIkdYgJwALyXclvBXZPGUpdbXeye+1nibNIkqQ2NCF1AEmSpGHoA9Yv/TwTGA9MASYTRY9nSo8tBdYCK4iZtDU6WwA/Bl5Stu1C4BRgSZJE6gV3EL+/E4BdE2eRJEltqC91AEmS1NO2AHYGtgI2Ky2blP6cDWxaWh/t/1mWAY8CjwGLgUfKfn4QuJMonqwe9RF0n5cC5xHXAGKG8n8G/o0oGkvNdBuwE3HfzSC+lJAkSQJskSlJkppvHLALsAdRtNyZKFTsRBQqmmkasENpGco64F7gdqLr9O3ALcB1wHNNztdu5gFnkk3i8wRwAnB5skTqNbcQfzeMB3YEbkwbR5IkSZIkdbNZwGHA6cDPiWLYQAcua4GbgUXAacA+RHGl02xI1rpyKNOBH5E//muBbZobTarwObJ78E2Js0iSJEmSpC4zA3gdsBC4i+YVFpcATwEPld7nVqLY9pfS+t2lx58C1jQpw7PAL4FTidZineBHZOOLVrMj8Ffyx7kQmNT8aFKFk8nuw9PTRpEkSe3GMTIlSdJo7AG8orQcRNYVeaSWEF257yDGrhwcz/Jx8mNbrhnl/jeg+ribmxGtDXcCtmb0/ye6C/gVUdz8LbB8lPtplncTM4/PGuLxVxMtTgcLnSuB9wFnNT+aVNWLgT+Vfj4PW2VKkiRJkqRR2Bn4FDFBzkhbMj5JFPu+SIzD+DLqd3dulanAXsAbgY8B5xDj9PUzsmNcDvwAeBWjL+w20u5EYfKmKo+NJ1q7rSPLfx9RRJJSmk72u+f4mJIkSZIkadg2JAqPVzH8wt4asrEl5xEFtXGtDt4AM4jWpvOJsT4fZ/hFzaeI4z+MND1gJhNFoAEie7mNgEvJ572YuNZSO7ifuC9X4uSkkiRJkiSpjpcDFzD8sSZvBb5KdDWf0vq4LbMb8AGiELiS4Z2b24kxNZs9Q3u5r5W9/5ll2/cG7il7rJ/oet6JhWZ1r0vI7tHnJc4iSZIkSZLa0CRioo3rqV+cWwH8jBiDcbsUYdvANKIb+ZnAvdQ/Z88QXeu3aXKuo8m3nv1QafvJRPf38smTXtfkLNJofJXsPn1N4iySJEmSJKmNzAL+GXiY2oW4dcCvgbcy9OQxvaqP6Ib+TWI80Hpd788DXtSEHFsAT5Mf+/LNwLcKGf4C7NCE95ca4e/J7tX5ibNIkiRJkqQ2sB7wj8Ss4LUKb38BPgxsmSZmx5kEvBb4MbW7n/cTBc2dGvS+44DfkC9iDhCT/ZSvn0O0JpXa1cFk9+vZaaNIkiRJkqSUxgHHAXdRu/Xlz4nJajR6mxItyh6kdgvNhcDmY3yvj9d4j8H3sXWbOsEmZPftHxNnkSRJkiRJibycbDbrasuzxPh02yfK163WA95G7XP/HPBJYsbxkToQWFtj388AHyUm+9l4DMchtcpgS/ElxNANkiRJkiSpR8wiWv2VTwJTviwBPoZjX7bCkcC1DF10vIUoTA7X+sAD1C5kFpflxEzzFwHfILrCS+3kSrL7dYvEWSRJkiRJUoscDdxP9YLWaqLAOTtZut7UR3Tvv52hx89cCMwYxr7OY+gCdb2u5ucCL2vQMUmNtJDsXnWIC0mSJEmSutz6wA8ZegzMRcC2qcIJgInAe4BHqX6d7gUOqfH6dw7xuqGu+QAxO/3pxPidUrt6P9m9+w+Js0iSJEmSpCbaBfgb1Qtad1C7OKbWmwWcQeWM4wNEl/H5VI4TuCPRRbxWa8z+suVyohXo+OYeitQQR5Ldx19PnEWSJEmSJDXJ8cTEMdW6Ei9gdJPJqDVeSoxdWa0oeSHZGKbrATcxdBFzcLzMx4hrvk3LjkBqjK3J7ucr0kaRJEmSJEmNNgH4EtULW9cBe6aLphGYCnyB6q0zbwJ2IsbFrNUCc7D15cQWZ5capQ9YSlaQlyRJkiRJXWIS8BOqF7cWAVPSRdMovQJ4ksrrubTKtgFi5vmFwG4pwkpNcC3Z/b1R4iySJEmSJKkBJgEXUFnYWklMBqPOtTVwDbUn8bkGeDPR3VzqJt8ju88PTJxFkiRJkiSN0VTgEiqLWw8A+yXMpcaZDHybymu8GnhHwlxSs/0T2f3uvS5JkiRJUgebDPyWygLXrcAWCXOpOX5H9W7mB6UMJTXRMWT3+pcSZ5EkSZIkSWNwFpWFrb8Bm6cMpaaYAOwBzKfymj8B7JAumtQ0O5Pd5xcnziJJktpAX+oAkiRpVD4IfLGw7c/AEcDi1scZtcOB9Yd47G5itvVGOJrohl/NbcCNDXqfVvh/wOcL224EDgCWtT6O1DQTgOeI8V/vBbZLmkaSJEmSJI3YEcBaKltibpAy1CjdwNCT2NxOY7503Rbor/E+/9aA92i1j1N5HOfjl9TqPjcT93c/MD1xFkmSJEmSNALbAk+RL2A9BeyUMNNY1CpkDhCtDMfqk3XeoxMLmX3AeVQeyz+lDCU1wflk9/feibNIkiRJkqRh6qNyhvJ1wFEpQ41RvULmwjHuvw+4q857dGIhE2AKcA35Y1kD7JMylNRgnya7v09MnEWSJEmSJA3Tu6kswn0oaaKxq1fIXMLQY1sOx9/V2X8nFzIBtgEeJ3881wPjU4aSGujNZPf2v1Z5fGNgt5YmkiRJkiRJNW1AzE5dXrC6nM4fE7FYyFxFTPJTvu3NY9j/osK+bqW7CpkAr6HymN6VNJHUOC8ku69/UuXxC4AdW5pIkiRJkiTV9BXyhapnga2TJmqMYiFzJXB6Ydslo9z3dGBpYV8fpfsKmQA/JH9MjwMzkyaSRm73KtumEENoDBCTmpV7LzEJ0OQm55IkSZIkScM0G1hOvlA1P2mixqlWyNyW/Czj64CtRrHvdxT2fRNwON1ZyNwCeI7uvEfUO+YC3ySKl+UGW2mvBiaWtj2faMH9eKvCSZIkSZKk+j5PvkB1H93TAqlaIRPgisL20czGfVVhHx+kewuZUNmS9TEqC0JSO5sEPEC0vNyjbPtFZPf1rsR9/bfS+jUtzihJkiRJkoYwCVhMvkD1nqSJGmuoQubcwvbbGdl4oDuSb9W5BtiM7i5kzgKeIX9sJyVNJI3cB8jGy30f8Xv/RbJ7+vXA18rWz0sTU5IkSZIkFR1PvjC1mO5pjQlDFzKnEeOAlj92wAj2+5nCa39W2t7NhUzIF3wGgN+ljSON2DTgKbJ7+FKy4uYA8F2yLynWAV9IE1OSJEmSJBVdQL4w9ZW0cRpuqEImwHcKjy0c5j7HAfcXXnts6bFuL2TuRL4laj+wZdJE0sh9ivwYuU+Wra8C1patn5oooyRJkiRJKjOFyglc9k6aqPFqFTJfVnhsCTB1GPs8ovC6J4H1So91eyET4H/IH9/fp40jjdgmwAryxczi7+3g8tpEGSVJUgLjUgeQJElDOpDoZjnoAaLw1yuuBO4qW5/J8IoWcwvr5xCtuHrFzwvrhydJIY3eYuA/iRbFUPszy/3NjyNJktqFhUxJktrXfoX1S4kWSL1iAPheYdspdV4zi8pi59mNCtQhLius758khTQ2XyIrZNZyX7ODSJKk9mEhU5Kk9vWiwvofkqRI62zyxYzDga1qPP948t3PbwKub3ysSGFt5wAAIABJREFUtvYXolvuoC2AOYmySKN1H/BDahczlxMTA0mSpB5hIVOSpPa1Q2H9xiQp0rqP/Mzb44C31Hj+3ML6dxodqAOsAW4ubHteiiDSGH0W6KvxuN3KJUnqMRYyJUlqX9sV1u9JkiK9swvrc6le3NiJfHf8tcAPmhOp7RXvlW1ThJDG6BbgYmKyn6J+4O7WxpEkSalZyJQkqT1NBKaXra8lJsDoRT8Blpat70T1cR/fRr7AeTHwaBNztbPicW+UJIU0dguA8VW2D2CLTEmSeo6FTEmS2tO0wvryJCnawzLgx4VtxUl/xgMnFrad3axAHeC5wnrxfpI6xVXA/1LZKnM8TvQjSVLPsZApSVJ7KrZAqta1spd8t7BenNTnSGDLsvUniRaZvapa0UfqVJ+j+j1si0xJknqMhUxJktpTsQXm1KrP6h2/B+4sW58FvLZsfW7h+d8HVjU5UzsrtsBcliSF1Bi/AP5GdCcvZ4tMSZJ6jIVMSZLa0wpgddn6evR29+ABYFFh22D38g2B1xQeK7bg7DXFMTGfTZJCaowB4PNUTvJli0xJknqMhUxJktrXA4X1bVOEaCPfId9l+nBgK+AEotA76CbghhbmakfFGe9tuaZOdy7wRNl6P/BwoiySJCmRCakDSJKkId0N7FC2vhtwc6Is7eBB4Arg0NL6OOAtwOsLzzurhZnaUR+wa2HbPSmCSA20Bvga8InS+tPATsCmwGbAbGATYPPStqnEEBQAM4kxNqcAk8v2uYxo+b6KGM6jH1hCTJb1MPB4aXkEWAw8RnzB1MvDVkiSlJSFTEmS2tdfiFaHg/ajcvbuXnM2WSET4B+J4sWgNcAPWhmoDT0P2LhsfSlRFJc6zQRga2D70rIJ8Ts+kRg+4W9j3P8Go3zdI8SXSneXlr+V1u8liqGSJKlJLGRKktS+/lRYf3mKEG3mJ8B/AOuX1jcpPH4R0YKql72ssH4NznqvzrANsD/xpc2+wN5EK8p2M6e0FD0N/JH4u/uPpeXJFuaSJEmSJCmZ2UQBaqC09ANbJk3UeDeQHd8AsHIYr/nPwmvKl2PqvPbwKq/5t9EEb2MXkD++f0kbRxrSrsAHgAuJVo5D/V538nI78H3grVQvfkqSpBGwRaYkSe3rMaJFz0tK633AG4EvJ0vUHr4LvKPK9ieBi1ucpd2sT344AoCfpQgiVTGdGBriFaVl2zHsay3RlfsxYvzKh0t/Do5puZQY77KfGP9yFdEt/bmyfaxP/L06DZhUWqYRQzNsWlrmlP28eWkZyYSpO5aWE4nC5l+AXwK/Av6ndBySJGmYLGRKktTefkpWyAR4OxYyrwJuBXYpbP8eMXFHLzuJmORk0N3AjYmySBAT7bweeDMx7MGkEb7+UeAWomXj7cTv/rPEpD2XjzHb06N4zWRikqHBZefSsgvZ5EJD6QP2Ki0fJQqtFwPnAJcShVZJkiRJkjrWbKI4V95V8cikiRprNF3LIVpHbV9Ypg/jdd3ctXw8UegpP7Z/SppIvWo8cBiwiGgBOdxu2KuBa4EzgJOJ3+uh9DUp+1hsT+Q+g/jCZSXDP/angIXAQbTnsUmSJEmSNCznkf/A+/u0cRpqtIXM0ermQuZg19XBZRWOyafW2omYjOsJhle8W050sX4/0Uqx23qLTQVeCpwO/IH8mMe1ljuAfya6uEuSJEmS1FFeSIzzVv5B97VJEzWOhczGmAosI39c30qaSL3kIOILl7XUL9LdRbQ8PA6YkSJsQhsRx70QeJD652ol0ar1+SnCSpIkSZI0WhdR2WJnctJEjWEhszG+RvVWXQuJrq7bpYumLrUeMRP3X6hfkLsb+DSV49r2snFEa81vEhOV1Tp//UTL1SOx27kkSZIkqQPsRkwEUf7h9vNJEzWGhcyx25nKcVSrLfcQEyK9i2jhNZKZl6VB44hWhXdRf8zHRcRYmRbfapsEvJrhjSn6R+CQNDElSZIkSRq+r5P/QLuW6NLZySxkjs0kYuy98uNZQczsXK+w+SxwGTF+32HAlNZGVwc6GriR+oW24xn57OQK6wMfIL54qHWeLwL2TJRRkiRJkqS6ZgL3kf8wu5jO7jZsIXNsvkXl8by59NgcopXXAmIG5VVVnlu+rCGbMfo4YJNWHYTa3ouB3zH0vbMW+DFwQKqAXWg88AbgaoY+7+uIVtZbJcooSZIkSVJNR1A58c91xGQvnWhLYPuypdlF2SmF99se2LDJ79ksp1JZ2PjvGs+fRrTgnQ/8HHi6yuuLy11Ed9d5wO7YRbjXTAO+wtCT+KwkCt/bJsrXK/Ylfrdrta5+Lw4XIUmSJElqQ6dT+UH2fCwy9ZLDqBwz9TZggxHsYzxRnJxHFCvvpX5h8zGiCDqfKIquN+YjUbs6GLidoVsCnkd8EaDW2Rf4DUP/fl4N7JosnSRJkiRJVfQBP6LyQ+xCbJHTCw4EllDZImv3Bux7c6Jb+RlEN/N11C5sLiO6rS8gurF3autWZdYHvk1ly+/ysRn3SJZOUHus0hXAR4gvKiRJkiRJagvTgb9Q+SH2+/gBtpsdTBQti63jXtWk95tJtP48nZgYaAW1C5vVuqOrc+wB3MnQ1/XwdNFUMA54N5VfagwulwEbJ0snSZIkSVLBdkRX3+IH2O9hMbMbHUG0gCxe7w+1MMMEYB/gNKJr8eIqeYrLw+S7o09sYV4N3/HAc1TvRr6Q+PJE7WcO8FOq/+49QHRHlyRJkiSpLexGFIqqTfoyI2EuNdYJVG8N+dGUoUq2B04mil03M3SX5MFlKfnu6LNaH1llJhDXotq1ugnYL100jcBxwONUXsOVwDsS5pIkSZIkKWc74G4qP8Dehl17O91QRaZ+4AMJc9WyGVGgXEAULFdRu7C5liiALiQKolu3PnLPmgpcQvXr8g1gUrpoGoXZwO+ofj0/nzCXJEmSJEk521B9bLtngTckzKXR2wT4NdWLmKcmzDVS04gu5fOJLuZPMbzu6OcRXdj3ISa4UmNNo/r9ZQu+zlarhe038HdJkiRJktQmtgKup3rh69+AyemiaYQOAx6i8lquAE5KmKsRxhMthecRkwLdQ/3C5hJi8pLTiXPjvTw2s4CrqT6mol3Ju8NQY55+i5goSJIkSZKk5CYD/0X1YtCdwCHpomkYZhFdrKuNM9nNE3dsTozxdwZwLTHBTK3C5urS884ovW6j1kfuWBsA11B5Tq8nWgGre+xN9Qm5vo3FTEmSJElSG5lH9bEJ+4lCmRMBtZ+jiWJltcLdFcT4d71iBtHy8nSiJeZy6rfavIto4TkPx4Ydynjgl1Seu2uxGNytdqV66+7PpAwlSZIkSVLRAcD9VC/63Ed0PXS8tPR2AH5M9eu0DvgsUYDqZROIsTJPI8bOrDY7c3F5hBiTcz4xRqcT10QL1uJ5ugqYmTLUCE0Etq+xNGrYgVa9TyvsTOWXJP3ACSlDSZIkSZJUNJMoXgzVVfevRNdctd7GxKQcK6l+be4AXpYsXfvbnpjlfCEx63m17vjly3NE0W4BMav6+q2PnNRbqTwnvwempww1CttT+zq/v0Hvc2Kd93lJg96nVXYgJtEqP4ZlRPdzSZIkSZLaykHArQz9ofwy4EXJ0vWW6UQrwSVUvxZriGLbeqkCdqhNiQLlAqJgOVSBeHBZSxRAFxIF0W1aH7ll9qfyfNxLZ46JWa+QeWOD3ueyOu/TaYVMqH4f3E/87kiSJEmS1FamAF8gJkqp9sG8nxg/7wjsct4MWwGfB55i6OLINcCeqQJ2malEAX8+0cX8Sep3R3+Y6Lp+GtGVvRsmRJkK3E7+OJcTx9eJ6hUyB4C9xvgeWxKF7m4rZAKcROWxXJg0kSRJkiRJNWxLtEKrNTP07UQxZ2qaiF1lb2ISmqEKyAPEmKXzcCzMZhpHTAI0j7geN1O/IPYs0TLvdGLyoSlNzDed5rQK/Q8qv7Do5OEkhlPI/MoY3+NfhvEenVrIBPgylcdzfNJEkiRJkiTV8SLgcmp/WH+c6Ko7nFmgbcWZmUmMSXg1tc/vo8B7cSKaVDYn3x29VrF5gOj2fy0x7uxxNL5r9ndL+27U+J17Udmy8D8atO9UqhUyi5M/Pc7of6f6iPFpa+2/0wuZE4A/kj+eh+i88VIlSZIkST3ocOoX3AaAG4APEsWfotOAC4gPyL1qEvAa4EdE191a53Ix0erLwkF7mU6+O/oz1P+9uIto4TmPKPiPpaC/GfA00Q3+fcTM2WPxG/JZ7wSmjXGfqVUrZJ5HFOLKtx0zyv0fXNjPM8DZVd6zkwuZALtROV7mvyZNJEmSJEnSCOwH/JBodVarcLOO6G77LmA7otvt4GMLWx06sanAUcA3gCeoX/S6GXgnze2irMaZQL47+n3Uv8aPEkXQ+URRdKSTNr2rbF93MPqC3KFVsh01yn21k2qFzB8Q48+Wb/vvUe7/rMJ+vgl8vcp7dnohE+Cz5I/pOTpzAihJkiRJUg+bQxQnhzM5SnE5k+7vYr49Udg6D1jK8M7LVUQX5m4/N71gc6Jb+RlEN/N+al/7ZcT1X0DcAxvU2f84sm6/g/v+PfDiEeYstsb81Qhf366GKmTuXNi2Bpg9wn1PI8ZFLRYsu7WQOQN4hPxxfTppIkmSJEmSRmkaMcPtr6g/g2950eYS4JPA0cDGLU/dWBOJsUTfS7TIu5vhF3XvAj4F7NTy1GqlmcREQKcTLZVXUPu+WEu0zF0InEy0ai56AfmW0WuJouZ5RCGvnhdUed8DRnFs7WioQiZUjvv4/hHu+5TC628jvnzo1kImxHAh5cf1BCNvRSxJkiRJUlvZjCgKXMPIW2neCZxDjKN5GLBVi7MP1wxgH+BEYlbfq6lflCouTwBfI4pGtr7sTROJ++g0ovA4nCEHHi4997TSa8cBX6jyvHXEhET1JgQ6o/C6yxt4fKnVKmS+u7D9xhHu+7eF13+ktL2bC5nTqGx9/6akiSRJkiRJapBdqZxUYzTLc8D1wLlES7a3AIcQ4xFu2qTsU4kiyAHAa4nC7DeAXwMPjuFY7iaKl0cx9slZ1J22J1pfLiRaY9a7p5YSXcOfIYqXxcf7S4/Np7L13HhiMqny5x/XtCNrvVqFzFlUTrS11zD3uy35c72O7EuXbi5kAnyV/LH9Im0cSZKG1sszi0qSpJHZn5jEZLC7+KPAG4nJIV4JvALYcpj7mga8sLRUs5Yoxjxe+nNpadtyYBXRKm1Z2fMnEC0qIRuDcGZp22yiRWmjZmteCVwJ/LK03Nag/ap73V1aFpXWNyPGvDyQmBDoxcSs94OmE4X9ofQR9/cC4B3Ah4ALS48dQH5Ih6eBn40tfsdYQhxreYvCU4A/D+O1c4mWsIMuBR5oWLL29m2iNfCgQ4m/L5dVf7okSZIkSe3tGPItne4EnlflebsSBYGvEy0u682A3inLQ0Trt5cRLTulRppGFDTnE18WPM3w783BVoRXEuO5fq7w+PdbeBytUKtFJsSXKuWPPUG+SFxNHzGmbfnrji97vNtbZELl8b8qbRxJkiRJkkbn7eQLkn8gWmEOx1SiQPNBojXaH4kusakLk9WW1UTryguJlm4fJF+8/cQwj1kaq8OA+xndfVwsgp7Q4uzNVq+QOY7Kc3dMnX0eUnj+M8CUssd7oZB5Jvnj+0zaOJIkSZIkjUwfMX5l+YfbC2lMi8TZROvGdwJfBH5MtCi7lege2oxC5Uqiq+g1xBhw/wl8GHgNMbN4tfEtjyXGIxwo/dlNYw2q/WxGjBs7QAyl0Ij7fttWHkAL1CtkQmWr1P+us8/vFp7/jcLjvVDIPJH88V2WNo4kSdU5RqYkSapmAvFh/h1l274N/D1RYBmrx0rL74Z4fDLR6nNzYpzL8nEvxxPF1PVKOQfzDLb0LB9PcykxxuajREu1kfoJMXv0/yMKu98GbgFuGsW+pKGMI363Pk821uv4Ybyuv/RnX2mp9vjRwLeIVtW94jtEN/3Bc/Iq4suTx6o8dzrw+sK2s5uWrH1dW1jfOUkKSZIkSZJGaBpwMVnLnH6iZWavGke04Bw8H/eQn0xFGos9gf+lfsvKJcAjwB3E+LOXAxcA5xCzoX+xtF7+mtuBlwObtupgWmA4LTKh8py+f4j9vb3wvGqTd/VCi8z1qJy1fb2kiSRJkiRJqmM20TJn8MPsWmBe0kTtYSZwM/lul/Zs0VhtAryPGGLhOGKimoOBfYiC3Wzii4XhOo58se2njQzbJoZbyPz7wnNuHGJ/VxaeN7/Kc3qhkAlRKC8/xi3SxpEkSZIkaWjbEy24Bj/EPkd0S1XYmfxERV9MG0eqMJd8Ieq7SdM0x3ALmbOAZYXn7VV4znZkY+AOtkLcssq+eqWQeQf5Y9wpbRxJkiqNSx1AkiS1hRcTXTF3LK0/CRwBXJQsUfu5DXgTUeyAmNX8beniSBUmFdZXJ0nRHpYQk5OVO6Ww/jbyY4teAjzYzFBtbmVhfXKSFJIk1WAhU5IkHQH8mmwcvbuBA4D/SZaofV0CfKJs/WvAvomySEUrCuu9Xogqtkg9iazYOw44uc7ze83UwvryJCkkSZIkSRrCXKLV1mBXwj/RXRODNEMf8EOyc/YwMbu6lNrryHcN/lnaOE0x3K7lEMXK+wvPPab02OGF7c8AU4bYT690LX+C/DFuljaOJEmVbJEpSVLvmg98B5hYWr8MOBR4PFmizjAAvBW4rrQ+BzgfZ/hVeg8X1rdJkqJ99APfL2wb7F4+t7D9HCpbtPaSGcBGZetrgMWJskiSJEmS9H/GA98k3/LmbLKCpoZnG6LoW34OpZQ2If97/Rzx+95NRtIiE2Lc3/IJfdYQk9gUJwKqNUREL7TI3Jv88d2RNo4kSZIkSTEG2s/Jf2BdQH7CCw3fQcAqsnP5nrRxpFxxfQB4Qdo4DTfSQibA1YXnX1dYv7nO63uhkPku8sd3Qdo4kiRVZ9dySZJ6x4bApcCrSuvriMLbR4gPrhq5q4APla1/FTgkURYJYpzbcgclSdFeipP47F3n8V5UvE/+mCSFJEmSJEnAdsCtZK1tVgJvSJqou3yL7Nw+QbQak1IY/GJicLkobZyGG02LzJlUdiUv72peb7Kubm+ROZ7KiX5emjSRJEmSJKln7QE8SPYB9UlspdVoE4Eryc7xn4FpSROpVz2ffEFqBbBB0kSNNZpCJsRkPtUKmb8Yxmu7vZB5KPljexKYkDSRJEmSJKknHQosIfuAeg+wS9JE3Wsz4AGyc/0THHtUadxBvjD1vrRxGmq0hczDq7xuADhuGK/t9kLmD8gf26K0cSRJkiRJvegkYDXZh9MbgS2TJup+ewPLyc75R9LGUY/6GJWT2XTL2PijLWSOI4bTOK6wrDeM13ZzIXNzYqiR8mNznF9JkiRJUkudBvSTfTD9NTAraaLecRLZeV9HNrmS1CpbEGM/lhenjk2aqHFGW8gci24uZH6Z/HHdji3JJUmSJEktMh44k/wH0+8Bk1KG6kFfITv/zwK7pY2jHrSI/N8DN9Ed4x5ayGycramcBOnvkyaSJEmSJPWM9YAfkf9Qegbd06W0k4wHLia7DrcC6ydNpF6zK9EiuPzvg/cmTdQYFjIb51zyx/Qgw+tqL0mSJEnSmGxAftbsfuBDSRNpQ/KTrvyKKHBKrfJt8oWqJUQrvE5mIbMxDiM//MgA8PakiSRJkiRJPWEL4C9kH0ZXAm9KmkiDdiU/a/xn08ZRj5kDLCVfrPolnT0GooXMsVsfuJ/88VyPrfclSZIkSU32fOABsg+jTwMHJ02komPIWj71A8enjaMecyqVRbjTUwYaIwuZYzMO+AX5Y1kL7JsylCRJkiSp+x0CPEP2YfQhYM+kiTSUT5Jdp+XAPmnjqIeMA35LvnDVT+e22p4JzCsshzb5PQ+q8p6zm/yezfIlKouyn0uaSJIkSZLU9Y4FVpCfkXirpIlUSx9wHtn1uhfYNGUg9ZStgEfJF6+ewy8+es3JVBYxr8YJfiRJkiRJTXQa+dmIf4szYneC6cBfya7b74FJSROplxwIrCJfxLof2CFlKLXMK8h/+TVADEuyWcpQkiRJkqTu1Qd8nvwH0Z8CU1KG0ohsBywmu35npo2jHnMKlS3yHgF2TxlKTXcUlUXMFcCLU4aSJEmSJHWvScSEFuUfRM/AWWY70WHAGrLrOC9tHPWY/8BiZi85lsqWuP3ACSlDSZIkSZK61wzgEvIfQk9PGUhj9iGy67kaZ5pX64wHzqaymPk4zlzdbU4m/6XJ4L8f70kZSpIkSZLUveYAN5B9CF0FvDlpIjXKWWTX9VFgy7Rx1EP6qN4ycyW2EO4GE4AFVF7fdcDbEuaSJEmSJHWx3YD7yD6ELgWOTJpIjTQZ+CPZ9b0emJo0kXpJH/AVKotdA8AiHHu3U20C/JrKa7qWGCNVkiRJkqSG25/8pDAPAy9MmkjNMAd4iOw6fy9tHPWYPuCzVC9m/gHYJl00jcLB5P8+GVyWA8clzCVJkiRJ6mLHEB88Bz+E/g0LCt3sJUSX3sHr/YG0cdSD3gQ8R2UBbBkwnxhXU+1rKtGVfC2V1/B+nJ1ckiRJktQk7yPGMStvFbVx0kRqhbnku4C+Mmka9aJdiC9NqrXOvBrYNV001XAwcDvVr9tvgU3TRZMkSZIkdas+Yiby8g+hF+A4db3k62TX/ingeWnjqAetD/yM6kWxFcDH8O+kdjEb+A4xC3nxWvUTQwbYklaSJEmS1HATgP8i/0H0rNJ29Y6JRAuqwXvgFmBm0kTqRX3AycCTVC9oPkjMbO7fT2lMJ7r7L6H69bkbOCxZOkmSmqwPOK+J+z+F+PZWkiRVN534t3iwK/EA8CmidaZ6z0bAn4DtS+sXAq8nWlhJrTQHOJO4/6q5Ffg4cD7x95aaayLwVuLfh9lVHu8nvhD7IDHeqSRJXavaN3mNWma08DgkSeo0mwHXkf27uQZ4Z9JEagd7kp945fSkadTrTgAeZ+j/7/8BeAN2Y26WGcCpwL0MfQ3+CuyXKJ8kSS1nIVOSpNbbAbiD7N/M54CjkiZSOzmWbOy7fuC4tHHU4zYEvkj0tBrq//33AB8AZiXK2G22Ic75Mwx9zh8D/gGYlCijJElJWMiUJKm19iXfwulRYJ+kidSOFpDdI0uBF6SNI7EV8G1gLUP///9Z4KvA7okydrJxwMuJ4UbWUPscf5wYmkRS5xgPPB94HfBuYrzb+aWfX0/8O2/rdqmOPirHtFkBfLJB+/8KsLpB+5IkqRu8BjgXmFpavxt4BdE6Uyo3jhgj81Wl9XuBFwNPpAoklexGzIr9GuLzxFBuAL4P/BB4uAW5OtXzgbcAbyaKxUNZBXwT+AywuAW5JI3dJOAY4ETii4p6k/gtJSb+Owe4AOspUlXFb/ieShtHkqSu9VbyrWz+BGyaNJHa3QzgZrJ75jKcLVrtYzfgW8ByavfSWkvcu3OBjVMEbUM7AB8G/kz9Xm6PEWPlVpvkR1J7GkeMe34/o+/heh/wdmp/YdTu/pGs5el84LC0cdQtLGRKktR888n/e3spDsGi4dkZeJrs3vlS2jhShVnAacAD1P9gvg64lhg64TBiNu5eMIU43gXE8Q+niHEbcV6nJMgrafS2BK6kcUP2XQFs3soDaKBnyR/LGWnjqFtYyJQkqXnGAwvJ/1v7HXrnw7sa40jy4xK+LW0cqapJRPfJS6g9jmb58gQx3Ma7gBfSPS2OpwEvBT4E/JL6rVYHl+eI7viHtj6ypAbYhxj7vNaXOX8Czgf+vbScD/yx9NhQr3sI2LOFx9EoFjLVFBYyh9YHbEv8ZbQnsFmL3ncjYJfS++5A8wfy3gB4Xun9Bo91G3pjBsSZxKDK+xDjE9Ubs6Sa8cS98XziP+BbAZMbFVBSR5sG/IL8v7ML6OwuQkrnn8juoxXEpFFSu5pDdCkcbuvDwWUZ8Htixu7jgO2ILprtbCIxudFbiTEsb6D2ZD3FZQ1wMTFO5rQWZ5fUOHsATzJ0IfIfqN2ycg7wvtJzq+1jMZ03kZqFTDVctcl+ngY2bPL7nkrlL+AVxLexI/VC4hvccsuBjxADYhdtTAyQXe57wFWln/uISRfeSXT/KHb7W0oMvL+IGOunEaYCbwKOBg6nejHtIeI/OBcQ3+oWr9tITASOAk4A9ieKltX0l973z8Q3RBeVfq637zML235KfDM/UicDB5atP0l8iKvnm+SLBL8kztugHYH3EOd7xyqvH0f987srcCzwSmA/KmeXGwCuL7332cD/Z+++wx0py8aPf89WFlh6B0E6SlN6EwEBQcGGWBEUFbvga8GCoq+IqEixAxaKiCJYKC8oIgiigA3pIB2ks7CwS9s95/z+uJNfksmkniRPkvP9XNdcu5lMudPmzNzzPPdzexNxSxouywLnANsVHo8SJ6c/SBaRBt0I8DPgLYXHDxCD//w3WURSYxsTx7+3EOdeL6X1mznPALcWpluAmwv/3kXvBr+aStQ0Xpco97Be4d8NiGRrqy1JnycStucAvyDqYEoaXMsDfyP/2vr7wGeAuU1ua0nga8D7cp67nbiROSgN0J6kMqfyLaJkhjQhKVpkvpjoNlG+32eJVnGtWIIY5TX7Gt5dZ501c5Z/T+G51YA/5Dxfa/oVcdekXSPAATRXT6h8+iulC+NW7Uyc/LWyv/LpvAbbXyRnnUPbjPVHme3c2eR62Sb5xcT1IsBxNO7qVO+u/+pEYrJes//s9BxxwF6qyfglDb41iYvs8hZGeyeNSMNiFpUt3P4CzEwakVTbq4kk5LfL5q1EDPrzc2q3XGplep5I5v+TuOl/MpEAOBQ4BDiIuDbYtzDtWpj2KJt3UGH6BJFsOJbo3v0H4DoiyTjWgVjvIm5mvZbu97iS1Fu/Jv93f9gEtnl4jW3+fEKR9pYtMtUV2R9FrzL778zZ92201rX3jJxtnN5gnVqJzLVoPaE4XlhnrRZiLppOnCC1eyK0kOp+sazVAAAgAElEQVSWqI28i+brFdWarmqwj35NZC5G8wWXs60ri15J9YG4lel6IlkuaXA1071xCyprIz1GZetyaaLWAB6m9B07OWk0Ur59iSRj8Xv63pxlphI35z9L9Hh6gPbPs/pxupXo+fURoiGHpOH0OvKPAV/vwLaPrbHtPTuw7V4wkamOS1lM+2SideD+ZfPWBk4iulk3chClrlVFt9J6cg+ia/d5VCaZnia6u99N3IFdA3g51V3NVwMuBnag+a5d04iuzq+q8fz1REuex4EViZaq2VoaU4km6sVm541sDfyQ/Ivwm4gk4SNE95xFide5HlECYNBr9YwQJ5Evy8x/hkhEzyfe55WI92cKkQwttz/x/uUNzvE4keB9mKhxtDywbeHfchsSLWe2xO5D0qA6kDgW1LIrcDalm3J3Eieat3Q5Lk0udwNvIM4/ZhC9O/4GfDdlUFKZtwGnULrWuIC4gZ81Spwb/aVs3hrEeevWRPfJF9P9sledcB/RevMqYiCPqxicrp+S2jcF+HLO/BuYWGvMos8QeYP1MvO/DFxIJAdVsjIxZsWyRMv3MaJRwe3EtX+nTCM+k+UL00Iin/IgvS0rtxqRZyheezwI/JtIIDdrKjFmyrpEz5+ngfuBa6nOi3TKVCL/twLx/k2h9P4Ve13Xlc3s9/IP7mJEEi0bQ6Nk5CZUj/z3DPCSJvaZ1yKzvCXms8TBIq+7xyyiaHneqIONulyX+3zO+uNEd5i84r1TiAvh23LWGQV2bGKfV+SsewrxftQzhUjKHUkkagexReYtVH9Wr6A6KbksUb812yJzGypbFBSnSwrbyUsOTwH2Iv/7fSEO9CENopcRv99a9qfyWHEtsGoP4tLk9WFK37cFxA1iKbX3Unkudg4TL3+wPNFo4N3EDfzfADcysZ4y7UyPEF3Yfw58kag3vxl2E5cmsz3IP150sjfOLjX2sVOD9ZYkxvUon9otdfT5zHZOrLHcxpnlsj1C78mJKTvV2nYtOxKNvPLKDmZzXb8m8k3tlH0bIW4kn0k0Zqq1n3uBE2h/UMZtqH5PXlT2/KLAx8nPNYwT7/m5RC+xelYnSr+U9/LJ/s37NpEn6YQpRG+NM6hfWuZ+4MeN4s/7cHtpY/KTkpvWWL5W8vNDTe4vL5FZnJ4mTpIa2YIY9Ce7fraFaJ6Xkp8UO6KJdRcD/pSz7l3UbzW5BtV1fY5tYn9ZM4hWqfX0YyKz/Ad9QIsxLEW8v+XbGSNeUzPJyNnk1119V4txSEprJlFf+Kwazx9M5XH2D7RWKkVq14mUvneP0l65G6lTPkjlsfAM8nuzdNIs4lx3G+IC/T3EBfdRxAXYCcTN+zOJ+vblF4a/LMz/UWG5YwvrfYo4Z3wVce6+Sg9eh6TB9Auqr/Wu7MJ+/pWzn9MarLN8zjofbHP/Z2a2c22N5XbM2WerU61tZ21Ja2OcZHM/+zS5H4heAn9tcR9jxN/BVhs27J2zrW0Kz21FtPhsZv+jRCO9PB8h8m7NbOcRqnu3tmp3YuDoVt+/n1Hd0xVyFn6exhnyZqatW3hR782J42by726ekrNsrQvLPPUSmfu1sJ3X56x/fRPr5dX1zOtqU8uS5LfMzBvRrGifzLKjxOjt3dDPicx2DtpfzNnO4S1uY0mqDzY3YKtMaZB8kfjt/igzfyrwPSp/36fiBa96ZzqVNzmvYfBLwmgwHUrlsfA00paxkqRum0F+y/CDurCv8l4Yxekxao/vAMOdyHwv+Q3EWpkOae6l80ai5267+7mP/J63tdRKZL6cKIvX6v7L8zEjxADIrW7jSVofnLvo80xswLz/EN3QK0z0S1ZrenWLL+5nOdvI3mF4Z84yd9Bas+Baicw/0Xpi6YKc7dRrQr4y1T+2OdTIMNfx6pz9Xldn+Q9mln2kxf21ol8TmZfR+uc7i+pm1tfQ3GAfWW/MiekVbWxHg2F1IvG1M3GCo8G2PvAc8bs9pmz+TKpP6o6nvWOENBErUlkm52y8WabeyiYxv4/HQimlGcAniB6DneoWqmovo/oab4zu1PVdMWdf49TvvpwikflS4O9lU7Zr+UOZ5/OmRqOy/y/578U40eX7LOBo4NPAF4jE3SVUJ52bSWS+Jec1FKdbib93nwe+RDS6e6TGsnNoPpmZl8h8S2Eb5fNuIwa7Po7InVxNfsJwIfG5QPQ4KH+uWKv65MJ2TiPyLnmv4d+0foPyBzW2taCw3+OJ9+9zwLeo3Wrzv8Rv4P+r9QWY6NRqInM28UXIbufAwvMvAuZlnnuO1usO1Epkvr7F7QDslrOd79VZPu8uyjfa2C/Elyi7rY1rLHsQ1e9bt2r59Gsis5Vm40X75WznTW1sB+JO2f2ZbX2lzW1pMOxC3DGbR3RlO4goPK3BMgJcTum48sXC/KUL88tPEJotcSJ1w2ZU3qWv1ZVI6qQR4mIxe25rIl1Kb3ki2bSQ6Or8eaJEmjcZOucQ8pNb3XJXzv4+UGf5FInMrE6PWr4P+cm624hr9XqJtulETdNTiZxIo0TmC4G5Ofu6g+gqXWsfH6I6dzVOJOmaqRmdl8gszyVcQe0e0C8hegpn17+Q+P0Xk7KjRBJ25RrbeRv55RQPrLF8nvfnrL+QaBiSHcy63FbEIJZ5r+H/n190I4k5TuuJTIgscbaf/nziDb82Zx//08Y+8hKZc2mvCPlUYlSl8m39s87yp+fsu1Yt0EY+mbOtWk3Y98xZ9sg299tIPyYyn6a9VnHZ/c8nCuu2K/v5/2kC29Jg2IXqY9qNwNeJ1pp2P+5/76Py8/sYcVJTXqv5Wdq/ySF1UvkNuFFi0DmpW0aIC9LyY+RRSSOSlLU81UmNx4iWY7bWnLiTqL72/VkX93dWzv6+W2f5YUtkrk5+YvE8Wr9OfwGlVoq1XJyzr+torsXt5uSXHcgb4T4rL5FZnH5K41aRKwIPZNYbozQI8gLg7U3EkdcT+LIm1gPYiOrr4CeJ6+NmzCIGw87u/63FBbJPzCcSYhOdVm8ywKxsN+hxSl36yqdzae9ub14i85I2YwX4bWZbC6j9I8rWSXyC9u9Yb0P16/hJjWVnk1/I9VxiRPJO6sdEZrvFlrMthP/a5naKsqPVd7OLv/pHMZlZvHM4Vvb/+dhas5+tRJwslR9TPk9lF945RC0gqV8cQ+UJYys1maRmTaX6PO0LSSOSVEteMrO8VdZV2FqzXRdSfe3bzWPhV3L2d06d5Yctkfltql/Pn+hO45Atcvb1JK1ds702ZxtzaFzLvFYi8yqaf60fqrGNcVob8yP7HR8DVmhivez3ZYwYQK8VSxKtX8u387fik3lvbGq/pPabPg7cQ/t3j/ISmRP5MeXVZ8i7aJhCdYKt2Wx2nkWprtVQb3t5P/ridDtRu+ANTHwQoH5MZJ7Sxr4Xpfp1XA7sO4Hph5ntLcSThcniZUTSMq++Svm8O4nj0a6010pcnXUm1V1Xyrvu/hfYJFl0Ur6pVN7BvoXWaolLjUylcvDNMZofMEFSGsVkZl4JrlFK5ztziPOfg4gbuqovbyTxg7u4v7xemVfVWX6YEpnLUz3QzVPAGm1ur5GfUP3efbyN7ZyXs516AzVD7UTmFi3sdzmikV12G3fTWp3Ld+Rs45UN1lmT6uvedlsqH5Cz/y36dSTB9xC1ntbKeW4h0V//sQ7u78EOr7t0zrwlqE5a3T+B/T5NtOgsT+jm7bfoM0QyJa8r+1rEj6n4g7qFuLNxMXA+ccAYZHPbWCevufgOhalTphJ3GR7v4DZr7WeJLu9D9V1PJLPPIsoclI8uWP7/NYCPAB8lWqJfCfyOKEtwT08iVdFexGeWVWxxfwNRtuPenkUkNWeU6C50NbAOsB7wC+Iu+GjCuDQcZgBnEDe/IS4oPkL9ro1SI0thXdVuW0iMCXEO8Xeh/Lq0/P9LE7/vfYmGLtcRvRd/S1wfjvUi2AGS1xOznWvPZuVteyKlzwbJ66l+racQiblOmwK8LjPvaeDENrZ1DNVlF98AnNDidq4mBkJq1qPEaN8vysw/iTgeNCuvV+pGxDVqLe+k8hoX4Gst7LPcz4hGebPL5u3Ur4nMuUSz6R/lPPd14M8d3t9TE1j3yZx5eS0f8ublrduKuVQmMuu1uJgHvJz44uZdnJdbvzAdRCQxfwl8le4WLu6meW2s042R5vLMovuJzG2J1qTqf+Un8TOJ3+zLiZq2NxI3Fi4gjoELeh7d5LE4cfI+SvUfYYg//ksTrd4eJ47lc8v+vYc4sRr0m0AaXI8DryFuhixBFKQ/AgcA0sTMJFrkvKbweBR4N+31fJHKPYa9lPpJ8dxnhOh5sgnRynAO0brsAuD39EdPztTyxmHo5vlf3rYnSy+uvFJOJ3VpX+tTnVs5l/byRpcSvbhWLZu3NXHMa+XGwB/a2HdeIvPiFrdxJ3HtU547bNQ7OvtZ3UYMVt2OBcTgRnuUzdu+X/9gLEtpZNisvYjuy500kRYKednsvARxXi2DibaMyO67UWJ6LjEgxQ5E64xnm9jHYkRG/QaiMG3eRX2/G29jnUZ1K6ReezHRneTXxB281dKGM9SOIEbSq3W8m1Z4fiOipfuriZ4CbybqoZ6BSUyldxOwP6WT5EMpK5AutWgxIoFRTGIuIAYKMYkpTR7LEH9XTiNqvO+cNpy+8FzOvNk58zpl8Zx5zVzTD4NscmwOzXdvb9U2OfPaHS9jjOoxO5akOsHYyDVt7DuvBW+r2xmlOoG7ZJ3lZ1A9onorLUnz3J55vG4/tsgcIeoR1CqiuglwLPCBDu5zIgebvC67TzQ5b6LdfbNfoLx95LmiMC1OXITvTCQ3t6B28dhpwGHE5/Iu2ksODpK8VpIn0l5z8np6MeDPk8BferAfNbY6zScgxym10LyBuAt4AfFZttIdQK3Zkugm2UwXtzHibuocosvDsXS3O5HUqt8SNyEPJ77TPyLuzk/0hFKTy5JEC/TtCo+fJ27c/CZZRBo2VzCYjSUG0RLEzfERWuvO/wRxM/0C4CKav+4cdk/nzKuX5JmovB6Yk+Hm+XSqr6H+TvdyEnm5qIkkTf8N7JOZtzpxjdesdlpAP5PzODuv2e2UlzGcVWfZNXOeX5n2x02B6qTvMv2YyDyEKG5az/uBPxJdnjuhXm3JRvK6IOclwfIO9u0OWATxxz4bd6tdlOcRf4wuKDxenOjGuhtR+229nHUOIEau+nmL+2pHylaReTVYpwH/6HUgHXAtsH3qIMT/EiNC1lL8QzxC/LG4mFLy0jqMvTGN6KLS6KSoPIF5NJHEzDuRlfrBl4iL1n2JE8uziYT9wymD0sBYmvg7VGxd8TRRp+z3ySLSMMrrMqrO24GoazdO7a78xXOcMaLl1kVEa+wrGP6GLO14AHhpZl43B9jLy1tMZLyPQbEM1Yn3h7q4v7zPcCLnTXnrtvo9aSdhnf3Ntpv0buW3n5cfK5ZL65Rl+q1r+ZbAUZl5TxH1d7ItkE4ifzCgduSNMt6sjTKPx4H7cpZbQPUXOLtuK9anuibHfyewPYjE5vlEMnl9IqF5U85yn6izjeep/qK3WwpgIoneiZpD9d2KVpt/S0W1kpjl5SXuJhJiuxF/2PYmWgCbxOydjxMDotVqFVL8vP5LHCdfQBSuNompfjYOHEgMOgbRAuBX5Nf1ksqtQAzyUUxizif+NpnElAZPMYk5k8rznPIafXOIm10HECMebw58mqjNbhIzX955+kRyC428OGfeZBgQNC851s1WwZ0e3yQv1ok0putnvRhrZGY/JTKXJFr5ZU+sPwD8mOqambWWb8fmtD9S3haZx3dQe0T1qzOPVyXqrLVjqya2P1F/IP7oZUdX34zaScYxqmsotHtXauM21+uEUaq7Y29F7wYB0vA4glISc5zSieA8oovOe4hjwZpEIfU/EDcE1HlLEScNqxX+XZpSbeEXEi3X8hQTmPcA7yNuoh3P5KlJpME3j0hAPVp4vD1RCkGqZSWiZ8CmhcdPEDfa/pgsIknt2pG4AbEokcQsJi9Hidp9nyOu75YjxlM4le4PRjosrsuZl80PdFLetrtVJ1Kd026uaRD1ZPCpfupantfC8ifA6YX/f5Wo5fiKsue3LMz/+AT3vQpR0LXVAq4bES0Xy9VLJv6VGKyo3L7EBXGr3lxj+502h/gcPlc2b4Ro0VErYfs4lfU/87qoN7IhsGIb63XSxVR+36YSf9x/kCYcDaAjqPztXE+0er6Q6KJjrcv2zCSOK6sDyxN1V1YoTMX/L0O0Bp9Gc3WQi12pyhXrlD5G1Ic7h2iNuRjWwtTguYsY7OcC4nfxQaJmU6drP2vwrU6cA61TePw48Ergb8kiktSuHYnzzmLNukeIruIXEt3GTVhOTF7N6TWIm0Gd7vK9OnGem9WrY3PKZFxe3qGbXfg7Pb5JXt3UYf3t5X1WHydySh01npnaKSI6UR/IieMGqmskFg8I5cuNUZ0crGfNnH2NEwXwW/WtnO28qc7ym+Ysfz2tF7dej+iqXr6dB+leN7EDqY47bySvovMzyz5C7UGEavlezj7vbHLd0cx6X2lx30XrE4mm8m3dTY/uMmjgHUH8ETyLKI/RbuvryWxFYFci4XI8cdJ9B9W/8RTTg8ClRBLoE0Rrt7W78i5InfVxSt/j57E2nSqtSRxny491KXvISGrfjsS56OXAZ4lajpOpZVgvTCUSN9nzxP/pwr4+nbOfB6j/mS6fs84H29z/BZntNNsS9MnMeu004ppO5H3Kt9PNMidfoPp927nD29ujzvJ75yxfL/9Syw8y22h3oOH7Mtv5cZ1lN6I69q+2ud+6UicyNyVqEZbH8DS160fuRvVF7KM0PxJwrUTmQkrdZ5qxPvBcZhsP0TiZeFnOvj/cwn6h+iAyTtTg65Yv5uxv9TrLfzln+de1sL8tqH5vUyQyAX6WE4etV9TI6kRB435q9d7vphNlPg4mujTdQPUJyyBMTxC1pI4iTkKW6+SbJHXIjyh9Zx8kf3ROTT4bUHmxcg+wbtKIJLVrBvAqhrcOXz85merzwevrrdCGEeDWnP006im4WM469ca7qOdvme30MpEJ0bMkm7fqVmL+nVS/bwdPYHtn5Wyv3vgbg5zIXIrqfMx5be63ruwb1MtE5uLAzTkxvLfBel/JWecymksa1EpkjgO3EHctGlmaqIeRXf/wJtZ9fc56z9B8hv/InPXnEzX2avkEsA+1R6erZwmqDxr3NtjWVjkx3kJz3TvXJRKWeZ/PnU3G3MlE5ouIOnjZWL5G6y1pi9YlDiqLTyAuadBNJ5K9XyVKchQHCuvm9BTxN+7Rwr+P92CfY8SgaScQN3SaOQ5K3bYIcBWl7+k/idppmrxeQgxKWX7OZStzSWpsR/LPAffu4D7eWGMfWzZYb4TqnpztXBvPpLrxWbOJzLmZ9b7dxv4hGjpkX/9L2txWIxvk7OuMNrc1hepE4OPUz6cMciITouRCNl/V7gDQNWXfoHlE3cZOTHmjapU7LWf/zXxBplEaPa18OqKJdfMSmeU/ypuof0DYlKgpld3GzTT/4fwyZ/15wEeo/YVeivy7PeM0bh7+08Jy/yEGHdmgyTjXIQa8ye6vmabB1+as9xdqnxRPJbrfPlK2fDbJcGeTcXcykUkhrrz3/c/ALk1uYylgf+DcsvgmUmdDGkSrETeqzqb6pKaVaZTo+ngR8Xfkm8QNm/2BPYnj9AsoDehTy66Fbf2SGPxkaaL20IbEzaW3EXdfjyTqupwH3Eh+i/Fmp+eJwTI+hd01ldbKVJ6YnpY2HCW0OXGDp/ycttmeTpKkGKsie853N51puLIkUaM9u/1mB197KLPer9uIYdec/TebyHwgs1675xt51+TfaXNbjYwQjR6y+Zps6cNm7Ex13Bc0WGfQE5lfpzr+/dvcd03tXow1M32e2t6Vs/x/aD658wKq61GMEj+yevISmV+mssnzQuIC+WCiBc1riCTj/1F9R2OcaLG3Q5NxQ4z4fXfOdooHvGOIkXHfXIjhNKIlUd7y59C4SXUxkVk+3Up0m/4k8A6izujuxEAAnwV+V+O13klzLYr2rBHvs0QNzcOBjxb+PZXq2qfnEj+Q7L6b0elEJsTorrW+5/8Bfkh8Vu8gkvjvBQ4luu/9k/zWZiYyNRmsQtQJ+get/w15jmiteTJxXHojsAmduaM3nTj+1OvWUcs04kbPq4jX9gPgEqq7zjQz3UXcHKpVTkXqpm2p7HUw0cETNXh2oPLG0o1Y01mSWrUz+SWRTqW9XpFFU8gvdTYKbNfkNi7KrPswrZe/Oi8nhmYTmdku8Re2uO+ipak+136KGFypG8rL8BSndrqXn5Oznfc0WGfQE5mbUP17+A8dHtOl1YuuVqZaicwXExntbIJr8xZjfw3Vb9AD1B/tOi+R+R4ikZeXuGs0PV+Io1UvIE4WJ/L+nklzA8/kJTLbmf5Laxfb2R9Os9OVxN2r7MHjzib3241EJsSBq53vSK3JRKaG1SwioX8urf1m7i+scyhxcT0ru+EBsBZxx/F4otV2K13mbyBqEq/Z66A1qR1A6Ts4StyI1OTwcipvlP8D6/pKUruyjXCK0w9or5bjCHBSjW1+r4XtHJWz/htbWL9Wt/ZmE5nZROpNLew765s5cVxCd8YleGnOvuYQA1A3Ky8h+RiNy/kMeiIT4FdUv4aT2tx/rk4lZfKmvETmLPLrS7ZbPDWvpdzvqX3no1YiE6J1TbYJcb3pXmLwoXYtS7S2bHUE3qeAz9V5jVnHt7j9vOk86g/wk2cK0dy7lf2cRakJfr8lMiG6n+aVNWhlepwYMKhbo8xLqWwGnEIM2NbMb+HBwvJvpf4NqEE2mxiV8DiiVnAz78so8XdsLyZ2F19q1ncpff/mEC2ONdz2pPJYfTWwTNKIJGmwzSYG+ck7tzuf1loOrkH0Bs3b1r9ora513vgVd9Lc2CC7UrvHUbOJzG/krLtV8+FXWIX8UeLPpfVa32sQ1y71/D5nX/8kuvs3sjn5pbQOb2LdYUhkbkx1XdVx4pyzmcZ4WVOIcV8uKs74exenvCazH8lZ7oe0P+LUDOC3Odt8Q43l6yUyIWpGfYsYebbWReZ9xGAvnWpRtzlwOpX1ifKm24lu5yu3sY81gPcTNUjvbLCf4vQk0Ry+2YGIatkZuJz6CdsriYGQyn2Bys/03Cb3d3VmvQ9NLPxcuxDvZbZLfK3pVuD7xPey44VupYSmECU4LqXx72AhcSz4HHHc69ZIg/1sLaKu8TlE4etG79ktheXbqckjNWs6UWur+L27ieZO0tWflm3w/N5UlhT4Ew5EJkmdsBZwD/nndPOAo4GtyT8HHiESfEdT3Xu1ON1Fe12p88b4uJEoMZNnBaL1Y7Fn0RjRiKudROZOOft+DDiMaEi2MfG+lU/16jTvRX43/tuJ0nz1WmdOJcrpnVp4bYc0iH0N8vNCt1G7pOF04MPkf4b/pLnGTMOQyIT80d/HiYaNb6VxS9qpxG/i65RyWE/C5LyIXJMYJKLce4lkarnpxKA/GxB3KxYSSav/EMmxsS7ENpW4uH8h0TppUeKH8yBxMXtzB/e1LDF69tpEvYniRfJcojXGtcRrHe3gPpcjWjSuSpwwLyRqgl5FHBgH0QhRKmEd4j1dlkjsPEX8yG4lPrcnUwUodcmixE2gj9J4ZNt/EiUufk6U/1BYnEgCv51o3T+1zrLFltzfIrrgS522LHEjcK3C498SN9/yzndmEzfl2j0hVvdsDqxP1FTL8xaiN1Dx4uFC4nN+pvuhSdKksDbRmnK9Oss8TFwHFxvFrEz0fqzXQ+lG4NVEMrNVOwMXk5//uZHIbzxB5AXWI/Ig5b2CjiaSevuWzbuOqIXYyAjRinTTFuJttO3DiHFO8swlbs7eRTQUm070ONiIaIG5VNmyHyN6TdXzJuJvat55+n+I9/UB4u/qWkRPrLwbinOAlxHvdyN7E40eym1LNP5qxQ+IsVeKHqW5lrhZ9xE5nKKfAAc2ue5XgU/XeO4pYkDom4n353ni81meyMNtSnVjjqeYpCX6GrXIlCTVNoWoAZk3emL5dA9R1uIlacIcOMsABxGlK/LuMhen+USto6XyNyNNyKZUtiD4Yo3lNiQG4FJ/GSEuct5c4/n9iJvIxc/3POwlIkndsCTRe6+ZnnvNTKcz8eTNV9vc92nE+f+ZmfnNtsiEOG94vIV9NrPtd9NaLfq8qVGLzKLXk99NutnpHqLxU7OGpUVm0TuZ2PtXPk3aBmImMiWpPbsSJxb1/rj8mbhbW691oep7CXAC9f/gP0YMijSIAyKpvxVbYY4X/n1TzjK7EzczpvcwLjV2APG55ZVXej+VJX5+gZ+fJHXbK4neh+0mba5kYmNyZB1G8wNxLiBuaBZbcU4kkQnR4vTnVJY2mUgiE6LlaHlpnFamx4mEYbO2oPWxMkaJbuytlgYctkQmRIvYs6nfYKPRdDXwgTZfw8AzkSlJrdkG+Cu1/6g8S/xBa6Z7iZq3MnAEcdJR672/h2hlJXVSeauNp4j6VeWKCbNWRj1Vd80GHiI+l+yF2SepPG6cTndGeJUk5duG6MJ8PfUTOWOFZY6jvcRVMzYgSj7VGuT4CaLr8IaZ9f6HSGYWp6Pb3P8yRG3MTxMDN/8ws912tv0KogTT3dRPhD1EdBN/O60PDlT0msI28gYdKk53EoPaNBpMqJbNqX5P1m1jOwdltvGjNuP5QWY7H2xzOxDnlF8D/kZlL5G86X7gN0TN0XqlGiYFE5mS1JzFiBOpWgN1PU10cx7WEcf7xSzihOF+av+hv5D2ir9LeaYQA+yVn5AvV/b85wrzL+99aKrha5Q+rz3K5h9K5bHiBCrrnkmSemsJ4KVEMm/fwvSqwrxe1v6bSiSVXlmI4TVEo4RBb63/AmJApb2I1/VqYAfaqw1Zz1SiJvUORNfz1xB1LFfv8H6G2VLEd3An4ub4G4kegJvjoI/0e1gAACAASURBVJNVTGRKUmMvJwarqtVN4kxiYDL1zqJEUqJWjaH5heft1q9OmA3cQOn79QdKrfi+WzZ/8yTRqdw6VNYJe0Vh/v9SeYz4DpNzoE9JkqSBZiJTkmpbguhOktftZQz4FdElReksCxxD7RpDfyYSG9JErUdl4vyYwvxfEzc0RokuYUrrPCpbzr+c6K5Xflw4Kll0kiRJmhATmZKUbz0qW2CVT3fQ2WLjmri1iFZyeZ/Xk0Q3F2miXkllDaN3A/8oe/wskVxXGrtS/fs/K/P48GTRSZIkacJMZEpStb2J4t553chPABZPF5rqGAH2J7/g+BjRCst6eJqoz1D6Xj1D9fftU+lCm9SmATdRu1j+GDE4gyRJkgaYiUxJKplKjFCc15X8OmCrdKGpBasC55CfzDgfWDpdaBoQ04jRLT8F7EcUXt+AGPRrBDiD2smye7E2awofo/YAYGNELdNdC9P2RD3TzYGXEC26F+l9yJIkSRMzGQt+zwQ2ysy7G3g0QSySlNIM4GfAPjnPnQx8gOg2qsFxMPANqkecvJFIZjzQ84g0SBYhbmwcTOU54jziu7MitUdUfQNRO7OfzABWAFYiRsaEUlJ/ceJ3MpMYSOtJogX6M8RxbwHxuseAR4CHC/+O9Sj2RlYAbiNeR6vn8zcBhxGf13iH45IkSZIkqeNmULqILZ8WECNfa3DtANxPfp3TF6YLSwOkmPTOa+WX1/pvIXBpgjhXA3YB3k8MRPRz4E9E4j6v3MJEp4XE+3It8DvgVOArwAHA1vS25fOJ1P48ak33AQdi61lJkiRJ0gCZRVyEZy9y/wtslzAudc4qwF+o/ozvAtZOF5YGyDLALyjVym0mUZbt8dIpSxGDDh1GdHH/B/BUkzH1enoYuIwYzf2jwDZEq89OeinNfybjxMjzhxLHfkmSJEmSBsYs4I9UX+jeQrRu0vCYCfyW6s/6bqJetNSMfYG51B5QpjiNAj/owP6mAZsRpS1OJrpBt9rysN+m54ArgeOBtzOxmwkjRKK00ecxBjxNDPi15AT2J0mS1FcmY41MSZrMTgfelpl3M/AKojuyhstUIhm0X2b+zURLsbm9DkgDaXWiG/XLiSRZrfPHZ4mBp+a0uP1VgD0K0650rov2GKXalsX6lvOIEhrPEvUwR4nWncUamksQv5tZRM3QGcDyRJ3NTnYdvwu4ELiAuLk0r8n13kYcx2sZK0w/Bg4HHmw/REmSpP5jIlOSJo9DidY55a4Bdicu8gfFy6jdVfOOwtQJOxGtw/L8h2jZOAimAj8i6viV+y0xQEu/DF6i/jZCdJX+BjCF2nUWPwF8s4ltbQ+8mkhebkp756RzgVuJFuW3EMnBh4jkXTcG55lJKam5YmFaF1gPWB9Yh/a6kT8PXE4kNn9LHF/yzAJuB1bOeW6MeA/PAj5N546DkiRJkiT13J5Ud0W8nqiDN2j+S+3ulL/v0D42qrOPcSJZM0imELUFs6/jyymD0kDaiDh21Ppt3EvtJOeGwJFEwrGVrtkLiQF2TgQOIlqGrtjpF9YBU4G1iOTsx4kao3fTelf0q4ik8QqZ7X8rZ9lircw/EAlhSZIkSZIG2trEYA/lF7+PMbiDvtRLZI4S3WAn6pg6+xjERCZEa66/UV1H7w0pg9JAWoRo3T1G/qAze5ctuwqR1PtXznK1pkeA3xAtC3cCFu/y6+m2lYDXEkncS4gu7c28DwuA/yPqaq5feJxd5q/Ajr17KZIkSZIkdc8U4E9Ut27aM2VQE1QvkTkOfG6C258GPNBgH4OYyIToknofla/lUSLRIrVqV+K3kh2M5xpiwJ5TiW7TjRJ2o8DfieTortQu6TAsZhGv8yjidTeT1My+jzcRZUEkSZIkSRoaH6P6gviQpBFNXKNE5q1MrAb0axtsf5ATmQDbEQOdlL+es5JGpEG2FPBTWu8+PR/4GfAWYNmeR91f1gY+RAz6k9fCNduK+ipg6ySRSpIkSZLUJSsAT1B5EXxu0og6I5vIfJQYdbh83vYT2P6vM9u6i+pkwiAnMiEGfsq+plcmjUiDbDpwGo2TlwuJAW3eweB3F++W1YBPAv+m8ft5BbbKlCRJkiQNiROovOidQ9SrG3TZROY9wE8y805sc9vLAs9ltvVlqhMIg57InApcTeVruo7ag7RIeUaANxOjbNdLuN1AtAS3hEFrNgbOAeZR//29GNgyUYySJEmSJE3Y6lQn5D6aNKLOyUtkvjwzby6waBvbznbFv4IYaGPYEpkQNQyz3VjflDQiDZJdaVzf8Q5i4J+JlHqY7KYCM4H9iZsN9d7vi4hR5SVJkiRJGijfo/IC9xai++cwyEtkjgC3Zea/vY1tX5PZxkEMbyITqrsDX4NJJ9W3BnA+tZNpTxOtwV8HnE0MOKbOGCFKQFxI9QBLxWkBcAzt3ciRJEmSJKnnFqe6ZuQBSSPqrLxEJsDhmfm/b3G7m2XWf4YYyGSYE5nrEHULy1/bjkkjUr8aIRL7T5KfQHueSGCuXLbOLGCx3oY5aWwEnEn91rC7JYtOkiRJkqQmHUR1om9a0og6q1Yicw0qu0qPEl3sm/WtzHZPL8wf5kQmwC+pfG2npg1HfWgd4BLyE2ZjREJt3WTRTW6NuvifiaPCS5IkSZL6WDbh8KW04XRcrUQmwB8zz32uyW3OAB7JrFtszTTsiczdqHxt84BFkkakfnII8Cz5SbJLgS2SRaai4qBLd5L/Of0X2ClVcJIkSZIk1bIMUSOt/CJ27aQRdV69ROb+medupbmaj/tk1ruP0gjew57InEK8h+Wv71VJI1I/WAQ4mfzE2BPAwVj/st8sChxFdbmI8cK8Q7EGriRJkiSpj7yJyovXa9OG0xX1EpmLUV3Db7smtnluZp0jyp4b9kQmVA8OdXzacJTYOsSxIy+JeR7wgnShqQnbAjeQ//mdgXVLJUmSJEl94ptUXrR+PW04XVEvkQnwo8zzJzTY3orEQCXl62xQ9vxkSGS+lsrXd1XacJTQHsAcqr/zjxPdlzUYZgJfobJucPkNrjXThSZJkiRJUriMygvWN6QNpysaJTJ3zDw/l+hyWcsnMsv/OfP8ZEhkrkjl63sOmJ40IqWwN/n1MG+iMrmvwbEn+YnpB4ANE8YlSZIkSRL3U3mxulbacLqiUSJzBLgts8zb6mwv24X2PZnnJ0MiE6IuaPlrXCdtOOqxN1NdX9euyMNhbeDfVH+2DwIbJ4xLkiRJkjSJzQLGKF2kLmA4W9U1SmQCfCGzzO9qbGvLzHJPA0tllpksiczLqXyNu9VfXEPknVQPEDMKfDxhTOqsxYBfUH0sexjYNGFckiRJkqRJalUqL1DvTxtO1zSTyFyDytpwo+QPUPLdzLZOy1lmsiQys0mOt6YNRz2yH9V1FBcC70gZlLpiCnAi1cezx7B0gCRJGhBTUgcgSeqYbPfPeUmi6A93A5eUPZ5CdWJmBtWDl5zSzaD6XPb7Ynfi4bcZkdgqPx8cJVpo5iX1NdjGgPcBx2fmLwP8Fli65xFJkiS1aFrqACRJHZM9pi9MEkX/OAV4RdnjdwJfJVogAbweWLbs+fuoTH5ONgsyj4exLIFKViKSV7PK5j1PtMT9VZKI2rMi8L91nv8+cE0H9rM+8D91nj+SuIHS78aBQ4CngMPK5q9HtMp+Ff7tkCRJkiT1wJpUdhe8K2k03dNM13KIBM0TmWW3K3v+gsxztZIhk6Vr+WlUvsb904ajLpoJ/JXKz3uM+oNi9av1qP59lk8ndWg/xzbYzxYd2k8v/ZDq13F00ogkSZIkSZPGclRekM5NG07XNJvIhOoL9RMK81ehenCT9WtsY7IkMs+j8jW+Pm046qLvUP2dPippRO1rlMicCyw6wX1MI0b4HrZE5kzgCqpfyz4pg5IkSZIkTQ4jwHwqL0iHseZZK4nMHchPanw6M/+yOtuYLInMG6h8jZunDUddsgvR+rL8s74QmJoyqAlolMgcJwY0mojXNbGPQUxkQnTNv4fK1/JIYb4kSZIkSV11PZUXpNumDacrWklkAtycWf5twI2Zee+us/5kSGROB56h8jUuW3cNDaJZwJ1Ufs7/AZZMGdQE5SUys4naiya4j9802P4gJzIhYn+OytfjYE+SJEmSpK47g8qL0UPShtMVrSYyP5dZ/q7M43nA7DrrT4ZE5lZUvr5704ajLjmMys95FHhZ0ogmLi+ReRnx2spf5+ptbn8FYhCk7PaHKZEJ1d+NMaJFuyRJUl+ZkjoASVJHXZ15vGOSKPrLKUQio2iNzPNnEyP4TmbZZNZVSaJQNy0PHJqZ9x3g8gSxdNtdwJ/KHk+h/cGr3k60WC76C9HKe9gcBVxb9niEwa2bKkmShpiJTEkaLtlaj7sDi6QIpI/cB1xS5/lTehVIH9sr87hezVANpo8Bi5c9ngN8KVEsvXBy5vEBRHKuVQc02O6wWEh1C/7tgZ16H4okSZIkabIYIboFD/MItK12LYcY7CNvgI67aHxTb9i7lq8MLKDy9a2VNCJ12mzgCYaz7ERe1/JTgcWAJzPzW+0qvVlm/aeBpYATc/Y56F3Li86j8nX9Lm04kiRJlWyRKUnDZZwYmKLcgSkC6TNnE4mcrFOIWnCT2buAaWWPrwXuSBSLuuMtVA7o8yjww0Sx9Mp84JeZednWlY28M/P4V+QfR4bJlzOPdwPWSRGIJElSHhOZkjR8Tso83gNYP0UgfeQZYBui1VT59M2UQfWB6cD7MvOGPcE1GWVvZnyfGORq2GXLRrwJWLTJdWcAb22wvWF0FfDXsscjVCd0JUmSJEnqqL9R2T3w5KTRdFY7XcsnYpi7lr+Xytc1H1g6aUTqtFWJVsflo1GvnTSizqrVtRwiCXdb5rn9mtzuGzPr3QtMLTw3zF3LIRLf5a/tprThSJIkldgiU5KG09czj/cDNkkRiPrWLOCwzLwTgccTxKLueQ2Vg9xcCdyeKJZeG6e6FWWz3cvfmXl8MjA6wXgGxVnA82WPN8Du5ZIkqU+YyJSk4XQ2la1opgLHJopF/ekzwOplj5/DrvbDaKfM43NSBJHQT6hMQO5C5fc+z4rAKzPzftrJoPrck8DlmXk7pwhEkiQpy0SmJA2nMeDTmXm74MA/Ci8CPpmZdxxwX4JY1F3bZB5fnCSKdO4DLi17PAV4R4N19qdyAKw/A7d0Nqy+d0nm8dZJopAkScowkSlJw+sc4A+ZeccAL+x9KOojM4HTgEXK5j0EHJkmHHXR0lS3ur02USwp5XUvH8lbsCCb6Dy5o9EMhiszj1+aJApJkqQME5mSNNwOonJ04iWB3wCLpQlHfeA7wOaZeR8mupNquKyZeXw7kcycbM4C5pY9XhfYrsayWwEblz1+prD+ZHNj5nH2uyRJkiRJUld8mOoRdn9J/RZJ/WwTIhFXnDauv/iELZPZ3+ZEDb1B9DGqvwtnJo1I3fR6Kj/r89OG0xX1Ri0v98PMMifW2N73mtjWsI9aDvH34RkqX+MSSSOSJEmSJE0KI8AZVF94fyFlUOq53YAFVH4HbgaWShmUuuoAKj/vbBfrYdBsInOHzDJzgUUzyywCzMks94qcbU2GRCbAf6l8jaulDUeSJMmu5ZI0GYwD7wauycz/IvD+nkejFLYjWuGWD2DyBPDawr8aTtlE3TNJougP2QF7liBarJZ7LVFXtCg7UNBkMy/zePEkUUiSJJUxkSlJk8PTxEX6w2XzRohulB9LEpF65WXAhUR91KIxYD8m30jMk8145vGglpPolJ9mHh+QefzOzOOfAKNdi6b/Tc08nszvhSRJkiQpgR2BZ6nuFnloyqDUNbsC86n+vD+eMij1zDuo/Nyzibxh0GzXcoiu0QvLlhulNKr7qpnnxoC1a2xnsnQtf4DK17hK2nAkSZJskSlJk81lRHfKbBfTo4Cv4t+FYfJG4Fyquxd/Hvhm78NRAnMyj1dKEkX/uA+4pOzxFCLZC7A/lS0QLydGeZ+spgHLlT0exzIUkiRJkqREXg48RXWroguIUbo1uKYS9U/HqP58D0sXlhLYkMrPfxgTc620yAR4e2bZW4ku9zdl5h9YZxuToUXmmlS+vgfShiNJkiRJmux2IEbuzV6Q3w1smTAutW9Z4PdUf6ZjwCEJ41IaixLdp8u7Ui+RNKLOazWROQt4PLP8JzOP5wGz62xjMiQyX0fl6/tz2nAkSZIkSYKtqa6DNk7UVazXIkn9Z3siCZ39LJ/Fz3Iyu57K78Mr0obTca0mMqE6Efl85vEpLa4/jInMr1L5+o5PG44kSZIkSWEF4I9UX5iPA5cC6yaLTM1YlKhxWj5QSXG6D9g2XWjqAz+m8jvxtbThdFw7icztc9Ypn3ZusP5kSGT+ncrX99a04UiSJEmSVDKNSIblXdTPJ0Y1n1pzbaWyI1HjL+9zuwwHdxG8hcrvxU1pw+m4dhKZUF0TszjdReNBz4Y9kbkqlTV2R4kbXpIkSZIk9ZX9icRl3gX+FQzXxfogWxU4mfwBfcaAY4jktLQk8BzDm3RrN5H5uZz1xomBshoZ9kTmp6l8bX9JG44kSZIkSbWtCVxE7W6X5xLJA/Xe0kTL2afJ/2zuAHZNFp361blUfk9OSBtOR7WbyFyNGMU9O63VxLrDnMicAvyHytfmQGGSJEmSpL42QrTOfIz8hNkCIhmySqoAJ5kZwEHAw+R/HqPE57F4qgDV17IjUD/D8Px2201kTsQwJzLfSOXrehZYPmlEkiRJkiQ1aVXgHGq3zpwPfBdbaHbLssBngfup/RlcR4w+L9Uyjaj9WP69+WbKgDrIRGbnTAH+ReXrOjllQJIkSZIktWNXqkexzbYIvAjYm2jNqYlZFzie2vVKx4kRyQ/CWphqzoeo/P48R3zPBp2JzM45kOrj+oZJI5IkSZIkqU0jwJuprp+Wna4B3kfUc1TzZgKvBy4gfxCf4vQ4MRjHomnC1IBaBLiX6nq3g85EZmcsDTxA5Ws6I2lEkiRJkiR1wHTgA1QnRbLTs8CvgX2IJIqqjQA7EvUt51D//XwS+BqwTJJINQz2o/p7tV/SiCbORGZnnEJ1HdUXpgxIkiRJkqROmg68Fbia+gm4YivCHwJ7YkvCqcA2wFeprluYN90FfAJYsvehasiMAFdQ+f2aA6yTMqgJMpE5cW+i+vV8KWlEkiRJkiR10eZE8mABjRNzzxD1NA8trDcZLA/sS7S8zHbfrDX9nRg53hqY6qT1gaep/K7dxOAmytcEbs9M3R7I6MicfW7c5X12y0uAeVR+H27EVvSSJEmSpElgDeDzwM00l6wbB+4gWmu+B9iEaLE46F4IvAU4FvgH9Wtelk8PAd8Ctux5xJpMPkr1d+8cYtRqTR4rAndTXRJkstxgkiRJkiTp/9sSOA54kOaTmuPAU8ClwFFEfc0NgBm9Db1pI0Ty9pXA54hk0EO09nrnA6cDr8LWl+qdU6n+Lh6VNCL10iLA5VR/B96VMihJkiRJklKbCuwBfI9ofdlKkq84LQBuJUZZ/gbwXmAn4EV0f/CbxYC1ge2AtwFfBn4B/IvqLrrNTo8SIwK/HVi8y/FLeRYBrsRk5mS0KPB7qj/7Y1IGJUmS1IyR1AFIkiadDYhBf/YgRuzuRC2254FHiNafDxb+/zxRj/PZwv/nE1285xIX8jML06JES8jZRNfa5YEVgJWIrpedGJxoDPgncEFhuhoY7cB2pYlYGfgbsGpm/tHAJ3sfjnpgMaLl+C6Z+RcRrcIX9jwiSZIkSZIGxKLA7kRdzfOAh2mvhWO/TfOI7vFfB95AJEalfrQVkdzPfoePwxvew2ZJqketHweuBZZOGJckSZIkSQNrLaIL93HAxcB9pE9M1pseA/4CnER0eR+WAYs0eWwFzKH6u/1zogWfBt86RMIy+xn/A1guYVySJEkt8U67JGkQzAbWK0wbAOsDqxFdv1emu8mW54iWog8UpluIep23EKOzP9rFfUu98hKie3E2qXUz0ar4pp5HpE55FfBTqltd/oMYqOyxnkckSZLUJhOZkqRhsCiR1FyJ6Ma9PFHvcjZR/zJbE/N5YqCevYjE6LPAp4mk5eOU6mw+CDzRw9chpfQiohX0ypn5TwEHAL/ueUSaiBHgU8CRxPGw3BVEgvPJXgclSZIkSZLa81tKXSw/mDgWqR+sDfyb6i7IY8So1p0Y/Erd90KihW1eOYyfYckASZIkSZIGzo6ULu5vx9qWEsAiwE/IT4LdAeyWLjQ1MAIcRLS0zH52C4BD04UmSZIkSZIm6kpKF/qvSxyL1E8OIsow5CU0zwSWTReacqwNXEL+5/UI8Ip0oUmSJEmSpE54C6WL/csTxyL1m5cRrTDzkmMPAB8ApieLThAJ5aOJWr95n9PFwKrJopMkSZIkSR0zlehWXrzo3yZtOFLfmQUcBSwkP1F2F9F6MzugjLprUaKr+OPkfy5PAAfj5yJJkiRJ0lD5GJVdZiVV2w64kfyk2TjwN2D3ZNFNHjOB9wP3U/uzOAdbYUqSJEmSNJRmU2rVtJCoNSep2kzgS8B8aifRrgHeWVhWnbM88AWiS3+t9/4eolyGJEmSJEkaYt+glAw4LnEsUr9bnuhu/hy1k2oPFZZZJVGMw2I94HjqJ48fI7qZz0oUoyRJkiRJ6qFVKY3QPA9YJm040kBYjyjHMEbtJNszwBnAXjgwULOWAt4DXEr993Y+cGRheUmSJEmSNImcTilB8KnEsUiDZAvgLGoPCFScHgG+A2ybJsy+NhN4PXA2tUcgL05zgWOwtaskSZIkSZPWppRaP90HzEgbjjRw1iQSbHOpn4gbB24DjiUGCFokRbB9YDngbcCpwBwav2d3AIcAS6QIVpIkSZIk9Zc/UkoavCNxLNKgWoJIuN1G4+RcsYv0+cCHgXUSxNsrU4GtgS8CVwKjNPf+XAa8obC+JEmSJEkSAK+mlDy4FhhJG4408DYnBqt5iOaSduPA/cC5RMJvb2DpXgfdISsR8X+ReD2P0fx7cBcxWNJ6PY5ZkiSpb3lxJklSpRHgeuDFhce7AhenC0caGjOAPYH9iIF/WulOPgrcCPwduKUw3QzcDizobJhtWZxIOK5fmF5MtLxcvcXtPAz8gqjXe1UnA5QkSRoGJjIlSar2XuDEwv8vAF6VMBZpGM0mbhLsUZhaTfgVLQTuJBKbdxCDCd1f+Pfhsv8/O8FYVwZWAJYnBthZvjBvbSJxuVqb2x4HrgEuJI41fyVekyRJknKYyJQkqdpMolvnSkSi4SVEN3NJ3bEh0VpzD2A7YFaHtz8feJ5IaD5DJAufIn7fT1EaPKfYhX0Joh7losTxoJMeI1p5X1iYHujw9iVJkiRJ0iRzOKVadT9MHIs0mUwjEpv7AycAN9D8oDj9Ni0oxH9C4fVsiA0JJEmS2uaJlCRJ+ZYF7gYWA54D1sSWU1IqSwFbEonA9SnVo1w1ZVBlil3cb6ZUw/M64F9ES1BJkiR1gIlMSZJq+wHwvsL/jwA+nzAWSdUWp5TYXJnqWpbF/0+bwD6eJeptzgGmE13DLyvMuxe4iajP2Q+DDkmSJA01E5mSJNW2HpGkmEIkMVYnau1JGixLEr/jxYlk5Cxi1PTphXlPUKqXuZCoo/ks0ZpyPrAWMUI6wL+JurmSJEmSJEl95RxK9e4+kDgWSencTOlYsEbiWCRJkiRJkqq8nFLy4laiVZekyedovKkhSZIkSZL63FWUEhivTRyLpDR2onQcOD9tKJIkSZIkSfneSimBcVniWCSlMY2olTtO1NBcLG04kiRJkiRJ1aYBd1FKZm6TNBpJqZxB6TjwmsSxSJIkTTpTUwcgSdIAGCNqY76y8HgJ4Kx04UhKZBbwhsL/nwLOSxiLJEmSJElSrtnAE0RLrIXAWmnDkZTAMsAC4jhwPzCSNhxJkiRJkqR85aMWH5s4FklpXEbpOLBZ4lgkSZIkSZJyrQY8TyQwngSWShuOpAQOpZTIPDxxLJIkSZIkSTX9jFIS45OJY5HUextSOgZcnTgWSZIkSZKkmjanlMS4D5iRNhxJCdxGHAPGgFUSxyJJkiRJklTTJZSSmfsljkVS732L0jHg3YljkSRJkiRJqmkvSkmMf+PIxdJkszulY8CvE8ciSZIkSZJU0whwI6VExi5pw5HUYzOIAb/GgXnAImnDkSRJkiRJqu0gSonM8xPHIqn3zqZ0DNgjcSySJEmSJEk1zQQepDTgx4vThiOpxw6klMj8duJYJEmSJEmS6voipUTGSWlDkdRjKwCjxO//7sSxSJIkSZIk1bU88DSRyHgWWCltOJJ67EpKNzM2ThyLJEnS0JuaOgBJkgbY08DqwObANOAZ4JKkEUnqpZWBnQv/vxf4c8JYJEmSJEmS6lqPUvfSx4DF0oYjqYdeSqlFpklMSZIkSZLU986llMx4f+JYJPXWXcRvf5SomylJkiRJktS3dqKUyLwVmJI0Gkm99H1Kv//9E8ciSZIkSZLU0FWUkhmvSRyLpN7Zi9Jv/xeJY5EkSZIkSWrobZSSGX9KHIuk3pkFzCd++3OBGWnDkSRJkiRJqm8acDelZObWacOR1EPldXJ3SRyLJEnS0JqaOgBJkobEGPF3dffC49nAWenCkdRDSxBdzAHmAL9LGIskSZIkSVJDs4EniFZZC4G10oYjqUdWIW5mjAO3JY5FkiRJkiSpKd+k1MX0mMSxSOqdf1L67a+fOBZJkiRJkqSGVgOeJ5IZTwJLpQ1HUo/8L6VE5scTxyJJkiRJktSUMyglND5RY5lZvQtHUg9sRel3/8fEsUiSJEmSJDVlc0oJjfuAGTnLHIKD7knDZArwAPG7XwAsnTYcSZIkSZKk5lxKKZn59pznbwRe2MN4JHXfjyj97t+SOBZJkiRJkqSm7E0pofFvYKTsuW0L83dPEJek7nkDpd/9aYljkSRJkiRJasoI0eqymNTYuey5EwvzPpQgLkndszjwLPH7fgyYN0GB1QAAIABJREFUljYcSZIkSZKk5ryPUiLzvMK8WcRo5uPAcYniktQ9v6P0u98hcSySJEmSJEkVZgLr5cxfBHiQSGiMAS8G9qOU5Pi/XgUoqWc+Quk3flTiWCRJkiRJkqp8GrgCeA+wRNn8L1FKapwI/BFYWHh8R49jlNR9a1D6zV+fOBZJkiRJkqRcJxPJi2eAU4CdgBWBpwvznyNaZhaTHAuB6QnilNRd11P6na+dOBZJkiRJkqQq04FLKSUwxoH7gWsy88qndVMEKqmrjqL0G/9I4lgkSZIkSZJyLQfcCYxSqo1ZK4k5Drw6TZiSumgHSr/x3yWORZIkSZIkqaYXESOTF2th1psOSRSjpO6ZCjxCqaTE7LThSJIkDYcpqQOQJGkI3QTsU/j/eJ3lRrFruTSMRoELC/+fAeyWMBZJkqShYSJTkqTuuAh4HzBSZ5kRTGRKw+r8sv9bQkKSJEmSJPW946jftfyedKFJ6qIlgeeJ3/lD2IBAkiRJkiT1uSnAudQe9GcMWCRZdJK66Y+UfutbJ45FkiRp4HlnWJKk7hoD3gbcUPh/1giwVk8jktQrdi+XJEmSJEkDZw1iFOO8kcxfmzAuSd2zPqXf+T8TxyJJkiRJktS0LYBnqO5m/smUQUnqqlsolZFYLXEskiRJA82u5ZIk9c7fgf1z5m/Y60Ak9cx5hX9HsHu5JEmSJEkaMF+iskXmbWnDkdRFu1D6rZ+bOBZJkiRJkqSWjAC/p5TceDBtOJK6aDrwOPFbfxpYNG04kiRJg2skdQCSJE1SiwB/ArYiEhxLAPOaWG9pYHEiObIIMAuYWli/3JPAKJE4eQ5YADwFPNGB2CW15hfAmwr/35tSd3NJkiS1YFrqACRJmqSeJUYr/zuwKrABkWjcAFgLWAlYEVgeWLns/9MnuN8FwMPAQ0RL0EcK/z4I3E4MTHJnYTlJnXEepUTmXpjIlCRJaostMiVJ6q1pwCbECObrAVsCOxCtMqcmjKvcAiKZeTOR2LyVSLheDyxMGJc0qJYhbiBMBe4nRi8fTxqRJEnSADKRKUlSd60CbF6Ytge2Y3Br5M0HrgH+UTbdkDQiaXD8mTgGAGwG/CthLJIkSQPJruWSJHXWUsCuwB6FadUOb38e0QX9GaIG5pOF+U9QauE1BViy8P8liFZgs4DZRH3Ndi1GJGK2L5t3L3BhYfpDWTySKp1P6bezFyYyJUmSWmaLTEmSJu6lwJ5E4nJb2r9ReB/RjftW4L9E3cqHiDqWDxBdU5+ZYKyziHqbKxM1N1ck6nGuSnR1X59oRdqOBcAVRFLzAuDaCcYqDZONgOsK/78K2CZhLJIkSQPJRKYkSe1ZF3h7YVqnxXXvJxIZ/6ZUg/JWmhu1vBdmE0nN9YjBhzYFtiYSnq24BTi9MN3RyQClAXUHsCYwRtw8eDBtOJIkSZIkaVgtAxxE1LobI7pyN5qeJwbKOR7YH9iw51F3zirA3sAXgYuAp2nuPRgn3oODgRV6HbTUR75N6TdxYOJYJEmSJEnSENoG+AXRdbqZpN31wNFErcyZCeLtlUWA3YFjgJtoPrF7OjFquzTZ7EHpt3B24lgkSZIkSdKQmAa8CfgrjZNzzwC/Jlprrp4i2D7xQuD9wDnAszR+3y4H9iEGI5Img5nEgFjjxKBdw3yjQ5IkSZIkddks4H+Au6mfhBsFLgHeTWmEcJUsTSR2L6NxN/w7gI9iUkeTw68pffd3r7GMyX1JkiRJklTTVCIpeS/1k243AJ8CXpAmzIG0BvBZ4Gbqv7d3AQcAU5JEKfXGuyl957+V8/xs4PU9jUiSJEmSJA2MXYkRxGsl2MaIwW32BkYSxTgsdgDOpX4rzRuBfVMFKHXZCkSL7mLyPutI4IO9DEiSJEmSJPW/TYArqJ1Qmw98H9ggVYBDbEPgJKK+aK33/zIGe5R3qZarKX3Py7/jqxDHnY+kCEqSJEmSJPWfmcCXiRG08xJoTwNfB5ZNFeAksgJwLLUHB3oOOByYkSpAaQJqteA+nNJ3/NCy+ScW5h3S5bgkSZIkSdIA2Jaoc1lrAJ8zgTWTRTd5vQA4AVhI/mdzHbB1suik9qwAHEe0tCy3GaXv9uWFeetT+v5/vFcBSpIkSZKk/jODaPlXrE2XnX4LvDhZdCraBPg/8j+jhcDXgGnJopNa9x5gHnAEsERh3gilgcUWAssBv6FUO/ZTvQ9TkiRJkiT1g1WoXQvzAWCfdKGphr2B+6hdO3OldKFJLRkBLiW+u3OAg4kbKydQ+k5/icrv+GdSBCpJkiRJktLaHrif/JHITwWWSReaGliKSPbkjXB+H1EmQBoE6xH1Xovf37uBo8seP0JlWYXD0oQpSZIkSZJS+TD5A/rcAeycMC61Znci8ZP9HJ8FDkoYl9SKz1JZj7f83+z0hUQxSpIkSZKkBLJdNYvTBdgKcxAtC/ye/M/0qIRxSc2aDlxPZcvLvNbGxa7mkiRJkiRpyI0A3yS/K/lRwNR0oWmCphKfYV7yx2SmBsFW1G6FWT4dkSpASZIkSZLUGyPAd6hOCswFXpswLnXWvsBTVH/OR6cMSmrScdRuiVmcjkwWnSRJkiRJ6onvU50QeAzYImVQ6optgSeo/ry/mTIoqQmLEjVfy7uYZ1uPfy1ZdJIkSZIkqesOpjoh8DCwacqg1FWbEaM9Zz/396UMSmrCntRujTmKrYslSZJyjaQOQJKkDngFcCEwrWzeQ8BuwHVJImrPTsDWNZ57Cvheh/azJ7BJjeceAX7cof30wouBPwArl81bQIx0fmmKgKQmnQG8CZiSmT8GfAv4WM8jkiRJkiRJXbUW8CiVLZqeAF6UMqg2HUX9unkbd2AfI8Addfbx7w7so9c2obpm5kPA6imDkhpYDpjD/2vvzsPkqOs8jr9zBxLCmQOQOxAwICCInMolCHIoCwqIILCgsIoCq8DyqIi7GJb1QFcFXBUQhICAAko0HFkEQSGoYJQbOZMQroQk5GAy+8evZ9NV1d3TM13dv+nq9+t56mG6uqr6U109Q+rbvyM7+U8XoZApSZKklPQ3wJIktZMRwE3A2mXrVgAfB/4eJVFzHZPDMfYBNsnhOAPJw8CxhCJQj3GEz8awKImk3r0CnEnlf48PaXEWSZIkSZLUZP9JtkXhF6MmakxvLTLnkOw+3x8/7eU12rFFZo+vkD2f82IGkuownWSrzBXAJVETSZIkDVCOkSlJalc7AveTbLl0LXA0yZZ57WQKcFYv23wI+HU/jz8GmE2YNbmah2nfCZIGATcAHylbtxzYgfYaK1WdZWNCC/KRZetuB34BrAGsVlq3Zum/owktjUcAywh/7xYSPutLgcWEYujrhAnP5hG+BJlbery8aWciSZIkSZIyBgG/I9ny7gVW3ui3q0otMv+cejy1geOflDrWI2TH52vnFpkQhhmYTfKc7oiaSAqGAJsRJts6g9Dq8k7gGVYWJFuxzAP+CtxI+JtzArArYcxOSZIkSZKUsyPI3px/MGqifFQqZH429XgpyTFB++Le1LHOoHiFTIDDyL6PB0VNpE4zHHgvcBpwNTCL8LvbqmJlf5dXgXuAbwFHUrzxdCVJkiRJaqlBhO7P5Tffv4iaKD+VCpl7An9KrTu1H8fenNDdtOcYy4EJFLOQCTCN5HnNxCF11DxjgaOAi4H7gCXEL0rmtcwFbgbOBd6PE2hJkqSIGp0wQJKkVjsY2KbscRe9jyvZ7q4Atit7fBzw/T4e43iShbxfEcbNK6ovAvux8pzfTZix/fZoiVQkQ4CdCN3EP0gYh7XS7ON9sYTwBcNqwIuEYvwrhLEuYeU4mEuAt1L7rkH4rI8itAYdUVo3DhhP+NJibGnpa85xhL+7B5ceLyD8Hk0rLc/38XiSJEmSJHWM20i2Fro6bpxcVWuRuTbZbqnbVD5ERYOB51L790yIU9QWmRBa6paf2w1x46jNrQJ8FLiGUGDsT+vGOcAM4DLgX4FDgK0IE3FBKEQ+Q2Nj4dYyhFDU3Ak4Bvh34DrCWLyL+3lOjwAXAts3KbMkSZIkSW1pA+BtkjfRRbp5rlbIBLgptf7CPhx3/9S+rxBabEGxC5m7kTy3ZYTWaVK9BgN7Az8G5tO3At+zhCLhGYTP4up1vubewPW5nUH9BgEbEb7kuJBQcH2Tvp3zLODfSseRJEmSJKmjnU7ypvnBuHFyV6uQeWhq/RzqHyLmmtS+3y57rsiFTAizM5ef36fjxlGb2BK4CHiB+gp4KwjjsF4IfBhYt8HXP6zB/fMyBNgWOBm4kvB3p9734+7SfqNanlqSJEmSpAHgTpI3y6fHjZO7WoXMocDs1HMH1HHM1cl2GS0fb7PohcyzSZ7frXHjaIDbndCKMt3yu9Lyamnbk4H1Y4SNZDJhXOLp1DcT+3zCJEgbxggrSZIkSVIMIwldg8tvkDeLmih/tQqZAN9MPXdtHcc8JbXPw6nni17InEzy/BbiZIdKGgmcSBjrsbei3EuE38OdCa0VO90awJGEWc3Tf5/Ty3JC6/CdoiSVJEmSJKmFdiF5U/xU3DhN0Vshc+vUc0sJEwHVcn9qn8+nni96IRNC8an8HIs0rqr6bzhwGjCX2gW4BcAVwH5YvKxlbeBU4F5C1/Ja7+kMwt90SZIkSZIK6TMkb4R/FjdOU/RWyAR4KPX8KTWONym17XKyk910QiEzPVHSP8eNo8gGAx8HnqZ2se2+0narxonZ1jYlzIg+j9rv8U2EWdslSZIkSSqUdLfqc+LGaYp6CpmnpZ6/v8bxLiRbNEjrhELmV0me4wVx4yiifQmT8lQrrHUBt5S2U+NGAMdSu9t+F2Gs0U0iZZQkSZIkKXfpVnVHxY3TFPUUMtcGlqS22brCsYYCL6a2+3CF7TqhkHk8yXO8Jm4cRbAe2b8h6e7jF+GENM0yCNgf+A3Vr8FCwtAXdt+XJEmSJLW9u0je9O4dN05T1FPIBLghtc2UCtt8KLXNXGBYhe06oZB5IMlzvC1uHLXQIEKLwFepXDxbBlwKTIgVsAPtAtxN7S79k6OlkyRJkiQpB38kebP73rhxmqLeQuYhqW1mk52J+/rUNt+s8pqdUMh8P8lzvDtuHLXIJsB0KhfLVhC6M0+Mlk77Ag9TvcA8hdA1XZIkSZKktpMuZO4UN05T1FvIHEooXpZvd0DZ82uR7X6+bZXX7IRC5h4kz/GeuHHUAkcRuipXKpL9HmeuHyiGAp8GXqfytXoAu/tLkqQyg2MHkCSpTotTjzt5JuG3gatT644r+/loki2Z/kQxC5T1Gp16vDBKCrXCUMIXAj8DRqWeWwycTShs/6nFuVTZ28AlhJnLf17h+R2Bh4APtDKUJEmSJEmN+iXJljpHxI3TFPW2yIQwhlz5dksJEwEBPJh67nM1XrMTWmR+guQ5To0bR02yDnA7lVv2zQA2j5ZM9ToYeIHs9XsbOIsw5qkkSepgtsiUJLWLZ1OPN44RYgCZBcwsezwc+CihwLlD2fplhNZpnWyT1OP0Z0nt712EVpb7pNYvAU4mfCHwRIszqe9uIVzLX6bWD2FlS9vhrQ4lSZIGDguZkqR28XTq8dZRUgwsl6ceHweckFp3KzCvJWkGrvQMyOnPktrb9sAdwDtS618gFDB/2OpAashrwEcIwwB0pZ47EvgFMLLVoSRJkiRJ6ov0zNOPxo3TFH3pWg6hK3l6Up8FqceH9PKandC1/DmKP1FUp9oReJXs783/AuMj5lI+9gLmkr2+d5Ed+1aSJEmSpAFjFLCclTeyK8i2wGp3fS1kAlxfYZ+eZS4wrJf9i17I3Izk+S0hORGS2tceZAv33cB3CJP+qBg2JXxxlb7Od5Kd0EmSJEmSpAHj9yRvZE+JGyd3/SlkHlRhn57lG3W8ZtELmZ8neX63x42jnGwDvEn2M//1mKHUNBOAR8he72mE8TMlSZIkSRpwziF5E3t33Di5608hcyjwUoX9uoFt63jNohcyH6D+GdzVHtYCniT7eZ8SM5Sabk3gj/TvCxtJkiRJklpuc0KX8vLu5ZOiJspXfwqZABdV2G9mzT1WKnIhczuS59YFbBQ1kRo1lDCxT/rzfnbMUP30Y+CpKstNOb7OjBqv880cX6cV1iT75UQ3YaIzSZIkSZIGnBkkb2AvjZomX/0tZDaiyIXMq0ie221x4ygH3yX7O/LfURP136+pPixEN/l8SbNLL6/x0xxeo9XGk53A6y1g55ihJEmSJEmq5CiSN7BLgQ2iJsqPhcz8TCQ5OVQ3cGjURGrUoWR/P34HDI8ZqgG9FTLPz+E1Lu3lNdqxkAmhtfVCkufyD2C1iJkkSZIkScoYAjxOMW7G0yxk5udGkuf1N2Bw1ERqxDrAHLKFq7ERMzWqt0Lm8zQ2kc1I4PVeXqOd/3b+E8mhRrqBS6ImkiRJkiSpghNI3ryuAHaNmigfFjLzsQ/Z9/GoqInUqB+RbYm9fdREjatUyHwr9XifBo6fbr2ePna7FzIBvkX2/wW7RE0kSZIkSVLKYOBBkjewjwKrxAyVAwuZjVsVeILkOT2IrTHb2fZkP6dfipooH5UKmdemHl/ZwPF/mzrWNRVer90LmSOAWSTPaSb+vkuSJEmSBpi9yHYr/E7URI2zkNm4y0ieTxe20Gp300he01nAsKiJ8lGpkHlA6vFiYPV+HHt94G2SvwcHVni9di9kQvh/Qfq8Do+aSJIkSZKkCtJFq27gpKiJGmMhszGnkn3/vh01kRr1brJfWHwwaqL8VCpkrkUYz7V83Yn9OPa5qWNMB7au8HpFKGQC3EDyvP4MDIqaSJIkSZKklDHAMyRvYJdgC7xOtCewjORn4XFgVMRMatxUktd0RtQ0+apWyDwrte7ufhz70dQxjqHYhcytyH4ps2/URJIkSZIkVbAdsJDkDexs4B0xQ6mlNgHmkfwMLCAUbtS+1iFM6lN+XT8QNVG+qhUyJwDLU+sn9eG4u6f2nU8YO7bIhUyA60me29S4cSRJkiRJquwwst1PnyYUuFRsGwFPkrz2K3CMvCI4jeR1fZRidReuVsgE+FVq/fl9OO7/pPb9YWl90QuZ7yN5bkuBNaMmkiRJkiSpiq+RvUn/B7BZxExqro0JBev0dT83Yibl506S1/WsuHFyV6uQ+dHU+ueBIXUccxShBWb5vruVnit6IXMQ8BjJ8zs6aiJJkiRJkqoYDPyc7I36s8DEiLnUHFsCL5K93ldRrFZ7nWoNst2ri/alRK1C5nDgldRz+9RxzGNT+zzOyt+HohcyAS4geX5Xx40jSZIkSVJ1w8iOk9YNvIQTABXJ+4C5VC5iDo2YS/nZn+S1nRU3TlPUKmQCfC/13JV1HDPdirW8dXInFDJ3I3l+z8WNI0mSJElSbUOAK8jesC+neF1TO9HJZGcn7yli1tP1Vu3hKySv7w/ixmmK3gqZ70k9txhYvcbxNiI5c3cXsGHZ851QyBxGeJ/Kz3H9qIkkSZIkSerFEOBHZG/aewpeq8aLpn4aCfyEytf0UsLQAiqOqSSv8Ylx4zRFb4VMgIep/31IF39/m3q+EwqZAPeRPMcizXQvSZIkSSqowcDXyc5m3g3MJNzUqz1sDzxC9jp2AefhmJhF9ADJa71H3DhNUU8h8wup5++ucqxBwJOpbdMT3XRKIfNKkud4ctw4kiRJkiTV72DgDbI38MuAKcCIeNHUi2GE4QCWkr1+84GPxIumJktP5LRR3DhNUU8hczzZSY8mVTjWnmR/P9ItzzulkHk+yXM8P24cSZKUJ7thSZKK7hZgJ7KThfQUyWYCO7c6lHq1C/AXQrF5eOq5RwnX7KZWh1LLrJZ6/EaUFPHNBX6TWvfxCtt9MvX4WsJYkZ1ofurx6CgpJEmSJElqwGqEm/tKYyy+DXwfmBAtnXqsB1xGctKS8uVKYFS0dGqV9IRORWw5XU+LTIDDU9v8g2RjhFHAm6ltdq1wnE5pkXkKyXO8JG4cSZIkSZL67yDgeSoXyRYRWgCuES1d5xpNaCG7gMrX5iXgsGjp1GqLSF7/Iraqq7eQORyYl9pun7Lnj0899ziVx43tlELm50me48Vx40iSpDzZtVyS1GluBd4FXE64yS23KqGY9gThZriIrcAGmlUIE5o8Sygip7sUdxNaaG4F3NjaaIpoYepx+nPRSZYRWpOXO67s50+mnvsJ2b9tnSRd9F4UJYUkSZIkSTnbj+xMv+XLbOBLwNhYAQtsAvBV4GWqv/+PAXvFCqio/krys7Bj3DhNUW+LTIAdUtstAsYAmwArytZ3ARtUOUantMi8lOQ5fi5uHEmSJEmS8jMMOJnQdblaQW0JYWzGbSJlLJJtCYWGxVR/v18mtIxNT/KjznEzyc/Ex+LGaYq+FDIhTH5Vvu2JZGfonlZj/04pZE4neY6Hxo0jSZIkSVL+RgNfpvoYjd2Elk/TgWMo5ph9zTKG0BX2Lqq/t93A68A5hC7+6mz/RfKzcWHcOE3R10LmGalt7yFM/FO+7qga+3dCIXMQ8CrJc9wqaiJJkiRJkppoLKGIMp/aRbeFwFXAAcDQKEkHtmHAwYSx/Wq1vuwpYE6hdhFHneVjJD8jd8eN0xR9LWSOIzube/nyBrW/BOiEQuZWZP+2OCeAJEmSJKnwxhAm/Hma2kW4bmAu8D3gQ3R2a8LRwCHAJWRnWa60PAF8Blu3Kmsjkp+V5cCaURPlr6+FTMh2uS9fLull304oZJ5O/V3tJUmSJEkqnMGEloXpcdeqLcsJXT7PIkzQMaj1kVtqU8IYo7cQxhKt5z26BzgCGBIhr9pHesKfT8SNk7v+FDIPq7BPz7JzL/t2QiFzBsnzc6IfSZIkSVLH2hr4OvAs9RXsuoHnCN2rTwd2A1Zpeer8rArsDpwJXAe8QP3vw9PA13C8OtXvApKfoTvixsldfwqZwwmTYaX3e4zevzQpeiFzIskZ3LuBzaImkiRJkiRpABgMvB+4DHiN+ot53YQx7h4kdEX/FLAnMKGl6euzHrAXIeP3gYcIrU37cq6vlPbdjeK3TFX+3kny87QCmBQ1Ub76U8gEOAG4NLUcXsd+RS9kXkTy3O6NG0eSJDWDNxWSJDVmBLAvYdKfAwhdrftjPvA4oWXVY4TWji8Txt+cQxhzclmjYUtGECY1mgCML/28AaFINAnYgjBGaH88AdxWWu4kv8zqTPcCu5Y9/gmhkFcEvyb8zSi3NuHLkWbYGngkte4qitFlfy3CDO6rla07Hrg8RhhJktQ8zrIqSVJjlgK/Ki0QioA9Rc33UX9X8tWB95SWal4jFDYXl5alpWUx0AUsKG03hjD+5ChCV9SRpRyrEoqXeU6aspgwLt1thIk1nszx2NLFJAuZxxCGd3giThwNUGeSLGLOAaZGyiJJkiRJUlsaBkwmTIZzJTCL7Bhu7ba8RJjY5yzCeJntPOanBr4hwKMkP4O/jJooP/3tWt5fRe1avgGwiOR5fSFqIkmSJEmSCmIccBDwZeAaYCbwJvELlOllPvAAcDVwLnAgodur1GpHkv18Hhw1UT4sZObjZrJftoyOmkiSJDWNXcslSWqtl4FbS0u5d7ByfMqJhILneEJX8HGEcSwH55RhRSnHPGA2obv6XOApVo7R+VJOryU1airwWZJdzC8ljJ/ZrPEk1R6OIVvUPhdYGCGLJElqAQuZkiQNDC+UljuqPD+EUMwcSxj3sto4mFB9/MxlrCxgrmjGSUhN0E0oZN5PGK4BYF1Ca8KD8bPcqd4JfC+17vfAFRGySJKkFrGQKUlSe+giTGAxJ3YQKYKHCJP8fLls3YHABcDZURI17jbg+dS6pU18vdeAy1Lr/tDE12umNQljpY4pW7cUOAkL25IkSZIkSYpsOHAf2XEej44ZSi03FPgt2c/BqTFDSZIkSZIkSeXWJQzBUF7AWgzsHjOUWmYQ8AOyRcx0S1NJkiRJkiQpuncDi0gWshYB+8YMpaYbBHyXbBHzXmBExFySJEmSJElSVceQLWgtBvaLGUpNMxj4Edlr/iwwLmIuSZIkSZIkqVfnkC1svUWYyVzFMQy4iuy1ngNMjphLkiRJkiRJqtuZhFmqywtcbwPnEVrxqb2NBe4gW8ScjUVMSZIkSZIktZlPAV1ki103A6tHzKXG7EjoOl6pO/nEiLkkSZIkSZKkfjuJ0BIzXfT6G7BlxFzqn1OBpWSv51PAxvFiSZIkSZIkSY3bE5hL5UmAzgKGREumeq0L3Ej2GnYDd+HEPpIkSZIkSSqIDYA/ULkQ9hCwfbxoqmEQcCzwKtnrtgKYgoVoSZIkSZIkFcwqwOVULmYuBb4CjIgVThkTgTupfL0WAIfFiyZJkiRJkiQ133HAa1QukD0DfAJnNo9pHHAxlcfC7AZ+B2wRLZ0kSZIkSZLUQuOB66lcKOsGZgFHREvXmUYRxix9g8rXZBGOaSpJkiRJkqQOdTgwm+oFzRnAPrHCdYgxwOlUnpCpZ5kGbBQroCRJkiRJkjQQrAV8h+pdmbuBvwDH4xiaedoY+AYwn+rv+zPA0ZHySZIkSZIkSQPSpsBVQBfVC2tzCJMCjY+UsQh2B34OvE319/ll4HPA8EgZJUmSJEmSpAFvMnAd1Yts3YQi3HTgWGB0nJhtZUPC+JaPUvt9XQhMAVaPE1OSJEmSJElqP7sQJgSq1XKwG1gAXAHshxPRlFsH+BfgXmAFtd/DOcCXgbFRkkqSJEmSJEkFsB5wHvAqtYtx3aVtrgNOBtaPkDW2yYSWl9OBZfT+fv2Z8F6tEiOsJEmSJEmSVESrAacBj9B7ga6b0ApxJvAfwJ4Uswv6BsBRhBapc6jvfVkC3ADsHSGvJEkqqEGxA0iSJEkD1GTgCMIYmZvUuU8X8BihuDkTuIfQIrGrGQGbYDSwHbBDadmDMPN4vWYCPwV+BszLO5wkSepsFjIlSZKk2gYTWlseAxwCrN3H/d8E/k6YBOex0vJ4aVmaW8q+WQeYBGwJbFFatgIm0vdLWTl7AAABMklEQVQxQP8CTCUUL5/NMaMkSVKChUxJkiSpfkOA9wAHlJYdCIXO/ugCngdeAl4GZpf+O6+0bh7wFqHYuZgwIdGbhK7bbwDDgVGlTGNKx1yD8G/8tYDxwDhgQunnscC6hNnF1+pnZggTH90O3AZMA15o4FiSJEl1s5ApSZIk9d9YYH/gA8DOwOYU79/YbwEPEbrJTyPMUL48aiJJktSRivaPLEmSJCmmMcC7gN2A3YH3Eoqd7eRpQrGyZ5zPB4jXBV6SJOn/WciUJEmSmmtDwniUW7ByTMpJpfWx/j3+FmGMzp7xOsvH73wzUiZJkqSaLGRKkiRJcaxCmBG8Z+zKcaWl5+exwDDCTOLDgJGlfXrWrQDml441v/R4IaHb9yJWjrn5curnF0tLd3NPT5IkKV//BwbS5MddGBmCAAAAAElFTkSuQmCC" width="75%" style="display: block; margin: auto;" /> --- class: center, middle # THE SOLUTION --- ## NetCoupler: A "causal structure learning" algorithm that... - Finds most likely network structure - Allows inclusion of exposure and outcome - Identifies causal links between and within network .footnote[NetCoupler was developed by Clemens Wittenbecher.] --- class: middle, center ## Algorithm (steps) used in NetCoupler --- ## 1. DAG skeleton of network is estimated <img src="data:image/png;base64,iVBORw0KGgoAAAANSUhEUgAABwcAAAJnCAYAAAByAlwjAAAABHNCSVQICAgIfAhkiAAAAAlwSFlzAAASdAAAEnQB3mYfeAAAABl0RVh0U29mdHdhcmUAd3d3Lmlua3NjYXBlLm9yZ5vuPBoAACAASURBVHic7N1nuCRltbj9e0+CyQMMDEnyIYggQckIKkFJophQQUTFLMa/etRjzseAGFAPglnBjAQTDkklB0FAchxgSAPD5PB+WLvfXV1VHaq7d1f37vt3XX05VXvXU6u7R2rNE9YDUm3bA7elXs8tNSJJkiS1ajzZ3O7NpUYkSZKkTjgSWJ167VBqRJKknjYBeFnq3DLgdyXEot6zBrBF6tyUMgIRAOOAbYDtiM69pIuBeV2PSJLUiw4CZqbO/QFYXEIs6i1DZHO7tcoIZEDNBJ4P7A7sBGwNrANMB54CHgPuBS4FLgHOApaUEqkkqRfNAN6SOvcQcFoJsUiSpDEgPavkkXLDUQ/Zhezfj0NLjWiwbAgcDnyc6Bx6hOz34fciSUq7luxzYuNSI1KvmED278aHSo1o7FsXeCdwIbCc2rlc3ms+8BlgWtejliT1ok3IPiuubvLa9YgJQpXXJqMRoArbIvVqZ9KWKwclSYW1Ojj42dR1K0YlOhV1L9Xfy/fbaMvBwe7bHfgbsIBinUd+L5KkilYHB4/OuW77UYpRzTuH6u/kijbacnCwu75GVGUpktPlvW4D9uly7JKk3tPO4OCPU9fdPhoBqpDxZL/Pj7fRnoODkqRCxpUdgKQqWwL7E+VCJEmS1L92BiZ2oJ0tgL8CL+xAW5IkSZIkMaHsANTTFgFXps4tKCMQSZIktW012dzOPYu7awmxp+AFxGqPh4jyoVOBzYADgNeQ3Td0EnAm8Czgpi7FKkmS+sd9RK6QZB+eJKkmBwdVz01EB4TK9ThwFdGZdwXwUuBlpUYkSZL60UrM7cpyKXAa8HNqd9RdA/wW+DBwMnBM6udTga8DB41SjJIkqX9dDry87CAkSf3DwUGptzwFnE8MBFZetxEz/Sv2KyEuSZIkFXc+8DHg4gLXLACOBR4D3pn62YHAHsA/OxKdJEmSJGkgOTgo9ZbfDb8kSZLU344nJnm16r3A84HtU+ePxMFBSZIkSVIbxpUdgCRJkiSNQe0MDAKsAL6ac37vNtuVJEmSJA24oisH9wCmDf9589TPhoADmmjjCmIPtWZNA54HHAxsBqwHrA0sBB4EbgHOI0r2PFWg3Yq1gF1T564CHk0cTwJeQJTxeQawIfHZPQGcDXykwT02B54zfO32w+9hLaJU5GPD9/o3cAEwl2Kfz3Rg98TxGqmfb0jj72UxcEkTbUPshfJwgfiS5hCf43OBTYHZwGRg/nCb1xGf56XEnjit2J/qv9f3ADenfmdn4CXAbsAmxPf7OHA/cCHw+5xrJEkai54BrJ/4c9oewAYN2rgJuLfAPScTedELgW2G258NLCJygruBPxH53bwC7VZMJFuC+xbgrsTxOCI/OgR4JpEPTCXKOV4PHNXgHk8bvsczge2AjYEZw+/tUSK/u5XIK84H7iwQ/xCxWqxidurnM2gu5/4b2Xwq3TZEnEXiS1qbyNEPJvLddYfje4zI7W4E/gj8hdbydIg9Emcljh8jyq4nbUPssbMPsAUj3+VDxAq7s4nvoh/9Lefc+jnnJElKO5LIiyD6YJKmAi9roo2ziRytWTsSOd6BRH40h3iOP0Y8l/8NnAv8gdbyvE2Jvpyk84AnE8dPA15N9CU+fTiGCUSeeQrwP3Xan0HkE3sTufF2RC62FjFp57Hh19XARUT/0T0F4t+QkUk+eQs2tqfx9zKf6DtM2wjYK3XuT9Te67ie8cTncAiRt29EfA4Th+8/n9jj8Fwi11vYwj0mEn9Hk66juj9uDSLHexHRd7sR8V0+TPTh/RX4JfCPFu4vSSIGqJKvR+r87rU5v1/0tW+Tca0PfAtY0mS7C4D/BqY02X7FfjltHTj8syHgDUSHV637np/T5hCwJ/A1oiOqyOfzJPAVGnfEVexUsP281+012t4l53cPbTKupC2AnxMdVM3Ecw/wOlpb2bog1dZJiZ9tQyRGje6/EjidGMTtRd+kM9+LJGlsysvXNq7xuz/O+d2ir7c1GdcM4NNErtNMuyuA7xKdKEVUJmAlX+9N/PxwotOh1n1rTYLaCfgcMdBY5PNZCZwB7NBk/OMKtl/rNT2n7Qk5v/ehJuNKWo/I05c3GctC4ONER2RRF6Xa+lviZ+sDP6O5HHMu+QPgvW4K2ffyYKkRSZLKsgnZZ8LVdX7/sZzfL/ratMnY9gX+XqDdxcCXiLytiGNy2tpm+GczifxkWZ37np7T5nSi7+/PRP5Z5PNZNtzmZk3Gf0TB9mvlNHmOzPndZvPPpMOIQdxm43kEeBcx8b6IWTltvS/x8xcT/YPNxPAbmu9HlSQl5P1HvZZuDQ6eQHQitNL+rcDWTdyjYr+cNg4E1iRmn7TyUH5mi7EnXw/QXMmgXh8cfBuwtMW4LqP4zOhag4MvovjfqZuJ5LvXODgoSaqnFwcHDyFmGLfS/hPAQc29daD24OAQ8L9N3C8vFx5qMfbkawkx+amRXh8cPJxsvtXs616yFTsaqTU4+GxixUGR+z9O/5Xk3JTs+/hXqRFJksrSi4ODk4Dvt9H+XcTqvmbVGhzchOYGtH6Y0+bhbcRfec0nqmQ10suDg5No798G11JsUl+9wcHPtHD/W2h+MFuSRO/tOThEzMj+DrVnFq8mEpzlNX6+JVEic5c24/gRjUtKVX63qEopgsV1fmcOsSoxXRKgn3wS+Aa1Zw+toLr0Q9qzie9yyzbjOAQ4k+Kz1bcmSkRMbPSLkiSpphOA35Etj1mxguhQqZUTTCdKT72yzTg+RfUKwlpayY+XEKWyKp1wedYgOs/e1EL7veK1wK+JVaB5lhMrL2uVh9+IGNx7Xptx7EBUgyg6iWwmcBb9NbM8r8P07q5HIUlS1kzgHGpPflpK7L97JXAH+f14mxATgdrpw5tJlBbdronfLdqH9yDwH+I9/IfaJTpnD8fw7ILt94rJRP/Xq2v8fBHx/q+nehumpB2JPryt2ozlY0RluLTK9lIraly3FVGtY3yb95ekgZKeaVFv5eAexB4nBxAlfJLXrUz8rN5rVqbVEf+TE89q4iH8BrIPmC2Ad5K/zPyOBveq2C/n2jNTx7cTZbCeT/wDfVuiU+PDwGk5bSZXDj5JPJxOIOqip2dxTyf2yTuZ/FnYd1G/zMJ0qj/f9Kz882j8ndSaQd3OysHjcq5dTdQEfx/x3VU636YQexH+qsY1NxCJSjPSn+HvqJ4tdx/x92w3YgB2TaKj6qXE7Ku8+3+gyXt3iysHJUn1FFk5uDmxkmtX4h/h6etelvh5rde6dWJ5eU6bq4nOhXcQOVXSLOAVxB4m6WsWEfuwNJK3cvB3wKrE8YNEKasDhmN4GpHnvpHo5EpLrhxcPNzeW4i9mdMl7ScQ+xv/N/nl5RfTeBZ38vO9JHX9v2n8nexKfsdIOysH9yK/1NYCohNnW0Y63MYTeeRJ5Jf2WkDz5bfSKwcvJ/7+VI4fJyYX7kP8XRwi/h49F/gB+SVHf9HkvXtB3gz+E0uNSJJUlqIrBzcj+l62AH6buu6exM/qvSbUaHuImLyVjqdSZnMfshPFpwNHE3sSp6+7hdqTj5LyVg6enzo+hxjk2pTo85lMrC48AXh/TpvJlYPXEnnNfuT3KQ4ROc//EBPD0rHcRn71hoopjHy2/5Vz/Uk0/k5qrc5rZ+XgN3KuXQ1cQOwhmZ40vwtwKtX5deV1DTEprpG8lYN/TLU5l+ivS34X44mqdL+rEXOzWx5Iksj+R7Te4GDSZ1PX1Zq50az9yXY4LAKOb+LaKWQH9FYT/5huZL+c6yqvVcT+KM081JKeScymOZZIRJq1MfkDVJ8o0EZ6f8TvF7g2rdXBwc3JH+g8i8b15I8iOs3S1369yZjrlbk6hcZ7Un4857r7qJ0Ql8HBQUlSPUUGB5OOzrmumcG4WrYkBm7S+eJHaTyjd4jodEl3OFzXxLV5g4PJ1/8RM8yLGALuBN4OTCtw3RRi38R0DL8r0MY5qWuvKHBtWquDg9OJzq70tVcQA6v17Ez+/t0X0tzM7vTgYPJ1NrBOg+tfTHaAcgXN/X+ibJuSzYtX0PzAqiRpbCk6OJiUnmxSa3uZZr03J5bbaG4F4CTgJznXf7eJa/MGByuvBUTlqKIOAX5ObNtTxLrEPoXpON7d5PXjc679eMEYklodHHwh+YN8H6XxSstDye/D+1IT980bHKy8lhKLDhrJW2RyYxPXSZKGpf8jWsbg4AQiMUm2t4xiJYfGke08WUnj/Qf3o/bD6B0F7p80kdbKjUJ0IqVnyz9A8wOUvTA4+KOc6+bS/EDpkWQTk5U0V5qg1uDg15q8N8C5Ode/oMD1o83BQUlSPb0yOHheTntvLNhGOt9cTcwerqfe4OA3aD1Ha7XM+BCxgi2d12zR5PW9MDj4wZzrbqH+qtGk7ckOFK+muRL+tQYH/0Dzk7c+nXP9B5u8tkx5KzJOKzUiSVKZemVwcH2ipHqyvftoPGEoaRzZgbWlRGWnemoNDi4F9ixw/06ZQqyUS8ZyK82Vqe+VwcGrcq77aoH7HpVz/VIal3+vNzj4iibvPUT+IotnFYhfkgZa+j+gZQwOvjInjmZLHCWtR9SgTrbzrQbX7Jdz79XEKrey7JYTz4FNXlv24OB6ZJPERcRqwiLyNrRuJjnJGxy8hmIdenvltPGpAtePNgcHJUn19MLg4A5kJ/r8sIV2JhBlNJPt/L3BNbUGB/9N8WoQnbI22Rylmf0PofzBwfHEqsn0dUX3DXxHThtzm7gub3BwPrX3sMyzFtn8tMxcvxnHkX3fi4iOYUnSYOqVwcG8STcHt9DOJmRX93+mwTW1Bgc/2cL9O+WAnHh2bOK6Xhgc3DvnmtsoVgUNomR7up2PNbim1uBgM1XgkpJlYSuvtxdsQ5IGUjMzWbrhXanjeRSbpVLxENnZtC9qKaLGD7HRdBlwc+rc7mUE0oJjyHa8fZvYA7KID5PdrPo4Wivv+bmctur5B7FaM6mdzbElSRo076J6hd4K4tle1ArgK6lzexCTkYr6PDGLuQyPEispk/oltzuAKG+Z9Edif58ivk0MMibtR+y3U9S3gYcL/P5jxJ45STu3cN9u2Z14j2nvA+7uciySJCVNAt6cOnchkRsUdTfwy9S5VsqCLqa1PsRO+RvZvKRf8rzjcs59mphUVcRHiEG5Rm03sprowyvij2Rz/F7O8ySpZ/TC4OBsYqVc0hkUfxBVpGcBb0jzZZsqrieW1ZfputRxvyyJ3zfn3OkttDOPbHI5i9jPsYjFwG8KXrOaWHWR5CxtSZKa98LU8QXAPS22VSmtWDFEzHIuYhHZzqduS+d2u5YSRXF5ud0PWmhnBfkzwfPab+TnLVxzTep4Q5rb87DbNgJ+TXbG/m9oXBFFkqTRtivZ/X5/1kZ756aOdwBmFGzjHGIiUFlWAv9JneuXwal0Tr0I+FUL7dxCTLRP2oziezzfDNxQ8JplZBdYbFiwDUkaSK2swuq0fcju/fKnNtrLK7X0bIqVTLi4jfs3YxIwlShxVEt6pVsrM+TLsEfq+F7gXy22dTZwWOrcnsCVBdq4kkgUiro3dTyzhTYkSRpEWwEbpM79uY32HiD2sUl2LuxKsck/VxOdHaNlCjBt+DWL/H0Np6WO54xiPJ2Uzu1Wk10F2axziJnlSXtSrAz+Y8CNLdz7vtTxENH5WGZnYtq6xL+D0h1aNwKv7344kiRl5E3q+Vsb7aUn74wnSnIW6ZdLD0p10mRisni9HA+yE46a3Ze5TLOAbVLnLgWeaLG9PxHb9CTtCZxZoI1/tnjveVSXcrUPT5Ka0AuDg+kOB4Cb2mjvMWIwaFLiXNGHcnpmd7u2IjbT3RN4BtnSTM2oN5DYK+aQ7ehqtv59nrxrm6nbntRq6aV0MlR05pokSYMqL7drZTAn6UGqBweLTprqdG63MfByoizmDsTM6FqdRbVMJfZELlL6vAzpvWruovUBtWuJ2fXJDrRGe+Gk3Ue2bFUz8jq6ptM7g4NrE4PoT0+dv4PYe7xX4pQkDbZnp45XEs/lohW7KqbknCuyrzBE9a9OWYPYP/FlRJWzLWmt0sCsDsY0WrYlW1EuPVhbRF4fXjqvaeT+Fu/9ZOo4PSlPkpSjFwYH1885d1uH75EuedDIIx267zOBrwH7d6CtfhicWjvnXDvf5S1N3qOex1u894rUcS+U4JUkqR/krYj7XYfvUTQf6FRutxnwZeBIOpMbzKBzsY2W9AS1dnK7RUSnz9MS57qV263MOdcrZUVnEqsx0+Xz7yf2fEyvepQkqSzpyffjyZZ0bFfR3KATE2iGgGOAL5DfT1nU9A60MdryFiHc1UZ7dzZ5j3oWtHjvdJ5nH54kNaEXBgeLPvRbUXRgrdUl9EnvBL5C5zod+uHBljczqtUHO8T3sJrqmfhFE4v0IJ8kSRpd3ah2UDS3S88mbsXhwC+I8lKd0uv53TRidWNSu3nyAqoHBwc9t5tGlNJPr8R4mFgxWGRrBEmSRls3+vCKrvpqN8+bCPyIqPjVKUUrSpSh0314edcWzfNWtXF/SVJBvTA42ItL7fNmFhfxZuCkGj+7ldgX8W5iX7ulRDmphanfewNwUJtxdFteAtfO/j4riM9nzQb3kCRJvWMs5nYHAL8iO1AGkdP9g5gtfR+RuzxOtvTlgcAb24yj26bmnGt378b09YOc200Gfg/snTr/OFHS7N9dj0iSpPq6UdWq6MBauwNK3yV/YHABcAlwFXAPsQ92rTzoU+SX1u9leRPelrTR3uIm7yFJ6hG9MDiYfrAuofbAWqsu6nB79WwEfCl1bjXwHeB/ab4U0wGdDKpL8mZr5XUqNWsC1QOD0JlVnZIkafQ8lXPu87ReDjLPnR1sq5E1gVPIDgz+Cvgkze9nmFdutdfl5V3t5HaQHQwc1NxuDeDXwHNT558AXkB0REqS1GvS/T73EflQJ/29w+3V81zguNS5J4D3Az+k+cGy93cwpm7JW+nXTjnUvIHjQc3zJKkv9MLg4MOp4zWAT5A/46QfnEi20+PNxEykIrpRkqvT8uq8z2yjvbyVB52oJS9JkkZP3h56vwEu63YgHfJyYMvUua8C7ynYTj/mdouJlZBrJM61uzI0ff0g5naTgDOIQcCkhcAhwKVdj0iSpOak87zJFO/v6iXvTh0vA54HXFmwnX7M8/Im7rWT5+V9BoOY50lS3+iFfU7Sg4NDwBZlBNIhL0odX0JriVInNkDutrzOwK3baC/v2rx7SJKk3jE/59xWXY+ic9K53YPAB1toZ4MOxFKGdO7VTm43k+znkP63wFg3EfgZcETq/CLgMOLfDpIk9aoHU8drA+uUEUgHTCFbtes0ig8MAmzcfjhdl5ezb9dGe3nXPtRGe5KkUdYLg4OX55xLl9fpF1PJdpj8toV2JgC7tB9O1z1M7KOYtFMb7eV9Ble30Z4kSRp9Yym3A3hm6vg8YlZ5Uc/uQCxlSOdeG9J6idSdyO4jNEjlMycAPwZekjq/mBgsvKDrEUmSVMw/c84d2PUoOmNzsnvinddCO5vQn5PAbiS71VM7fZG75py7oo32JEmjrJ3BweWp4/FEiZyizif25EtKz9DuF+vlnLu7hXb2ovX9XNLfS7c3/00nihuQnyA0Iz2jGuAfLbYlSZLqyxvwaiWPuAF4IHXuULJ79vWLdH6XngjVjPVpfcJU2bldXu51WItt5eX4g5LbjQd+QJSpTVpKDBb+tesRSZIGydLUcat9TnnPq/Skl36R14c3r4V2Dm3x/iuBFalz7e7tXMRysqskd6S1am7jgCNT51bQv9sKSNJAaGdwML0JMcDsFtqZB1yTOvd8+nN2dd7gaCsdYW9vI4b097JuG221Ym7Oude20M6mRJ33pIeIDkdJktR5ncrtVgPnps5tALyuhbZ6QTq/a2Uy3Jtofa/vsZLbTQZelTq3Eriwhbb6zTjg+2Tf/zLgpbS2SkGSpCLS+cQ6tNYn+C/gjtS5lwBPbyWokuW9/ykttNHPfXh/Th0P0VrOfjCxgjLpH8BTrQQlSeqOdgYHH805lzfrphlfTB0PAV8H1mixvbLk7Zmyc8E29gWOaiOG9PfS7cTix2Qf/idQfK+hLxAzrJO+B6xqMS5JklRf3r6+reZ2/0v2mf0JoiRlv0nvx1K03NJmwHvauH/6e1mbbI40mi4hOgKT9qV4pY/3ky1HehZwX4tx9YtxRA57bOr8cuCVwB+6HpEkaRCl85nxtD4J7PM5bX2H1iZQlSlvz709CrbxDtobGE3H0Grp9ladSrZKxbuBpxVoYyLwpZzz32o1qD6yKVEtLfnaqMD1W+VcX+TvwLY51/frHqCSStDO4GC6kwBgnxbbOhO4KXVuD+Ih1U4Jqg2B3du4vqhHgftT544FZjZ5/UZEuaFOfi/b0d0HwwLgR6lzawA/pPkZWK8FXpE6txw4pb3QJElSHTcSK7mSntNiW/8m8ruk9Ym9mNvJS9YiZiZ303Wp4/2JkkvNmA78BJjRxv3Tud14YO822mvFN2uc26zJ658DfCjn/MmtBtQnhoiOseNT51cCrwF+0/WIJEmDKp3PQOt7BZ4O3JU6tw9wGrBmi21CVI/as43ri7qJ7OT2twLTmrx+f+BzbcaQ/l72KnD/Trgf+GXq3FTg5zRX4nSIyOe2T52/D/hV29H1vo8S+yomX+8ocP1Xcq5/TYHrT8+5/pAC10sacO0MQl0PLEyd+zCtPchXEgNCS1LnX02UMioyC2cckZScSpQ6eGEL8bRqNdmyQOvS3EN1F+AiYkPkdqT3/JtIzFZuZUZYqz5OdvbTnsTM6HorBoaAtwD/l/Ozz9LaHj/9aCrR+VnrlbeidlqDa/ptBp8kqfsWkS31fixRCrGVlWonkn12Pxu4CjioYFvPAk4i9nJ+UwuxtCNdInU8MfDZaEb1lsBfiE6eduTtyXcyMQGsW74PXJ46txHx/hqtpDwC+D3ZzsIzib3Hx7KTyf59XQ28iyjjVS93a/Rq599xkqTB80+yK8S+ROyXVzTPWwYcTXYfw1cBF1NsEtO6RLWpy4n9DLtZnnQZ2bKaGxODZfUm+Y8D3gCcw8he0OkJds26KHU8A/gpxatvteP9ZKuQ7UX0b9bro5xB5Ih5uflbyP59kyT1mFb3PgFYDJxB9UzY9YG/Ex1BdwMPkt1c92PEzPS0y4iHx2mp83sRM6bPAc4ebv8BosTSVGAWsY/NzsTy6UMot2TVl4mBzmRy9QLgWqLE1p+A24fPrwfsBrycGAit/CP/YSKhSm/m24zfEKv3konMi4HDgFuJPR7T5akeor0a6WkPEsldejb0c4lB5e8CvyYGb58gOpf2BV4//L9plwOf7mB8ve4Uis0UghiArudVwM9aC0eSNEB+QORTFROJlW/fBO4ZfqVnWJ8K/DGnrQeJPWgupHpgaJPh37+SyAcuJmafP0bkT2sR5XR2IAaeDqb5FWqj4XRiVnCyxOrWxEzvk4l85yZiktsc4BnE+z6ekQk9y4kqCq9v4f43ELlQcj/uHYnVmfcS38mDZDtgXkvk652wHDiGGNhNVoLYEriU+DtyxnCsjxOf1c7E4PKhOe3dT+T9Y93bcs5VZti3u2pyK+C2NtuQJA2O+UQ575ckzm1ATOJeQeQSDxKTWJIOJ/qR0v5BPOfSk7t3JXK7y4jBpcuIPqdHiMGkWUQuuDMx+WtPulsuPe2zRKn0ocS5g4l+y1OIiVD3DP+80nd1NLBT4vevJN5f0clvEAOBX6A6Vz58+LVo+N7phRlXEX1unXIfkZf9InV+H6I/9gzi786dxIDqxsDziX6rDXLa++7w70uS+sDq1Ctvv5laNhn+/XQb9V55gz9JryM6Moq0We/18Qb32y/nmlZLK1R8tkFMq4gHat7PlhIP2W+mzuclY7W8tcH906/b85thl5zfzevgqeXtxOypdr6/q8lPNmpZkLr+pALXJn0x1c4TLbbTih/R3meW9zq6i/FLksp1LdnnwMZNXrsmsXqwyDMmbwAkaW9iMKhTz7RfN7jfWjnXvLfxW6/rpUT+1mrMbwTenHO+2b2h9yYG6Ircc3pOOxNyfi+v3GctzyObaxV93UO29FQ9F6Wu/1uBa5OOyYml3YodjXQ6n0u+thzl2CVJvWcTss+DqwtcvzkxGavI82bTBm2+khi86tTzrdFEqrzn+TaN33pd6f6fonnNJsRgWPL8pQXuf2LBe86t0c6ROb+7Q4E43kQMFLfz/f2A5reHmpVz/fsKxJv081Q717fYTlH/R/Y9pPfkrOf3OdcX+XfLP3OuP6bA9ZIGXLvlaO4mynZ2ctbqacS+JOk9CFuxgmKDap3yEWJpfS1D5D8sFxCll/7a5v2/BXyAGIAs0zeIQanHW7z+bKKGexnfoSRJg2gJUW1gbgfbvISYGf6nDrS1iqg80G2/JDpu0hUxGllGlJ36Xpv3vwQ4iqguUabziUoQreb+lxNVQW7oWESSJKlZdxDP8fR+xu34OTGJKb3FTSvup5xV8R8k+tGKupZYAHF3m/f/OjG5Pl2do9u+Q+SbrfTBLSEqxR2H5UQlqW90Yq+Ky4jZv68gkoKbaL3WdsXlw20eTbFZUBCdMBcA7yFmyX+nzVhasYqY7fRasps051lBfHY7kF+WqxVfBLYgZoP/mSgPUYYziLJHXydKIjTjWqI87GHEgKkkSeqee4kVYs8hOkouo/3n8f1EiaZ9iFLxRQbZVhClqz5C5DbtrgJs1clEh1ozM8FXEXsV7kKUXe2E3xMz/o8n9uu7hXImgl1F7Af0HqLUfzNuJUpP7U7MsJckSeW4BngmkdN8hZicfjtRFSy9h2CzriXKgx5MTAZbUuDa24h+u4OJFXhzW4yhHauIShgvIcrGN3IfMSF/N6LUZrtWE9XDNiT6Ek8j8q27iZWe3fQ7onz+J4j32chConT+tsAnifciSeoTQ0TppaTVtL7SK2kKI/usJD1JQvnkcQAAIABJREFU8VnXGxKdVLsDs4l9TGYSpR4XEg+sm4hl4/+k2GybCWTLLi2kczNdJhJ1x/cnSh2sM3zPR4kk4nKi8yg9Myf9+a2iMwNl6e+7XtvjiZrwSe18NmsSn8MBxHe6AbF583yiBv01RIdhOzPFZlFdK34pzQ9KJk2muuZ7p/5/0YypwKQOt/kU5a8klSR1xwyye7csIJ737crLIxZRvDNpGjH4uDeRD8webnsJEet84GZiv5d/UKy89xCRDyQtplhHVSO7EWXgdyRiX5PIke4k8pnzyE4QW4Pq/fogcot2O1EmEblDWq22099hO5/NOGIA9CCizOW6RI7+GDEx7WYiz83bb7xZ06neJ30F8e+JovI+p079/6KWvP+/dMpoxy5JUqsmE5UCdiH679Yhcr8niGf4PURucAPtr7rrtCFiP8T9iElp6wyff5QYRP0HMXGu3UUR/WCIGEjej+jDm0PkZJU+vMuJvcXta5IkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZIkSZJUiqGyA5AkjRlrAOsC6wGTgGnEc2bW8M+nAxOG/7wYWAKsAJ4cPvcYsAx4CJgPrOpK1JIkScoziercbvrw+VlEjjcNmDh8bgmR360Enhg+9ziR280n8ruVXYlakiRJjaxJdZ43leo+vBnA+OE/LwKWAsuBhcBqIs9bykiet7pbgatzHByUJDVjArA5sDWwLbAZMAdYn0gm1mckgeiEVYwMEj4APDj8v7cCNw+/5nXwfpIkSYNkPLApI7nd5kRutwGR280B1u7g/VYz0nn0EJHHPQTcAvxn+HVPB+8nSZI0qCYSud22RK63KZHjzSHyvA2Iwb9OWclInlfJ8R5gJM+7iejXU49xcFCSlLY5sAewEyMdRlsyMjO8VzzBSGfSv4ErgUuJFYiSJEkKTwN2B3YlcrttgK2Iqg+95ClGcrsbGcnt5pcZlCRJUg/bisjzkn14m9N7fXgLiBzvZiLPuxy4bPi8SuLgoCQNtmlEArErsDfwHGImUT+bB1wMXEJ0Kl1OlDqQJEka66YCOxO53a7APkQHUT+bR+R0lfzuCqKMqSRJ0iCZDjyTkT68/YiyoP3sdkb6764kBgyXlRrRAHFwUJIGyziiw+gFwAuJFYLj617RvBWMlAJdRMz+rtQhh9hbcMXwnycT9c0nUL1/zRQisZndoZggVhj+FThv+HV3B9uWJEkq245EbvcCYjCwUzPFkyWiFlO9TzTEnjPLh/+8JpHfjWekTNVMIrer7GfTKYuAvwHnErndbR1sW5IkqVeMA55F9N+9AHg2nevDW051nvcUscVPZSXfE4zsFz2FqDgxkVhkALDW8PlOl6J/HPgLI31493WwbaU4OChJY99M4FAikTiY1jtn5hHL/yvlnh4YPvcgIwlFp0wk4qzsa7gesAlRBqtSDmt6zavru4FIMM4F5jKS7EiSJPWDaYx0Er0A2LDFduYTe8BUSjzNI/K7B4Z/Np/oJOqECUQ+ty4R77pEudNtGMnvWt2/+hYitzsHOB9nm0uSpP61NiN9eAfR+uT5+xjpv7uZ6Lu7n5FJ/Z0s2z6J6j68dYHNGMnxtmZkULGo6xjJ8y6ic7mpcHBQksaqSUSn0WuAw4jZ3M1aSJRrupR4CFeSiSc6HGO7NmQk0diFqLH+DIrNonoA+DnwE+I9S5Ik9aIJwIHAq4EjifKhzVrMyP591zEy2avX9mlej9gnZ2ui7P3uROmsIishHwHOIHK7vxNVLCRJknrZmsSA4GuAQ4g+vWY9SWyncynwL0b68J6sd1EJNmZkoPBZwG7A0ynWh3cf8DPgx8C1nQ5QkqR+NkSUkjqF6BhZ3eTrBuBU4I1EWapOlSgow1Si5vr/A35NzIBv9nO4EfgI/b8vjyRJGjt2A04iZns3m9PcDPwAeCsxgWpC16PunMnEnjrvBn4B3EPzn8PtwKeIyWSSJEm9ZBywP/B/xIStZnKbVcREr+8CxxMT5Md1Oe5Omg48F/gQ8FuK5bv/Aj5IVBmTJGlgrQEcSzwYm3mALgTOAk5gMB6i2wMfAP4MLKW5ZOvPwOG4wl6SJHXfJOBlwD9oLrd7ishdPkCsvBvrtiDy2LOAJTT3GV1MfKb9PAlOkiT1vzWJPrwbaC6HeZKRPryNS4i325J9eMto/PmsJD6fA8oIVpKksmwAfJqoE97oYXkL8FlgX/p79ni7ZgJHAd8nNjlu9LldR8zGKlKWVZIkqRWziSoG99M4R7kL+BLwPIqVnhprpgJHEJUzHqa5VZVvpVhZVkmSpHY9DfgC8CiN85UbieoHezLYE5vWAl4O/JDY6qjR53YlcAyDnRtLksa4LYlSUY1WwT0EnAzsUU6YPW9N4KVE6YJGn+WDRGddqxsoS5Ik1bIxMbi1iPr5yGPA94gS6v1cQmq0TCQqP/ycxp/lo8BngFmlRCpJkgbFtsQ+ecupn5s8AHyN2I9PWZOBVxKrBButKLyf2G5oSimRSpI0CuYA36D+Q3AZsR/LYUQHiZqzNvAmYgPnRsna2/CzlSRJ7Vsb+CL1B7JWAr8DXkKUkldzZgDHAXOpn9s9Aryf6HCSJEnqlI2IvQHrDQouBX4CvIDBXiFY1Gyib+5K6ud59wJvZLArqEmS+txUot72Amo/8BYAJzEYewiOtl2JkgX1Erg7iXrvJm+SJKmoScCJ1C8r9STwHQZjD8HRthPxWS6mfufRCdh5JEmS2jON6MOrVwbzcaIPbxD2EBxtzfTh3UTsPT1UUoySJLXkWKI8aL0H3Ftw35TRsCmxl0+9vQmvAHYuK0BJktR3XgzcQ+3c4g7gPcQ+yeqsDYj9e+rt1309sHdZAUqSpL41RKxSe4T6ecYbsWLBaNiSKMtab1D278AzygpQkqRmbQqcS+0H2q3A0bjfTDfMIDqSFpL/XSwHPkvsYShJkpRnfeBMaud29wCvx5Vr3TAF+CCxh2OtUq4n417TkiSpOVsC51N/Yv9RuHKtG9YCPk/tsv1LgY8RlTwkSeopQ0RJo1ozXeYT5Qncc6b7ZhMJxhJqD9g+t7ToJElSr3oZtVerPUrkds4g7761qd95dB/wotKikyRJvW4c0Yf3JPm5xENEKXknf3XfRkRZ+VrlRq8Hdi8tOkmSUjYCLiT/obWImNniDObybQX8gtozzb8CTCwtOkmS1CtmU7sSxFLgC8TsZpXracBpwCryv6vvYYUISZJUbTPgUvJzhyeBDxHVClSubYHfkv89rSAqgY0vLTpJkoB9gXnkP6wuBLYpLzTV8ELgLvK/s4uIfW0kSdJg2hm4nfw84Spgl/JCUw37EGW/8r6zq4HNywtNkiT1kP2BB8nPGc4jtgpSbzmc2vt+zwXmlBaZJGmgnQAsI/twepwoP+C+gr1rBnASsWIwrxTVXuWFJkmSSvIa4CnyK0F8AGcn97LJRKnRFWS/v4eBg8oLTZIklWyIyOXy8oTHiP499xXsXbOIUqN51SLuAXYrLzRJ0qCZDPyU/FkrZwMblheaCtoPuIP8kmFvKjEuSZLUPROAb5Of210AbFFeaCroWcCN5Jef+kCJcUmSpHJMBX5Nfp73a1x51k8OBO4l+z0uBo4tMS5J0oCYAvyJ7INoFTFb2dWC/WcdonxEXqL4PyXGJUmSRt8k4Ffk5wHfwf2I+9E04Azyv9P/xZUBkiQNimnA+dSeNGRO0H9mA38hv1/2XSXGJUka46YCfyX7AFoAHFliXGpfpcREXpnRz5cYlyRJGj2TgN+SX0b0tSXGpfYNEWX+l5P9fr+NE/okSRrrZgH/IL/c+IElxqX2TSD66vImgn2kxLgkSWPUTODvZB86/wK2LDEuddaLgSdxgFCSpLFuCvBHss/824AdSoxLnXUA8Aj5q0IdIJQkaWxaB7iS7PP/SmCTEuNSZx1NTOpLf88fLTMoSdLYMhm4lOzD5gpg7RLj0ujYm1gNmv6+P1NmUJIkqWMmAn8m+6y/EfeOHoueCTxE9vs+pcygJEnSqJgOXEP2uX8JMfFfY8vzgYVkv+//LjMoSdLYMAT8lOxD5nIcGBzLdiVKTaS/99eVGZQkSeqIb5J9xv8b2KDMoDSqtgXuI/u9v6PMoCRJUkcNAWeSfd5fSAwaamzah+wk/1XAy8sMajS5WaYkdceHgM+mzl0MHAo80f1wWnYCMeCV507gcx26zweALWr87Gr6a4b2TsSqgtmJc0uA/YDLSolIkiS16zjgtNS5a4i9Zx7uejStexWRk+R5GPhwh+7zNmDHGj/7D/DlDt2nG7Ym9g/fOHFuJXA4cG4pEUmSpE76NNkcaC7xrF/Y9Wha93Zql7m/GfhKh+7zUarzoqR/ks2Ze9muwJ+oXsixkKgOdl0pEUmS+tphRIdBcubJdcC0MoNq0S/I36h3NfEeayUDRcwBltW5z686cI9uezawmOr3cQ+uLJAkqR/tAyyl+rl+O9UTgfpF3urH5KsT+yZOJSbD1brH3A7co9u2Ax6n+n08jHuIS5LU715GrBZLPuMvI7YK6jdnUTv/WkpnctfNyPZ5Jl8/7MA9um1fxk6uL0kq0aZkOw7mA5uXGVQb6g0OriZW/LXr/Q3u0Y+DgwCvJvteLgDGlRmUJEkqZF3gAaqf50/SmUG0MjQaHPxiB+7xugb3mNuBe5ThEGAF1e/lGmBSmUFJkqSWbU1237n7gY3KDKoN9QYHVxMrC9v18Qb36MfBQYjKaen3ch5W4pQkNWkI+CPVD5LlwHPLDKpNjQYHb+7APa5rcI9+HRwE+ALZ9/OuUiOSJElFnEH1c3wVMcO8XzUaHHwAmNDmPS5ocI+5bbZfpv9H9v18stSIJElSK8YBF5FdXbdvmUG1qdHg4OVttj8E3NrgHv06OAjwDbLv542lRiRJ6htvIPsQObHUiNqXNziYLrewexvtP7tB2/0+ODiemGmUfD9PEStMJUlSb3sxY28gKG9wMJ1/HdpG+1uk2svL7ea20X4v+BnV72cZ8IxSI5IkSUWdSDZHeUOpEbUvb3AwnYu1k7Ps36Dtfh8cnEh2ktsCYP0yg5Ik9b7pwDyqHyB/of+Xn+cNDl6YOv5WG+1/K9VW3kzzfh4chNhn8BGq39PPS41IkiQ1Mgn4D9XP7yuJToN+ljc4mM7tzmij/U/SOLeb20b7vWAmsZd08j39tdSIJElSEWsTewcnn+W/LzWizsgbHEzneV9qo/0f0DjP6+fBQYg9FdN7Z59aZkCSpN73Gcbm6rC8wcHjU8eP09pGzZPIJmN5e9T0++AgZD+zVbS34lKSJI2u91D97F7O2Fgdljc4+NrU8VJgdgttDwG30zi3m9vOG+gRLyL7vg4pNSJJktSsr5Ht1xoLq8PyBgfT/VEP0Npkt2nEvtuN8rx+HxwEeAfV72klsGOpEUmSetZM4DGqHxwfKzWizskbHNyMbMfPK1po+xWpNu4gSlGNxcHBccBlVL+vs0qNSJIk1bImcB/Vz+2TSo2oc/IGB3cFrkqde1sLbT8/1cYjwLo595vbzhvoIenS8f8sNxxJktSEOcSE/uQz/H2lRtQ5eYOD65OtdHZYC22nBxlvICbOjcXBwfHAdVS/rzFRAWxc2QFI0hj0dmBW4vgh2lum3+tWAz9OnXttC+0clzqulCcYi1YBH0ydOxTYoYRYJElSfccBGyaOF9L/ew028oPUcSdyu58QqxDHqg9SnbvuDuxXUiySJKk5JwJTEsf3ACeXFEs3rAB+mjrXiTzv9FaC6RMrgQ+nzr0U2KqEWCRJPWwccCfVs0k+UGZAHZa3cnBTYHOqNx5eCWxcoN2NiAQlWWZzy+F2x+LKwYqLqX5vYzkBlSSpX6VX0X2h3HA6qtbKwXWAJanzRSYxzSA7C3+X4fNjdeUgZGfopzvfJElS75gA3E/1s/vtpUbUWXkrB2eTXeFXtIR8ug9wObBBTrtjZeVgxZVUv7fPlxtO+1w5KEmd9Xyq9xZcBJxSUizddAdwUeJ4HPDqAtcfSyzTr5gL3NZ+WD3vq6nj1xClyyRJUm/YBdg5cbwS+HpJsXTTI8DZqXPHFLj+FVTPwr+eGGQd676cOn4JsHYZgUiSpIYOIwa1KhYA3y8plm66nhjoqpgEvLLA9ccTe0tXnEeUKh3r0tsKHEcMMPctBwclqbOOSh3/kkguBkG6/NTxBa49tkFbY9XvqE6gZhEDzJIkqTekc7vziP0HB0E6HzuW5jtAjksdn9Z2NP3hAuA/ieM1aG0fH0mSNPrSed5PiUn+g6DVEvLjGNw+vDOBxxLHc4B9SoqlIxwclKTOGSL7j/8xsUFtk84g9uCp2JrYa6WRvYBtE8cLGVulQ+tZAfw6de7wMgKRJEm50s/lQcrtzgEeSBzPAQ5q4rqtgT0Tx3l724xVq4ky/EnmdpIk9Z4JwCGpc4OU56X3gn4WsGMT1z0f2CRx/ChRvnQQLCYm+Sf1dZ7n4KAkdc7WxN55FQuB80uKpQx5g3rNzDw6LnWcHmQc6/6QOt6/jCAkSVJGZU+WihXEgNmgyBvUaza3S5aaSg8yjnXpDrL9qf48JElS+XakuvT3I8AlJcVShrxBvWZKyB+XOk4PMo51Y6oPz8FBSeqc9Cq5SxisByRkSwm8Cphc5/cnAy9r0MZYdyHR+VaxNbBOSbFIkqQRu1M9qHMN0ZEySNLlQI+kfp4yjthDOen0TgbUB66keluB2cAWJcUiSZLypfvwLiD2lh4k6f63Y4CJdX5/BpEL1mtjrPsbUSmiYkdgakmxtM3BQUnqnGemji8vJYpyzQVuTxzPBI6o8/svIfbZq7gDuKjzYfW0RcRm0BVDNFfKQZIkja50bndZKVGU63pisKtiEvCKOr9/EPC0xPEjDNZqS4BVwBWpczuXEYgkSarJPrzYS3te4ngOcHCd338lMCVxfAPVeeIgeBS4NXE8Adi+pFja5uCgJHVOekbwDaVEUa7VwI9S5+qVnzoudXwa1TNwBsW/U8eblxKFJElKSud26ef1oEjPCC+S2/2YwaukAeZ2kiT1OvvwoorVT1LniuR53+9oNP1jzOR5Dg5KUudskjq+q5Qoync6MWO64mBg45zf2xh4buJ4NdGBNIjuSB1vVkYQkiSpirldSO8lsxuwQ87v5VWMGLRSUxXmdpIk9TbzvJAuIX8EURI9bWtgj8Rx3t7Ug2LM5HkODkpS58xIHc8vJYry3Unso1cxDnh1zu+9DhifOD6f7AN2UDySOk7/XZIkSd1nbhceBf6QOpfeVxCye03/C7h6tILqceZ2kiT1NvO88G+qS6pOIsqHpr2O6r24zwEeGMW4etmYyfMcHJSkzpmSOl5UShS9IT1L/PjU8RBwbINrBsmTqeO+3cxYkqQxxNxuxOmp49cSe6wkHZc6HtRSUwALU8fmdpIk9RbzvBGNSsjnTfq3D29E3+Z5Dg5KUuesSh0P8n9jz6T6Ybk1sHvieF9gq8TxQuA3XYirV6U711aWEoUkSUoytxtxHjAvcTwHOChx/HSi3GjFCuBnXYirV41PHZvbSZLUW8zzRvwMWJI4fhawY+L4YOBpieNHgLO7EFevGjN9eIP8l16SOi09y2haKVH0hqeAX6bOJWceHZf62c/JzrAeJOlZRoM8Y02SpF5hbjdiBbH3YFIyt3td6md/AB4c1Yh6m7mdJEm9zTxvxKPA71Pnjkn8+bjUz35M9X7Ug2bM5HkODkpS5zyWOp5TShS9I11ioLIPzVTgpQ1+d9Csnzp+tJQoJElSkrldtdNTx0cC6xCzp9OlptK/O2g2SB2b20mS1FvSeV66X2bQpPvljgEmAjOBwxv87qAZM314Dg5KUufckTrevJQoeseFwG2J45nAEcTA4PTE+VuAS7oYVy/aInV8ZxlBSJKkKuZ21W4ALk8cTwJeAbyA6sGwh4BzuhhXL0r/XbmzjCAkSVJN6TxvszKC6CF/BO5NHM8hyom+mpjoX/Ev4OouxtWLxkwfnoODktQ5t6WOd8z9rcGxGvhh6txryZYj+MHw7w6yZ6aO03+XJElS95nbZaVniufldj8Blnclmt5lbidJUm8zz6u2kvwS8selzn2/K9H0tvTfFfM8SRIvJga5Kq+/lxvOqPgF1e9xNbBpnd/flEgwKr+7gtj0uXK8EtikzvWb59zvV229g96zLtWfyTKqZ2VJkqRyPIfqHOSmcsMZFd8km2vtWuf31waWpH5/eeo4PTCWNCPnfnPbeQM9aE1iH57kexz0UmWSJPWa11D9rP5zueGMirPI5l2z6/z+dtTP8ZYB69W5/hk590svGuh3m1L9/p4iSuz3JVcOSlLnXJo6fhawVhmB9JC7qO7wGQ8MJY7/CtzdzYB60AFUfybXAYtLikWSJI24kpjYVLEN9SdFDYJHgd+nziU7RK4Gru1eOD1pP6LkasWdwAPlhCJJkmpI9+HtDUwpI5AeciPVn0t60Otsonz8IDswdXwF1f9e6CsODkpS59wPXJ84ngi8sKRYesnpLf5sUKQ3dv5TKVFIkqS0p8jui3xEGYH0mNNb/NmgOCx1bG4nSVLvuYXqfQcnkx34GUSnt/izQTGm+vAcHJSkzkrPpD6mlCh6yy+BBTnnnwB+2+VYes1M4EWpc+m/Q5IkqTxnpY7N7eCPxKS4tGXAz7ocS6+ZBLwidc7cTpKk3mQfXtbPyK9m9RBwTpdj6TWzgYNT5/o6z3NwUJI664zU8UHAZiXE0UsWA6cCt6de3wUWlRhXLziG6rIVdwKXlROKJEnKcSaxN3DFs4GdSoqlV6wEvk02t/sRML/EuHrBUcR+0hWPEmX0JUlS70n34R2B+wQvIFYIpvO8U4g9CAfZ64E1Esc3Av8qKRZJUo+6nOrNaU8uN5yO+gXZzYVHc++dzXPu96tRvF83jSfKWCTf20dKjUiSJOU5j+rn9U/LDaejvkk219p1FO83I+d+c0fxft12BdXv7avlhiNJkhq4gepn9+fLDaejziKbd80exfs9I+d+PxzF+3XTGsC9VL+3d5caUQe4clCSOu+bqePXAxuVEYh62jHAVonjpcQKS0mS1Fu+lTp+ObBtGYGopx1O9cDqKmKWvSRJ6l3pPO+tjO4AmvrTG6ju213IGBj4dHBQkjrvJ0R5yIrJwGfKCUU9agrwidS504B5JcQiSZLqOwu4LnE8HvhSSbGoN00APpc69yvg5hJikSRJzTuV6r6Y6WT7azTYZgAfTZ37DvBICbF0lIODktR5y8l2DhwD7FlCLOpNHwE2SRwvBb5QUiySJKm+1cAnU+cOAw4pIRb1phOB7RPHK4FPlxSLJElq3hKyk75OwD2mNeJTwJzE8VPAl0uKRZLUB8YD11Jdi/omYhVhP3PPwfbtDCyj+j19ttSIJElSMy6g+vl9H7BWqRG1zz0H27c1sIjq9/SdUiOSJElFTCJW+yef5dcOn+9n7jnYvr2AFVS/pw+XGpEkqS88l9hrJPkA+W6pEbXPwcH2zCQGiZPv5x6ibIUkSeptOxMVIpLP8V8CQ2UG1SYHB9szGbic6vfzCLBemUFJkqTCDiWbo3y11Ija5+Bge9YBbqf6/dxG/y/8+P9NKDsASRrD/gacArwlce6NxOyjb5YSUfseIh6MSStG8X7Lc+734CjebzSNA34MbJM6fwLwZPfDkSRJBV0NfJ4oD15xFLEHSbrsaL94mGyutXQU77cq5379vOfy94Bnpc69k8iZJUlS/zibGMg6NnHuXcANwP+VElH7HiCbd60cxfsty7nf/FG832iaAJxBLFqoWAUcDywuJSJJUt+ZBvyH6lkmy4hVhRosXyA7g6rfV5JKkjRoJgFXUv08XwW8uMygVIoPMLYqXEiSNOjWAu6k+tm+BNizxJhUjm8w9laSSpJKsB2wgOoHynxgqzKDUlcdSzap+DuwRplBSZKklmxCVDJIPtefBHYqMyh11RHEzPvk34F/Yal4SZL63U7AQqqf8fcT+Z8Gw1vI9uGdD0wsMyhJUv86mOwGtvOA7csMSl3xSrL7E90PbFRmUJIkqS17ETPJk8/3R4HdygxKXXEIUU4qvc+gE/8kSRobXkJUhkg+6+8EtiwxJnXHG8lOALsDWLfMoCRJ/e+DZGeePADsUGZQGlWvJ5tULMaOQ0mSxoLjyOZ2jwF7lBiTRtfhZAeFlwPPKzMoSZLUcZ8im+fdDfxXmUFpVL2ZbB/ek9hvK0nqgCHgVLLJxUNYhmoseivZmWbLgZeWGZQkSeqoL5LN7RYA+5QZlEbF0WSrQawC3lBmUJIkaVSMA35KNs+7n9g+SGPL+8h+10uBQ8sMSpI0tgwBJ5N94DwBHFViXOqc8cDnyQ4MLiVKU0iSpLHlE2Rzu8XA8WUGpY4ZAj5AdouAlfgdS5I0lo0HTiOb5z1CbB+k/jeB6MNLf8dLiD2mJUnqqCHgq2QfPKuIB9L48kJTm9YB/kT+bKMjS4xLkiSNrg+Qff6vBr4DTCoxLrVnOvBLst/rCuC1JcYlSZK6Ywj4BrX78MaVF5ratC5wPtnv9ingwBLjkiQNgLz65auBc4G1S4xLrdmZ2KQ4L6k4qMS4JElSd7yf/NzuYmCDEuNSa7YBbiD7fS4HXl1iXJIkqbuGgK+Rn+edBcwqLzS1aG+iRGz6+1yIe0lLkrrkHcAysg+jO3GWSr+YQKwWWEz2e7wH2K280CRJUpcdCywimxPMA15cYlxq3jhi7+gnyd8r3A4jSZIGzxAxESxdZnw1cAuwX3mhqYCJwP+Q3xd7O7BTeaFJkgZRrdkqq4EziFKV6k07AJeR/91dAKxfXmiSJKkkOwG3UXt2+UblhaYGtiS/vNRq4Epgs9IikyTZL3ZPAAAgAElEQVRJveA5wAPklxn9IVYC62U7EfmcVdwkST2lVp3r1UTScWx5oSnHRGK14FJq7y80sbToJElS2dYGziM/T3gMOIGYga7eMAE4kSgllfed/RCYXFp0kiSpl2wM/JP8nOF+rBbRayYT+0Pmrfqs7B05vrToJEkCJgFfIv9htZroYHJ5e7mGgFcQJSPyvqP5wEtLi06SJPWSfYEnyM8ZVgMXAnuWFp0qDgf+Rf539DjwuvJCkyRJPepl1J4wvhr4PbB9adEJolT8McAd5H9H84g8UJKknrETcBX5D65VRKnRrUqLbnAdAFxB7cTvDGIFqCRJ0glUdxgtp3YO8Wdgx3LCHGh7AHOp/b2cAzytrOAkSVJPGiIqSa1kJGfI279u9fDvnAFsUUqkg+0AavetVvrwZpcWnSRJdUwCPkbtWUhLgZOBDcsKcIDsAfyF2gnFfcARpUUnSZJ6yZrA96nOFa4iJna9F3iK/HxiBXAq7mnXDTsRez/Wyu3mA68qLTpJktSr1iFbNv58YjLRZ6k9GWwx8GVgTvdDHjj7EtU5auV5dwIHlxWcJElFPAP4G7UfakuBHwG7lhXgGDWeKBFxCbU/+2XAN4FZJcUoSZJ6y1bAtVTnC+m96rYEzqZ2frECOBPYu2tRD4ZxwGHAX6n92a8ETsdKEJIkKWtn4HaqK3ul96rbmfr9SEuISWRWjOisicDRwGXU7z/9KjC9pBglSWrZAcCV1H7IrSZKXh4LTCgpxrFgOnAiteuRVxLAM4D/KilGSZLUew4FHqW68+fEOr+/J3AB9XO7K4ncbuKoRT32rUl8hjdQ/7P+M+7tLUmS8h0LLGIkb3gCeEmd3z8AuIb6ucfFxKT08TXaUGMziHz7LupP/jqDmKAnSVLfGge8muqZSrWWyH8a2K6UKPvPeOBAYqb4k9T/bP+EqzQlSdKIvH1n7iHKkjdz7YuBf1M//7gP+F8cvGrWEPAc4DvAY9T/bC8hyk9JkiSlrQGcRHXucCPN9beNB14H3E39XORW4BPA1h2OfayaALwQ+DG1y/VXXmfjKk1J0hgzCXgTcBP1H4KVGefvBjYoJdLetivwFeB+6n+Gq4BzgeeVE6YkSepRaxM5QjJvmEvx/WTGA68lW5I073U98CFgk7ajH3u2Bz5H/dnjye/psFKilCRJ/WBj4J9U5w8/BaYWbGdN4B3EIGCj/ORS4J3Aeu2HP+bsDnwdeJD6n+FK4PfAPuWEKUlSd4wjSlj9mcYJxgpi78L/x+DOmlmDKO3wZZobWF1EzDh3BaYkSUrbCbiN6slEJ9F+effnA2dRvRKxVsfHJcCHiQlPQ23etx9NAPYjBgSvo3Fut5TYA3LnMoKVJEl9Yz/gAUZyiOVEpYh2jAOOJCYoNcpZlgN/Ad5LTH4aRGsCBwNfo7mB1SeBb+AWQJKkAbQDcCrVNdDrve4FvgccBcwqId5u2Rx4CzFraCHNfzb/DcwuIV5JktT7XkN1GaMngJd2+B5bA98cbruZ/OUB4AfAK4F1OhxLL9kYeAPwS+BxmvtsHgQ+hZU0JElSfUPEHnbLGckjHqLzlaR2ISYsLaa5XOYu4BRicHF6h2PpJVsBbyfKgTYqGVp53UkshFir++FKktRbJhMbGp9FdTLT6HUbkZicSCy9n9TtwDtgGhH7icR7abQ3Y/K1YPiaw2l/xr8kSRqb8vaduQl4+ijec00iPzmDWPnWbG5z//A1ldxuzVGMcbRMJFZFVnK7G2j+/S8m8uGXDbcjSZJUz3TgTKrziYsY3clFM4FjKdaHt4LIicZSH94ZwDyaz/MeI97/AcSKTLVoEEuPSNKg2ICYPf4aYlZSEU8R+xVeD9w8/PoPMVtpVQdjbMUaxGz6rYFthl87Ex1z4wu0sww4D/gJkYgt7myYkiRpDNmIWK22R+Lcb4m9Ap/oUgyzgVcArx6Oo8i/55cAVwH/IvK6m4jc7k6iRGmZJgFbAtsykuPtTFTGKDJpayVRgusnwG+IqhGSJEmNbAP8muoJX98l9gpc1qUYNgaOJvrwim4HtBC4ghg0vImRPry7icG0Mq1JfL6VHG9boo9yW4oN7C0FzgF+NPy/Szsb5mBycFCSBsNmwAuBFxB72RTdQLliCZFg3ArcR5RXmDf8v/OJWeoPDf9eK2YQg5rrAXOA9YF1h/93UyJ5eBqtzwy6lxgQPI/oPFrQYjuSJGlwPAf4BZGPQMzY/gjwRcrrcNmIkdzuAGLmeSuWEXndf4g8aT5RovSB4T/PI8pytjqJahojud16wIZEbjcH2IToJNqM1is3PAj8ETiX2If7kRbbkSRJg+lIojT7jOHjJcT2NKeXFRAxaeqFw6/9gSkttrOYyPFuIfrrkv12yf68VgfaZlHdb1f58wZEfrc1MejZah/eXYz04f2V2FdQHeTgoCQNnjWIpfsvIDb33Z7RWYb/FNHhtJTYC3EF8SAfYmSfw5nD957G6JR8WgRcykgycd0o3EOSJI1NQ8A7gS8xkqfMJ2Z1/7WsoHJMAPZiJLfbkdEpk76Y6DBbRuR5qxiZaFXZ52UGUclhCpFzdtpS4HJGcrurKH9GvCRJ6j8TgE8T+9VVxkhuBV5CVFroFWsSE9VeCBwIbMfo9uEtIXK+Sh/eOEYmoc0iPqvR6sN7CvgHkeOdC/x7FO6hBAcHJUnTgJ2IPV12BfYlZvj0o9uBS4iSqFcSnUeWGpAkSUVNB04l9qyruAI4iijR1MumEmU5d028RnNfxNE0D7iYkfzuClqvUCFJkgSxuu1nRGWtirOBY4j97HrZdOCZRH63NzFwOKfUiFqX7sO7jO6VcRUODkqS8m1C1ABP7uu3DbHXTdlWEZ1y/2Fkz5ybiETCMqGSJKldWxP7zmyfONftfWc6bUNG9nep7PmyHVHqsxfcw8j+OJW9cq4AHi0zKEmSNOY8C/gV0e8FUYHgi8B/E/1N/WgzRvrwKvv6bQOsXWJMFauIPa4rfXg3AzcSfXiWCS2Zg4OSpCLWJhKMLYjOpPWJGUrrpv48qcX2nyL2jqnsc/PA8PHRwH8N/842RFIhSZLUaUcAP2SkfNIS4G3A90uLaHTNIjqRtmRkz+cNGNkXsLJ3TKslQhczktsl97ep7Gd4G5HXPdXyO5AkSWrOCcDJjPRZPQK8mti/eCyaTfShbc7IXoBzEn9eb/jPrZYIfYqRfQsfYqQP7zj+P/buOlyO8vz/+PucnLgHIrgTvLhDsFK0OG3RQEvaUqHU+NXTUkG+XygV2tAWgksF9yKlUNwpDgkeEiSuJ+f8/rh3vqPrs/vszH5e1zVXsnJ279md3b3nkfuBNQr3WQVb51BERETayCBs/ZlxWGfi+ljZgy0Ll9cGVijcp1wi8ntsNFcvtp6OiIiISJr6AGdgo5u9nOMNbHS5mIFY3jYWy+PWxS9b6uV2Kxbu04j1BkVERERqMQArF98b2J7AOs3EDMZyuJUIt+FtgZ/njaKyNryp+K/zLo0JV0RERNrFV/ETi1MdxyIiIiL5siJwJ+EGo5uxxg8RERERya7VgUcJ53mXYAPapTFOw3+tJzmORURERDJuT/zE4gLHsYiIiEh+bIWtg+LlGT3YDMJOhzGJiIiISP32w9Yv9vK8xcApTiNqD5/Gf83PdRyLiIiIZNzK+InF/Y5jERERkXyYBCzBzzHmAAc5jUhERERE6tWBzV5bjp/nvQVs7zKoNrIe/uue1/UcRUREpIk+xhKLD10HIiIiIpk2APgz4fJST2JrqYiIiIhIdg0DriWc592LrZkszdEHWIS99m86jkVERERy4CH8xG6M41hEREQkm1YHHiHcYHQpWndGREREJOs2B14jXC7+PKDLZVBt6ln892Co41hEREQk4y7ET/AmOI5FREREsmdfrAKB1p0RERERyZdjgAX4ed5c4HCnEbW3q/Hfi60dxyIiIiIZ9x38xOJLjmMRERGR7Ehad+ZtYAeXQYmIiIhI3bqAMwhXhXgR2MhlUMJk/PfjWLehiIiISNYdgJ9YnOc4FhEREcmGYcA/CDcY/QsY5zIoEREREanbKsB/COd51wHDXQYlAHwG/z35peNYREREJOPWwU8s7nAci4iIiLS+TwCvEl93pq/LoERERESkbrsC7+HnecuwShEdLoOS/7MZ/ntzreNYREREJOM6gYX4pcBEREREijma8Loz84AjnEYkIiIiIvXqwNaMXoqf580E9nQZlMQMALqx9+clx7GIiIhIDjyNn/ypTISIiIhEJa078xKwscugRERERKRuQ4BrCOd5jwJruAxKivIqeCwD+juORURERDLuKvwEcFvHsYiIiEhrWRl4gHCD0fVoQJGIiIhI1q0PPEc4z5sC9HMZlJR0I/57pYF6IiIiUpef4CcWxzuORURERFrHLsC7+HlCN1p3RkRERCQPPg3Mxs/zFgEnOo1IKnEm/nt2uONYREREJOOOwE8sznAci4iIiLSGSYTXnZkF7OU0IhERERGpVx+s7acHP897A9jaZVBSsYn479uP3IYiIiIiWbcJ4TJhIiIi0r6GEC453gs8htadEREREcm6FYE7Ced5NwMjXQYlVdkO/727wnEsIiIiknH9sIWMe4FXHMciIiIi7qwHPIvWnRERERHJm62A6fg5Xg82g7DTYUxSvWH4sz6fchyLiIiI5MDL+GsJDXAci4iIiDTfgcDHhNed+bzTiEREREQkDZOAJfh53hzgYKcRST3ewc/X+ziORURERDLuevwkcVPHsYiIiEjz9AEmA8vxc4E3gW0cxiQiIiIi9RsA/IlwVYgngbVdBiV1+yf++6n3UkREROpyBn5icaTjWERERKQ5VgTuINxgdAswymVQIiIiIlK31YBHCOd5lwGDXAYlqfgt/nu6v+NYREREJOOOx08sfuI4FhEREWm8LYFpaN0ZERERkbzZF/gQP89bBpzmNCJJ08n47+23HcciIiIiGbcNfmJxleNYREREpLGOAxYSXnfmEKcRiYiIiEi9OrBOwGC5+LeBHVwGJanbDf/9/YvbUERERCTrhmAzBnqBpx3HIiIiIo3RH7iAcHmpp4B1XAYlIiIiInUbBvyDcJ73L2Ccy6CkIcbiv8f/cRyLiIiI5MBbWGKxGOjjOBYRERFJ12rAw4QbjC4HBrsMSkRERETqthnwKuE8bwrQ12VQ0lAfYO/zx64DERERkey7Az+JXNdxLCIiIpKe3YH30bozIiIiInlzFDAfP8+bBxzpNCJphgfw3/OVHMciIiIiGXcefmJxoONYREREpH7eujPd+L/x7wA7ugxKREREROrWBZxBeLbgS8AmLoOSpvkz/vu+h+NYREREJOO+hJ9YfNdxLCIiIlKfYcDfCTcY3YdGFouIiIhk3cqEZ471AtcDw10GJU31Lfz3/iuOYxEREZGM2xU/sbjIcSwiIiJSuw2A59G6MyIiIiJ5swvwLn6O1w1MxipGSPvYF/8Y+J3jWERERCTjRuMnFg85jkVERERq8zni6858xmlEIiIiIpKGScBS/DxvFvBJpxGJK2viHwd3uQ1FRERE8mAWlljMQaPOREREsiRp3ZmX0bozIiIiIlk3BLiKcJ73ONZBJO2pE39A4LuOYxEREZEc+Dd+ormK41hERESkMqOxEcPBBqMbgBEugxIRERGRuq0HPEs4z7sEGOgyKGkJT+AfEyMdxyIiIiIZdwF+YrGX41hERESkvJ1JXnem02FMIiIiIlK/A4CP8fO8RcAXnEYkreRy/GNjB8exiIiISMadip9YfM1xLCIiIlLaJGAJ/m/3B8DeTiMSERERkXr1wQZ7LcfP894EtnUYk7SeH+IfHyc6jkVEREQybh/8xOJ8x7GIiIhIsoHARcTXnVnLZVAiIiIiUrcVgNsJ53l3A2NcBiUt6TD8Y+Rsx7GIiIhIxq2On1jc4zgWERERiVsXeAatOyMiIiKSN1sCr+PneD3AGdhMQpGoDfGPlZscxyIiIiIZ1wHMwxKLGY5jERERkbD9gY/wGwEWAyc5jUhERERE0nAcsBA/z5sDHOI0Iml1fYGl2PHymuNYREREJAcew09GV3Aci4iIiNjgndOIrzuzncugRERERKRu/YELCFeFeAGbFSZSzgvYMbMcGOQ4FhEREcm4S/ET0p0cxyIiItLuVgBuI9xgdA9ad0ZEREQk61YFHiKc510BDHYZlGTKP/CPnc0dxyIiIiIZ9338xOILjmMRERFpZ1ugdWdERERE8mg3bDkXL89bhlWKEKnGL/CPoc85jkVEREQy7hD8xOJ/HcciIiLSro4FFuD/Js8FDnMakYiIiIjUyysX342f570P7O4yKMmsY/CPo585jkVEREQybgP8xOIWx7GIiIi0m/7AeWjdGREREZG8GQr8jXCe929gJZdBSaZthX8s/dVxLCIiIpJxXcBiLLGY5jgWERGRdrIq8CDhBqMr0bozIiIiIlm3AfA84TxvCtDXZVCSeYOA5djx9JzjWERERCQH/oslFstRg6SIiEgzTEDrzoiIiIjk0WeB+fh53kJgosuAJFemY8fVEmzAv4iIiEjNgmUutnQci4iISJ51AKdgnYHeb+9MYA+XQYmIiIhI3bqAMwjPFnwZ2NRlUJI7t+IfX+MdxyIiIiIZdzp+YnG041hERETyaii2Nkiwweh+YGWXQYmIiIhI3UYDdxHO824ERrgMSnLpHPxj7GDHsYiIiEjGHYWfWPzccSwiIiJ5NB6/jHdw3Zl+LoMSERERkbrtBLyDn+P1YDMIO10GJbl1Ev6x9j3HsYiIiEjGbYGfWPzdcSwiIiJ5cxAwG/+3dhFad0ZEREQkDyZha795ed4HwN5OI5K82xn/eLvEcSxtT4s+Sh70x6a/rwSMKWwrAcMKWx9gIDAA6AsMCfztUmAB9oU0u3DdvMJ1s4D3sHVUZgIzgDmN3RURqcFL2Mi2TmBDx7GIiEj9+mG53VhgXOH/44CRWGnLLiyvG4jlecMCf9uN5XIAHxf+nY+f272P5XSzCv9+jBTThc3I/y621iDAq8BhwDOughIREZFMG0C4DW80VqJ8KDAcO68fhLX1FWvD68Fvn5tLvA1vFvAufk4ocQOA84ETAtc9ieV505xEJO3ihcD/1YbnWEf5u4i0hOHA+lhJo/GB/69Bc+tfL8aSjVewDokXscV5XwbewjoZRaT5XgfWApYBgwv/iohI6xpCOLfz8rs1gBWaGMdSrJPQy+deKmwvA29gjU/taDRwJbBn4LqbgWNRh6qIiIiUNpJ4G976wJqEB3U12mKskzCa43lteO1qXazq0maB6y4FvohViBBptPexwQHzse8EtaeLyP9ZAdgP+ClwG9Zg05uBbQHwGDby5nhgA9QBL9IsN+N/FjXySESktQzHyhP9CLgJeBv3eVsl2yJsBPWfgM8Dm2AzFbOsfwX32RrrGNW6MyIiIlLOGOAA4HTgTqzR33UOV8k2D3gE+B02+Gn9tF+YJuugsrWg9wc+wn8dFgOnNDAukST34h+Dq7sNRURcGw98FRul8jLuE4Q0t4+xDs6fAntRWYOMiFTvf/A/d4c6jkVEpN2tjY08vgh4Hutccp2TpbXNBe4CfgHsi5U2zYrDKN/wlbTuzKcaHJeIiIhkx8bA14HLgddwn5uluX0I3AL8BNgdK2uaFd/EyrEW0wGcBizH39+3gO0aH5pIzB/wj0OdazikNQfFhSFYiaJ9CtuadT5eN1ZP3Ksp7q0R+AHWgLMcWIg1dCzDpix7+mElCDvwy5MOxaY0R9e5GUfpH9okI7AvOe+LbgFwN9ZheBtWClFE6qea5SIi7gwEJmCdZftQ/8jrHvz1Yt7DRqB7ud5cLPdbjM3sW164ztOF5XJgeVgHlnsOxUa2r4y/nuFKWB5YjaHAHoWNQhz/wvK6W7FyVa1oNWBK4d8kA4DfAycGrtO6MyIiIjIMG+z+KSzPq3eWTzd+u917gf9/iK0j2IPfhuetMejx2vA6scoUXnxD8dvtvPUMx1L9IK5RWD67b+HyPGxQmNeG90aVj9csWwLfAM4pcvsorDN3n8B19wKfxfJskWaLtuHd7iqQdqeSh9IsY7AfnYOAnalsqnvQUmxEUrBG+EvAq1gS0ZtapKUNxkbDe/XSx2PlQ9fHaqpX62WsHOKVwKMpxSjSjnYEHij8/3LgGIexiIi0g5HAEcAhWMdgtY0vy7BOp+j6Ly9jjRTNWutvELbOoZfPBdfHWbGGx5uGjTi/CvtdalaOWkoXcB+wDtZQFrU6tu7M1oHrLgW+hDXOiYiISHtZCb8Nb0eqn0G3BGuvC+Z4L2ID5JvZGTWUeBuetwbiiBJ/V8zz+G14T6YUY72GAE9jnavbJty+OZbnrV243AucBfwAG2Qn4sIngTsK/78AqzojIjkzGDgK++FcRuXT+JcBj2Ojl4/DGmuyMMt1DP5aibcSruFdyfYithbP2tEHFpGyRuB/lh53HIuISF4NwGaSXYvNmKs0x+kGnsFO/E7E1u7LQpmmUfhrJd6IPyCt0m0a8HPcz2j/KRZP0kC0/dC6MyIiImIdacdhM3i6qTzfWYqt3fdbbJDu+mRjjeZx+Gsl3gHMpro87znge9ggM5emYvH8NeG2Y7CZl17Mc7FcXsS1VfGPy/scxyIiKdsNuASbfl/JD+pCrDPtW9iswmpLd7aqDmxE0nHAn7Ba3pW8Hj3YSPMvUX2pK5F29h72GVqAlRkREZF0bIflMh9TWS6zBCvB9D0sLxzS9IgbZx1s8Nv52Oj3ShuQHsM63YbHH7KhdsFfW+aawPXF1p3ZvsnxiYiIiDsd2AyeKwh3IpXaFgA3AadiswoHND3qxujEBnRNBC7Eli2qtA3vX8Dnaf5a1EcG4jg7cH1/4DzCcb4AbNTk+ESK6cDvkP/AcSwikoJ+WGmph6jsx/M1bN2TI/DXhWkHa2MNQ3dS2Yj7OdgPuuuRSCJZcDf+Z0efGRGR+nQCB2I5S6Wz5LzcrtkdYC6tDUzCOt4qGRg3D3udmjGbcBTWsOWN/vcajYZhsz+Dcd1LcslRERERyZ/+2ED2Z6iuDe9A8tMZWIlgG94Syr9Os7E2vGJrPKcd2zysc7IX+Frh+lWBByNxXYkmH0jreRj/GK1lOQcRaQGjsVJL3oydUtvTwHdR2UzPEOBwKivNtQxbu0ajuUWK+z3+Z2bfMvcVEZFkI4DvAG9QPrfzSqJv4CTS1jMQazS7EquMUeq1Ww5cD+zewHiuw28w6gW+iq0781rguh6sESsLZV5FRESkPuOAn1FZqfTHgW+igbeeYdg6jDdgpVRLvXZLsPWbt2pQLF3Y5Ixg+dcDsXXAZwSuW4ZViuhoUBwi9ZiKf6zu6jYUEanWKGz0cbmGj7eAM4FN3YSZGSOxUef3EW7ESdruQZ2EIkm+iv85+abjWEREsmYoMBlbi6RUHjID+DWwjZMos2MocDy2hk25dXseBvZI+flPTnie/yW+7szhKT+viIiItJ7R2GCgcgPTpwO/QOUny1kRy7UeoHwb3u3Alik//xkJz3Mm4U7LmcCeKT+vSJpOwz9ev+g4FhGp0CDg/1F6zZlu4O9YI4fW/areGpQfydWDvcYaqS/i2xP/M/Inx7GIiGRFP6wMUbm84xZgP2ykslRnZWz9xXco3Xh0G7BFCs+3Mdb4V6qx6sXC/URERCS/hgA/pvTgr2XA1djMHc0wq97awK+ADymdS18JrJvC800gvGZ00nY/ln+KtLJP4x+zv3Yci4iU0QWcBLxN8R+fOcC5qGxoWgYAXwCepXQS9ydgFUcxirSSlQknwyIiUlwHcBThEpPRbQHwR5qzPl476AscDTxK8dd8OXA5tefTA4EXKD1b8Traa11IERGRdtMX+ArhEpPR7WPgLGB1RzHmzSDgS9gArGKv+VJsOZRa13keDbxP6TxvCjb4T6TVrUd4hq2ItKhPULoR4y3gVKz+tqSvA/gk9kVZ7D2YiyV+mqkp7e4j7DPxoetARERa2PrAvyieV7yPzXRbwVWAbWAX4msCBrdFWLWOamdq/qHI4wVHrr+PlcG6Ahvp/iVsVuhGwOA69klERETc2xp4muK5wDRsSY4hrgLMuQ4sr7qH0h2zn6e6mZodwM2UnjW4HJvU8W/gEuB0bNLB3sB4bBKCSKvog53z9AJvOo5FRBL0xer/LiH5R+ejwu0DXQXYhnbE1iUslgg8gEb3S3t7EP/zMMZxLCIiraYLOAWYT3IeMR9bw0SzyppnG+Auiud2T2ONfJU4pMTjlNu6sc7KfdFgMxERkSwaiOVxxWaVfYC14amDqHn2Ah6jeP51HzZorxKnlHicpMFgwcuLgcuAnVPYJ5E0PYN/zA51HIuIBOwA/JfkH5kl2ELGI51FJ3vhf4EmjTSfjMoISHu6EP+zMMFxLCIirWQz4BGSc4elWAmicc6ik72Ax0l+f5ZhjX2lGvNWA2ZTfg2aYGdgL/Ae8FNg1bR3SERERJpmZ4qXs9TgL7c6gCOAl0l+fxZinbZ9SjzG5li+Xmo96aQ87zXg28CKKe+TSFquxj9uKx0QKSIN1AH8gOINC1eh9e1aRRfwZawhKOm9+jewkrPoRNz4Dv5n4MuOYxERaRVfoXgliJvRetGtohM4DphF8nv1JLBWkb+7l/Idgz2B7Z9YQ1XfRu2MiIiINFwn8HOKdxpdjAZ/tYq+wDeAeSS/V3dhawpGDcY6FivJ83oL97sTOJDqypaKuDAZ/xg+zm0oIjIU+BvJPzLvAoe6C01KWAn4O8nv20xgd3ehiTTdAfjH/28cxyIi4toA4C8k5wgfAZPchSYljMRmcia9b3OAgyL3n1zkvt62PPC35wHrNHoHREREpOFGAbeR/Nv/DvF8QVrDmsDtJL9vbwHbRe5fLJePzhKcgc0QXb3ROyCSos/gH8u/chyLSFsbT3IZ0R5sEdtR7kKTCh2BdQZG38NlWIkCkXawDv6xf6fjWO3/XpkAACAASURBVEREXFqN4mVEryF5ZLK0lv2AN0nOz8/AZgvsSvHZAl6n4BNYR7DWCRcREcmHzbGSkUk5whS0dlcWHIGtAxl9DxcDJxXu87mE24tVg+hqYuwiadkM/7i+znEsIm1rX5Kntc8E9nQYl1RvNMVHjv2Z0jXMRfKgE6vZ3wu87TgWERFXdgY+JJ4LzAYOcRiXVG841pmblNtdT/x99joK52ONg5s1P2QRERFpoEPwz3mjswV3cRiXVG8ccA/Jed6FWEdhUp43CxsollRuXiRLBuDPfn3JcSwibWl/YBHxH6HHsanukj0d2EzBpHrkV6O1ZST/nsI/5rXouoi0m12BucRzgBeBDR3GJfWZBCyldFmpXuAx4ERgkJswRUREpIGOJDkfuB9bckaypw/W0VesEkRwuxutGS358wp+5bv+jmMRaStHkJxUXILKDuXB/sDHxN/fG9GXreTblfjH+7aOYxERaaZPkTyS/Do0WCIPdgXeI7l86G3Alu5CExERkQY7Cms8j+YBU4B+DuOSdByErQ8dfX+XYm0c492FJtJQN+Af7xs7jkWkbRyDP2032LBwssugJHXjgWmog1Day4/xj/WJbkMREWmaA4iXHuoFfohVFZB8WA14lfj7/C+0vpCIiEhenUS8OlQ3cILLoCR1m2IlQ6N53jVotqDk15n4x/oRjmMRaQt7E+8Y7EaN6Hm1Gv4U7eD2N9RYKPl0BP5xfqbjWEREmmEHktckOdVlUNIQncBPSM7t7gC63IUmIiIiDXAI8ZKT3digf8mXAcD/AG8Tz/MudhiXSCNNxD/Of+w2FJH8G0+81GQ3cKzLoKThxgHPEU8uvu8yKJEG2QT/GL/BcSwiIo22EvEGhB7g6y6DkoYbATxEPLc7x2VQIiIikqrNgPmEf+uXAIe6DEoabnWSB4Kd4jIokQbZDv8Yv9JxLCK5NgJ4kfAPyzKymVTcB3xUZLsipecYBLxV4nl+lNLzNMtY4L/ES8l+2mVQIg3QD38thlccxyIi0kgDgUeJdwx+3mVQNbqe4jnX7Sk9Rx8sFy72POem9DzNUqyDcKLDmERERCQdKxJfJmYJsJ/LoGr0OMXzr7+k9BwjgBklnuebKT1Ps6xKvINwGfBJl0GJNMAw/NnRTzmORSS3+gC3kJ9RJ08R35fgYr2jU3iOY0s8Ry/wixSeo9nWIl6/fA6wkcugRBrgZfyZ0QMcxyIi0iiXEc9PfuY0otrdRem8a3wKz7FvmeeYksJzNNsY4A3C+7EY2N5lUCIiIlKXvsA9xHOVL7gMqg5Js+C8bSHWsVevL5V4jl7gByk8R7MlVX/7EFjXZVAiDfAOdnwvwvowRCRl3yVf9apLdQ6m1elZrpEqi52DADtjo82C+/JfoL/LoERSdh3+8b2p41hERBrhROK5yXXYmnRZVC7vSqPT8+oyz5HFzkGAzYmXHJsODHUYk4iIiNRuMvE85TcuA6pTqc7BXuCkFJ4jqZpC1jsHAfbGBj0H9+VxrANZJC/uxD++13Yci0jubIj1vAd/SB4g251B5ToHn6nz8dfASm7msXMQ4MvkZ6aBSJJf4R/bRybc3oWVLhARyaLVgNmEf8efJdudQeU6B9+ivlGkI4nnw3npHAT7rfPK8Xjb+U4jEhERkVpsgVXECv6m34Wdw2ZVuc7B++t8/PXLPH6WOwcBvk18f05zGpFIun6Df2zvn3B7f2BwUyMSyZG7Cf+AfACs7DSi+kU7B5dhJZSC121ex+P/OPJY0dHYWe8cBLiE8P4swRIqkTw4Dv/Ynpxw+8+wNRxERLLor8TzlPWcRlS/pM7BaP61Vx2Pf3KZx8565yDAOYT3ZzmwjdOIREREpBodxGfAvUf2z12jnYOLsXY873IP9bVHnUn5PC/LnYMAfye8PwuB1Z1GJJKe4CSW7yTc/r/AoKZGJJIThxH/Qfys04jSEe0cXAhcE7nu3BofuwN4NfJYU4m/jlnvHByOjcIP7tMNTiMSqU1SOY1t8I/rqyO37QrMxT7rIiJZsxPxGWJfdRpROpI6By+OXL60jsd/hPK5XdY7B/tjpeKD+/Qg+r0TERHJiuAgV287xGlE6Yh2Dn4A3BS57uc1PnYf/PXKSuV5We8cXBF4n/A+Xek0IpHqdZA8C3o3/OP6wsht+2DHvohUqRN4nvAPx61OI0pPUufgfsSTjX41PPaEyOPMBg4lnlhkvXMQrARVdL92chqRSPV+SHyU4RD8xvNgmeFR2InDi80JTUQkdQ8Q/t1+hOyuMxiU1Dm4O/F8b0QNj71x5HGWAPsmPF/WOwcB9iC+X592GpGIiIhUoi8wjfBv+N+cRpSepM7BwyPXvU1tJeSjbYEzsEkReescBDiB8D71UF/VNBEXfgqsFbluLOHBjcHrZwGPNSc0kXw5gvCPxjJgI6cRpSepczBptNDBNTz2RZHHOJ/kBqQ8dA52APcR3q+bnUYkUr3tgHnAiZHrvZmxi/FHJnmlOG5rWnQiIumJdpb1ADs6jSg9SZ2Dg7EBHsHrvlDDY58deYxrCM8wz1PnIMC1hPfrYbfhiIiISAUmEv79Xgys7TKgFCV1DvbDGv2D13+yhseOVhE7m3jHY146BzuxTpJoXiuSJXsAc4CjItd/gB3THxcud2Btd71YW56IVClap3yq02jSldQ5CPE649dW+biDsXKDwcfYjvx2DkJ8pmQP+elElvbhra16FVYyF+AO/ON6XWBS4PIFDmIUEalXtPzSTW7DSVWxzsHvRq77d5WP2wW8G3mM/cl35+AmxEvP7uI0IhERESknOiDq927DSVVS5yDAbyPXX1bl447COlGDj7EZ+e0chPhMyeXEZ2GJtDpvyYepwNDCdcEKOSsB3wxcPqf5IYpk2ybEfyzGO40oXcU6B8dHrl+GTUGuVHSKvld6MM+dg2ANbcF905euZM2n8I/fN7GZNL8OXHcysAi/sfSHbsIUEanZqkA34d/rHZxGlK5inYNjsXwueH01Oe2Bkb+dgXUY5rlzEOA6wvt2idtwREREpITtCP9uLwXWcBpRuop1Dm4dub7aEvJfjfz9o4Xr89w52AE8QXjfTncakUj1gst3vYZ9F/wpcN0Xse9Brw3vG27CzL88rE8iyY6PXL4LeMlFIE32EuHSSV3A56r4+4mRy9FFUPMqOiLtWGqr9S7iyu34awuujJXLDZZg+RG2hkNH4fIbzQtNRCQVRxP+bX6S8HoMefU+9h0fdEwVfz8xcvlirJM1734XuXw4th6viIiItJ5oG94ttMc562P45/EAA7Elkio1MXJ5ap3xZEHSrNLj8ds6RLLgOmzQANhAiAeBcYHbT8f6rbzj+s3mhSaSDy8THkVSTQdZFhSbOQjw5chtz8T+OtlahEswdWOj9CH/Mwf7Ax+i8lOSbUmLjhfbJjiKUUSkVv8h/D32NbfhpK7YzEGIr6P9FpUNYlqBeKmpTQu35X3mYCcwjfD+HeI0IhEREUnSAbxN+Df7004jSl+xmYMA34rcdn+FjxmtmLYEWLFwW55nDoIN+JpHeP+2chqRSPU+T+VteDq+G0QzB/NpPLBe4PIS8rUmTTlXYOUDPZsCm1fwdycQHmlzO5agtYMlwM2R6/Z3EYhIHf4KTMc6+ctph1GYIpIfo7FyU55e4B+OYnHhesKNSKsCu1fwd8dgA6A8jwDPphhXK+shvva2cjsREZHWswWwSuDyfOAOR7G4cClWQt6zE7BBBX93QuRyNF/Ms6RjRHmeZM2l2JIPlbThaeZgg6hzMJ92jFz+FzaipF3MAW6IXBct0RDVQbxE1dS0AsqIaOfgTk6iEKndcuBMyv+29QDvND4cEZHUbE/4u+1J2ut7bClwdeS6crkdtGepqSDldiIiIq0v2oZ3J1b5oF3MBG6LXFeuhHwXcFTkuotTiygbonle9DgSaXVLgf+hfBveYtqn47/p1DmYT9tGLv/HSRRuTY1cPhboV+L+e2JlRT0fEe9gzLsHIpe3wtZoE8mSi7CTi94S93mf8MhEEZFWF83tor/Z7WBq5PJhwIgS949WjlgKXJNyTK3uIcLrK44HRjqKRURERJKpDS+e502kdAn5/QmvT5a0RnXeRc8HtkXrDkr2/BGYTek2vLfK3C51UOdgPm0UufyokyjcugP78vCsAOxX4v4TI5evxEpttpO3gXcDlwcCa7oJRaRmS4BfUzwp7gVeb144IiKpiOZ2jzmJwq3HCK8jPRBbT6aYz0cuX4utr9xOFgAvBC53UFmZLhEREWketeHZUkizApdXAfYocf+JkcsXEx4Q1Q5exjpVPCOBlRzFIlKrBcBvKd6G14Pa8BpKnYP5tHbk8itOonCrB7g8cl2x8lPDgEMi101NO6CMeC1yea3Ee4m0tt/jL84dtRytNygi2aPczlwSuVwst+tLvNTU1NSjyYZXI5eV24mIiLQW5XlW4eHKyHXF8rykwf+XpR5R60sa+Kw8T7LoPGBRkdt6URteQ6lzMJ/GRS6/lXiv/LuIcOfAAcDYhPt9BhgUuPxf2nNEPsS/cFd2EoVIfeYC5xe5rQ9KLEQke6KjgNs1t7uUcFnonbFSmVEHAKMDl9/F1u9pR8rtREREWtcAwiW/lwEzHMXiWnTNwENJLiEfXTboEeDZRgXV4qJ53ipOohCpz4fAX0ge4N8HeLO54bQXdQ7mzwBsYV7PEtprIeOgl7G1VjxdwOcS7jcxcnlqg+LJgjmRy4OdRCFSv3Ox0YdRHSixEJHsif4eR3+v28VM4LbIdcck3G9i5PIl2MzxdqTcTkREpHVFf5fnYZWw2tETwNOBywOBIxLuNzFyeWqD4smCaJ43KPFeIq3vLIqfr6kNr4HUOZg//SOX27Vj0BMdeXRi5PL6wA6By93Ey5G2k4WRy0osJKvexz7/SSdWmjkoIlmj/M4Xze0mYiNKPWOAfSP3iZYjbSfREj3K7URERFqHcrywaJ4XLS26BfCJwOWlwDUNjai1qQ1P8uIt4CrUhtd06hzMn+hMmWii0W6uIvxjuSmweeDyRMKLnt4GvNf4sFrWwMjldk9MJdvOLHK9EgsRyZpoftcv8V7t4UZgVuDyqsDugcvHYmsOeh4CXmhCXK0qmtsVW89DREREmk9teGGXEy4hvxOwQeDyxMj9r8VKErYrteFJnvyScBu9RzMHG0idg/mziPA03AG0dwPSHOC6yHXeyKNO4qWopjY6oBY3LHJ5gZMoRNLxOvA34nXL23WtLhHJrujvcfT3up0sBa6MXBccVX5c5LapDY2m9Q2NXI6OMBcRERF3ojneUJIbx9vFTOCWyHVeu10/4ksFRWcathu14UmevIB9/oNteD3AO27CaQ/qHMyn9yOXV3USReuIJgve4sV7A6sFrv8IuKlZQbWo1SOX23kWpeRDdOTRImwdBxGRLJkRubxa4r3aRzS3OwwYAWwNbBa4fjFwdbOCalHK7URERFrXImB24HI/rER6O5sauTwRKyF/IDA6cP27wB3NCallKc+TvDmDcBveXMKziSVlXa4DkIaYBqwcuLwuNoOmXf0Tm4Ls/WiuAOwHfDZyv8uBJU2MqxWtE7k83UUQIil6urB56xJ8HLl9MLASdgI2BhiC1envBIYX7jOc8GCahdh3xVJsZF4vdkI3F0vGZ2EjHostpiwiUq1phDu91gUecRRLK3iC8Hf7QOBwwqXjwUpNzaa9KbeTdjMIGAuMwxqRh2L5Xgc2iIDCdcG2kEXYYIJu/EFkHwPzsdxuJpbfqXFKRBphOuEcZl3ig/7byU3Y/o8tXF4F2IN4SdFL0Dn32pHL010EIZKi+4FXgPUKlz+K3D4Evw3Py/MGYgMIvJm00Ta8BVj7ndeG14NVGpyD34Y3izb9PlHnYD69iNXl9mxNe4+m6QEuBX4QuO4UYPvI/dq9HME4wjMRltLencqSDytiHf9eA3IH8B/seB9L4xbs7sXvJHwfS9JfLmwvAa8RX19CRKSYF4GDApe3Aq5wFEurmAqcG7h8EtaYFtTuud1AYJPIdS+6CEQkRaOA9YHxhW09bGDsGKyxaHADn9trPJpJPLd7BQ00FZHavEi4c3Br4AFHsbSCbqyE/DcC130X2C1yv0uaFVCLWgeb/OCZB7ztKBaRtIzB2vAmFy73Ax7Eb8OLrrOZlh78PG8Gfp73UmF7nZwOElPnYD49Anw+cHkHV4G0kIuB7+NPTd4tcvtzwOPNDKgF7Ri5/BQ6wZXs6MIaQLcHtgE2whqORkXut1Jha7QO/NmI0YZZsBOe6Viy8ST2vf0w7T1CVESKi84SjP5mt6MrgLOAvoXL20ZufwerHtHOtsF/fcBOamc5ikWkWn2ADYHtCttGWGfgig5jGl3YNkq4rQd4A8vtnsbyuofROjkiUt4jhCtb7QCc5yiWVnEh4c7BvSK3P4StT9bOoucDj2G/RSJZ0BerjLMd4Ta8EZH7rUpzlkvrxDofx1K8DW8a1lEYbMPL/LmVOgfz6T+Ry3tgPeuLHMTSKl7BXpeditx+URNjaVX7Ri6380g1aX1jgZ2xzsDtsFk0jZoF2Ahd2AyXdbEyx57p2InOw4V/H8OSEBFpbw9iM5K9QU7bYA3UmT8ZqcNMbMH6g4rcfjFtWhomIM3crhMYGdkWA/fV8ZgiQStg52rbYQ3jW2OlorKiE1irsH0qcP3b+Hmd12Go6hEiEhRtw9sbazjP5SyVCj2LDViPloz3TG1eKC1LbXiSJSsRbsPbksbNAmyELqxaxXrAAYHrX8PP7x7CJh5l6hxUnYP59Bw2anGNwuVB2AnKdc4iag1TSe4c7EalubqwxZ2DbnERiEgRXVhD0T6FbQvCixTXajHWwOytJzMHW1NwObaGIIXrgiPwBgH9sRO2Ifhr2KyIX/e81kXk1yxs3sjR2djMl1uB27BF10Wk/byHrbO3VeFyH6xT7M/OImoNUyneOXhpE+NoVQdHLt+O/UZ5nXsjiHf4BbcVA/cbEnicXmxg3bcbGLvkXyc20MHL7bbBvtvqtQQbOPEeVpFhDuE1osFKrwUHXw0EBmD5ptchOQKrQOGtXTiG2nJPb8T7YYXL84G7sLzuNrQ+lIjAo9i5qHcOORKrdnWnq4BaxFTg1wnXLwaubm4oLWcA8c5BteFJK+mLtcF/CsvzPkE6bXiLiLfhLaJ0G95grDxpP8LrUHv53Rhqr0yxTmE7qnD5I+y728vzZtT4uCJ1+w12AuRtN7gNJ3VPEd6/hRX8zTD8E8PgVkmn6b4Jf/eLqqNuXQcT3rfZ2JemiEsjgBOAvwIfE/8MVrItwKb8Xw3cCPwE2AUrSzWMxunC1r/ZHNgfW+f0fKyj780a96UX++47g/iaqSKSfz8m/H0QHWWedXcR/84rt3ZYF3bCFf27+yt4vm0S/m5KLYG3oA7gZ1T/G7Mcm6XQjZ1QJ91nOrBn0/ZE8mYI1nhyBdaBV0sutBh4Bvgb8EtgIrArVoJ0ZANj74ONet8MOzf8CvBbrNN9Gvb5qWV/XgDOASZgHaYi0p7+Qvi74Uq34aTuFcL790EFf7MC9p0f/d6sZHD/4Ql/94Oqo25dRxPetxnoN0TcGwV8AfgH1kFXS140HxsUexXWufZdbMbheBpbUaIv1oa3BdaGdyrwB+wc9a0a96WnsC+/xM49W5JmDubXpcDXApf3xWYSvuEmnJYwF/g6NgU46FoHsbSakyOXr0TlbsSN/tj31THYD/KAKv72Pfzp/I9ha754P+LeY/fHH03USN3YLL93sQ69qMFYPfWN8NfS2ZzynfKfKGynAa9iCzVfjp1siUi+XYYtzO6NuNwB+95I+o5pF93YejTRklN3OIillfRiZaur1Ulyw9Jy7Lg7F+ukrmRQnoinCyuRdzQ2ILGaMvCzCOd2L2Hnsy7WVFqO5ZrvYZ2TUQOw3C64TuKWlM9lNyhsp2IDyK7AcrvnUolaRLLiEuDEwOVDsJnL7Tzr5EPsu3GNyPXXOIil1UTb8C5D6w2KGwOwMpvHYG151Uw0eQe/HOcT+G14Hi9nbMa5xzL8NrwngZsjtw/B8ryN8fO8TxBe3z2qA+ts3AL4HpbHem14r6cYu0hR0dl157sNJ1W1zBysR55nDiaNnN/SaUTSjnbGZmx8ROWzGx4G/hc4Ali9+SGnrj/W2H8K1kH/PpWPSHoIGxCyQtOjFpFm+ifhz36eGkdqmTlYjzzPHFwX6zgN7turVP6bEt2eA7Zt6h5IHmyNVbOpNJ/xRlefh80uXLv5IaeuL/Y6fAVr+H+Hyj93T2Gle8c1PWoRcaEDaxgPfg+c7TSidNUyc7AeeZ45uBvx388NXAYkbacDOw7/glWeqySv6cYq35yNlVpftdlBN8BArHTqqdh5eaVVMXqwNUJPxiqmiTTM8YQPvsXYAul5oM7B9DxJeL/udRqNtJN+WMfeQ1T2AzoL+8GdhE33bwcbY7ME78Rm85Z7jRZjjU+buAhWRBpuf+IDJaKz5rJKnYPpuZTwfj2DzQg8Gngb/6S00k6Kd7HS3JOBvahu5pe0l05sHfM7qezYmocdW5PIRyNRJdbGBoLdia2PWO41WoLlvyopL5J/JxP+/M8nP+e96hxMRx/gecL7FZ3hJNIo/YHjsHOLSvK8mfhteO0w2KkT2Aq/DW8Z5V+jRVgb3kYO4pU20Bebpho86PIywlydg+k4ifh+nY99maleuTTKaOCHWFmmcj+UL2ONkVujY3IUcCS2BuMiSr9uPcCt2OLPaSz6LCKt4zHCn/d73IaTGnUOpmM74h1/nwvc3hf4Mv5srmo6Cb1tGfBf7ER2EjaQpd1/o9vdCGyW23TKHz9vAL/CqiX0cRBrKxmGlQ68DOsEKPfa3YuVZtXnTSSf+hOfYXyR04jSo87BdHyX+H6dh9rwpLHGAT+lsmoQLwA/wkpptntb1IrYedi1JK+fGtyWYwPmtLa7pO444gfcvk4jSoc6B+s3mNJf7POwkQ6TsVHiA51EKXkyDvgd5Tu2ZgK/RSOkSxmOrUlxD5ZElHo9n8NOjto9MRPJi/2If86PdRpROtQ5WL++WFnG4D49RXIHzCBsVOtcSncQvkxls5vmAvcDZ2Azx8Y0YP+k9YwCzsLOG0odHx8BFwC7osbLYgZj6/XcSrwscHR7FasSpNdSJH++TLzBeDeXAaVEnYP1G07pZVjmojY8SdcqWP5WrmPrPWxt8q3dhJkJI7FBlfdRfnDmU8BBbsKUPOrA1uUKHmRvYwdllqlzsH6/oXxDT3BbCjwI/A82YlWNPlKpYcDPKD0iejnwD6xkXpebMDNrVayBdzqlP8MPA3u4CVFEUnYr8Yb3VZxGVD91Dtbvx8T3afcyfzMK69BbTPxEdTnwMbYswc5YKcRLsFmDleSOXjnS0wp/PyCVvZRWMBB7Xz+m+PvvVTE4BJsNI5Ubi33eXqL0Z+xZ4ABHMYpIY/QhXrLvNWCIy6BSoM7B+v2F6trwFmMDt84EPo3NYhKpxEjs/GAhxY+vZcDVWLWqdq8EUa01sO+jtyj9GX4AO4cSqdvWxOvc3kK2P7zqHKzPZ4g3AJ2PfemchjXklBqRFGz0uQY7ed0KzUySsL7YyJgZFD+G5mKNsVpAu36VrvNzJ7CloxhFJB3rAQsIf7b/Q7Yb4NU5WJ+9ic82uqyKv18V2/9uwjPSlwNXJdx/Jew3ZzKV541eOdIpWHWTjVHumDWd2HrR0yjdGKn1j9PRgc3+uJHSo8wfBCY4ilFE0rcz8eowfyfbv5nqHKzPicT352zCbXilBuwEO5qDZeGzfExJ+vphx8ZMih9Dc7BStms4ijFPvDa8Byjfhre5oxglR04n+Yckq07BRjF42+kNfr7xkec7AxsdkUWbE29QfJ34SLQ+WLIwCUseoutXFvuRCJYx0Ajx9rU78ROA4DYNOBWbVSjp2xa4kuILIC8H/oiVJhGRbPoG8c/2JU4jqs9JxHOtfg18vlUTnu/QBj5fI61PvHNuJrbGb7U2xGby9xDubPx0mb/z8sbjsAaD+7GqE+Vyx9mEy5FqVHvr2pb4bJbg9i7wfWAFVwHm3GbAhZQu7XU5+gyJ5MU5xD/jk10GVKfvEM65ftzg59uUeJ5XrppCq9qB+Hf/88Tb26JteNMpn4fNIFzlIcsDDaU++1B68NcrwFfJ/izmVrUT8FeKl5ZfjlUA1OsvNetHck90HtaokcqNBd4kfAwsArar8O9XxkYLnwc8Rvm1zpYW7nde4e/UWJB/w7H3u9ixMQtLPJV0Nsd4bHZvsdHm75LdxnCRdtcJ3Ez8c/0Nl0FJ0w3HGoiCx0A3NpOwHtsCdwce831gRJWPMYRwOdJSDQ7R3yavMoUaqtwbiDWqFmus+AjL7bS2UXOshj/Lt9j7MclZdCKSloHA44Q/3z3AYS6DkqZbDVvPLXgczAc+UeHfV9uGt4DwoK1RKe2HtK4RWF5RrM3obSyv0PI/zbEhpdvwppHdyUrSAsaR3DGU1dEzUp3hwEPEv1iOq+Mxh2IzBCdjMwZL1aP2tmgZA8mP/Yl/xwQT2DPQTEFXtiG5ZJ+3XUNtM0xExK3hwAuEP8/LsDW+JP8GAncQ/07/ZorPsQ/wdOFx/5jC462MX470TuLVLMoNNlM50ubaheLr3i3B3pOsr2WfVV7jUbHPzc1Yo7KIZNdqxJfomAfs6DIoaZpRwJPEO4gPr+Mxq23D6yZcEn7tOp5bWs+BWOdf0ns/D2vDG+osuva2HXAPpdvwVC1CapJUUnIx9oUg+TWC5I7BtEvLdmFrD56CfVG9n/CcSaPDg2UMGllCTBpjIHARye/vcuD3qOOpVexDvCPB22ZiJwoiki1JJSW7gWNcBiUNN4jkNWYbUVq2E/gcVk5oQsqPHS1HWsmo9l5sXR2vlL1GtqevL/Brkkct9wAXY6V5xb1dgCdI/pzMRhUiRLIuqaTkfGBPl0FJw40BniL+vT455eeJtuHN0EAHZQAAIABJREFUSnjOpDa8YIWHvinHJI03BFuGpliH8Dkot24VBwKvUvyzuKu70CTLDiN+orcEONhlUNIwI4FHiH+J3I41yDTa2liDzxRsxFGxqdHBRDdYxqDaElbSXKuRfHz1Yo2IuzmLTIrpi3XGLyE5EZyMNQSLSHZ8kvgao93ARIcxSeMMJlzy09v+Q2NLcPbFjrVGG4pfjvQa4rMmim1edQqvsUoDzmozmuLVBqZRf8laSV8XdtzPJ7kz9zxUDkwky44n/tlegL6P82os8Czx9/xamnOeXm0b3jzCbXjDmxCj1G5diq8h/Qy2tIC0llIl/pdh7XsiVfsGyR2ER7oMSlK3MslJxb9xV95xDOFyUtFRcEmdFcEyBms0PWIpZjeSZ4cuw364tEZQa9uM4h27N6CkXiRrjiN+wrAcrT2VNysADxL/3n6KfM/S98qRnoE1QFVSyt4bcBYsRyqlbQ28Qfy1XI7l4kPchSYVWIfkgQO9wL1Yg7OIZNP3iX+uF6EqYHmzBsnlvO/Eqka4MJZwDlauDW8Z4Ta81ZsfshSxP1Z9I/qeLcXeXw2sa22bE1+L1tuuxAaQilRlEvGyPT3YF0IzZpRJY+2MTTGOfmH8i9aqGd2XcBmDDyjf2BMsY7AVmuXkwndJHrXyX6zTSbKhC/ghye/l82hNAZGsORI7uYt+nqegk7082BybHRd9fx/HOg3bSRd+OdJKR7Z7OeSN+OVItV6e7ySSqwq8DmzvMC6pTifwdZLfy+nAps4iE5F6fYv4b53Xhqc2keybQHK1hFuxmUOtYhDW3ngallN9SPVteFo7uvl+THKu/CSwgcO4pDp9gdNJXobhadQZLzU4mngZql5s0csxDuOS+kwi+YTwNlorqSgmWsagXKIxF3/dmb3Ixj5m2WSS34fr0WyzrCp2IvIemmkhkjUHkDyi91F0spBlRxNfN9x7X7UmiBlGuLFqJuVzSK9ChVeOdCvac5DkySQ3GN2NzgmzaiusMzD6nn6ESoaJZNkXSW4UvgktyZJVHVgOktQ2exMwwF1oFfHWj56E5VOvUz7/mkO4Da/V9zHLOrA1BJPeh8txNyNV6rMfltNF39M3sEoSIlU5muKjRLd0GJdUbzBwGclf+teR3TKPKxEuY5A0KyK4LQMew8pIHQGs2PyQc+vnJDesnYZGf2XdqsDDxN/fGWiUuUjWHEByR9J7aNHyrOkP/I7kfOduWqsaRCtaGcsFz8NyyEWUb7Dy1s7xypGu2eygm+y7xF8DVZPJhxWBfxJ/f2cDOziMS0Tq8wWSO5JeAjZxGJdUbxjwN5LzkSuxWUJZFMy/HiO5Qzu4LSXchtduFTEapQN7TZPaTLVOXfYVWz/yXWAjh3FJRhVbX0Jrh2XHLiTXJs9jmYkhhEeGJ9XMjm6vYaOYJmGjmtSRVZ0O4LfEX9ePgT0dxiXpGkDyAIOZwCccxiUi1dsMeJXkvEBrh2XD9sBzJOc1KhVbm76ER7fXUo50L/Izynoy8X1dABziMCZJVxfJAwzmYOdTIpJNu2KDvpI6WbR2WDbsQ3I7bDf2+5y3Nry9sP26ERukUm0bnlSnE/gz8df1A/T7nyeDSR5goA5Cqclo4C6Sv5RfAXZzFpmUMhxrIEpq2JhDe5zcR8sYTKd8ojEDS0pOw34Y1QFe2tnEX8MPsZJFki+d2HdKUhK5rsO4RKR6w4BrSf4dnAbs7S40KWEg1rCXtB7sIuAEd6Hl0nDCDVaVrH+9DL8cqddolbVGvNOI79c8dM6XV78k+f3ewmVQIlKXVYAHSf6degaVEG5VI0g+3/bOudshP/fWjvba8N6kfO71HuE2PHWAl/Z74q/h+6gqVB71AS4mud17DYdxSUb1JXnKcS82Dfw8rKFJWsOhwDskv1/PAuu5C825assYLMBKSJ2BlTDV+j2+Y0lOKjSTLL86gHOJv+8voHUlRbKmE/gZyYOIeoALUfntVrI3NlK62Ahq/fY2h7f+tVeONGkdz+g2F78c6RG09lp9nyLe+ayOwfxL6hB+AxjrMigRqUt/4AKKD2Q5C1WLaBUdwOewtpSk9+tx8l/KvJRoG165yg7zCbfhac1N3wkkd66q7HB+dZBcKeIpbHahSNUOonin0yzgVDTTyqWdsR/BpPenG2vUz0u5o7QMwx8Vfifl15zpxkaET8Eah9ZqesStYVvir1UWp6dvhr3vxba0Rk/tW+Z5stbpfBbxz8aNZG92hIhYCehinU5zgB+iEweXtgRuJ/n96cHKAqnRw52+WLWEU/DLkZbrLPRypuAo94HNDjzBhthnPhjnR2RvhskalM65dkrpeXYq8zxrpvQ8zZLUQfhvNANDJOuOpHin0wzgK2R3/bo82B14mOKduL/ClvgQX71teGs2O+AWsStWXjj42rxJ9iaPbEvp/Cut/TmkzPNkqW29WAfhNWhpLanRcGzERrEZV29iU8C1UH3zbIh9qIv9GL4CTHAWXbZ04TfyXIN1elfSwHNN4W+2Iv8dJOOAtwi/BouAbVwGVaMJlH5vz03peW4r8zzjUnqeZkqqU/9LpxGJSK1Klav0BoCdhgaANdMaWCNGsXx7GvBJZ9FJKeOwkeqTsQ7ADymfS3rlSL2Gq2avgz0SeDkhpt2bGENaNqb0a31JSs+TtBZzcMviKPxziO/HRU4jEpE0lCpX2YvNFFYbXnNtQuk2vGfIZvuKC8GBWtdQWRn4dmvDW4P4IIF5ZDNX2Y/S7+3PU3qe+8o8T9aqJ3YAVxHfjx+5DEqybw+KjzTvxcpXHoVGITXS5sClFG84WoqVDFNjXn28ElJTsIabcmUM5mEjSSZjjUN5KrfYCdxLeH97gM86jKke5ToHP6D+EdOrULzBPcudgwOAh4gfC/u6DEpE6rId8BzFv6teAb6ARjA30gZYGbDoyF5vW44NXNFszuzw1sAOliNdQvmGqzmEy2KNbmCMf094/q828PkaqVzn4ELqz82HYUsP5K1zsA/JM5WPdRmUiKRmX6wjsNj31hPYTMMuVwG2ga2BqyneprQY+AFqR61XtA2vXM41F78Nby9ao6JDWvoQb7dZjlUGzKJynYNvU/9Ah7Uo3+6btc5BsPPHJ4kfC7s5jElyYBDWyxwtQRP9YP4/slc2r1V1Yg0Ed1P6i+pG7ORY0jcWew/OoLL1ZqKjwVdrQoyNSuhPIb5/WZ4tVq5zsBc4uM7n+F4Fz5HFzkFInkX6DjYDQUSyqR/wTUqPup0J/JTsfne1mg6sIeJmSp+I3oU1Kkn2DcbKiXrlSF+nfK7QS3ik+86k01F/dMLzXJTC47pSrnOwF/h8nc9xUgXPkcXOQUieRTobWNVlUCKSmqHA6di6bMW+v6YD3yZfg5xd6gMcipVqLvaa92ADdcY7ijHvvKoOXhteuUFay7D1Db31ohs5QMvTqDa8pPaoHzbouZqhXOdgL/VXVzm9gufIYucgwOrEZ5FOw34bROqyIjaKuVQnyQLgD6jDqlbDsXrw0ZO16HY/1lggzTMIe81PwzplP6K6xp2tSL901DBgKul+3lYlfhJxO9kuwTCB5EQwePm6Op/jxTKP30u2G9i3J75Pv3cakYikYTjwC0o3Hi3GOhG2chRj1g3GOimeoXTO8DjwKUcxSvOsTLgc6ceUzyeXUl850pHES+g/TLbXmUvqHIzmKf+u8zkeKPP4We4cBFtDfCHh/bnaaUQikrZx2DlbsUoFvVhFpN+gDqtajQS+QfkBQHdj1TukebxBWl4bXiU512vYgK5JNKb8+wrAhcA6KT7m2sTb6a8n2+vMJXUORvOwy+t4/E7iM6yT8rysdg6CzRSMViA822VAki+rYSen5Uro/Rc78V2zyONsgJUzaHf9sEaCSyjdONcLPI+NaMnyl3xeeOWjJlH5aPA5hMsYpDES/DNYsn8elmjUK7q2ymyaMwuykZI6B6+LXF6GzRatxY7E3+d/JTxnljsHwToQgvvTDWzqNCIRScuK2CjbcrPkn8d+w4qd0K4BTGxsqJnQB2uMmIKVMCr1mk7HcoksD8KR2kXLkT5G8aUEovlZsLx9qRzw15G/XUz2B3ImdQ7eQvz8tNbG7vUJz/BdBtyR8JxZ7hwEOJX4Pu3qNCIRaYQ1KL3GcbAN7zSKn7duBhzS6GAzoD9+G1658tPPYW144l60DW8a5fOt97GOxdOw3D6N5ZxOxHKxs7G1Quv1D8Ixz6L2tq1WkdQ5eGPk8kJqf/32ijzWB8CDCc+Z5c5BsAlewf1ZguW4IqlZD/gtNtKo1JdpN3byOhH/g7sdVq6qm+zWQK5HH+zE64/Ah5T/Qbob+DRqOGp1K2OJX6WNO0sJlzGotXPvtsLjzQW+Q+0Jy+bES51NqvGxWskE4q/9mdio+eB136jx8S+IPM6fsAaq6HNmvXOwP/FZzTc5jUhE0rY6cBblR9b2YAu4fxHrWARrJH+rcFs7rl3VieW35xEv45K0PYjW/JFkw7BGix8ANwAzKH88LcdmukWtSXzGyPcbGn1zJHUO/pl4/vWzGh//l5HHuRZrSIw+Z9Y7BzuBRwjv00NOIxKRRtoQa4Mq16HVDdwKHIPfOL47NjBlCfYb1W66gD2Av2CvQ7k8+XZs/UcN7G9t1bbhLSC8VnQtS2t1YOdRvdg51ynUvv7kDgkx5uE8LKlz8CfEK7GcVOPjRydFnAfcm/CcWe8cHIQNRA3u0zUuA5L8GoZ9mU2n/IlrN/Aq4em6j9MenV6jsR+dKcB7lH+tlmAfWpUeyK4hWOI8Gesgj5buSdqiZQwqES0j8CbwWapPRP8aieUp6l/ktxVMIP46nwmcHLnumRoeeyDxRvSdyWfnIFgCHN0vlRoUyZ8h2O/Q81TWKfES4d+hV8h2ycJKrYCf271NZa/VjbRno5rUJ1iOtFhOeXvC3/0hcp/XSadyhWvFOgePjFz3FtXnsp1YLh18nIPIZ+cgWMNidHCgvqNE8m041oYX/a4r1ob3OuGZ2fWWbc6KMViedwmVLSuzuHBfVdfJrqFU14a3HJtxW20b3kaEB2+9hq1ZWa2bI/E8RD46pIt1Dn47ct39NTz2MOIDJLYgn52DYNXmosds1iuISAvrwg662ylfctTblgJXYiOS1mt+yA01Als75ufYCJToSVex7RXsSy8PHQkS1oV1pJyCdfxWMrPgPcJlDIo1tgYXIPZGOz2OdYxVYl3io6T2q2bnWtgE4q/rmdhJUTTZ27zKxz4m8vcvY8lYXjsHwRKw4H5d6TYcEWmgTqxywQ2UXq8m2oh0LVYxYkPyNQBsKDZq/MfYzL9K8903sJlIqzc/ZMmpLvxypFPwl3MIGk28VPBxzQuxoYp1DvbDSkMFr9+zysfeJ/L372Mj+vPaOQjxcvt3uA1HRJqkH3A0cBeVlbTuxQaxXwochQ1SzpNR2Gy/XwFPUnkb3ovYrPzRzQ9ZGizahjeT8sfDu4Tb8IrNCvx54G+8z98jhb+pxKbEj9Hdqti3Vlasc3As8XPSDap87EmRv3+2cP29Cc+Zh87BDqxtOLhff3EakbSNFbAPXLQRudw2h/AU7az8uCadoFeaXPVio5CmYD8CeRjlIZWLljEol4DOJ/wZ8Ur1dmE/asGGSu///6T8yJBfRZ7ncfJzLE4g/jqeWbjtqsj151b52P+M/P0PCtfnuXNwX8L7tRi/rKCI5NdILM+5k8obS3qxktf345fQzsoaGLWuBedts7HOhAPJxyx8aX3RvO1bhI/J18hPGdtinYMAv49cf0mVjx3NDc8pXJ/nzsFtCO9XD8XXlhWRfFoZ6wCptg0vugZuVs4LvTY8b/25/1JdfvshasNrV2sTbvstd9zMI9yGN7zwOP2xySHRNrwerKpXuc738yLPU8ssulZVrHMQ4msP/rzKx/5P5O9PLVx/b8Jz5qFzEOAw4u3Kedk3yYiNgSeoLsEIbu8C92BfvN8C9sdmObk4uR0N7AJ8AVuT5zpslFClo+mD2wdYneP9qb2+tOSPt87MZCzJXkTp46gbS0imYD+KSYnJcqyU7xSsNEZUJ/Y5C/7NxPR3zZkJxF8Tr3Mw2tH1AZWXwluVcCK3HH9WSJ47Bzuw773gvtW6XqOIZNM6WEmpWnO7mdhaG3/C1so9CBiPm3xoFFZW7wRsoMzfsd/VJVS/X7Oxk/lDqX39X5G0PEf4+Py223BSVapzcNvI9QvxG+LKSaoq8YnCbXnuHARbs7KexjYRyY8tsFyo1jzvbeBurLT1qdg59zq4GSw1FmsPOAk4G+tYeJnwUkfV5K9TsRnmGvglnrFYp98ZWAddtGpDqTa8s0rcZynWATiCuH5YB3Xwb45sxM45Uqpz8PDI9W9T+edxfcJtpsvw2+nuTXjOvHSgdWKVbIL7NgnyM2pQWttg7Mtui8B1t2EnXXtR2QdtpcK2W+T6bmBWYXsX+6GeiZVhnIk16iwpPFc3NlrDa7gZhDXa9CvE2KcQSwfWATgG+4IfV/j/uMLlQZXtdqIebDbWrYXtUawzQSRoLjYb7Z+Fy32BzbARaTth5cxWCNy/D1avfKMSj9lZ2E7CymCejSUuiwu3b499xoIx/LWenciQ27FkYtXC5RWwROS6Cv72RMJJyF3Yug1514uVITgrcN3BwK/dhCMiTdYfazQOlry5E1t/dS+ss62c0fgDroJ6sBxuFn4+F8ztFmEnyguwHGpu4e9mF+IaiP1uDsF+97wOAS+3G42NjB+N5XUrYXlgrXqBp7Hc9lZsJGp3HY8nkpb1CVeNWEb1M+iy6hGsmoa35tNArCGpkhJKRxfu73kS+4y3gz8DOwYuHwz80FEsIuLOUKytwGtf6MUGvnZjZZqHVPAYqxS23SPXdxPO7WZhpZu9/y/B2igWYb9b84m34fUv/L+rEGsn8Ta80ViON5bwd3q1lmPtdl4b3uNYrioS9D7W6Xxj4fIgYEus/W5n7Lc1eH5USRten8L2NeB4bADjr7HPCMCukcf8ELi+np3IkBuwQf3eLOVVsHbSOyv42xMIz/S9GZiRanStqQe4kPASBAcDFziJRtrKWKwEU3DkwxcDt/fFOvxOxz7Es0keMZHVrRt4ChsNcjTZKZEqrS9axqCa49IbJfMm8FnshzFaUjRva8hNIP46nBm4/YzIbddW8JgdwKuRvzsqcHueZw4CrEl435ZhJQdFJN9GEB5V2UP4JKMPdgL8E+x7MDqiNetbDzYb60Ls5DI4sEaklURLilbSYJIlpWYOgs1IDt52X4WP+0jk774euC3vMwdHEK+Gs5bTiESk2VbCBkV43wFLCJ/j9sM6CH+BzQycS+U5VBa2bqzq2flYW0lwULZIrbylCrwStq9R/flHb+HvDi08ZrSkaN7WkCs1cxDgN5HbLqvgMTuxdtDg3x0cuP3ehOfMy8xBgA0J79si6hskK1LW2thUfe+gm4+VzyylExs5MRErP/AktZV0cr19iI3i0IdMmmUVbBbbs1SfYLxCvOzU55obfsNNIL7/wc7BpNIC5dbEij7mHMIzi/PeOQjwDOH9K/cdLyLZtjKlG4yKWR+btf5brOG9XKmdVtreJVwafzl2sirS6q4nfCx/zW04qSvXOTiWeMm48WUec6PI/ZcQHtyZ985BsMb+4P4d7zYcEWmijQiXnZsHfKrM3/TBvgdPxGagPE1tS+642qYTHmy9ENiq8pdMpGarYZNnXqD64/Z5wu3tvdgSDXlSrnNwq8htC0kuvxpUbkmhexOeM0+dgxCf4BCd3S2Smm2wadXBD9yOJf+iuC6so3EvbJTFedjI19eobrHgtLYlhee+EZttNAmbJv6VwH1UWk+a7QisDEdan4lVmht+w00gvo9nRu7zYOT2cmvoXRS5/5TI7e3QOfg7wvv3M7fhiEgD1dJgVMrK+LndGVhe9RrW+dYKud1ehAeJBL/vPgLWq2PfRZohupb0Zm7DSV25zkGAmyK3n17mMc+O3P/vkdvboXNwMuH9O99pNCLSLNtj7QneZ/9dwksDVaPV2vAWYx2A1xBuwxtaiLejcJt3/+mo6pc0VgdWCexD0vlM9JC/Wa7lOgfBKvUFbz+pzGNeHbn/uZHb7014zrx1Dl5IeP++pzUHpRH2Bv6G/0M7DVus9+UaH68beL2wRfXHryU+prB5/18RGwEQrUfegY0mWIg1BiWtW/MhyevdzMBmByU5NfD/W6reS5HarAv8ESvt0UO4dnat5uOvRdhOLsZOijwnUryjfzBwWMLft5uHsYERnrw1PIqI2QHrOPNOOt/DZgo/WcdjvlvYovrhrwforRkz7v+zd99xk43nH8c/z/bGWmurtqutHiVK1NUJogQhUTYkQkSk/ggpSxoSQiQRJHpJJHoNghCEkBBEZ/VlWWubLc8+z++Pa05OnV7uU77v12te+0zZmevMnJlzn7tcV+kyEhhEvF40WNtuIeEaNT347bYPsIlrXm2b6aW/p5fuq+Zr2ADpdlgK5RuxY0a5dqGIS975kWc+Nsu8aC4mnNVgCjb4lVTzvR9WBiL6/4vm4ch1te1E8m8f4Ar82nzPYCtsXm3w+Sr14Q2ifB/eSKyNN4hwDeloH97C0t/d2GS1XpL78GZgbc05VeLtxc7918ImfKwMXIv1sSyqb9NFqloLW2W7Fa3rw5tFctsm7y4FzghcnwJcUOaxw4E9I7dd3PqQUu9hrDSGZ71yDxRp1BTCKQT+iR3k864/fr3EuVhjRqSdBgDfI5yard4ZR/OwTs05kdunY6vqliM/tiW+/dGVg8Ox9yT4mA3KPN+UyOOeI96oK8LKwc0Ib98TbsMRkTbYB+uA8b7n/8U6TYpoJOEaITdgKfFF0qYIx+daVg4OILwSphdbyZJkj8jj3sHO8YKKsHJwVcLb96bbcESkzY4lnLXhIfLVD1CPCYSPGec4jUbyZjCWwSDYZ15vH95crO852of3JvATqqfVzJJaVg6OJp7GeM0yz/flyOP+lfCYexNeM28rB7cjvH0PaeWgtNLx2BJ9z11YodRqs3TyYGtsYAFsu4u46ko6Z0WsAPg44DasgTAPayR4f8/DVkJ4f3uNCO968Hv5JWz1oedW7PtcNB9iHb3BeouHYakKoqZErl+MHViL5pXI9ZWcRCEi7XIstoLaGwD7Bzbj8j1nEbn1Pta2fQBbufgpbBXS9x3GJJJkhcj1Rld/ZN0i4A/AVwK3HYadr0VNiVy/DFuFXDSvYQMFfUvXx2GDpEV8L0TyrAvraA92tnvnwh85ici9adj234atJv8K8B/Kr0QSqdVErA9vFJby3OuX8/rwau3T83wdODNw/TrgxLZuQTq9i31fPxW47RDgpITHTolcL2LmL1AfnrRJX+BcwiPPlxCfaZlnZ+Bv+5GOYxGp17cIf3/PqPzwTKpl5SBY/azgY6IFisEadsEZXkuwAduoIqwc7E94+9RxJJIPXcTrTl2Hn26q6PbFPw70AAe4DUckZgrxc7O8qWXlIMDGkcfMJz6zfiThbBy9JKfTLMLKQfAz4niX4ZUfLiIZMwC4nPjvpxaQmG/ivy+LgG3chiMS833C399qNZWzqJaVg2BZboKPeQN/gpMn2mZcSHJd0XsTXjNvKweHE96+D5UGR5o1BLgeOCpw22nYCWmROok/Gfj7dmdRiDQmmgZ3vpMo0uFO4PXA9ZGEv99gv2/BFKLR/1MkiwnXYeiH1XkVkewagK2YCZ58/RrYj+LOJI+6Fvhp6e8u4PeoLpeki9p2vsewlR+ewdjvWdDnCLdfHo38n6KZF7k+zEkUItIOw7AVgl6N1V7gZOALWA0/scnSF5b+7g9cTXxFvohLauf5bsbSAXuWx+qFBk2p8n+KJNrGG6rBQWnGssAdWH0GsNUzXwZOoFjp9Sbi5zR+AkvFIpIl0cLFRZ4x2IPNogw6LPB3F3Bw5P6L2xlQynUR3l+0elAk28p1GH2FYha5r+R7wE2lv4cBN5I8A1XEBbXtwi6NXD8scn1K5HoeV1rWI5o1Y1Hio0Qka8YCfwN2LV3vxkqMTHUVUIodAzxS+nsM1s4b4i4ckZDoQH6R23mLgSsjtwXbef3wz209RW7nRdt4izU4KI2aCDwIbFm6vhA4EEsvWjR7BP6+1VkUIo3T7OCwiwhPcNgDOyEA2B5YJXCfV6ewqIbi1yIDS8nV4ygWEWnOWOA+wh1GR6IOo3J6sBPNp0vXVwauoVhp9SW91LYLi9YP3AqYVPp7XWDDwH1encIii+4v0f1JRLJnVeB+YKPS9XlYnS7V00u2ANgbeLN0fUPgPHfhiISonRf2+8j1ffFTyO+G1U/2eHUKiyrWxtPgoDRifaxR4Z1QzQR2BP7sLCK3gikHb3EWhUjjosvpl3cSRXq8ADwUuN4PK0wO8ZnlV1HsNHvRfaWoqRlEsm41rG3ndZDPxTqMkup3iW8OdvI5q3R9a+Bn7sIR+Z/3IteL3rZL6gjyMkEcEbn9BuLvX5GMJJyubD7FbuuK5MFm2PntaqXr7wDbUuwO8lq8DeyPLYYAO258w104Iv+jdl7Yk8DjgeuD8GvCT4k8NjphrGjUhydN2xFbKeMVrnwFP6VmEQ3GTph6sUHSIi/lluz6BOGCtI9XfngmbUu8sPBpFR5/ZOSx/8EKEc+L3L55hee4NeE1xzazESm0G+Htu9dpNCLSiM2wjnPve/w2sLHTiLJnZ2ylpfcefsFtOCKsTvj4/IbbcNpiHeLtrEoTGvaNPPZ1rM7g9Mjtu1d4jksTXnPdZjYihTYhvH1PuQ1HRJq0F+Fz2BexY4TU7jD8968bOwcWcWkHwsfqB92G0xafJN7m+kGFx38t8tgHsAlPCyK3V6oTf2/Cay7dxDak0acJb58yIEpdDsXSrAQ7y4s+O2FP/PcjmuNYJCuWI3xwWEi8wHHW1Ts4mDQQ+KvI9eewmnvlFGFw8ETC26e0NCLZshf+JCevw2i1iv9Dyjke/31chK0iFHFRdxjaAAAgAElEQVRlANaeCx6jRzuNqPXqHRwcQHgiRFLbbjqVJ3sWYXDwS4S37zq34YhIEw7HVsh43+eHUX3kRgWPFzNRe1ncWpHwsXoO0NdpRK1X7+DgSOJt399Erj9a5TXvTXjNvA0O/ojw9p2ttKJSq+OAi/HrqNyN1Wp4s9x/KAilFJU8eA9bBewZQLj2ShHNJt4Zckzk+oXYwbTIPhG5/k8nUYhIIw7HUsIPLl1/BPtOv+gsomw7DX+iWH/gamAFd+FIwS0CnojctpmLQFJkEfHJnNG23aXYqpAii2bFUNtOJHu6sJrRv8ef8HAHlglMKeQa8zXgntLfI4CbgOHuwpGCex1LD+wZRv4mK9XrfeL98kdHrl/SoVjSTH14Ure+wK8Jjypfhg0eCEzD3pMl5G82rhTLlYS/56e4Dafl6l05CLBTwv/xLkuo3umb95WDA7EZasHtW89pRCJSC6/DKPjdvREY4jCmvBiEDbJ67+tj+IOvIp32S+Kzp/Ok3pWDYJPfyrXtamnH5H3lYBc2+Te4fds5jUhE6tUPOJ/w9/giVAKnFUYCL+G/r9cDWnQjrlxP+Ht+gttwWq7elYNgWXHKtfEWYlnTKrk34f/laeXgMOJpVld1GpGk3kBs1nNouSk6+HnWw39f8pjfWYrlUMLf9ehs86xrZHCwD/Bqwv/rpba83HkfHIzWG3ydymlWRcS9pA6jC1GHUSuthM3k9d7fS92GIwUW7VR5nXydxzUyOAhWWzupbfdwDf8374OD0XqDs7E+ARHJhqHYypng9/hUpxHlz8eAufjv71Sn0UiRHUX4u/6Q23BarpHBwX7A2wn/rxf4Uw2veW/C/8vT4GC03uDzbsORtBsB3Ie/w/QA33IaUfqcgP/+nOQ4FpFmLYelUQoeKPKUWrSRwUGI5+P2LgfU8H/zPjgYnTxyrttwRKQKdRh1zhaEa14c5zYcKahBhDswe7GsCHnR6ODg1xL+Xy/x1FNJ8j44GK3NU0tHmoikw0hs0rr3/e3GaohK6+2L9ZF6faW19A2ItNoK+Puhty+u6TSi1mpkcBDgzIT/1wvsXsP/vTfh/+VpcPBmwtt2pttwJM0mAM/g7ywLgM+4DCilgoOneRpEkeKKDmblabCn0cHBVbBBsODlKqzDrZo8Dw6OJV7seWunEYlIJWOwAuzBDqMjnUaUf8HZvN3Arm7DkYKKDmblabCn0cHBUcTbdldjk2OryfPg4FLALMLbto/TiESkVqtgK0C87+5cYA+nEeXfj/Hf7zmovIa4cS/h4/YZTqNprUYHB9ci3sa7jNoy5dyb8Jp5GRxcmfiCkI2dRiSptS6WcsbbUWYC2ziNKJ1GAIux9+gtlEpP8iG6xPwjYLzTiFqn0cHBZuR5cPBnhLfrWfQ7KJJWSR1GtcyclOb9Fv99fx9YzW04UkDR9k83MMlpRK3T6OBgM/I8OHg84e16G+jvNCIRqcUmhNOZv49lMJD26oPV7Pbe91eoXs9MpNUOIXzsnkt+9sNGBwebcW/Ca+ZlcPDXhLfrcbfhSFptT3i24JvA+k4jSq8D8d+nCxzHItIq/YFphA8YZ7kMqIU0ONg6Y7DZkcHt+rLTiESknGiH0Xuow6iT+hM+yfwv+TnBlGzoIl5j73KnEbWOBgdbZylgOuHt+r7TiESkFjthtUG97+3L5GcCSBYsBTyF//7fhep4S2cNwhasBI/fP3EaUetocLB1VsQWfwS3a4rLgCSd9iO8ozyF7TySLHhiqHQrkidfJnzAWASs4TSi1tDgYOtcQHibpgODnUYkIkl2Jt5hlIff86wZib333udwHVppLZ11AOHjdg+wqdOIWkODg63zU8LbNJva0qyKiDtTsHN173v7T2wSp3TWGsAH+J/DL9yGIwX0LcLH8I+wcmFZp8HB1rmC8Da9BgxwGpGkznHAEvyd5G5guNOI0q0P/iz8hdhsIZG8GISlxAgeOP5C9jsyNTjYGpsRz1P+VacRiUiSKcQ7jEa7DKjgNgDmoRU54kYf4AnCx+5HyP7qBg0OtsaaxGeTn+I0IhGp5nhsoof3nb2TfHRgZ9XOhM+Rj3AbjhTMMOKrB69zGlFraHCwNbYlfLzoBb7gNCJJlS7gdMI7yJ+xwQEpb3PCjTCRvInOMO8FjnQaUfM0ONi8gYTTpvRiKfJUj0YkXaJ1o+5EE5nSYD/8E7MeYH+34UjBbEe8TXKi04iap8HB5vUB7ie8PW8AQ10GJSJl9QXOJfydvQSdj6VBsP39ETapVqRTphBvnxzsMqAW0OBg84YALxDenn9jxxIRBgJXEd5BzsZOEKSyU/Dfs687jkWkHbqAvxL+fZhLtmuQanCwedF0oj3Ajk4jEpGgvsBvCX9PL0YdRmlyGv5nM4dsDypI9vyR8O/DQmBrpxE1R4ODzYumE+0FPuM0IhEpZwhwE/E+vKxn+MmTK/E/m7eB5d2GIwXSB3iA8O/DLCw7QFZpcLA5XcTTiS4BtnQZlKTHMoR3+B5slovU5jH89061eySvViZcq6oXmEZ209KtDVwdubR7JtWJCa+5TJtfs12+SryR9FunEYlIUFKH0amowyht+gA3439GLwPLOY1IimQ5rE5w8HfiPWAVl0E1YUXi7ayj2vyaX0l4zRXb/JrtElzN7F1ucBqRiJSzLPB3/O9qN3C004gkyWAsbbf3OT2ELcwQ6YQ1gPmEj+svY/XPs2hD4m2u/dr8mlMTXnNwm1+zXU4g3od3ptOIJDXGA48TnjF6kNOIsmUs/knUS45jEWm3KcQPJveiVShFswOwmPB+8BzZnUElkjfqMMqWpYGn8T+vO8l+7TfJjj0J15r30gspjWSxfJx4B+IbwBiXQYlIoonAs/jf1QUoNXmarQS8g/95XeI2HCmYLxPvw7sdpZEsmt2Jt/efxCYUS8GtA7yGv2PMxgrnSu0Ox3//znIci0gnnE28cXEBWo1SFGsB75Ov9BQieZLUYdTuGZXSvEnYb6n3uf3cbThSMN8j3rb7MxqkLoqVgNcJf/4fAZu4DEpEEq2PDdx739WZwFZOI5JabIEtxPA+t6+6DUcKJloOphf4pdOIpJM+BnxIfjKFSAtNBj7A3zHeAjZwGVBGXYP/HmpgVYqgL8m18y5ANUrzbi3sWBHNUb6Hy6BE5H8+TjhF4PuowyhLdsFWeXqf3+Fuw5EC6QL+QLxtdyNKf5Z3K2PZb6Kf/eddBiUiiXYg3MH7CpqgmSVH4X92i4Ht3YYjBdKf5Np556JJ/nm3ITCD8Oeu3x8BYF9sNqC3YzyNzRiU+vTHn+U9FxjkNhyRjlkWeIF44+JKNMs8r5IaFb3At10GJSL/syPqMMqDkwiv3NnUbThSIIOBR4kf529G5zh5tQbxFYO9WH1aEUmXQ4BF+N/T/wArOI1IGvFbwpP4VnUbjhTIcli9wegx/3w0yT+vPk4861cvcIzLoCQdvko4z+xD2I+E1G97/PfxesexiHTamsRXkfUCV6EahHmzGZayJvpZ/8plUCLyP4cS7zBa3mlE0qjoCq630GcpnbMSyavIbsMGDyU/1gHeJnmin+oQiaTLcUAP/vf0r8BwpxFJo6IruJ7Gak+LdML6wLvEj/0XoWN/3myNlY6LftY/cxmUuNcFTCW8U1yHTvSacQb+e3mk41hEXFid5BnH9wPjHMYlrfNZYB7xz/jXKAWFSBpEO4zuQp0MWRddwfUgSu0onTMW66yMHvf/jdU0lezbnXB5Ee9yBcoAIpImfbHJmMHv6WXAAJdBSdNGEl7BdS06r5bOWRN4k3gb4F5gjLuwpIUOBeajzBASMQBr7Ad3inPQ0uFmPYP/fiotqxTVysCLxA88b2KFtyWb+mGNh+jn2guc5jAuETF9sUH6aIeRVm7nw8qEZ/Ze7DQaKZox2Ark6PH/Paw2pmRTF3A84SxCSismkk4DgT8S/p6ejb6nebEB4Qm433UbjhRMubTir6OSBlk2EDiP5D48DQwW3DDgdvwdogdbQSjNmYj/nj7uOBYR11YiuQbhQuBoh3FJY8Zhqz+TGhU/cBiXiJiBwNWowyjvtsKOo95nrPoQ0kmjsHOcaDugG/gOWuWQNcsCt5Lctvsl+jxF0mQEcB/hPrxvOY1I2mE//OwfPaXrIp2yKjCNeJvgI2CKs6ikUSsCDxP/PHuA/3MYl6TAOOBf+DvFYuAIpxHlx7H47+tPHMcikgYjCU9ECF5uRgXTs2J/4B2SG4mHO4xLRIw6jIol2N5cDGznNhwpmGHEJyJ4l79hM88l/XYDXiX+GS7GVhKKSHosDzyB/z1dAHzGaUTSTqfhf9azgXXdhiMFsxxWkiKpnXcTMN5daFKH/YEZxD/DOcABDuOSFFiL8CyAOcCuLgPKmdvw39stHccikhaV0hXNwmpzamZyOo0FriG5YfgasIm70ESkZALhlObqMCqG8wmndVzFbThSMF1YbdPFxNsH87F2X19n0UklIyifXupdNNlAJG3WJZzqbyawjdOIpN36YBOpvc/8ZWzStUinVConMxPrw5N0Gg9cT/Jn9zyabFB4mxMeNX4b2MhpRPkyGL+450xUuF0kal9s5lvSQeou1LGZJl3AF7HB26TP6zYsFZWIuKUOo+Lqj63S8j77x4GhTiOSItqR5FnJvcBDwDruQpMEBxKuWxq8/B3LMCQi6bEd4fOxN4GPOY1IOmVp4Gn8z/4ONOlGOu8gYC7J7YZbsVJCkg59sHIT5fpcbwCGO4tOUmFv/IGrXuBFYDWnEeXPnvjv75WOYxFJq0nAgyQfrD4CzkCz4lzbCXiU8p/RCaiGmUgabE+8w2h9pxFJp40hPDh8DVqJL523EtZpmdRuWAz8FqWgcm0Lwqmno5/RT7AJByKSHp/Gzr287+pTWP0oKY5JhNv6p7sNRwpqXeCfJLch5gE/BZZxFp0AfJLkmuDeZ/R1dI5YeEcQTvnyD6yYvLTWufjv8eccxyKSZl1YGoJyM1rmYCkMlnYVYEF9HLiT5M+kF3gAS00tIu7thzqMxGyEnfR5+8KJbsORAtsfS3FbrmPiVNR51GlrYfUhe0j+XJ7A2n8iki7HES7JcQ/6/SyqXYBu/H3h827DkYLqh/0ulVtFOBNLKT/YVYAFtSlwN+X78O7HJhlIgXUBU4kvIx3iMKY8m4a9x0uA0W5DEcmECZSfad4LvAV8Bf1mtdv6wB8p33E0BzgWrRYUSYtoh9HdKEVI0R2Mvz8swbJZiLgwDriW8m2794D/QxPA2m0N4CLCHcrRTBAnotWCImnTha0OC35frwEGuQxKnDuJ8O/3Jm7DkQJbHbiX8u2814Evod+sdtsIuI7yn8MsbEGGVgsWXD/gAsI7x+9RHbx2WQ//fX7QcSwiWdIFHAK8RvkD2/tYqoLlHcWYR32APbBaj+Xe9x5stvkENyGKSERSh9Gf0cmXmDPw94vZqNabuLUv8Dzl2xizgbNQvelW2wG4mfAEkujlFmBNVwGKSFkDgasIf1/PRhM0xc4B/oC/X7yJ0nWLO13AF7D9sFxb413gFGCsoxjzqA+wD+Ga89HLEuAKYAVHMUqKDMMKgwY7eKe6DKgATsB/v09yHItIFg0Cvkn5dFS9wCLsQKeZco0bCnwZeI7y73MvNmioNFMi6aEOI6mmL+H2/3MoBZm41R84GssEUakT41pgG0cx5sFALM3cE1Ru2z2I3meRtFqG8Goc9eFJ1GDgUfx95AHs91/ElSHAd4APKN/2WABcDGzgJsRcWAr4KvAildt5t6L3WUrGEj5gdANfdBpRMQQLvG/oOBaRLBuG5SovV4/QuzyNnTBNdBJltvQBtgLOAz6k8vv6JFYzSETSI6nD6HiXAUlqjQBewN9X/oINGoq4NAT7zarUedQLPIO17VZ3EmX2bIxNEnmX6u/r/ii1lEhajQf+jf+dXQh81mlEklYrE/7Nv8htOCIALIvVlZ5P9T6847EU9FJZX/w+vGp9ow8D27kJU9JoVcLpW+YCuzuNqBhGAIux9/wtdOIl0gpjgR9TeSWhN+P8buBwVHMramPgTCrP2Pcu9wB7od8vkbQZDzxOuMPoIKcRSdqtidWZ8PaZU92GI/I/I4HvUr1d0gP8Hct0MNJJpOm1LpZq/1Wqt+0eBg5EEwRE0mwdwt/nOcAuTiOStNsKOx/w9pmj3YYj8j8rYCUwqk0G68YmMB6KrYoT32bYxK93qN7OuxP4JOrDk4BNCc8geQ/4hNOIiuNA/Pf9AsexiOTNQKzR8BTVD46Lsc6k47GBsaIdJAcDO2Idwf+l+vu1EKspuLmLYEWkqnUI12OdDezsNCLJir3wa471oAFlSZcB2Eq2f1C9rdKNZcWZSjHbdv2wjuBTCWcHqjRp7iasPSgi6fYJwhNh30Ip4aQ2x+LvN4uAyU6jEQkbBhxJbX1SH2GDXF4fXtEMwdpsZwPTqK0P71JgfQexSsrtCczD31leQulYOulS/Pd+H8exiORVF7AbcAv+St1ql1eAc7FO0jzOPO+LzSD/OnAH1rCq5X15A/ghSucgkmaTCc+6VIeR1OsHhE+8Va9X0mgycA3hVRCVLm8CvwP2A8Z0Pty268JW/x4D3Ez4HL/S5R3gZ1jKORFJv30Ip+D7L7CS04gka84nvDhkFbfhiMT0AT6FrRLsprb2zEvAr4A9sCx9edMX+BjwLeAuam//voqd243ufMiSBZ8n3FH+CNpZOqkP/nLfhWhJtEgnLIvNRPo7tiKiloOp17l+EzYraStgUKcDb9I4bDLIVGw73qf2bf8Qm8iwJzYLXUTSa1/Cg/1Pow4jqV8X8EfCJ5U6R5C0WgbLFHEnjbXtpmIzrwd3OO5mDcfinoptxwxq3/aPSv9nf6B/h+MWkcZ9FX91fy/wELCc04gki/oD9+HvR48DQ51GJFLeeOA4rA+v1naO1867uvR/t8Iyi2XJeMJ9eDOpfdtn4ffhKUW8lHU84R3nDjQ41Wmb47//dzqORaSI1gBOJlxvtdbLAuCfwBVYDZwDsJU5Qzq6BXErANtj9QPOAm4jnFqwnk6jG7DtylpnmUhRqcNIWmkY8B/8/ekBLKWjSJqtCpwEPEn9bZ9FwL+Aq7COmIOwNFXDOrkBCcZiqySPBH6OrQp8mca273bgENxvk4jUpwv7XQp+p69D52nSuLHA6/j7058pXvptyZ71gB9hqwTrbQfNxxZFXYa1FffHVuK5/h1dEZvs9WUsRejtWMauRrbvWmyycOoGQvXjki59gd9gJxeei0vXF7sIqMBOAb5X+vsbwC8cxiJSdKsDu2LpRyfTeAOhF2tkv4qtDJ6O1XSdXro+o3TpAeZiv7sfYYONUctgx9ClsBV7Q7CVG2NL/44J/D0eWI3mOntewhoitwH3YI0LEUm/LixVyA8Ct10PfBb7fRFp1ARsIow3yHwuduIqkkYrAbdik2BvwdJk7oa173aguTbSm1jK+RnYTPQZWLvubfy23RIsleciLCtMUjtqOJY9Zhi2cmMQMAprx40O/O39u2rp/zTqdaxddzuWgmpOE88lIm4MAC7C2nWeX2GrYXqcRCR5sRFwP/4E5+9gdWpF0mhtLJ38IVgt5UlYO283YBsaz+7Vg02ofw2/D28G1sYL9uH1Yu2obsr34XnpTJfGxl+GEO638/4ehU3sX5XmVu2+gLXzbgP+hs79pQZDsZmGwZHlU9EAriuP4X8OaziORUR8g7GOpLOwGeS11ils9rIIS+HZidfqxQYtb8JWG6nWrEg2DcBWMAe/2+dgnc8irbAD4ePgl9yGI5JoVWAa/szpbSP3D8AyK/wMeJja67Q0e1lMuAZsuy8zsQ6ibwHrNPA+iki6DMO+0953vAdbQSjSKgfj719LsHptImmzCX5pnBnE2zhDgN2x8+DH6Vwf3kI624c3HcvwdQzW9hUJ2bzK/SOxdEDeDtUNHNXuoKSssfg1MV5yHIuIVNYfSyt1HJa3+2k6d/Bv1WVRKe7zsLo866CJISJpt1mV+4dhq0HUYSTt9nXCx5Nt3YYjEjKJcPqlV6k+6akf1hY6Er9tV0+9wjRcFpfivrS0HeugiSEiWVKtD28cNlE1+J3/QruDkkI6E38/m42t0BJJiy0JD8A9jf0+VjIUqzOY9T68R7FUo+rDk6pGAjdWuH8i8Bz+DjYPK0op7hyO/3mc5TgWEanfGGA7bAXFGVgaqxexiRcuGxCzSM6jrjpRItkyFBv4Kyepw+iIDsQlxfU7/P3tPez8QsS1tbE0n96++QqwSoPPNRJLS/UFbIXhjdg59CI6145LuszBOoeuxNJHHwhsSOPps0TEvXFYjbdy1sJfDe39DuzW/rCkoPpi/Rne/vYsVuJExLVtsQFrb998DL/cQb3GYVkkjsLKet2GLdZx3Yf3AfAP4BLgRGA/rLZi/wa3M5U0qtl+l2H5andNuO/jWM2F0aXrM4FPYasIxZ1rsCKhALsAdziMRURaZwC2vH8s1vgYXbqMw36nRwHLEq43M5jkDp5Z+CeD3ViarJmE6xcG/36ldF1Esu8sbJbkJgn3rYWdzKxcuj4XmwhQaTBRpFmDgHvxV7Q+ju2jqk8rrmyM/e55nUTPATtiqwhbqT824JjUtvPqBC6Lda4OxdqCA/FrOAV9SLju9AKsbResUx2sd/MqVu9QRPLlWuw879MJ922GlQPyftumY+ny/tWZ0KSgRmCTjFcrXf8Ltt8tcRaRFN1uWN/54NL1B7B98sMWv85AbL8fi18PMFgn0GvndQFLYdknyvXhfVD6dzb23fH68N7G2njvBv6eAbxc+lukKTtjjYo/Jty3I+ER9pextCviVn/8Tv+5aNaniJj+WOFiESm2TbGTibsS7tscvyB6L3ZysVHnQpOCG0c4feOf0ERQcaORFFMu9EOrL0QkbG/sd+vChPv2wjqTvd+2F/EHa0TabU3Cx9afuA1HCmxP4CP8ffFebGAubQagPjxxbAi2UqQXOD9y32GEU6D8B1i+o9FJOdvjfy7XO45FRERE0qMftiKrF5spGbQ36jAS9zbHVjt5++H/uQ1HCqiVKaZERDppafxUyL+I3HcEtqLY+217GFuxItJJe2GTFHuxle4Hug1HCuhAwuMZt+KvHhSRiNPwvyw/D9x+POGC6nehkew0OQP/sznScSwiIiKSHsfjtxEuCtwe7TD6B+owEncOxd8Xl2ApfkQ6YTfCkyT+Dgx3GpGISO1+hd9Xd3Lpti5gKuEaVDeQnJpYpBOm4u+L87FyVSKdcDDhc96bULY9kbLWJ/yF+R5W5+A3hBsVl5KzIpY58Az+57OS41hEREQkHSZgJ+A92IDLWajDSNLrl/j75IfA2m7DkQLISoopEZEkmxGexP8NLGPE+YTbeReWbhdxpQsrXeXtk69itddE2uko/FWrvcBVaDxDpKw+WIqBbsINiz8RblScjeqApM1E/M/nccexiIiISHrcSbht90PgAsJtu9+jDiNJh37AX/H3zWfRCi5pH6WYEpEs6wc8Sbid92XgFsLtvFNdBSgSMQwrTxVcqT/AaUSSZ98m/Ft4GTrnFano64S/NL3A84G/u4FjnEUnlRyLivuKiIhIWDBNo3cJZhrowVYQiqTJsljtS28/vR3LZCLSSkoxJSJZ913i7byXCPfhqeSMpM0EYAb+fvorp9FIXgXLavQC52KLokSkjJWAeYTTEQQvC4ADnEUn1dyG/1lt6TgWERERcW8k8B7hNCrBSzfwRWfRiVS2PjCX8IpXkVZRiikRybrVgYWU78Obi2r3SnrtQHiCjgaxpZVOIfx7+DOUATGX9KG21i3AriSPoi8q3f8cMAur/zG7dPH+fgGrZyOdNxh4v/TvB1jO7m6nEYmIiIhrlwCHkNxm7sbSjf6HeNvOa9+9BMzpSKQiyfYBrsH24V7gIKxWjUgzvg2cHrh+OfB5dP4kItnRBdwDbEXyyvqF+H14H2JtvWg77wWs3qqIK18Hziz9vRjYEbjPXTiSA13YPvW1wG2nASe4CUckOw4geaZRcGZ5N8kzkh7FTty1NNedPfE/jysdxyIiIiLu7Uj5meTV2nZPA59D9RgkHX6Iv2/OBzZ2G45knFJMiUgefJ7a+vCS7vsH1oekBReSBr/D3zenAyu6DUcyrAtLURv8vfue04hEMmIE8C7lU05FL97jHgR2cxCvxJ2L//l8znEsIiIi4tZg4GXKdwqVa9s9DuyLOsolXbqAP+Hvr9OAUS4DksxSiikRyYPR2ErAevvw/oZNHhNJk0HYgLW3v/4LGOI0IsmivsDF+PtRD+HVgyJSwflUnlmeNCi4p5NIpZxp+J/RaLehiIiIiGM/pbbOIm/w8CE0g1zSbSngKfx9935ggNOIJEu6gF8Q/v071WlEIiKNu4LaBga9dt4DwPZOIhWpzTjgDfx993K34UjGDCA8kbAHOMZpRCIZsg3VBwa9BsVdwOZuwpQK1sP/rB50HIuIiIi4tR5Ws6OWtp0mfEmWTATew9+Pf+k2HMmIPoRTlinFlIhk2a7UNijYA9wMbOomTJG6bQ4swN+Pv+k2HMmIgcB1hH//DnMakUiGDASep/zg4JLSfTcBH3cUo1R3Av5ndpLjWERERMSdPsDDlG/bBSd8fcJRjCLN2Ilwutwvug1HUk4ppkQkT4YAr1JbH95GjmIUacahhPfnT7oNR1JuCHAH/j6zENjPaUTScUVMfdQXq7ExClt2Pbp0GQsMB4YB/bGczYNLj1868P8/wmZidGOzbyclvEZv6d9bge9i9Wckve4Dti79vRHwb4exiIiISH26sLbcKGAM1qYbVfp3BDAUS5UyoPR3H6zN51kIzMdOoMcB6ye8hte2uwf4PpZeSiSrvg2cXvp7MbADlmZUJGgAlnrP6yTqBY4Ffu0sIhEpor747bxxgb/HYymzlwL64ffh9Svd5vH68BYDqwOrJbyGN/nheuBk4Mk2bIdIp/wSO/vUx3wAACAASURBVF4DfABsBrzgLhxJqWHAjcB2pesLgQOx30EpkLwODvYBVgbWKF3WDPy9Qun+Tnof+yF+rnR5vvTvC9iXT9wZAbyLNSDfBpbH7wAUERGRdOgCViS5bbcS1nHUSbMIt+289t3zWCeUSNpdCHy+9Pd0YBOsVo0IWIacPwJ7la4vAY4ALnEWkYjkWV/8PrxJpUuwD6/TfZfvYe28Z/H7754DXgQWdTgWkXr1A/6CXyfzWSzl6IfOIpK0WQa4Db/02Xxgb+BOZxGJNGkN4GDgHOARrFOmWg7xNFyWYA2Ny7FZHZtiMzSlcw7E/zwucByLiIiImInYMfoXWF2/ebhvt9Vy6QFeBv4AfB3YApvJLpI2g7DzJm/ffQxLLSSiFFMi0k5d2CSvQ7GVyI8RrpOW5ks3NtByKXAMsDGWeUwkbZbFBrO9ffcGOr9QRtJpNJYxz9s35uCvHhTJhL7AllhKp1uwGT2uGwitvCzAOsF+AexDOKWptN6l+O/9Po5jERERKaI+2Kql72AnrtNx3x5r5WURNghzDnAAlrVAJA3GA2/i76uXuQ1HUmAYcDfhc9O9nUYkIlnXD9gGmIqtVJmJ+7ZZKy/zgb8DZwCfwn5HRdJgfWAu/r56sttwJAXGYmmTvX3iA/zVg1JQWUkrOh7YFdgF2InmO1Xew1JJzgDeKv39bun2OdhsoAXYCsQlwOzA/10BW+kHVlPwNmym0DAs//kY4vUM+zUR62Ksrs3tpcsTTTyXhPXBUomOxjrulsM+fxEREWmvUcDOwG6lf0c1+XwfYIOKM7Bju9e2m4GdFC/Cry3YQzitzkjgKqxOzV+wyWdgbTuvhqFXn9pr5zWT6WEJ8DDWhrwd+FcpJhEXtsAGgwaWrn8Dm6QoxaMUUyLSKitgfXi7AjsSrvXciBn4bbu3S9ffwUr4eH14Xm3BbsL9OhOxyS+LsDbe7djqea8Pz+u3G4+18UbRXB/eQmyw0OvDe6qJ5xJp1j7ANVj/fy+WmeVqpxGJKysBd2G1V8F+R3cGHncWkaRCmgcH1wA+h9U5WJ/6Y32PeA2YVuQI/w3WIPkFteVr9mrkeDnTg/nTV6L+Zd1vYYOSVwJ/Q51JzdgceKj0913YwLOIiIi0xwTgs1hn88bU3wb6AGvPefVfvLbd8zRXw/k07GT5dGw2ey1WwK+FMylwWZn66x++i3XI/wHrhF9S5/8XadYU4KLS30uAPbF9UopjNJZK9GOl63OxFTD3OItIRLJmbayd9ylgvQb+/wzCbTyvzfcSNmm+Ub/D6geeg016qKYP1lcXrXM9Cevbq7dv8nVsUPJKbNCwt87/L9KsHwEnlf6ei00Me9JdOOLABOCvwCql69OxgUHtB5I6Y4CvYrOp61nGPwvrTPkhsAd2ctMOfbFZ5a0yBNgK+CY2c+NV6tvu17EOrfVbGFORnIL/Xn7dcSwiIiJ5tCzwJeA+bEJTrW2cOVin9KnYYOL4NsXXBQxt4fMNxCYfHYd1Ar1EfW276cBZWJpVkU76Df5+OBNYzW040kFKMSUijRqH9aU8Rn3tnQ+wVXUnA5/Esji1Q3/8lfGtMBRLkfptbDXWG9S33dOAH2MDqSKd0ge4EX8/fIXms7ZIdqxJ+LfqVfzVgyKp0A/YH1sNt5jaDqjvYqk9P48dVPNUVHUc1gl2JvAMtTcy/oMNMqqOTe2CDdg1HMciIiKSF32wyVrXYSv6amnHzMQmSh2JTXqqd/Vdmo3C3o/TsPTwtbbtnsVm+bZr0ptIUH9sQN7b/55Btc+LYCVsdY73ub8DbOA0IhFJu/7AQdhq425qn/x0CXAY1lGd5ixm9Voe2Bc4m/DvabXLv7DJZM2mXBWpxVJYiltv/7uL5tLnSjasjWUgDA4Mr1Lxf4h00NLYgfAVqh80lwCPYrPHd6RYP2BjgUOxDrNZVH+vPgIuRTORqhmLv4LhJcexiIiI5MEgrM3yNLV1ijyN37br7yBeV0ZjE+MuxQZFq71PC0uPVaYIabeRhFe7Xk++JmEWTbVz5gmEP++3gXXbHJOIZNdwrA+vloxX3fh9eFtRrGPJRGyy29VYKaJq79Vs4Dxs0FSknSZiJbi8fe8st+FIk6q18zYm/Hk/i01mEHFuNeCXWLqoagOCdwOHYympBAYAu2AdRLW8fzcBOziJNP0ORwdEERGRVlgRWxVXbaCrB6u1cjRaDefpB2wHXICl2KrWgXQXtgqxSJ1s0lkfw+rRePvc1AqP7UdrSy5I6yyP1f0qRymmRKRWk4BzgXlUHxC8A5sotoyTSNNnILA7lmq+2vu3BLgW2NZJpFIUOxFe8fuFCo8dRr5W+ebJBlROAb8V4ckJT2OZCkWcWhlLI7CEygfEJ7D83Su4CTMzhmDFnm+hejrWh7GOJ/Fdg//+7Ow4FhERkSwai9UpW0TldsizwPdQCpNqBgGfxtKxLqDye/ofbJBQpB0+jZ9howc4oMzjVgSO6FRQUpcrgP3K3KcUUyJSi1WxQa1qNaMfA76BOp6rWQobOP0L1dOx/h3r3Bdph/8jnKFk6zKP2xIbTJT0uRfYtMx922Irkr3P+FEsO4iIM8th9fMqdXLMx5bRK11SY0YDxwOvU7mBcRuqIQGWusxL0ToX64wTERGR2iwN/JDw6qLoZSE2KazcSYtUNgL4KvAildt29wFbOIpR8u2n+PvZHGC9hMdsBjzZyaCkJltgnfl7J9wXTTH1DEoxJSJho4FzqFw3ei7wK1TOplHjsLrSwYkaSZcbUbpnaY8L8fezt0leoLM/tg9KunwG+9w2SrhvN2yMJTjRQDXExZkh2MGuUp28N4ET0Qh2q/QHDsRWClZKVXAFlmu6qLbHfz+udxyLiIhIVgwAvgbMoHw7411s4FCzx1ujD9bBfy+VO4+uB9ZyE6LkVB+sREFwddlykcfsU7qv3Ixz6bw+2CqeXiyVXZBSTIlIJcOwVNKVSti8hq06GuEmxNwZAByC/7uddOkGLsZW64u0yiDgEcIrgKOp4o/D+pCVXSA9hmBjKb3EJ+7tSXhh1r3YimURJ7an8kznJ4GDsQOhtMcWWFqqcikg5gHfBPq6CtChM/DfhyMdxyIiIpIF3gqhcm27F4Avovpj7bQRNsGrXIr+hVinntrX0ipLYQNI3j52F1Zn0POV0u1Xdz40KeMI/M8rmApsW5RiSkTK2xWYRvl23r+x1Sr9yvx/ad62hCflRC9zgGNR3WlpnfH4A029wKWR+08v3f7zDscl5Z2C/3lNCtx+IOFSH7ei83JxZDhwNuU7LV7DBmOKOCDlyibAX6ncyEtaipxnz+Bv/0qOYxEREUmzwcCplK+NMgNLbT7QVYAFtDY2GFOubfcUlQvUi9RjEvAB/v51ZuC+n+CvatCKBveGE17Z7dWcj6aYuh+lmBIRswxW4qfcpPJXUR9ep21O5YwRD6J0rtI6WxBebXZc4L4rSrfNBoZ2PjSJWJnwZ7Vq6faDCZ+r34jOzcWRvQjPOAhe3sMKFKu2mzu7YgOBSZ/PIuBkijHTfCL+dj/uOBYREZE02wF4ieS2w4fA99CJokvbYB1ESZ9PNzaIM8RZdJInuxDudDiidPvF2KTQHuDHTiKToDMId/BviVJMiUh5BwDTSW5HvIOtUitCH1FafQqb8JX0+SzAyjj1dxad5MkUwucQu5Zuvydw+5ecRCZB1xBejLUicFTktqvQ74I4MAArRJx0wFqCrSQc7iw6CeqD/ei/T/Ln9TD5n/V7LP72/sRxLCIiImnUB/gRybPIe7AC9tHaY+LOfsBblF9FuIa70CRHvoO/X32EpRq+C79DYiaaCOrSaoTTSfViHceLA9eVYkpEwH6rf0dyu2Ex8DM0iSAt+mIrN4Mr+IOX+1HtWGmNc/H3q/exVWneJNElWAa2LmfRyXbEv/8nR65fhlI/iwOjgbtJPkg9j+XMlvQZjeWSTvrcZgA7ugut7W4jPJtWREREfMsCt5PcRniZcA0rSY/hlE8LNhvY111okhNd2Gxkb796i/jK4imughNuI17aI3hdKaZEBGAF4B8kt/P+A2zqLjSpYCzwZ5I/t3fx00iLNKo/4ZWC/8UmgwX3te2dRVdsfbHf53JlPnqxwV3VI5WO25LkWcqLsNo0OvlIvz2A14l/ht1Y/aC8zQoZjF9vYyaaUSEiIhK0AclpRJdgA0/D3IUmNdoGeI7kFZ+noppBUl0X1sm4IdYZGTwfGAz8k/A5Q/Dvf3c0UvHsTvnOol6UYkpEzDYkpxH1+vCUQjT99iS5nNNirA9PpJoB2GTP9YhnghmJTQZNakt0A9d3LkwJOJrK7bzT3YUmRXYY8bQlvViH0gYO45L6LQPcQPIPzKXkawBtT/xtu9JxLJItSwMjgPHAKsAkrNNsldJludL9mhQhIlm1F/4EmuDlTWArh3FJ/YZiaWWS2nY3otSPUt0awCP4ncZvYfUt/4ylFZ5D+Q6KrGTmWApru43D2nKrAxvht+1Gle5PexrOAcCLVJ5NHl1J/CGWou4V4BvoN0GkCL5EOM2wd3kWWMdhXFK/5Sif5eM8tHpIqtsUfzLhQuBV4O/AH4DLS7cl7V9LgAmdD7chw7F23PL4fXgb4LfzRpKNPrxlsTZbNDtE0qUHa+N57bxnsTrheerXlzZoZGXYF4HfEj/g3AZ8DtsBJVu6gP8Dfkx8RvlNwP7YwSHrzsWKtQIcDFzhMBZxqz8wEev8GofNjB9V+ncMlnp3DNZYaMRiLEXvu8Dbkb/fBV7DDtTvNrwFIiKtsz92TIyuLHmgdN/bHY9IWuFI4BziKwH+hmWPmNvxiCRL+gHfB07EPz/oxs4By3U8LgH+BBzU9uji+mIdVl7bbjzWthtduu793Wi91G6sPTcDvz0X/Pt1rKPtrUY3oEHfpv4Z495Eye9jKwREJN+OBn5NvP/vZuAQYFbHI5JmeX14PyF+TP4D9rl2dzooyZShwM+xiQPeb0O1dl4vVpPUxSrV/tigXrk+PO/vZRp8/kXE+/DewVZbzwCmYe28GY1uQIPOAY6hvvGbbmyiwI+w+EVa6ijitUy8NEWanZJ9u2KFZ6OzD24hHzNKp+Evh2+0Y0CyZQg2g/2LWCPmRuyAnrTy2cXlA+BhbJXuSVgn/CTyl9JXRNLrQJJnkp+H0kvlQbkyAPdhK6dEqvkEtsIsqZ5l0mUxNjDXLgOBzYDDsXPQa7G6OOVmuXf6Mht4FH/w7UBsVU47zpXHYKs4a/lsvJWFd2HZL0SkGL5N8uoS9eHlwx5Yn0L0M74apZOW2uyODXjVsjKtF5tMMKSN8QwFtsYmOf4cm8TwPMnnqy4uM7G6rRdjE+j2A1ZrxxsBrE3tmSG6sd/2q9sYjwjfIr7zLQD2dRmUtNyqJNcbuo30p9WpZD38bXnQcSzSPuOxAbazsbQIC3DfeGjk8mEp/lOxdLijWvkmiYiUHE78RLAbSz8i+TEeK2IfPdY8gKXcEalmMNa26qV651EPcEoLX3s81hY6FWsbfVTl9dN6mVOK/2ysrTqmBe/NRVT/PLxOpX8A27bgNUUkO75P/DdhPjYYIPmxJpYWMvpZX4cm+kltRmH7Sy3tvF7gCy187VWAQ/H78NIy2avey4fAncBUrN3aigUpd1B9cFCTv6RjDiG+Ay4APuUyKGmbscBTxD/za8juiqYT8LfjJMexSOusgqVJuYHkGXOtPNDPxGpvvYSlBP1X6e+XsJlWM2lfh1VP6TV/jc0OHNqKN09ECm1X4icb3VibT/JnBLZSPXp8uRPVoZDa7YqlWKrUcdSDZSJpNOvIClin05+x9lW72nazsbbbW1hb7nngMfy23bul+5Nqsbbq8hJwPjbZduk636eNqbxi0PuMnsEGI0WkWI4m/rswD9jRZVDSNith9Wejn/mlLoOSzDkU+52oNCC1BHiaxvuGVwO+gmWom1XhdZq9zMLacW/g9+H9G7+d917p/nYtKOjBMlv8EtiN+hfbfLrK82vyl7RMLV/mjYH7Ce/I84G9sQ4Fyacx2Oe7XuT272F5i7PmPmxpOsBG2EFBsmcwduDbFTvArtHEc72FpRh9FcvD/Q5+7Rjv70ZrAg7E6toE85+Px697s1op9kYH+RZiv8u3Y6t6/9vg84hIMa2JnUgEV40tAj6LTQSSfBqOHTM+Ebn9bOBrnQ9HMmo08DtsVnQP5dPSHQpcVsPzDcDa6LuWLus2Eds7WNtuWulvr2bMdPyaMTNKcddrAH49m2idm7H4bbt6B/k83dhq3ttLlyewTp8kXaXHbk78fL63dNsbwA+B32MdeSJSHFsBfyW8amwe9rt9j5OIpBPGYauH1o7c/k3gzM6HIxk1AbgcK03gtSmSTMbqmFczBNgO67/bheZSXr6BTeZ6jXC/3Vv4taAbrQk4iHC/XbBm9Xj8dl6jKVUXYO+X14f3XJVYnscmzEXff6/t/QzwA6zWt0hbjcdWykRnG23jMqgGvUD5EfdWfZmWpfKsg6ytWBtFPA3VEmAvl0E1YAR+fuq3yO7qx6LqhzUkLgfmUv+MneeBq7Dl/QdhEx7SUmdpRWAHbGbnL7AB+UZmTz2HpY1ZtbPhi0gGjSDeJlqEdRhlzUOU/11sVefXICqvTM9aZ8vSWHr16HYc7jIoySRvdnnSKsIeKk/E64N1FP2exto9L2M1VX4IHAxsQnpS5I7DOsy8Wjm3YbPT693GacCPiXfygn1fy/2/94HjsYlqIlI8K2Od5MHfhdlYrdasSaqZ7F0ubtFrjKfyKqmvt+h1OmUctkIqusJoF5dBSeb0w9oSi0n+fiyh8oTS/ljGq6uoPwNDDzbw5dVu/gyWLjMN2bO6sFW6OwJfxiZY/hXLNFZvO++/2BjBhITXOSXh8V62iNexNmbf1m+eSNwg4umHerAvZhZVGhxcSGvyAR9b4TV6yd7gIJRvXK7jMqg6HYgf+wWOY5HarYPVl3mb2g+ws8lHrb5o3vV6Uh08ChxHa2rZiEi+9AX+Qvx34xiXQTWh0uBgD62ZMPHZCq/RS/YGB8GOjdMIb8cC4isKRaqZSPJgs3eJdkavg03WernC/4le2lGrz4VoXex6UtE/jb1vE4Bh2HsS/b2bi7V/0zIBTkQ6byjwOPFO/KyWA6o0ODiX1vzeBcvPJF2yNjgIsDqWLjG4HTNLt4vUYxNssn3Sd2MJ8YGtjbF2zvQy/yfp8iHhPrxW9M270EztRK8PbxSwPMkDspr8JU78jPjOmMV0kp5Kg4O9WM7jZj1W5TWyODgItpw8+sP2BNkpbnwpftz7OI5FKhuKzcB5htoOoguxmTrfxlLg5nVV6FDgk8A5VP8t8y6LsHo9WzmIV0TSKakD5CKnETWn0uBgL9aZ3qw7qrxGFgcHAT5GfDX+K2hgQeoXnF0erYF3M9aJcTi2krCW9stiLHX6iVgpgHJpS7NuEDbz/Axs8K+W96abeE2pRViH2ojOhi8iKfRr8tMHBZUHB3uBz7fgNar1O2RxcBBgJ/zsWd7lMWxFl0g9BmMDXj3E23m/xM4djqP8IGL0sgA7v/oGydkR8mIYNtj5G2qfFLeQ+ATOedg5fKPpTEUatgXxkeobyfbJWbUO9X82+fzrVnn+rDfMvkR8e05xGlFt+mB5qL0fWnV6pdMKWMdGdIZb0uU14Fwsve0wF8GmQLCIcy2rCv+JrX7RyYBIca1D/PfiPrIz0SdJtcHBV2iu7boClVNN9ZLdwUGA/Yif5P/GaUSSZZsCLxHen3qw2i/V2ilvY3UM9wOW6XTgKbEydr51HdYRVO0981K3TnQRrIikzvbEj+lXk+3Js9UGB+9t8vk/UeX5szw4CBZ7dHtOcBqRZNnO+H2r3mUxtaXUfAX4FZZqNA3pQV2YhNV4v53aVhUuwcpkZDUbmmRcX+KpCF4lPXUcGhUdHJxP/Au5bhPPf2bkuZLqZ2R5cBDCK/C8wbZmisl2wub48d7pOBaJ2wDLQ76IygfGD7CUsNuS7UkK7TAC+CJ2cpRU9yd4eQP4DhokFymiewj/HswAxjqNqHlJg4PR9tfkJp7/pCrPnfXBQYi3X5cAH3cakWTZMOAGqnd49GKpMS/F6iCpbkrYUlhqqr9QfYLCu8DJwLJOIhWRNOhPvM7c82S/Ez46ODiH8G9iD5bKr1HnRZ4/qZ2X5cFBsExCwe2Zh9VME2nEKOAf1NbOex+b1L8V2Z6k0A4jgaOx9KPRSR3Ry6vAt8j+77lkTHSFWA/5KF4bHRx8DyugGrztZw0+dz/iddGiDY08DA4Ox4qfBrfpBqcRVRcs5Jr1hl2erIIVGK40mLUQmz39aSz1klS3EjYb8CmqdyQdR7ZXDIlI7T5N/HfgAKcRtUbS4OD5kesXNfH80Y62pLZd1gcHBxJPafggOomX+q2Arf6rNJjVjWU9+CxKj1Srcdg5TLV0XR9gE8D0vooUT3SF2BJga6cRtUZ0cPBVbNVN8LapDT73YOx3s1o7L+t9SMsRr/92ldOIJKsmYYPNlQazFgB/wrJ8qa+pNhOB7xIfU4he3sIGFJUNTNpuILayJLgDXuo0otZJGhz8VOS26TT2Rds78jxvAJ8h/mXO+uAgwIHEt2sLpxFVFqwDuYbjWMQap6dSORXmLCyn+YqOYsyLrbBUMpU66V4DjkQz9kXyrAt4kvB3Py8r6ZMGB7eMXJ9LY6ult448z4dY7de8DQ6CpSKLbtfuTiOSLBmBte3mU7698RHWtpvgJsRcGA9sjJ2bV8q48S5WA1KdciLFMAT73gd/B37rNKLWSRocPChy2zQayyz0ucjzvICt2M7b4CBY3d/gNvVgNX1FajEKa8NFa1gGLzOxtuDyjmLMg/FYH95NVB6AnYb14SmjmrRNdNXgPGwWaB4kDQ4mrfjbo4Hnvj7yHD/GambkcXCwC1v6HNyuG51GVN5Y/B/VlxzHUnQDsNkwcyh/kHsaS4852FGMebUacA6V3/vHsZStIpI/0VWDi8lP8fekwcEu4JnIbVMaeO7fR57jPCxFaR4HByGeCvJBt+FIBvTFaqdUqhf9InAZcLejGPNqJeB04qtegpdnsQkNIpJv0VWDHwKjnUbUOkmDg4OI//Zt18Bz3xl5ju+S38HBPoQnzfcCf3AakWTBICwTW6U6yE9gKwWvdBRjXq2JTfKo9N7/E6ubKtJSXcTTJ+WlwwOSBwcBzojc/qc6n3c08dmba5LfwUGIzzDvIZ2r8oIzpM5yHEuRbUp81Urw8m+s80IpzNprGeAHwGySP4ceLCf80q4CFJG2eIDwdz0vGSGg/ODgiZHb7qnzeYcS/63cgnwPDn6M+CxVnXBKOesBD1O+bfcMdi7kzWqeRPbr16fRMODbWF2fcp/FZVjmDhHJnz7YgFnwO/8jpxG1VtLgIFinefD2i+t83hUIZ9dZgk26yOvgINgiiOB2daPag1LeVsTHB4KXR4CdAo/fEKW7bIeR2OKjuSR/Dkuwvm7VI5SWiaZPWoAta82LcoOD60ZuX0h9J1DfiPz/B0q353lwEOIdcqe7DSdRsKbkzo5jKaLBWGqBcmktX0UpLV0YiX0uH5H8ubwF7OMsOhFppbWJTwJY02lErVVucHB5wseeHmDVOp432jn0XOl5Jye8Xl4GB8HS2AS37UK34UgK9cdSVpZLD/8G1rbr5yrAghqGfS7lJoC9j30uIpIvuxD+rs8jX5MByg0Obh65vd4U8t+N/P87SrfneXCwC8sWFNy2qS4DklQagvUVLSG5PfEcsD+a2N9po6hcnukV1OctLfI7wjvXH92G03LlBgcBHo3c95U6njd6gP1i6fa8Dw4eQnjb3iZdOY/7Y7XrvMbiILfhFM5WWCrXpAPXu8CxqBaKaxOx2eTlGn6Xo1WEIll3KuHv9V/dhtNy5QYHAW6P3D61jue9O/J/v1O6fXLC6+VpcHA3wts2B6X6Ft9GwFMktxk+wAantL+4NR5LgVyuLtANWAeTiOTDlYS/4xe5Daflyg0OAvw3ct/n63je6Iqoz5Vuz/PgIMTLSL3sNhxJmR2Ir0T2Lm9h+48mf7m1OjZWk1STsAcri6FVhNKwPsB0wjvWrk4jar1Kg4PHRu77Z43PuXHk/83HUvdB/gcHB+MPvnmXzZxGFBZMfXq941iK5kjiqXa9y9XYyjVJjy2In1x5l+ex1dUikk3R7/bBbsNpuUqDgwdFbn+F2iYxTSA8aWIJsGLpvskJr5enwcG+wOuEt293pxFJWhxM+bonN2GrdSU91sfOZ5M+r9dJ1zmbiDSmP/Hae9s4jaj1Kg0OnhC5794anzOaMe1DbLUU5H9wcGnix/L1nEYkadAFHEfyxKIerCTFss6ikyRbY6s4k9p5zwBruQtNsmxTwjvTLPK3qqfS4OCyxJfn1nKQPCfyfy4P3Jf3wUGwIsbB7TvFbTghwVqSSqPTGYOw2YpJB6hX0DL3NBuErapZSPyzmwMc4CwyEWnUKoS/y4vwJzDlRaXBwUHEO822reE5p0b+z+2B+yYnvF6eBgcBfkV4+37jNhxxrB/xFcje5W3sfEfSqR+2mjMpjfwC/Gw3IpJNkwl/r98hXZmcWqHS4OB4Gksh//vIc54fuC/vg4MANxLevhPchiOODQP+RHI770Vs0YWkU6UyTrOBfd2FJln1NcI70tVuw2mLSoODAH+O3F+tht4AYEbk/+wYuL8Ig4PRxtNdbsMJeQY/LhVabr9VgSeI7/NLgF+gpe1ZsSHwb5JnjJ2O6kOKZMnBhL/Hd7sNpy0qDQ4C/DZyX7Uael3YiXDw/xwYuH9ywuvlbXDwk4S373G34YhDywMPktxhdAH5m2yQV2sCD5D8OZ4HDHQXmog04UTC3+eLnUbTHpUGBwFujdw/tcrzDSVem3WLwP1FGBw8ivD23eg2HHFoTcL9pt6lGxt0Uqr4bNiM5LT/PcDJ5G/SiLRRNFf519yG0xbVBgf3iNw/ncr5lKODx2pZCgAAIABJREFUf68T7jgvwuDgqoS370PS8cMzEXVqddLaxBvu3v6wt8O4pDEDsRmUSZ1If8JS2IhI+v2S8Pf3R27DaYtqg4ObR+6bi82QLWe7yONnET4xnpzwenkbHBxBuI5FN5rgU0QTsFpE0f39I+Awd2FJg/piHX1JNWruofLvooik0/WEv8t5zJZUbXDwM5H7p1G5P+qwyOOfJ9xuLMLg4HqEt+8dt+GIIxthfeLR/f09YCeHcUljBmGTYJP68K5EtSKlRv8ivPNs5Tactqg2ONiPeOOjUp2VmyOP/WHk/iIMDnYB7xPexhWcRmSCNSR/4jiWvNuQ+AraXmwG0poO45LmHYrVUY1+tjdjjQ8RSbc7CH938zhZo9rgIMTrLlYa2Lgk8thzI/dPTni9vA0OgnWWBbdxA7fhSIdNAt4gvq+/gGoTZd1e2OS96Gd7H7CUw7hEpH7PEv4eb+Q2nLaoNjg4gPgAx3YVnu+eyGNPjNxfhMHBvsTrDo50GpF02ibE+3F7sXGBiQ7jkuYdSXKpoD+iSf5Sg2hNlrFuw2mLaoODAD+LPKZcetUxxIu1Too8pgiDgxAvdr+123AAuA0/ni0dx5JnG5PcqLgKrTLIi42wepHRz/g2lGZCJO2i6THXdRtOW9QyOHhC5P5y6VWHYTVWg4/dPPKYyQmvl8fBwWA7qhfYx2040kFrAW8S389vwVaVSvatSXzSRC/wd2Bph3GJSO26iNcTHe40ovaoNjgIVhs5+JiLyzzXBKzkife4JcCKkccUYXAQ4GnC27ix23Ckg7Ymnlq3F7gMGOIwLmmdrUjO7HYTSiUvFQwivMMsJN6xkge1DA6uQ/y9WC7hcf8Xedz9CY8pyuDgNYS38QC34TAYf7XTTLR8ul02J7lRcQb5/P0oshWA54h/1neixoVIms0l/51GtQwOjidcqL0HWCXhuQ6PPM9zCc81OeH18jg4eB7hbfyy23CkQ9YlORvEhajmcN4sRzxzUC/wMEoxKpIFyxL+7s5yG07b1DI4uGnkMXNJXgk9NfK42xMeU5TBwWitxj3dhiMdMpn4qtFe4BSHMUl7TCR5kv9NqI9cEvQhvsLH60wqoqeBRwPXB2B5zKOmRK5f3KZ4suDDyHXXK8Z2xF/RdDvWISitNR4bFI42uk8Dvklxfz/y6g1gG+DJyO07YrUJRSR9ugiv7vU6S4roLWwyg6cLOCThcVMi1726DUWUtradtN+yWP2q6KTI84EvYCssJD/eA7YH/hG5fVPgctJRQ15Eyosel+c4iSIdHiF8njoUm6gflNT2u6SdQaXc7Mh1rRjLvwnAn4h/1lOB73c6GGm7V7A+vBcit+9BPie2SpP6EV/5scBFIClyMfDxwPXDgF8Hrm+GpdzxfAT8uf1hpdZHkeuuUw1+MvD3Lc6iyK/BWOfR+Mjtp2Gp27JiWWCHCvffhaVbbtY4KtdwvRWbvZV272Dv153AxwK3H4rNPD/bRVAiUlZ/wp27iyl25/7FwK6B61OwWbLe4N9Ewr/VPcAVnQgspdLWtpP26oedy6wauf1c4BiyM0g+DNitwv33A9Nb8DqdakO22yxgJ+BGwjW69kKdhSJppz68sMuA0wPXDwMuClyfTDhrxIfADe0PK7XUziuWYdiKsegEsO8CP+58OA0bDWxb4f6/EB/4bsQKwCcq3H8jlmUw7V7H0sjeRbi8yLHAU2iiv0SMJLzM9H234bRNLWlFwU74FkQeu17g/nMj911a5nmKklb0IsLb+Hm34TCtFEc3ySlhpXFdwJXE9+sfugyqQZsR347g5Vstep3Tq7xO1go+Lws8S3gbFmOdSyKSHl2E66r0kM+0gLWkFQVLoT8z8rjgyeUPI/fdWub1Jie8Xh5nX0ZrcH/bbTjSZr8ivl//xmlEjVmVym2un7bodY6v8jofL/9fU2kYljknuA09wP4ugxKRilYk/J193W04bVNLWlGAMdg5afA3LDjh5ZLI85xb5nmKklb0j4S3MSlbmuRDH2xyf3S/ztLEfs92VG5/Hd2i1/l1ldcZ3aLX6ZQxxFOMLqTyIgYpINUcjLs68lhvFlJS51K5maNFGRxMU83B9QJxPOgwjrzyUoYGL9eSzd+LaoODT7XgNfqRXAg4y4ODAJOwGfHB7XgfWNllUCISM4fw93QZt+G0Ra2Dg2CDHcHHXVi6vQ/W2VRLJ8nkhNfL4+Cgag4WxxTi+/Td2OrjrKk2OPgmrZkk8d8qr5O1wUGwWfJvE96OucA6LoMSkbKiNQej6cDzotbBQYCbI4+dWrp9GPE28eZlnqMog4O3Ed7GPdyGI230feL79OVOI2pctcHBaKr0RgzAxgryNDgIsD7x38HpWKYzkf+JDnjlcQepZ3Dwk8S/NP2AgyK3T6N8TYaiDA5GZ5q6nH1wAvl+r11aC0s/Efys/wss7TKoJlQbHGxF584eNbxGFgcHAXbGVucGt+WvZHOgWCSvnif8HV2v8sMzqZ7BwU0jj5uLdRjtFLl9FuXTK01OeL08Dg7eTngb93YbjrTJyliHcvCzfgUY5TKoJlQbHOwlnF64EZvX8BpZHBwES6EVzZ7zL7I5UCySd13AfMLf1zxOAqtncHD/yGOnYX11R0Ruf47ybcWiDA5GJ7ls5DYcaZMNgUXEj+tZrTFZbXCwl3AJsEZ8pobXyOLgINj5XDCzUC+WblbkfwNbr0RuX73TgaTMX7CGiGcMsAs2uzboEixlQVF1Ea9PMs1BHJ5gvcFyKcGkfn2w9LmDAre9D+xOa3J6p9VhTf7/Ka0IIqXuAL4TuW177ORLRNJBbbuwR4AnA9eHAp8m/lt9BfFaLEWzWuR6dF+SfLiA8CSvecCngBluwukIte3Ke4j4KuENgW84iEVEKksaKIseu4vmBsILAFbGUshPiTzuQuz9K6p+wITIbdM6H4a0WX+sDy84wWc61s6b7ySizlA7r7zriZeE2gNbBCUC2LLi4OhxHk8C6lk5CHBa5PF/J7xSJprHPKoIKwfXID7bvtxKynYbgZ9n/i20gqmVvkB8X3aZPrYVklYOPkP8NyJa7L1WI4nPvo4+fy/ZXTkI9h2Lri55l3zOWhXJorMIfz9/4jactqhn5SBY7bzgYx8hPvN+0wr/f3LC6+Vt5eBIrI3rbd9isjvDWMrbh/i+fJTTiJqXtHIw2vb6iMbbKYOJp1VPattldeWgJ1qLag4w3mlEIpLkWvL1G/7/7N13mFxl+f/x9246SYBUktBCC73Ll45IDSCCSJHeBFG6fhUUUURQRPhRlK5UQbog8KUJKFWkCwgEQlFCICGUJKQn8/vjnnM955xpu7Nn5jnl87quc+3O7OzuPeecOfPMU+67mu6sHAT4bezxjxJt0ywAlq7z+0VYObge0ef3od9wpEWOp/JcznomkGorB+PtsJ6kkF+ayj7/16v8z6yuHATrr3+U6PN5H5s0KwUWDOQ8Hbt/s3YHkkJXxm5vTvQi8xgwsX3hpNLmsdv/xN9Kyh2xWVAA91Ds2WBJGkTl7JJbsbqcefMQ0WLuw4Bdm/xb+xEdWHwaS9+RJyXg21hqvsAI8jcJQiSr1LardC32oS+wEdEUov/G2jJFthnRAdaXyfcM4yLqg6unHngYqzWZN88DL4Vu9we+2eTf2p3owOI7wBNN/q00O4boJNpBwBmeYhGR2tTOq3RN7PaWRNs0D2CDB0UW78NLok6bpMsQrNZg2DXYyrG8uYdoxosxWAmcZhxItM//EfK3qnYRtvhjTui+pbEJtFJgweDgk7H7t8cKcRbZG9TvILq6TXGk2c6x2z4/ICulaGscBYwK3Z4N/K+nWFptIXBd7L5m0xIcErsd/6CSF+9hq6zDvku2Z1OJ5EW8bbc59mGxyD7COoZqubpNcaRZmtp20hoHEE0/txA3wzyPro3dTqptdyX5LC8xlcpOxYNQykKRtIm383ai+RUzefEs8K86P8/rZ/LuUDsv/44DhoZuzwBO9hRLq80Dbojd12w7L/57eb1eTADOi913IsoAJthsmklEl5bu4jWi5HU3rSjAd6hcQlzCVsoMbvC7eU8rOhCrN5eGNDqdWIdfCZhL42MjXdOPylQeZ3qNKDnV0opegNXkiqdTG93Nv71W7O/Oxjrkb6vyP7OcVjQwAFtxmcfzRCTrXib62mz2w1JadTetKMBeVX6nq9f7rav8Xp7Sivai8vPAeK8RSdI6qUyRdInXiJJTLa3o9diEpXmx+1fv5t+Op5paCCwHXF7lf2Y9rSjYteDfRJ/XpV4jEpG43sA0oq/Tr3iNKHndTSsK8H0qr8slrATOgDq/B/lPK7ok1jcRfn5reI1IkjaIyuvCj71GlJxqaUV/idVHrtb/1h2bxv7GDGxfxsvoZD2taGAwVhIoj+eJNCFYOVgC7or97LA2x5JGN2AXlrhbsYtFke1NdBBuEvCcp1j+B3eBfhQdm6TsQbSj9AsqZ5jkzZtYZ3OgN7B/N/9G/Np5J1ajJq9mA+fE7juS5us1ikhy7ozdVtvO9km1CWL3A5PbHEva7Ey0ttgMLKWO5Md2wKqh2/OpzACQN1OAe2P3HdTNv3EI0RU5DwP/6UFMabeQyjq1BwCLe4hFRKpbgKXUC1M7zzIBza9y//VU79srkv2x9NqBt8hf6ZOi+ybRVYOfYLU48+wF4MXQ7WZSyB8Su30z0fI5eTMDOD9233fQ6vPC6gx9H18yuyuwbBtjSaPPsXy8J8e2eJ2OIjo6dvta/KUjUkrR1jg8dvv3dG3FbdZdHbt9SDd+tzewb4O/l0dXEB0AHQ58zVMsIuLE35u3BNb2FEtazMMmMMTbdqd5jCkt4m27G7GMDJIf8bbdjeSvnko1V8duH0T3OkAOaPD38uhGogOgA7Ga2iKSHvE+vG8AS/kIJEWmAEdQ2c6Ld4QXTSdW/iMsr2kTiyw+QeBSirF4In4udydbzgBs8UvY1T2KJhsuwhaABJYBdvQUi6TMK0SXleZphkEzaUV7Is9pRXck+rwWAit6jOe5UCzjPMaRJyOx4xo+zut4jShZtdKKgs2K/iL2s66miNo99nuTcJ1PeU0rGriQ6HO71W84IlL2d6KvzXhdhixrJq1oT2xd5f/lJa3ol6h8bht5jUiSthiV7ZutvEaUrFppRQH6UJk+qaspc7eI/d7n2L6E/KYVDfyM6HPTSmKRdOkEJhJ9neZpNXgzaUV7Is9pRb9O9HnNx1JmS34sT/QYL8LaRnlRK60owDBsQmP4Z11NIb9/7Pfexn2ezGta0cAfiD43TRgoqM7Y7XgHx7eAse0JRTKiAzg9dt8d2AXUh1FYjmnKMUzwFEfe7EL0+vAC9Yt758l04M+x+7o68+iQ2O1rsUHWIrg2dnsHlFpUJA3Ojd3em3xN9pBk/CJ2+1HgGR+BSMtshxvUAutkfcxTLO02n8qJEc227W4CZvU0oIyIt+22oPt1fESkdRZRWfbjuzSuoSzF0gv4eey+G7GJzJIfu8ZuP4FNHiiCaVRmkWu2nXcl/rLitVu8nbczSi1aSPHBweuIzsbpD5zVvnAkAw7CavyF/cpHIGU742Z1xOtmSvN2iN2O163Ku6tjt/ej8UDXSKIpbqFYM2+ew2Z3BgYDm3iKRUScu4hO7uiFUitJ1C5UrqI600cg0lLxtt1fKE7nB1S2yXan8UDXQGCvBn8nz97BMgsFemMz90UkPX5PtGbyIPQeLlFHEi0rkLcVpmLUhxd1II0HupYh2q5ZROWAWZ49gdWlDAzHLb6RAokPDs6nsubKPlR2eEsxDaeyEXEL8KyHWAK7hL5XvcHkxAeAH/AShT8PE62zMpTKmVhxB2BpqwJPAa8nHFealYAHY/dpcFDEv2ppzb+CfWASGUjlYPEjFO99vwiK3rZ7AXgxdLs/9jm3nm9g6eYDbwJPJhxX2qltJ5Juc4AzYvcdAmzf/lAkhUZTeX5cS3Tih+RD0dt59wAfhm6PoXLANO5QogOI8X7AvFuAPecwtfMKKD44CPZGER/suRylEBH4HdEC13OAkzzFAjYQs235+y+wFFjSc0OJ1pCch3WoFEm1GUON0hIcFLtdpJnlgX/Ebuep7o5Ilt1N5QfEC1CtEbFJXyuHbi8kP/V1xOkNrBu7758+AvEs3jZr1LY7JHb7aoq12hLUthPJgsuJDvZ0AJdhmVykuILzYGjovpnAj/2EIy20DNG+2pnAq55i8WUB8KfYffXaeR2oDw/UzhOqDw4uAr6DvbACS2MvMuWeLa7jqZxdewaWbsaXLYElyt//FRuslJ6LFy2eQDH3bTzX+Hhq12/YkGin2xysJk3RxOtS5qkAtkjWHUu0TtYQ4DZs9YwU0/7A0bH7fgu85CEWaa3lgL6h25OBKZ5i8ek6bNJbYBNg9RqPXR74cuj2ovLvF038eqC2nUj6LMD68BaF7lsBu2ZV6/OTYjiZyuxHPyVaCkTyYeXY7VexCX9Fc2Xs9teBYTUeuyXR/TYduL0VQaWc+vCE3jXufxarNfiT0H07AmcD3291UC3yG6KrH2e3+P+9jL0Zh2V1Zdt2wDmx+17AzgeflFK0NcbGbvscAPbpHeBxrNEAdr3cn8rXAlTOLL8d+KxlkaVX/FxZwUsUIlLNBCy96Hmh+zbGZpvHZ01mxcXAHbH7Wrmq520q23bPtPD/tdIG2LEPe5to21/yY2zsdlHbdtOwzwy7h+47CPhRlcceQrRT/a/Af1sWWXq9hw04BPtiaSx7y3xvEYlINY9jacK/F7pvN+DnwKleIuq5M7EaioHpLf5/z1PZznu8xf+zVcYDv4jd9xRwoYdYpPXGxm4XtZ33CvY63qB8uy+wN3BJlcceErt9E9GJtEWhPjypqy/wGNbJEt4apV+RfFkZ+yAdPgc+BVb1GVTZa7iYlvMcS54cSfR4xzsP82BjKq9tF1R53OGxx1TLzd8XmBp7XLXc5rdV+Z95e+PtxDqQgucX7kwSEf86gbuovBZldeKXNGc08D7Rc2AWSiOTZ98gerxv8xtOS6xE5bXt+iqP2y32mElUZsfpAN6KPW7fKn/r8ir/M4+vo/hnweF+wxGRGvpjKaPDr9dFWOe4FMcawOdEz4Op5K/vQZwTiR7v8+o/PJO+QmWb65dVHndc7DHxtJlgNdenxx63eZXH3Vflf47syZNIoYFEn98XfsMRH+p12s4D9qSyGOelqLhxUYzG6hSFc5QvBPYD3vASkbMCsFr5+5coVtHYVhsYu13kN4ebsHztgTWp7PT5GtFOkknAQy2OK60WEZ1t1QEM8BSLiFRahK2Afi12/6+BvdofjngwBBsgDtebLAHforLmuOSH2nbOPcCHodtjqPxs+2WiaZU+B+5scVxpNjN2ezEvUYhII3OwNHqTQ/d1YKn2tvISkbTbclg7b/HQffOxAeKiriYrArXznOuBuaHbGwNrxR6zJ9GarG8CT7Y4rrSaRTQl9QA0wb9wGh3wj7DZleELS39swGi3VgUlqbAs8HcqVwieBNzb/nAqfDX0vVKKJqsjdntR1UcVw0wq847HV08fErt9NcXM7x6IP/da6atFxI/pWFruj0P39cJqSys7RL4NAe7H6uSG/Rq4of3hSBvFP/MVuW23ALvehTVq291IMVNNBdS2E8mOScAeRDvHB2J9OJrkn2/LA48AK8buP758v+SX+vCcadiYRdgBsduHxG5fTWtLU6RZsMI80IEGB6WGPYimiithjY09fAYlLbM8lal0SsC1PoOKuZf6y7+led8hetyr5efOuq6mFQXYJva4aUC/8s+WwmbihX++WpW/AcVIK9qBdbqFn6M6kETSaTsqr18LgcN8BiUtMxIrOB9/H7qPypSKkj97Ez3uN/sNpyW6mlYUbAZ5+HGzcbXpBwEzYj/ftMbfKUpa0SnkO6WWSB4dSOX1aQ6W9UbyZxxWFzd+zK/0GZS0zf8SPe7n+A2nJbqaVhRg19jjJuP6pcZin3nDn3+XrfF3ipBWdACV7xMiNR1N5QDhfCw9leTH6liKzvgF8M9YbbU0GIDN3i0Bn6DBh6TtT/TY3+I3nJbozuBgBzAx9tg9yz/7Yez+egXLizA4OJTo84unoRKRdNmPygHCRcAxPoOSxC1PtE5zsD1MZRoiyafxVB77vOnO4CDA87HHHlW+/7DY/W9QOSM/UITBwV5YuZHwe4RSxotkQ3zAoIRN8v+Gz6AkcesAH1B5rP+E+sqK4kiix/5qr9G0RncGB3tjA4Lhx+5U/tlpsfvvr/M/izA4uCzR5/eR33AkCw4nOsIebJcBfTzGJcnYFfiMyuN7M+k6vuFZIEqDlbwtiB7/Z/yG0xLdGRwE+HnssXeV74+vwvhWnb9RhMHBDYk+v3/7DUdEumBXbIZg/Pp0LeoAzoMvY/XV4sf3PnR8i2R1osf/bb/htER3BwePiz32qfL9j8buP7nO3yjC4OBY1GkkkmVHUdmHtwg4C2UOyINvYhNy4+9FN6CBwSLZkejx/5vXaFqjO4ODYKsnw4+9keoT//et8zeKMDi4Ffnv/5UW2JfKWeYl7IPUKI9xSfN6YbMnqg38pnG20SW4+LRyNXlLEz0HppO/Dw7dHRwcS3Tl9Hxg99jvzwKWrPM3ijA4eADR5xfP9S4i6bQzllYvfo16nvxdp4qiA6sxU63Nfg9WQ1yKYwDRdv4CLH1mnnR3cHAYlRMjdiba3quXagqKMTi4M9UHUUUkO46gel/PI+Svo7soemMDvPFjWgKuQDXDimYc0XNgit9wWqK7g4Nrxh47F9grdt9nwGJ1/kYRBge/TfT53eQ3HMmSfajeifRfYBOPcUn3jQAeoHqj4lLS2ah4F9exMdxvKLkVX2Wwlt9wEtfdwUGw2Vfhx0+L3f5jg98vwuDgRUSf3y/8hiMi3TAemwwSv05NweoTSnYsAdxK9bbdTaQnTby0Vzy17Jf9hpO47g4OQmXbLN62u7fB7xdhcPB0os/vYr/hiEiTDsI6x+PXrLex7C+SHaOp7JsItvOpnQpb8qsT+Jx89zV1d3AQbBVcvXbeZQ1+vwiDg1cTfX71MmaIVFgfeIfKF8pC7AWWt9moebQXlQXmS9gs85M8xlXP2rg4n/QcS57dRfScONZvOIlrZnDw0Cq/E94adZ4XYXDwVaLP76t+wxGRbhpH5eu4hK2kuRarKyrptjPVa0cvwLJEpHHSl7TH1UTPidN8BtMCzQwOhksVVNv2afD7RRgcjKdZPdhvOCLSA5sBk6je/3MWyiqQBXsBH1N5DOdQv8SJ5N/DRM+Jw/yGk7hmBgePqfI74W3TBr9fhMHBd4k+v694jUYyaRi1V51NRDPN02o0cDvVj9skGl8gfToZF+spnmPJs+8RPS8e9BtO4poZHBxI9VU1JWzVdKPUq3kfHFyR6HObR/00qyKSToOpversA2APf6FJHUOwyXnVjtvHwA7+QpOUOJjoefG833AS18zgYG9gcpXfK2GpphrV5cz74OBQKlMTL+81IhHpqRHAQ1S/7r2JOobTaixwP7X7Iv7HW2SSFqcSPS/u8BtO4poZHBxKZQr5YHuDxqts8z44uBbR5zaL+mlWRWrqDZxNtD5DsC3CPjQN8xadhPUCvoN92K12cfw7sJS36LomPHt1fc+x5Fm8g2U+MMZrRMlqZnAQ4Koqv1fCUi41kvfBwVOIPreH/IYjIj3Qgb2mF1D9mncD+XpPyLIO4ECqZ4IoAc9hHUoiw6h8Ta/uNaJkNTM4CHBOld8r0bX0mXkfHDyK6HN70W84IpKQPsCFVL/2LQR+i006Ev/6YDWkZ1D9eD2ISu2IWY/ouTGbfGV9aWZwEODmKr9XomvpM/M+OPhLos/tL37DkTzYkcrlqME2A0tTsLiv4ITtgJeofnxmY6mF0l6DZghu9uoHKJd6q8XPlzzlnm52cHDrKr+3CFi5C7+b58HBTqxeRfi5He01IhFJwmbAv6nedvgCa9tphbA/22GDf9WOj1KESTXxlFO/8RtOopodHFyzyu+VsLZiI3kfHIxfX37qNxwRSVitVOQl4BOs1EyjFdTSGh1YCtEJVD8+s7Dj0yh7kRTLm0TPkzyVB2p2cHCXKr+3EFi2C7+b58HBPli/evi5HeIzIMmPxbCOiFozzadib2D9fAVYQJtRWSsivD0BrOYtuu75Ji7uKzzHUgTHEz1X/kt+XrvNDg52YIPU4a2rHeN5Hhzck8oPK5ptKpIPfbC221yqtyOmoc6jdtuI2inBStjknjwNUEhyDiB6rnxGfgb4mx0chMq2XVfbMHkeHIx3wi0ElvMakYi0wuLY5+CFVG9TvA8ciWUMk/bYDniW2u28R7E64SJxPyZ6rkwkP6/dZgcHO6ls4y3Rxf+Z58HBg6n8TDDQa0SSO5sDr1H7zexNrIGhjqTW6MAaFLVykpew2mlHYxfKrLgWF//XPcdSBEOBmUTPm297jSg5zQ4O9kReBwc7sLpF4ed1pdeIRKQV1qP2KrUSNvP8OKxmobTG5sCfqZ7Kv4RlgjiZ/HQCSPIGYJM1w+fNj71GlJyeDA42K8+Dg38l+rzu9BuOiLTY1lSuOgpvrwGHkp/JwmnTAYyncoV/vPP+CJRBS2obTWWNvQO8RpScZgcHeyKvg4O9qByz+a3XiCS3+gHfBz6m9pvbVOAM7AImPdcPOAz4F7X3+QLgD8AynmJsVifwEfYc5qLOx3Y5j+j58yH5SA+swcHkxFchLMRSdIlI/vTGalDFU5DEOy7OQXXuktIH2Bf4J7X3+SLgRrqW4lrkJ1ROGBzlNaJkaHAwOdtR+bw29xqRiLTDAGyS0afUbnN8CPyMfHSQp8EAbOFErTT+JWAecCnqN5WuibdN/oNl+cs6DQ4m59tUXmPy0DcpKbYEcCZWm6bWm91cbFVYV2o7SKVlsJqBweBZre3PwOp+QuyxTXDP40HPsRTJ0liKyHa+AbeDBgeTMQhLNxt+TjeMcMIUAAAgAElEQVR7jUhE2mEgcAo2EFhvMtItwFZohnMzlsLStcavsfHtfmADTzFKNi1JZcfvZV4jSoYGB5PRl8pO6r96jUhE2m0oVpN2NrXbH3OwSecbeoox65bH+knjq/njk79uAlbxFKNk04rYYE/4XDrFa0TJ0OBgMoZQOXZwldeIpFBGA5dQeZGKbxOwmUia/VzfksDhwCPUzg8fbH8DNvUSZXJOxz2fEz3HUjRnED2f5mP1jrJMg4PJuJjKiR768CJSHMOAc6nfeVQC3sU+vGlVcX2DgAOBe6ldvzvYnga28ROm5MCJVHZA7uA1op7T4GAyfkH0+SxEnf8iRbUsNgA4n/ptkn9jgw9Z/2zbakOxFTuPUjtFfLA9QPbfT8SfeAawOcBaXiPqOQ0OJuMaos/nC1RTWjxYBjgL+IT6b4Yl4CngGGCMl0jTZyBWa+9WGnfEzcdW8GR9UDAQrnOk4svtNQgrRB4+v14p359VGhzsufFUfqj5ldeIRMSXkdjErg9p3LZ7Hks7v7yXSNOnP7ALNohRL8tG0En/FzQoKD3Xh8paI+9iA/5ZpcHBntuUyom8l3uNSETSYCyWMr5exohgosnjWAr6pXwEmkKDgL2AO7CJtPX23zzsfSvrE7HFvyHAFKLn13PY546s0uBgz+1B5fP5ideIpPD6AwcBr9K4I6kETMQ677fD0p0UxYpYDvK7aDwgWMLqhlxAvjrdRuEGISZ6jqWotqdyIOjPWC3ILNLgYM+MozIl2RtYzQQRKa6+WNvuJbretrsM2BWrm1wUQdvuZqzd1mg/zcbS72c1Nbyk06ZUrlB9lOx+ztLgYM+MpnIy4AdYB6OICNhA15FUTi6ptb2KLQzYDpuUUhThPrw5NN5Pn2F9EVq9I0nancpz7TqvEfWMBgd7Zg3gc6LP5SWy2+6XnOnEZkzfTtfeOIM3z1uB72J1Vnq3PerWWQHYF0vX9w5d2x8l4B/A0cDg9ofccofhnuf5nmMpsj9Qed6d5jOgHuiNdXaEt1YXaR5U5X9mcXB1SeB1oufBAmBzn0GJSOpsC9xA49VwwTYTWxV3HDaBI08fVJYF9sQ6fibQ9bbdi8D3UOe8tM7ZVJ53v/UaUfM6qWxnDWzx/1ysyv/s1eL/2QoDgGeIngeLsM/oIiJxndjAw500Xg0XbJ9g9fOOAtYjX314KwP7YxPe/kPX9kewyvJIWt8PIcV1I5Xn3g+8RtS8an14rZ6cXq0Pr6PF/7MVhgNvEz0P5qK69ZJSQ4AjsBp5jerohbdZ2BvrucDeWNqDLBiC1fc4FZtVFF/23Wh7ExucyXuNr/CKq6zXQ8myxYimdw0atd/wGZS0VS/g/6i8FqkOqIjUMhhbTXg/jevohbc5WHr5C4D9sI6XLEyoWAKb2fojbIX9B3SvbfcelqJZ9RmlHfoAD1N5Hh7hMyhpqw5shWX8HPiFz6BEJDOGAd/B+uMa1dGLTwp7FPgN1p+QlVVzw7DyGj8D7gE+pnvtvNewNH5ZzSAk2TIYKwkUPgcXADv5DEraqg/wCJXXoiN9BiXSVcsBJwFP0r3OpHBj43ngT8DPsdV4G2KdNu3UD0vB9zVshsYVWCOouwOBwfYW1lG2STufhEd9cLntZ5LtHNl5MAaYROWMk918BiVt0QtLaxe/Jl3jMygRyZRRwAnYJLD5dL8NNAtbTXczcAZwAPA/tH9lXV9ssHIXbGXfZdiHru4OBIYHBC8BtiIbA6CSL0OxzxfxjqODfAYlbdEBXEjlNekOdC0Ske5bATgFeJruTfYPtunAs1jmiZ8B38RWtrQ7O1Z/YDVsdeQPsQxKjwNT68Reb3sDW8iQ1bTTkm1jqTx3v8DS/Uq+9aV6eaMLfQYl6ZOV5bDDsJpnOwE70vPixrOBj4DJ2EXyw/I2FSsCPAsb8Jhb/n4BMANrJAzABqwGYftvyfLXEVju4aWwzq8R5a897bCaDfwduLe8vdnDv5c12wAPlb+/E2ugiV+bYp2g4fpQ87AB+Nu9RCSt1gebaBFfJfoUtkJmbtsjEpGsWwL7UDq+vC3Tw783B5t4Nbn8Nfz9XKw9NQd7v/oCm93+OfZethg2AWLx8t8K2m4jcO27MeXvl8LapT0xD3gMq2NxL1aTR8Sn1bESBYuH7luIrSC8yktE0mqd2KTVw2L3v4ZNQp3e9ohEJE9GYFmfdip/HdHDvzcL67P7CGvbhb+fj7Xt5mFtvdm4PrwBWD9e0IfXibVBO3HtvFG4PrzR9HxBwSysvyTow3u7h39PpKe2xT53hNP5zsUy7/3FS0TSav2wVM7xhRyPYZ/B57U9IpEEdWArAH+ADUTEC6dnfZuOpff5FdaQanUO5bQ7F7dvtOw5PQ6lMm3IPKxxIfnSD2swxq9V72AfokREkrA2lqL4JuBd/LfHkty+wLJGnINlkhiUzC4TSdTuVGZrWYja33nUC7iOymvVh8CKHuMSkXzqBDYCTsZWJjebaSGt2+fAX7GsFjugbFeSTkdTee7OwT6bSL4sBjxA5fF+g55PchVJrWWAPYCzsVV2n+K/gdCVbQ6WHusK4HBgLbJZxL6VXsPtr6zkoS+KI6hMF7IAOM5nUJKo4VSvRfQWej2KSGuNwj6snollEOhuTRdf21ystsdVwFHAekRn6Yqk2V7YZK/wOb0Iq4+elYwzUt/iWDaW+LVrEpZGT0SkHZbD3nPOwVayBKVk0r7NxsoXXYpNmF4DpWGW7Die6pP8VWs6P0ZhaZDj165/YyujRSrk+UPeSOwDzjhg1dC2HO2dybMQm4X5BjCh/PX18vfvlX8u1a2AS8HwEtbBJumyL1aHLt7x+SesgfFF2yOSpKyP5SePF0t/A0tLMantEYlI0Q3D2nJB+25c+fvlsRmS7bIIS2U1Ade2C7Z3sIkyIlm1C3ArlZ+X7sHqfH7W9ogkKatimXfWiN3/X6yUw1ttj0hExBmF67cL2njjgGVpbx/eAiw1fdDOex3Xn/ce1g4UyaojsMHt+KD2dcC3sQFwyabNgZuxchhhr2F9eJPbHpFkQp4HB+sZjKsdMxIbPQ++74V1MPXD1aHpXf6dIH/5MrhVMzdgjYNPsI6ioIZh8P0UbJReuu9YXKHUX2LFrSV99gb+iOXxD3sJq1E3se0RSU8dCFxGZVrj17D85B+0PSIRkfoGYZ1KS+Fqxowsf98HV3OmLzAQV3MmqC89Ghtk7Ie9p83H0kQF9ak/wtWnnoI6hiTfxmODSPF2wFtYtpaX2x6R9NTXsAl98Vpa72EdRmqvi0iaLYG180YS7cMbgfXhDcTaeP2x966gD2821tYL+vAWYhOZS8A0XL9dUKc6aOeJ5Nl+wDVUTvJ/HuvDe7fdAUmPHQn8FrsOhr2ApTv+uO0RieTc33BLc/f3G0qu3Yvbz5t7jkXq24XqqUCmYelCJBsWB35P9RQqj2BpRkVE8uhW3PXueM+xiKTBVrhJjuFtBnAYxZ1kmjWLAedRmUasBDyDdZiLiOTdP3DXvq97jkUkDb6BtenibYMpqA5hlgzBJrZW68O7D1jSX2gi+bYr7sX2Evpw3AoDsJn8wQCT6vWk38rAv6j+pnQ36nxIu52A/1D9+F1G5cpQEZE82Qh3zXsHtTtEAJYGnqJ62+DvwCr+QpMu2AJLh1ft+F1L5cpQEZG82gd3/XvccywiabEq8CrV2wl3UZmeUtJlV+B9Ko/dIuAsbFW1iLRIB1bMM3jhbeM3nFwKD8Be7zkW6bpBwI1Ub1x8hi1112B6ugzBBv+qHbNZwMH+QhMRaasncNe/vT3HIpIW/YALqN1OOAl1PqTN4tgxW0jlMZuDVkeLSPH0wtInB9fCTf2GI5Iag4lmUAlvn2B9eJIuS2GTvKods+nYqlARaYMjcS++ezzHkkeXoNStWdMPm4HcAXwP63yolZ5yQ08xitMHOIrqKcNK2ASItb1FJyLSfnsQTbcnIs63gJlUbzM8ja1SE786gYOoPos8WBW9sbfoRET8OgF3PbzFcywiadIB/BiYR/X2w/3AOt6ik0Bf4Dhs0LbacXoJWw0qIm3SD/gQt2R3Db/h5M672L5dgOqcZUFf4E4sxdSg8n0rE63PGV/mfjNKR+VDB1YHcgLVj818LAVBf18Bioh40gm8ibsearBDJGos8ADV2w8l4EFgPV/BFdx2wAvUbndfhq0OEBEpqsHAp7h+ppX8hiOSOmsD/6R+H55eN+3XifXhvUX1YzMP68Pr5ytAkSI7DfdivMJvKLmyNm6/PuE5FmksGBgMjtl9oZ91YqmLas00nwdcDIxuY7xFtj3wLLU79Z4H1vcWnYiIf8fgrol/9hyLSBp1AFcCX1C9LbEQuBpY3lN8RbMFVj+rVtvuNWBzb9GJiKTL2bjr4wWeYxFJo97AD7HU8dXaFXOB84ERvgIsmF2wFYG12nn/ANb0Fp2IMAJ3wZwDjPIbTm6cjLvQneI5FqmvD9Z5Gm4ofK3K41bABg1rvaHNxgbY12p9yIXTB9iX2jPASlgH34+whqCISJEtBnyMG+RYzW84IqnSgZscOQ/LGFGrbTEP+CPwJR+B5lwn8HXq7/+5wJkoE4SISNjS2PWxhE1gHuY3HJHUWgV4mPp9SJegz0qt0BdLE/88tff/DKyck+p+i6TAZbgX5+meY8mLR3H7VKmJ0qsX8CeinUC7N/id7ai/cq2EzX7eC73J9dTi2KrN96i9r+dj17AxnmIUEUmjM3HXyYs9xyKSFoOAO4i2I84HNqF2Gvlgexbr5NAkpJ4ZhNW9f43a+3ohSvslIlLPdbhr5smeYxFJu+2oP0i1CEsrvys2iUyatwTWh/df6k++uwwtThJJlXHYh7ASMA0Y6DeczBuCDViUgA/Qm0ta9QJuwL1BLQC+2cXf7Sg/NlzXqVYapO9js/ukazqALYHLqZ3KNWjA3YRqPoqIVLMUtqI9mBWr2sdSdGOITu5ahK0gDLfTdwVepn7bbiKWFWRse8LOjY2BC4HPqL9/78bKM4iISG3rYO9jJWAStkpHRGrrBA4E3qF+O+Rl4AQ0cNUdncBXsJT9tVK5BpO/rsOysolICt2Fe8Ee5TmWrPsmbl+qjmM69cLSRIUHBvdt4u/0Ab4DvE39BsZC4K/AodhqOKm0JvBL4F0a78u7UIovEZFGrsJdO3/iORYRn9YD/oN7PcwG9qvx2F7AwdRf2RYMLj6GfW4a2sLYs2xl4GfABOrvyxLWTt7ST5giIpn0EO4aepDnWESyoh+Ns1MFfYT3YQOKg7xEmn7rYjVQ660SDPrwbsMmNYhIim2Ne+FOwEb+pTnX4vZloxSV0n69iKbhWEDtDqLu/M09iKaTrbXNwlIlHQiM7OH/zbIOYANs9v0LNN5vM4GLsJXOIiLS2Fq4WeUfobpdUkx7Yqtng/bEB8BGXfi9DmBn4AHc66jWNhe4EzgcpTlfG/gB8A8at+3mAH9AnUUiIs3YGXc9/RfKWCXSHb2Bfehae+ULLOvYfhQ7G0sn1ob+KY0zbZSA6Vj6/hV9BCsizXka9yLezXMsWdWJdcAFHQWD/YYjMb2IDt4uAA5I+H+sj+XPDtK5NdpeBc7C8qD3STiWtBmK1WK8DHifru2fyVjaLxVaFxHpvvtw19PDPcci0k4dwEm40gkl4CVguSb+1jjgAqKDjPW2ieXHb0f+U70NxJ7nBTSehR9sH2FtX6XdFxFpXgfwCu7aur3fcEQya0OsnzAoD9VoFdyzuD68vNeiHobrw5tE19p572Bt8CEe4hWRHtoX92J+1HMsWbUJbh8+6DkWieoEriE6MHhgC//fKCxX+TN07Q20hNX8vAs4FdgBWLKF8bXDOGzw9bfAP7F93pX98AVwPTYbMu+NLRGRVtoed219HWWGkGLoR7TNVwLuoeeT9oZh6eQfp/FqwmD7HLgXm+i0M9mf7LQCVkLhPOAJYB5d2w9zsJRSX8eOj4iI9NzhuOvsfZ5jEcm6pYHv07XMVsE2FcsecQqwLdkuI9QBrIalKb4IeI7oJLt62wxsgHVHbFGGiGRUb6IzPjf2G04mnY7bfyd4jkWcDuBSorN92pmXf1Xs3JhI1xsZJazj6TXgauBYrJN3edKXMmQAlm98L+DnwP9hA53dea7K6S4i0hrhD7g7eY5FpNWGA38n2sa4gOQHxlfAank2qk1YbXsTq319IjAeS7eUto6Uflhq4m9gk9b+gsuO0tVtIfA34Ftkf8KbiEga9cMy7QTX3XX9hiOSG2sCvwTepfttn1eBK4GjsdWFy5G+PrzFsKxn+wC/AO4HPqV7z3U+cDeWbnWx9oYvEpW2F1jWfR84p/z9TdjMUOm657A6amADQhM8xiKmA7gYOKp8u1T+/nJPsfwPsAvWQbsBzXVWzcbOrQnAG9iy/SnlbXL569wE4g0sAYwGRmArIkdhqwJXLX9ttrEzHfgrNih4F/BhEsGKiEjEwdgkE4CHsA+pInm0FtaeGFu+PQ/4Nu78b5X1cW27jWluoG8uNmj4Bta+exsbjJuK1Umcgq2+S8pgrD7iCKwG9hhgJWzG+DhsMlozz+ML4BFsteRdwH+TCFZERGo6FZuIDHAVcJjHWETypgPYDMv+sBOwHs31fc3CtfEm4PrwPsL6waZg7dakDMH67UZgfXlBH16wLUtzz+MzLEte0Ic3NYlgRXpKg4PJGox9iFsCm/EwDvtwKo2Nwj68d2D7bCW/4Qh2LH4HfLd8u1T+/lJvEUWNxNKH7lT+mmRx48+wRsbM8vdgg3ELsY6bcMNjMLZyeADQH6uPMwjXYZRk+qeXsIbEvcCT2GwjERFpnT5Yu2SZ8u0NsNWEInmyIzaxcYny7WnAntjKtXYaimV6GF/eRiX4t2dgnzVmYrO7g/sWYJ1O4YlhA7H2XH+sfdcHa9sNx9p2AxKM6zWsXXcfVpoiyQlqIiJS31DgP9h1fy62sn2y14hE8msUro23Pfb6S8qnuD68z8v3fY5lFJtJtO8s6MNbDOuv64ddA0Zi/XhJ9eGVsM+NQR/eP7B2p4jk3Dm4ZcLneY4lSw7D7bfzPccibmAwnKLzO14jqq8Tm/F+OLaq8SW6XqMvrdtnwANYmoJdsYaKiIi038m4a/N1nmMRSdqRWIdJcI5PwCY4+hbUbjkYV7slHGcWtxnYysCzsPqBYxLbWyIi0qyLcdfpMzzHIlIUvYB1gCOAPwAv0/UafWndPsEGAn+OZcVIcgGDSMto5WDylsFmmPfBPgAuh1t5JLXdBuxR/n5HbFBE/OgALgSOKd8ulb+/2FtEzRkEbIilIl0d62BalWRnJyVhIVav9I3y9hLwdPn7RR7jEhERMwSbVT4IG5xYCaX7k+zrjU1kPCZ034PA3qT3s8sAbPXuxsAa2CDmatgs7zQpYW27IIX9v7C23b+xdp+IiKTHKsDr2ITjT7E+vJleIxIppsHAl7B23mq4dO1DfAZVxQKsnuIE7NoR9OFNwNqAIpmiwcHWuAHYt/z9D4HfeIwlC/pguZaXwFI2DifZuiDSdR3Yys3jyrdL5e9/5y2i5A3HBglXxT4IBDVjxuDSCPRO8P99gaUk/RibODAZ62QOBgPfQimkRETS7kLg2PL3ZwMneYxFpKeGALcA24buuxwbKMxiyvIhuLbdOGBpXJ2YkeWtT4L/bxY2CXQa1o77CDcYGGyzE/x/IiLSWncAu5W/PwZbsS4i6TACN9l/FVz7bjSunE8ztZ5rmYm186Zg9Q0n49p5rwMTSbbGoYhXGhxsjQ2BZ8vfTwJWRBeOerYBHip/fyewu8dYiu4sXIdnCTge+K2/cLwJBgkHAEuW71sca3AEdWgCQb2a2dig9jxsQPDT8t95CMtlrnNbRCS7VgDexN4HpmOzyj+v+xsi6bQScBeWVQFsJdspwK+9RdQew4ClcG27DiprzgSC+tJzsPbdfFwNm75YzZjBwBPAFu0JX0REWmhLrO4r2GDAKmilt0hWdOAGCfvj+vCWwFYEDyI6SSxec3pu+ftpWP/93Vh78QZg/9aHLyJ59Qgu7/ABnmNJu3Nx++pIz7EU2a+I5svWqoie64t1IpewTqX+fsMREZEeuAX3HnmC51hEmrE5Ngs6OI+nA1/1GlH2dGBphUtYx5LqyYiI5MM/cO+PezR4rIjk00BsYlhQQzDJrGIiUjBfxTUsXkKrNOt5DbevlvMcS1GdSXRg8GS/4eTKrbj9upPnWEREpHkb4a7n76IPi5Ith2Ezo4Nz+G1gTa8RZdeluP14oOdYREQkGfvgru1PeI5FRPy5F3ct2MpzLCKSYR1Y0fnggrKN33BSawXcPnrRcyxFdQbRgcEf+Q0ndw7F7ds81W4UESmiJ3DX9L09xyLSFR3AaUTbek9gqZekObvi9uWNnmMREZFk9MJqiQXX9039hiMinhyDuw7kPe2+iLTYkbgLyj2eY0mrY3H76EzPsRTR6UQ7i07xG04ujcTqFZSwIsZaRSwikl174N4zn/Eci0gjg4A7iLb1/oTSnPfUAKwuYQn4jGgtahERya4TcO+Xt3iORUT8WA53Hfi351hEJOP6AZOxC8oilLqnmvBy7c09x1I0PyfaWfQTv+Hk2lO4/byO51hERKR5ncCbuGv6ln7DEalpaeBZ3Lm6CFtBKMm4C2WIERHJm8HAp7i6siv5DUdEPPkXrp23sudYRCTjfoa7oPzecyxpMwCYhe2baah2Tzv9kOjA4E/9hpN7P8Ht6x97jkVERHrmaNw1/Q7PsYhUszFugmIJmA3s6zWi/DkKt3/P9RyLiIgk52zc9f0Cz7GIiB+/xF0Hjvcci4hk3AjcANgcYJTfcFIlXK/jes+xFMn/Eh0YPM1rNMWwHipuLiKSF4sBH+NWY63mNxyRiD1xKS9LwCTgS14jyqcx2Ou/BLzlORYREUnO0sBc7Pr+BTDMbzgi4sHmuLb0A55jEZEcuAx3UTndcyxpcgluv+zvOZai+D7RgcGz/YZTKO9i+3whsJTfUEREpIfOxL2XXuw5FhGwmsYn4eocl4AXsbop0hov4Pb1OM+xiIhIcq7DXd9P9hyLiLRfJ/ARdg2YByzhNxwRybpxuA/q04CBfsNJjXdxudyH+w2lEL5HdGDwHL/hFM7FuH1/sOdYRESkZ5bCUjUGs8rVjhGf+gHXEm3n3YatcpXWOR23v7/vORYREUnOOrjV4ZOAvn7DEREPrsG18/b0HIuI5MBfcBeV73iOJQ3WRmkW2+kEoh1Gqo3Sfrvg9v/NnmMREZGeuxJ3XT/VcyxSXMOBR4m28y7AZjxLa22M2+cPe45FRESS9RDuGn+Q51hEpP32xl0DrvIci4jkwJdxF5UJ6AP7ybj9cYrnWPLueKIdRv/PbziFNQBXA+hzNPtQRCTr1sLNKv8I6O83HCmgtYF3cG28OSg7QTt1ApOxfT8fWNJvOCIikqCdce+v/8LSd4tIcSyOqz86BfXji0gCnsY1LnbzHItv4RnO63mOJc+OxHVcloDz/IZTeOEVxNt6jkVERHruPtx1/XDPsUixjAc+w51/H2OTEaW9wiuI9/Eci4iIJKcDeAV3jd/ebzgi4kF4BfEmnmMRkRzYF3dRedRzLD4NwWbXloAP0AysVjmC6MDgBWhf+3YkGqgVEcmT7XHX9dfRjFJpj+Oxmt3BufcGVuNc2m8P3HG4znMsIiKSrMNx1/j7PMciIu13Iu4acIbnWEQkB3oD76JZB9/E7YMrPMeSV4cDC3H7+XI0MJgGY3ADthM9xyIiIsl4Afd+u7PnWCTfegO/I5ou/n6UztKnQVg61xIwDTtGIiKSD/1w6aNLwLp+wxGRNlsJ9/p/0XMsIpIT38NdWG7yHIsv1+L2we6eY8mjw4gODF6BBgbT5DncsVnNcywiItJzB+Gu6w95jkXyawjR1EYl4DKgj8+gBLAB2uCYbOE5FhERSdapuGv8lZ5jEZH2ex13DRjrNxQRyYPBuPogC4AV/YbTdp3AR9jzn4vtD0nOoUQHBv+AUpylzWm44/MDv6GIiEgC+gD/xV3b1/cbjuTQSsBruHNsAXCc14gk7FjcsTnLcywiIpKsocBMXB/WaL/hiEib/QbXzvuu51hEJCfOobh1xzbBPfcHPceSNwcTHRi8Eg0MptFGuGP0N7+hiIhIQk5GdcekNbYApuDOr+nALl4jkrjlccfnFc+xiIhI8i7GXefP9ByLiLTX1rjX/z1+QxGRvFgGmIf7gF+kOiGn4y6qJ3iOJU/2wWaRB/v2KjQwmFYdwCTczP9hfsMREZEEDAFmYNf2ecByfsORnDgc95khqFe8hteIpJZXcMdpJc+xiIhIslbBTcT+BKs3KyLF0Bt73ZewOtN6/YtIIq7HfYD8oedY2ilcb22c51jyYm9gPm6/3gj08hqRNPJ73PHa13MsIiKSjAtw1/azPcci2dZBNA15CXgCGOkxJqnvLNyxOtZzLCIikrw7cNf5oz3HIiLt9Sfc6/9rnmMRkZzYAHdheR/o6zecthgFLMLNfJae24vowOBN2KwWSbfdccfses+xiIhIMsbi3pM/B5bwGo1k1SCiHZAl4Aagv8+gpKEtcMfrfs+xiIhI8rbEXeffRhOyRYpkf9zr/3LPsYhIjjyCu7gc4DmWdjgM93zP9xxLHuxJdGDwZjQwmBUDgdm4tCQ6biIi+XALSp8uzVuaaJaNRdgKQkm/XsBU7LjNBQb7DUdERFrgKdx79B6eYxGR9hmK63/9AMvyISLSY1/FNSxeIv8Xl9twz3cHz7Fk3R5Ea9DcggaYsuZe3PHbynMsIiKSjI1w1/Z30XuzdN3GwGTc+TMT+LrXiKS7rkOdxiIiebY30XTfIlIcj+Je/xt6jkVEcqIDeBV3cdnWbzgt1Qf4DNfZodRIzfs60YHBW1HnYxYdjWpTiYjk0eO46/s+nmORbNgLmIU7byYBX/IakTRjH9wxvNJzLCIikrxeWImc4Fq/qd9wRKSNfpnGEooAACAASURBVIh77f/McywikiNH4C4u/+c5llbaBvc87/AcS5btBMzB7cvbsIFXyZ7lcMfx355jERGR5Hwdd31/xnMskm4dwEnAQtw58wKwrM+gpGlL4CbwfQR0+g1HRERa4ASiGZxEpBjWQJ/xRKQF+uFSCC0C1vQbTsuci7uIHuk5lqwaT3Rg8B7s/JHs+hfueK7sORYREUlGJ/Am7vq+pd9wJKX6EU1DGWSDWMxnUNJjD+OO58aeYxERkeQNBj7FrvMLgJX8hiMibRR8xluE1QoXEUnEz3AfIn/vOZZWeQ33HJfzHEsW7QjMJrrKVAOD2fdL3DE93nMsIiKSnHDqaGVMkLjhwGNEBwYvQCvN8uD7uGN6uudYRESkNX6Nu9Zf6DkWEWmfC3Cv/W95jkVEcmQYVoevhK0MG+03nMStgLt4vug5lizagejA4L2oZmNebIY7rg96jkVERJKzGDAVN7N0db/hSIqsDbyLe/+fAxzkMyBJ1MpEU8SKiEj+LA3Mxa71X2B9eiKSf9ujCaAi0iKX4i4wv/AcS9KOxT23Mz3HkjXbEx0YvB8NDOZJJ1aTpoTVqFnCbzgiIpKgM3Dv35d4jkXSYTzwOe68+BjYymtE0gpv4I7x8p5jERGR1ginBj/Zcywi0h59genY634m6p8VkQSNAxZiF5hpwEC/4STqXlyjaXPPsWTJdsAsNDCYd9fgjvGenmMREZHkjMRN8JmFpZKU4joeq00UvOe/jGXXkPwJ11o/ynMsIiLSGutg2SFKwCRs0EBE8u9WXDtvJ8+xiEjO/AV3gflOjcf0al84iRiAG+CaBvT2G05mbIlLNVsCHsD2peTP3rjjfJXnWEREJFlX4q7xp9Z4TNbadtI9vYGLiNYXvB9lC8izbXDH+i7PsYiISOv8FXe9P7jGY9TOE8mXQ3Gv+995jkVEcubLuAvMRKo3Ivavcb9vtQb9dsU9p+vbF06mbQHMwO23R8nXSlKJWhxXr2AK6Xx9i4hIc9bCzSr/iOoTfb4GDGlnUNI2Q4GHiA4MXoYmy+VdH+BT3KrhxfyGIyIiLbIT0YwAHVUeo7rCIvkyEpf57z2qv+5FRJr2D1zjYvcqP3+MdKYg2ofq6bIuwT2f/dsaUTZtTnRg8DFgkNeIpB3CMw439RyLiIgkK5xe/VtVfv5n4EttjUjaYWXgNdyxX4DV4ZZiuBF37Hf1HIuIiLRGB/AK7nq/fZXHPI8NJohIfjyFe92v4zkWEcmZbxIdGApbBZt9vkO7g+qCo4Dzq9z/Lq5DRLV26tsMV9i2BDyOBgaL4kTccT/DcywiIpKs7XDX+NeBztDPRgLzgP08xCWtsyUwFXfcpwO7eI1I2u1A3PG/1HMsIiLSOofjrvf3xX62Xvn+LdodlIi01E9wr/sfe45FRHKmF/A27iKzSehnZ5bvO8ZDXI0chaVGXCl039q45/GEj6AyZFOiA4NPAIO9RiTttBLu2L/oORYREUneC7jr/M6h+79Xvu9nPoKSlvgWNuAbHO+3gNW9RiQ+DAXmY+fAJJRySkQkr/oBk3Hv++uGfnZ++b5DPcQlIq2zLurvFpEWCjqKSsBN5fs6sQ+WJaqv0PPtu1hsN4fuOxn3PE6p8Xv9WxxXFmwCfI7bV0+igcEiCqceG+s3FBERSdhBuGv8Q6H7g1RUf/QRlCSqF3AW0fqCjwMjfAYlXj2GOxfW9xyLiIi0zqm46/1V5fv6AtPK953pKS4RaZ13sdf3QmApv6GISFb1BZaocv9gXBH7BdiqovG4xsb/tSvAbjgGi20Rlh4T4FFczOvFHr8GcCGwZLsCTKkNgE9w++kpYHGvEYkvv8GdB9/1HIuIiDSnF7ZiKK4P8B/cdX4DrM5gcPuZdgUoLTEIuJPowOAfsLa+FFd4ouRPPcciIiI915/qE7mHAjOx6/1cYDTwDdx7wC3tClBE2uZi3Gv8YM+xiEiGnQVcB2xLtAZNeKDgfGwF4YLy7bfbHGNXHIubMfEYMASXSucDXCqdZbCZVAuBvdofZqqsj5tJVgKew/abFNPWuHPhHr+hiIhID/wQ6wTaBRssDJyEu87/EbgIaw+VsAwCkk3LAM/jju0i4DSfAUlqrIU7L572HIuIiCTjfOAa4MtEU0aHBwrOxD7TB314L7c5RhFpvV1wr/mbGzxWRKSmXthKwBI2o/w0YAVgaVy9ki+I1i5ZgM1AT5PjiM6WDg9uXoGthvslMAcNfoCtpAwPDD5P9ZUGUhy9gI+x82EOtgpBRESypwMb/CsBH2ITwVbDsiXMKN8/j2it4RJKP5lFm2DHODiGM4HdvUYkaRPUkl8EjPEci4iI9FwfLEV8CUsreCqwHLAKbtLXZ7iBwRIwC9WeFcmbAbgVw5+jjCEi0gOLA68SnXH8N6z2XKnGNs5HoHUcT3TwMlxD7wqiA2FfAMv7CTMV1sUNApWAF9DAoJgbcOfFbp5jERGR5vXDUoWH225P4TqTqm2bVf1LklZ7Y519wfGbBGzoNSJJowtx58jhnmMREZFkDAHeJNqH9yDwLLXbeUt7iVREWilcVmA7z7GISMaNBaZijYogPWetRkUJW76cJidSPc5qz+dETzGmwTrYcQ4PDA7zGpGkyf64c+Nyz7GIiEjPjATeo+ttO9WqyIYOLEVscFyD9tyyPoOS1NoRd57c7jkWERFJzirAp7j2QHilYLVtay9RikgrHYl7jZ/nORYRyYFNseLFjTqP0jjA9j3qDw4GjaXnidbfKZLViKaeehENDErUUKrX6hQRkWxaE0sl2qjDaBFwhqcYpev641LGBtstwGI+g5JU64dLITwTO4dERCQftsY+v4f7vWptR/oJUURaaAzu9T/RcywikhP70rhhsQArdpwm/0vjxtBCiptuaTVgMm5fvAQM9xqRpNWjuPOkqK8XEZE8GY+13eq17+YDN/kKULpkNPA00QHds4BOn0FJJtyOO2929ByLiIgk6zC61hf2G18BikhLPYd7ra/mORYRyYnTaNyweNBXcDX8kMYz4s/1Fp1fq2KrwIJ98RowymtEkmbh19JpfkMREZGEHEvjjqMXvUUnjawDvIs7VnOAA30GJJkS7jj+redYREQkeWfTeIL/n71FJyKtdBrutf4Dv6GISF50UJmyKL697y266k6i/mDm+8Agb9H5Mw6YhNsXr2Mzz0VqWQN3vjzjORYREUnO76jftvsCpZNOo52Az3HHaSqwldeIJGtG4spGvOc5FhERSV4ncAf1s0S87i06EWmljXCv87/5DUVE8qQ/8A9q1x9cRLpqVvyI+h1eX/MXmjerEB0YfAMNDErXvIl7nS/tORYREUlGL+Bu6nccKbNAuhxPtC3+MjDWZ0CSWeGUtGt5jkVERJI3AHiW2n1487C2oIjkSweu73cBMMxvOCKSJ8OBd7CLS7XGxZr+QqvwY2qvGrzFY1y+rIKtlgwPDI7xGpFkyQW4c+dbnmMREZHkDAZeoXbH0Zb+QpOQ3lh97/CxuQ9YwmdQkmmn4s6lkz3HIiIirTEGKylTqw9vrLfIRKSVfo97ne/rORYRyZk1gOlU70Ta3WNccadQfWBwBsVb+bQ80bo0E9DAoHTP9rjz5w7PsYiISLLGAh9TvePocH9hSdlQ4GGix+UybMBQpFnr486nxz3HIiIirbMBMIvqfXjbe4xLRFpnd9zr/HrPsYhIDo2negdSmgqdhmfDhrdv+wzKg+Wx1Z7B83+T4g2OSs/1xSYFBDWoBvgNR0REErYZll4qnGJ0EXCWz6AKYNcGP18ZqwkUHJP5wDGtDkoKoQP4Ly7l1HC/4YiISAvtQfU08t/1GZSItMxAYDb2Ov8ETSoUkRY4nsqGxZVeI4r6KZWrBv+JFWYuiuWAt3H74F2UNkKadyvuXNrJcywiIpK8vansOLrba0T5dijwqzo/3w74FHcsPgG2bUNcUhyX4s6vAz3HIiIirfUjKvvwLvQakYi00r241/pWnmMRkZy6iGjDYoLfcCJOIxrbAmBdnwG12bLARNzzfw9YwWtEknWH4M6ni/yGIiIiLfILou2nKX7Dya1lsIG/79X4+RHYSs7gOLwFrN6e0KRAvoo7x27yHIuIiLTe5UTbeS/6DUdEWuho3Gv9bM+xiEhO9QaewV1spvoNJ+ICoo2eX/gNJ1FDgJF1fr4s1omkgUFJ0ghcnYL/YOmoREQkXzqA+3FtiLkUK+tCO3QA92H794DYz3phqVzDbdjHsfdgkaQNwNLFl4DPsTTyIiKSX32BV4j2FYlIPi2He63/23MsIpJjiwOv4WrTDGribwzCBrxGAysCq5S3Fcv3DWni7/4mFNMnQP8m4kqrc4GVavxsGaIDg//B9qNIEp7CnVvreI5FRERaYwDwLO56v2wTf2Mg1n5bCmuHrAyMK38/pvyzxZMINoOOxO3bHUL3DwL+QnRg8A9owEZa6y7c+baN51hERKT1huPKzywA+jTxNwYT7cMbh7X1VgRGlX82MIlgRaRH/oVr563sORaRLlGBzOyZDuyCdSINwS42E7DGwQpYB9AIbKXb6PL3SwHDyo9vxqfAx1iqq6nAB+WvU8rfv4OtcAKbnb03MKfJ/5U2Y7Gl4ZdW+dko4AHcwOH7wFewhp9IEu4GNil//1WsoSEiIvkyG9gVa9uNwSZsTcEN7o3G2nIjyt+PxLX1lqS5leWfYZO5PsTadB8CH5W/n4y17SYAM5t8TmmxPPD/QreDrBvLYAOD65dvl4DTsTT5Iq10N9amA/tM97DHWEREpPU+BnYG/okN8o3F+tGCPrygbVetD2/JJv5fCWvnBX128T68yVif1QTy028nkhZ3A2uXv98Fy7InIpKY4diM52OB27EBuanYar14oWNf2yTgd8Ax5ViHtWRPtM8fsee1auz+pbBl4sHzngys1t7QpADWxZ1jT3qORUREkrcksC3wXeB6bEb5VFxaad/bf4GHgEuAE4DxWBsoCzqwSVzhfTkG2BQbDA3umwHs5ilGKZ4xuM9ub3mORUREWmsk1nY6HpuUtAgboPPdvith7aN3sPT2F2KT4rej+UUFIgKb4V5jD3qORUQyrjewJpYK6VrgVdI1CNid7QPgZqxBtAXZSTm6Hm6fhwcHR2LHI3h+HwKrtz06KYp3cY33rHTIiohIpV5Y2+4g4DKsLZGWQcBm2nZ3ASdhbbsBCe6npBxDNOZFwH7ArNB97wMb+gpQCut53DkYn4AoIiLZlOc+vH4J7ieRPOvEsrGUgHnAEn7DEZEs6QS+BPwEeBxLMdWqN/rpWCqpD4CJWDqBCeXvPyj/bEYL//9s4DHgFKxDpjOB/dcKD+FiDgb/RhItKP0hsIaX6KQoLsadbwd7jkVERLquA1sBfhKWOvALWte2mom13yZj7bk3gTfK308q/+zzFv7/eVid3NOxlXm9Eth/PbECtr/DnXJzYrf/gaWIF2m303Hn4fc9xyIiIs3pBWyMpSR/EphL6/vwJmFtuzewtt5ErO3X6j68WcDfgB9hKdmbSWkvUhTX4F47e3qORaQhXdD9GoGl3hwP7Fi+3YyF2OqiCVhjYTKWkuojbPBqSnn7pMm/PwxX22YUtnopyIe+NJYrfSzND/JNwVIZ3Istu/64yb+TpJ2Be0K318LifLj8PeXb22AzwkRaJXwu3grs5TEWERGpbwiWkml8eRvT5N9ZBPwH17b7ANeeC9p5U4Bp2AfP7loSV8dwKVyNm1G4tt0KNF+f/BOsTXdfefuwyb/TjE6sA2szag9S3gIcgnV2ibTb/wBPl7//G1azXERE0m8prO9uPD0ro7OAaB9e0G8X1IAO2nyfNvn3h1PZhzcCa5cG7bzlab4P70NcG+9Bmu9rFMmjvbCVt2ADhYf4C0WkMQ0Ott/iwDeAA4Ct6d6b8XzgX8BLuNV+b2D1KuYlGmX39QNWxlLjjCtv62GFWLvTsbQQeASr9Xc7Nvup3Xph+3lVXKfSVsBFuMKyU7GBwVfaHp0UTX9swHwgNmNwBP5f7yIi4iwG7A7sj3UUdbfd8yrwAq5tF2xzkg2z2/oAK2I1lYO23TpY+65vN/5OCcuKcT32QbnZjq6uOgE4r87P3weewwZZp2DvsUGH3NTQfSKt0ol1Bo/COohH0vrXhYiINGcItvrnACzFZnf78F7A+pcmYKv9Xgfexv9n+n649l2wrY9Nhu9OBoiF2ADh9cCfscwNIkW2OPaZom/562jsdSIiBdYLm0V+LZb2qatL94N6LqeVf3+xNsedhD5Y6tDjcXnXu5N+9C5s1kV3OqF66ltVYgnHPQU3SCjSDnfizr/tPMciIiLWMbQFVjtwOs217XbFOpyypic1deZiz/8gWtOuXZXK9KHVtgXlLd7uvBzVkZb2+APu3Pum51hERCSqH9ZOu5bupYXPQk3mRgZisQd9eBPpXh/ezdi+69PuwEVS5K+418WmnmMREY+WBX6NzT7uyhvpp9gb6WHAMh7ibZdlgMOxlE6f0bV9MxX4FZYCoZUGYDN5F9aI4xNggxbHIBJ3JO4crLcaQkREWmspbGBvMl1rv8wA7gCOwtJ05tUobMDvBrrX7j0fW52YhF5YHcH4oF+tLWjrfQz8DFu9JdIue+DOxes8xyIiImYF4P9h/T5daUtMA/4EHIytDsqr5bE+idvp+qS4D4FfoPrOUkwn4l4LZ3iORUQ82Bi4EUsh0OgN8znsQrEFzdd1ybLewJbAL7F0C4321zys42mjFsVzSp3/PRe4DVslUG07EZthJpK0MbhVEBM9xyIiUkTrAldhq9IatVVeAc7G0o+3M/NBWvTC2sKnYXXVGq3iW4h1Nm3Vw/97UoP/E2zB4OHb2Kz4LGbmkOwbhK2wCDqXi/g5UEQkLbYAbqVrE4z+Cfwc2ITupd/Miz5YiaSzsHSpjfbXXKzu2voeYhXxZSXca+Alz7GISJt0YLUEn6Txm+NE4HQs9ZFErY7NLnqbxvvxceDrJFc7cySW9rVWJ9aC0BZ+zDTgWJQ2QVrrWdw5p5RnIiLtsRPwEI3bJP/FskWs4yfMVFsROBWrsdOVSXP70716PmDvi3Mb/O2gw+9JLGV9ETv0JF3uw52fW3qORUSkaDqxtM7P0Lh98gbwU6zDX6LWwrJ8vUfj/fgI8FU/YYq03Wu4c3+s31BEpNW2pXGD4mPgImAzkhvMyrMOYHPgEmzwrdHMrW0S+J8X0fWaOQuwlaGXAcMT+N8ijZyGO/9+UOXnQ4AvtzMgEZEc2xR4lPptgc+wumFfofuDWUX1JSyV6IfU37cvY/VquqI38CK123CLsHbbzdiKRpG0OBZ3np5V5ecDgB3bGpGISDHshLUd6rVFpgAXorZDV3Vi/RFXYKnj6+3bJ7DVmiJ59hvcOf/dKj8fia1AFpEMWwvraKj3pjcBpSzqqX5YHZuXqb+vH6T5eoDj6Foa2GDW+d3Ayk3+L5FmbIQ7D/9W5efnYquXRUSkeathbbt6k4UmYiksl/QUYx70xVbvPUX9dtdTNJ74cl6V3wuO3wzgAqwOuEjaLI87Z1+p8vMfAd9ua0QiIvm2EfAw9dseL2L19QZ4ijEP+mN9eK/SuA9vPU8xirTa1rhz/Z4qP78UGN/OgEQkOaOxunf1ZijfC+yAVgkmqQObPXsftff9QuB6ul8M+vby79ZqtAQ/ewGtzhI/OoD3cYPUw0I/Wx6rW7O3h7hERPJgGHA59WvNPALshlYJJu3LwJ+p3w67g+rpeNal+uSu97AB3CVaG7pIj72CO2/DEw+HYSsvqs00FxGR7lkWqylYrx/pLySTkUqcTiyNaL0U/Quxut4jPMUo0iq9sCyCJaxu/aDQz8YB84BdPMQlIj3QARxO7SXyi7DZ5qoH1nprArdQu3H3CXAoXRuc3aTO3wm2ydjsMdWnEZ+uwJ2T+4Xuv6Z8374+ghIRybi9qZ/m8m6az0wgXbcS9n5Wa5BwJpaNIxic7QO8FHvMi9gqerXXJCt+hTt/jwvdf275vmN9BCUikhOd2CSL6dQemPojsIqvAAtkXeBOare3p2J1p0Xy5AbcOb5b6P7byvd9zUdQItKcFYAHqP1G9jhWI0/aayPqz0L6/+3dd7wcVfnH8c8tCakEEhJCkRZ6r9KC9P4DREE6iAUpagCpggKCCD8VsNACShcBQRCkg3SRKtIjvSfUQHpy7/398dz97cyZ2T67Z2fm+3695gXZMvvMnd3Zs+c55zn3U7mh9yjxycEebDbWGcDwJsQuUsqXgCOxchxBX6X4/ryq/7bVKL5/1ZgWEaneWGwUeak2xL+w9QSltVaifNn+R4CVgVMotteuAzbxEaxIlUYDPwHmd24fT/G9fUf/bUtho8n7sIS4iIjUbhzlS4jeBazlLbr82gBbJqXUebkVWMJXcCJ1WhY4ClsWK2gfiu/tif23fZliH95XWxWgiNSvAxuxOZ34L66n0ELx7WBbrORnqZHmh5V43t4xj++hOAtUjRLx5UzgDaxWf2GWxFAsYV2YHduNlTAuNCwOaHmUIiLpdAClK0G8COyKSsP7tgnwMPHnaDYwA1tPcGlfAYrU6GisGskh2MxXsFmuH1J8Xw8HLqfYtvtR68MUEUm1Dux6W/jdrMFf7WknwqW1g9tU4Dv+QhOpy++BV4A9Kf6OHElxCYT3+m9/gGI7b7fWhykitRgGXEP8l9V0bC0TlS1qH51Y6c8viD9nV2PJlYJu4K2Yx92HRpCJf4OBV7H35LPA9v2330bxvTqB8Hv3wNaHKSKSKvNhCaW4dsIcrFrAQG/RSZzdKa7XETe6fEF/oYnUpBsbWNqHdR7thnUSXUHxPX0k4YomR3uJVEQknYZTLNcXl3AKlicX/7qxczKN+HN2BTDEW3QitRkOvE1xItGW/bc/QPE9fTjh9/gerQ9TRKq1LNYhH/cFdR+qSd7OlqJ0CdgXKa4Jeahz37uEa0CL+LYZ4Q6ifwJnBf79HjAv8G+NrhMRKW0x7Doa1z4olKqU9jQWKx0ad+7eBNb1F5pITdagOIK8D3gC+Bnh3yPBtt1xfsIUEUmdFYAXiG8r/B1VhWpn4yi9XNBTqEqEpMcOhN+/9xAemDqFcDtvLz9hikglOwOfEf1S+gSV7UuLDuBbxJcM+xQbqVsYhf4ZNgt0QOyeRPy6hOJ7t1DyNq7R3Ad8z1OMIiLtbjNgMtHr5hfA99Eo8rTYnfjzOA39uJb0+CXF926hg6hU++4ETzGKiKTJN4ivIDUFK/En7a8DK7v9OdHz+CGwlb/QRGoSXDt9HuX78PbzFKOIlPEdrAPe/cD+G1jGY1xSn3HAM0TPZy82avdsVI5K2ttILJEdd11yt0M9xSgi0s52w0qGutfMScCqHuOS+ixK6bUIlUiRNBiCrSsdHDlequPop35CFBFJjUOIv4Y+CSzpMS6pz/LEr0U4DyVSJB3GYmWMq+nD0wQkkTZzMPEf3qtQnes0G0R49lXwR/gPPMYlUq09qdyo6EPvZxER156ES/gVtluABTzGJY3pxtaHjPsuPMljXCLV2pryI8kL2ym+AhQRSYGjib92Xg4M9hiXNGYY8eXk5wEHeoxLpFrfpro+vG/5ClBEon5E9AfaXKzcpGTDQURnDvRiC8KKtLu/UXnk0QRv0YmItJ+9iSYGe7GkksqIZsO+wHSi34dn+AxKpEqXUTlBeKq36ERE2tuxRK+Zs4Dv+gxKEtOB9W/EteUP8xiXSDU6gHup3Ien65VImziG6Ad0JrCjz6CkKXbGGozu+T7SZ1AiVVgCW1OpXCeS3sciIubbRH+MzcUShpItmxK/ztDpPoMSqcIoypeO70XvYxGROCcTvWZOA7b0GJM0xzeIH+R/sM+gRKqwHNb/XK4PT+9jkTbwVaI/yKZjpV4km7YFZhBtXOzuMyiRKkyg/Kijo/2FJiLSNjYBZhO+Ps7B1h6UbFoX+BiNxpX02YfS7boe4Ex/oYmItKV9iU8Mbu4zKGmqHbEJHG6J0R18BiVSheMp34d3qL/QREyH7wA8Wxn4JzB/4Lbp2Oyye71EVJ9VgZVK3DcTW1cnCWsD40rcNxW4M6HXaYWvYH+X4YHbpgEbAc96iUiksk7gYWA9oCvm/uNRKTURybelgMeA0YHb5gB7ADf6CKhOywFrlrivB7ghodcp14acBdyc0Ou0wtpYW3RU4La5wFbAA14iEqnOzVgHp1vuuBc4Gziq5RGJiLSntYEHgSGB26YC22N9e2mxBrB8ifumAbcl9DpfBpYscd/HpKvfczus/RtcS/JTYH3gv14iEqmsG3gC+80V14f3Q+B3LY1IRP7fQsBrhDP2s7GkUdqcSvmRCCsm9DovlHmN/yT0Gq20OdHyBK8S7lQSaTerEa27X9hO8BiXiIhvw7D2iDvzZmefQdXpcMq37TZM6HUeLPMa7yX0Gq20DtapFjyO94HFfQYlUsES2ABVt+xUD5YcFBERWAR4h/B1cgaWAEubsyg/a3yJBF6jA+vfKvU6aUqmFuyAzRgMHscLhCd8iLSb9ShdQv5wj3GJ5FoncA/RD+V3fAbVgErJwdMSeI0NKrxGGpODYPWd3WO5i+jIXZF28nPiP4c/9RmUiIhn1xG9Lh7nNaL6VUoOXpDAayxH+TUw0pgcBCsT7x7X48AAn0GJVHAE8R3Ev/EZlIhIm+jGKugEr5G9pHct6XLJwaQG/W5R4TXSmBwE+BHRY7nJa0QilZ1N/O+uH/kMSiTP4n58/dZrRI2plBx8h/jpy7W4oMJrpDU5CHAe0eP5vteIRMqbD3iZ6Oijkz3GJCLi035Ev8uvI70l9CslBz8jXFKrHqdVeI20JgcBfkb0eE72GZBIBZ3Ao4Tbdr3A730GJSLSJk4k+r3+C68RNaZScnASjbdhL6/wGmlNDgJcTPR4DvQakUh5Q4A3iCYIj/EYkwiQ3g6TRozDElnBDpV/ANtg09PT6FSssVTONtiMuHoMwkoyxigNvAAAIABJREFULVDmMc8Cq9e5f98GYH+bTQO3TQdWAd70EpFIZV8B7iN8Hb8MW7dmFJZAHIINDCiU2RhBcVbsNKw86WysHEsP8Dm2TukU7DP/Yf+W1mujiOTDwsCLwIKB254CxmPXtDQ6nMrlBPcGrq5z/53A65QvW/U+sGid+/etE7ge+GrgtrlY2VGtLS3tajXgacKDOm/E2ndjgIHAUKztV/hdNhybUQPWnpuNLZswneJAgtkU23ZTsLbd3CYeh4hIklbB2nUDA7fdCuyEdbSn0VnYpIVyxmOzJesxDLvmDyvzmEdJrkx9qw3C+kLWD9z2GbAydtwi7Wg7ouuJTsT6o0di7+vBVO7Dm4X9xg324U0GPqDYzutp1kGIZMGNREdef8lrRI2Lmzn4kvPvqxrY/17Ovv5LtM53mmcOgnV+fUL4mK71GpFI0QLYWgr7YzM9rsUa8+66Ss3YerGGxnPAHcC52KLJ2wJLoxK8IuKfO3p4JtY5kGZxMwfdtt3tDex/a2dfHwIfObeleeYg2Hfn24SP6V6vEYkUzQ+siyX5TwH+jM3i+Izmt+0Kn/nngbuxCjFHYms5LUsx2Sgi0g5uJ3r9WthrRI2LmznotvMuamD/366w77TPHATri/iC8DFd4jUikaKR2PJcBwCnA38B/kVx8FYztx4sUfgsdv38PVYdbxtgKdSHJzm3GdEPzQE+A0pIXHLweOffMyg/86+cO519nYiNVgjelvbkIEQbUH3YaC2RVloO2Bf4HfAAlphrRSdRvdss7PN/DVYvfTyNl7oTEanWGkQHLGVh7Ya45OBxhEvR9FD/ALc/Ofs+BxtpHbwt7clBgF2I/h138RqR5NFSwJ7YbOB/YJ8t3+23ctts4AVs9u1x2G/ocrNPRESaZUei16jdvEaUjLjkoNuHN5X6f1c/5OzruJjXS3tyEGzQcvCYeoC1vUYkedMBrIAN5D8X++x9iP+2XLltBvBvrALNEcBG2KxFkVy4j+iXYRZKq8YlB7fCOuyDt323jn0vRrjTrQcrQZXF5GAHNpIjeFx3eo1Ism4INgPvJODvRGdtpHWbCzyJNY4OwEb1iYg0w18IX39exMqFp11ccnBPom3Z4+vY9wiio1bXJJvJQbDSY8HjeppstP+lPQ0CNgdOAG7CRm77bpclsc3Dfu9NxAZUrpDUH0xEpIzHCF+L7vEbTmLikoMbA684t+1Tx76XIzyYbC6wZMzrZSE52AU8Q/i4/uo1Ism6YcD22FrmtxGtQJfWbQ7wODZJYT/smiE5kaeSIRsTXlMOrHxKn4dYWuUK4H8D/z6A2ksTfJPwuhf3Am81Flbb6sNmGzwYuG1rrJzjY14ikixaCas1vh22bmASI3R6sM/p+9h79UNsNFCwDjnY6MPCugzDsA70wtqEnViH8VBgEaxUy2hsjZtadWMj9tYGDu2/7WWspMFt2IzItK4DJiLtYyVgV+e2o8n2WlqXEW7PfhM4g9ras3sSHon+LDZ6NKuOxgbiFErorIn9qL/VW0SSNeOwdt32WGIwiQoKhbbdR1gH7hSs7RRcSxCsjVdYV2YI1q5z1yYcAozt30b3b7WWlOrC1kRcjeKA09cptu3u7Y9NRCQp2wDrBf7dh/XhZVUf1od3cuC2A6h9iaBvEh4E9XdsoEoW9QDHEC61vwuwKrYsikgSVqXYhzcea2s1qtDOewcbWF9o57l9eIVy81Dsw3PXJhyGLZU1hmI7r1YDsHL36wZuewH7bN2O9eHNrmO/Im3lKsJZ8Tv8hpOoUjMHF8ay/8HbV6xx325t8sLIpSzOHCy4l/CxXeo1Gkm7LqwRcQHwBvWN4pmFNW6vB34BHIgNeFgW6/xZHGs8/LYJ8XdjDY01sU74Y4E/0Fi5hBlYp+xhpH+9CBHx5zeEry1ZmhFWaubgUOx6H7x9wxr3/U/n+RP6b8/qzEGwtXqDx3aL33Ak5TqxUpu/BSZR/wjtl4AbscGc38EGja0ADAcWxErL/6kJ8XdhicLVgZ2Bo7AZgfdR/0zHWcBdWMf94k2IWUTy50bC15kb/IaTqLiZgxths3V6ArcVKndVqxMbzB/c765YMsN9vSzMHCx4mPCxnes3HEm5AVhJ44lEP0+1tIv+g1W5OR1L2m+EDSgbCiyD9Yud0aT4FwXWAr6GlRX+I/AI9VcrmwbcDBwMLNSEmEWabkEs+x58Y2/pNaJklUoOgn14g7efVsN+xzvPDdY8z3JycFvCxzad+tdrlPxaD1vDqdZOls+wcranAjthjYYuKjsUW2i41UZiHdM/wEY61tpJNhdLFO6DNZJERKoxCPiY8PVkD68RJatUchDgEuf2C2rY7/KES03NoTi6NMvJwXUJH9s86l+vUfJrNeBMau8o+gJba/AMrJN2Oaqr4PMN4M+JHkF1RmDt2EOw680LhK8blbYebLDlt/r3JSJSq7FE+5w28BpRskolByE6WP2EGva7jfPcj7DEYNaTg18j2qeSxCx+yZcNsbKaU6itnfcpNrvuFCypuDTVVWg4mnC1v1YZhU02mIBNpHqV2o53DpZr2AObwSiSCt8i/EZ+jdpLqbSzcsnB3Zzb36G6RAPAxc5zJwbuy3JysINorff9vEYkabEE8BOiM27Lba8AF2KzAVem/mtTJ/WtSdAMo4AdsMbRvVj5gWo7z67AftRkZfaPiDTHzsR3fmRFueTgZs7ttXSAFEqQFrbgKPwsJwfBSvYEj+8Iv+FISozFOm/cNY3KbW9hI7S/i83Oq/a3V5x9G3hukkZg7bOfYB1g7sDbUttMbObuzjT2dxCRfPkB4WvJs37DSVy55OD+zu2TqP638Z+c557df3vWk4PdWLs1eHxf8xqRpMXSWL/Vf6m+nTcJOA8r+7si9fdddQF7NxB7kkYD/4PlF+4j2udfapuKVdvbAvXhSZu7gfCb9xS/4SSuXHJwINHSf1tXsc+h2Ic8+LyNA/dnOTkI0b/ptX7DkTa3DnA50TK+cdsMrOzSsf3PS1K7fhkPwa5Jv8HWp6mmkfFfbCSTZhOKSJyJhK8ZWSsfVC45GDeIaa8q9tkFvO08b5fA/VlPDk4gfHz3+g1H2twa2OCtGVRus8zFyq0X2nZJtsfatW03GGvbnQE8QXVtu9ewv9GCHuIVkXS5g/D14xi/4SSuXHIwroT8xjH7cI2guC5tYVuz/76sJwch+je9xG840uYKfXjVJMGCfXgrJxxHu7bzhlLsw3uT6tp5L2O/tzRrV9pOJ9Ek19peI0peueQg2LTo4H1XVrHPSqOVsp4cXI/w8X1C+160xY9ubBq9u3ZT3PYpNhN3WzTtHmAl7AdeNaPwP8JqtC/mJVIRaVdvEL5WbOM1muSVSw4CnOTcd3sV+9zBec5kbE2KgqwnB5chfHxz0I9XCevEZri5Jd3iti+wTqWd0UAmsDV0JgD/ovLfbio2m2UZL5GKSLubD1uvK3jdWNFrRMkrlxwE+INz30R3BzEOdp4T7KPLQ3JwM8LH967XaKQdDcQqMlQzqOljbOmGrclWdZp6rQocDzxH5b/dFOBnWPUNkbawCuE36YdkL8lTKTm4jnPfDCqvoef+KP6xc3/Wk4MdRNcyylqDVOrTjZWJqjR6ZhbwV+Dr2NpYEm81bNR5pfV75mAlumpZkF1EsmlhotfbrP1oq5QcXBJb16twXw+V19C71tnfr537s54chGjJoPF+w5E20YmVdaq0brLWSa7O8lilHneGs7vNw9ZUXM5PmCLSpr5M+FrxhtdomqNScnAT576pVB7Q9KjznAmB+/KQHBxAdOak1pcWsPfGoVjCuFy7ZCZwHVZZZaCXSNNhTeBXVP57zsaqcCzqJ0yRIncG3C1+w2mKSslBiM7Q+W6Z/VXT4ZT15CDAbYSPsV3W/BA/OrC69S9S/gvwWWzU3kg/YaZWJzba70rKr1E4E2uIjPISpYi0gx0JXxce9RtOU1RKDkJ0INfxZfY3kugo/DWcx+QhOXgF4WM83G840ga2BZ6ifNtuEvZeGeMpxjTbELiI8msUzgHOBxbxFKOItJdDCF8jrvEbTlNUSg52EB3QtE+Z/S1P9Lo6OnB/HpKDAA8QPsZd/YYjnnVg1b4qrSf4FPAdrDSvVK8T2BIb6FWuPOt04BdUnqQk0jQ/I/ymPN1vOE1RTXLwSOf+h8rs7yTnsXfEPCYPycEzCB/jyV6jEZ82JNrQDG69WA3yncjezGQfxmKfN3e91OD2BfYZHe4nRBHx6IeErwfVlFpKm2qSg+4AuJcp/R10mPPYJ2Mek4fk4FGEj/H3fsMRj9YD7qZ8Z9FDwO7Yep3SmNHYWj3uuqdu59EZqPNIJO9+TfjacILfcJqiUnIQ4CfO/XeW2d+ZzmNvcO7PS3LQXVLpaL/hiEdbUb58aA/qw0vSIlgfnluBL7h9jLUFtdSStNzlhN+M5WbMpVU1ycEx2Oih4GPiymR2EC0Bs1fM4/KQHPwe4WO8zG844sEYbBRMqS+3GdhI5xV8BZhxQ7DP4UuUPgdvYetoiUh+nE34OlBuxlxaVZMcHAp87jxmwxL7c38c/yDmMXlIDn6d8DFmsaKIlLcANpOtl/h2xWzgEqIzayUZA7GBDf+mdNtuMjbSX0Ty6XrC14S9/YbTFNUkBxfHyi8HkxlxS2x0Ey3vt4vzmLwkB91JEef5DUc8WBRb3qdUG2Ma8FtgWV8BZtwwbFBqudmarxHNWYg01S2E34Q7+Q2nKapJDgLc5DzmtJjHbO485jPia5vnITm4M+Fj/JvfcKTFdqf0zLUebO2mpXwFlzOd2Pl4jdINjGsJl04Rkey6lPDn/0Cv0TRHNclBgD84j7kg5jHu+tuzgYViHpeH5ODGhI/xEb/hSIvtSOmZa71YW0KdRa3RgbXtyq3zeAtaL0okj/5B+Fqwhd9wmqKa5CBEZ7jHzaJ0y+1PxtZYC8pLcnAvwsd4td9wpIU6sMFHHxHfppiDrX+nEuat0Q0cRPl1Ca9FywVJi7jrsWzuN5ymqDY5+DXnMe8QLZNzmfOYuE4myEdycEvCx3iP33CkRSqNNLoLWN1bdPk2EGtgTCb+3HzSf7+IZNu1hD/7u/sNpymqTQ5+xXlM3KAutzzXX0q8Zh6Sg2sQPsZn/IYjLTKGaDUZt223trfo8m0A1nZ7j/hz8xkwARsoJiL58C/C14Ev+w2nKapNDu7rPGYS0RKI1zmP+XXMfvKSHNQA/3xaGmvLlRv8tZy36PJtCFZK9FPiz88HwG7eopPcuI/wG29Tr9E0R7XJwYHAFOdxWwfuH4at4xW8f4MSr5mH5OCmhI/xPq/RSCvsRrREW2F7nNLl2qS1FgD+l9KLHl+NXc9EJJv+Qvgz/3W/4TRFtcnBDqJlW4Ll4LuJJv3+p8Rr5iE56M6ifM5vONIC21C6EsTz2GBA8W8otu79LOLP1d+BBb1FJyKt9Djhz/+6fsNpimqTg4OJdqpvHLh/JNHrZlxZ7LwkB3cg+t0h2bYvtmZxXNvhEbJ5/UijkcA5hEslB7dL0VqE3uRhBN5M599xJTLzYg7wJ+e2AwL//w3CHeqTsFFbeTXc+fd0L1FIK3QBZ2AjitzzPhM4BWusZ7EBnUafAccA62DraLn27L99pVYGJSIto7ZdUR82Iyoo2LbbARgb+Pdk4I5mB9XG1LbLjw5spPKtRMvozgPOxGYLqjJIe5iOtbdXJX5A5g7YOoXq5BPJPrXzimYSrfgQbOftjSX+Cp4i31UR1M7Lj26sD+8KoteIGcBxwCbE9xdJ632CDX5dD3g65v4DsGTuMq0MSvLjGsLZ6P38htMU1c4cBFjLedwMbBYOwAPOfceVec08zBw8gOhsJMmeUcCdxI9eeRBYwV9oUoVurPNvJtHz9zlWTllEsuVcwp/1I/yG0xTVzhwEWJzwKMweiut03eDs45dlXjMPMwfddXnu8huONMlwojOMC9vTqIRou+vASo3GVfOYSTbXmRWRolsIf+6z+Huu2pmDEF0veSrFZMgTzn0/KLGPvMwcPJTwMV7sNxxpktHY4K64dt7twJL+QpMqDMD68OKqRXwMbOsvtHzKw8zBt5x/L+UjiDbyNDbqsmAwtlbP0sD4wO29wFUtjKsdLe38+00vUUgzrYaNrtvauX02cBi2ltPLrQ5KalIY/b8W0VGShc7BE1sdlIg0ldp2Ye8QnmnTiZXYGYXNtgm6okUxtSu17bJvHFaSzi03PA84Hpt59lSrg5Ka9AETsXb6I859g4A/Yh3r7rpbIpINaueFPQy8FPj3/MCu2EzrdQK3z0ED2tXOy751sD7tLZzbZwLfBrZD573dzcX68NYDXnDuG4mVAz6q1UHlWR6Sg685/1aZObjM+fcBwLcI/8C6A3i7ZRG1p5Wdf7/uJQpplrWAe4ElnNvfBTYDzsM6JyQdXsLWSL3Eub0Dm139q5ZHJCLNorZd1KXOv7+JVcsIlpp6nGxWeqiF2nbZtjyWKHerPnyEJcrPwGbWSjq8ia0Bf2bMfUdggx26WhqRiLSC285zv7vz6Ern3wcQnUV9M/Z9l2fubwK187JlXazq16LO7W9j7YU/tjwiacSzwPrAdc7tXVi1mzNaHpFk1oaEp6i+6jecpqilrCjYFOzZgcf2Yo2I4PP3qPCaeSgr+hbhY/yy33AkQetg09Xdz839hNdmknQ6iPA1rrCdTz4GxYhk3XKEP9ufkb3Pdi1lRcHKS011Hu+27Q6r8Jp5KCv6FOFjdGdWSnqtiA3wcj83T6FZJ1mwD7Z2lHt+r8ZKzItIdmxF9vuaaikrClYuvifw2B5sDa/g83cq8/w8lBXtAD4kfIyreI1IkjSe6G+dPmzA/xiPcUkyDsJmP7vn9399BiXZMYhoJ3HW6g/XmhwE+GvMc4KdbIMrPD/rycFxhI9vFuHR95JepRoVv0OdC1myCfEJ4AvJXhJBJG86iCa+1in7jPSpNTkIcFHMcwrbbGChCs/PenJwJOH2ay+V/yaSDmsCU4i+76/AfgtKNqxFfAL4Gmz9GhHJhhFEE2FZ6/yvNTkINmOqVDtvMuWvg3lIDq5O+Pimot/9WbElMI34xJEqCGTHlsT31Z6DSslLAu4n/Mb6vt9wEldPcnCXmOcUtvOqeM2sJwd/SPj47vEbjiRkDeALou/5030GJU2zBvGdhWf7DEpEEnE94c/1yV6jSV49ycGNY55T2K6t4jWznhzcm/DxuevUSjqNIzpYoA+4AHUKZtGyWLlR93xf7jMoEUnc44Q/424JzbSrJznotmOCW6UlNPKQHDye8PHd7DccSciXgRlE378n+gxKmmY9orOi+4Cf+wxKsuEowm+qh/yGk7h6koPdRDuBCls15TOznhx8lPDxHe43HEnAKKyssPt+Vx3rbCtVZuy7PoMSkYZ9k/Bn+mWyNaKwnuQgwIsxz+ujuvKZWU8O3kL4+E71G44kYBi2Xon7fj+fbF0PJGwJ4BWi5/0Yn0GJSKJOIvz5vsNvOImrJzk4CPg05nl92Ky5cvKQHPw3+r2fNWOx9QSVGMyXuIogvdgACZG6LYO9kYJvrCwtalxPchDiGyQvVfmaWU4OuuUIetBaJWk3ALiP6Pv9WI8x1esK4IkS21kJvUYXVru91OtMSOh1WmUF4B3C534WlX+AiUj7GkW0bPwmXiNKVr3JwRNinvcB1ZXNznJy8EvAPMLHt7bXiKRRncCNRN/vaVyb5HeUbnP9IcHX+VuZ1zkpwddphcWBSYTP/Txge59BiUhiViXbfTL1JAfBlshwn/d4Fc/LenJwA8LHNhdLLEl6DSI6aaMXq/KWNtdRuv2V1Iy4+YAHy7zO9xJ6nVZZFfsNGzz/M9DvN2nQvYTfVBP9hpOoepODA4EFnW1ola+Z5eTgpWR7lFoenU/0M3KO14jq55ZYCW4zsc9xo3Yo8xpp7Xhbg2id+g+wDmMRSadrCX+mb/AbTqLqTQ4OINq2G1bla2Y5Ofgrwsf2b7/hSAJOI/oZudJrRPW7jdJtrh5splyjvlzmNfqAPybwGq22FPAh4eP4HFjFY0wikpx/Ef58Z2lpiHqTg/MRbecNqfJ5WU4OXkd2fxPk1cVE37NpXQ7oBUq3v76g+t9q5exW5jX6SOcSHGsD0wkfx7vAYj6DknTbg/AbajbJ/NBqB/UmBxuR1eTg0sAcwsf2Na8RSaPcz34fcDfVzaJoR+WSg33AwQm8htvhnoXkIFiDyZ1F/gBak0gkrbYk/HnuBVbzGlFy6k0ONiKrycGFiK43fJDXiKRRWxH9Pn8cGOwzqAaUSw72YTOCG3VehddIY3IQ7Hsg7nfpQJ9BiUgivkX4sz0dWNhrRMmpNzlYrywnB1fCBtIEj21brxFJow4k+n69mfT225RLDvYB+yXwGu7yCVlIDoL9bdxjuRMtHyB16iJaeiQrC5crOZicPxM+rhdJ7xeQwGhgMuFz+jrWUZhWlZKDjTbyR2IlN7OYHAQ4hejxpK1MqogUPUb483yX33ASo+Rgcn5P+LjeJ71JJIH5gTeJntPFfQbVoErJwUk01gkyCPikwmukNTkIVmLMPR6tKSqSfgOIXu/P9xpRcpQcTM7NhI/raZQ4SLPFiLZZXgJG+AyqQZWSg3c3uP+FifbRZyU5CPBLosejNUWlbu7og16sNnXaKTmYjPFERyEnMYJD/LmK8PmcSfpnlbjJwV6is11XamD/hzn7ctf0SntysAP4O+HjmYZKE4ik1c5Er1E7eo0oGUoOJmNVom3WI7xGJI36DeHzOQ9rw6dZXHLQbX9t3MD+4yroZCk5CLYmd/B45tBYe1hE2sP3iV7z1/AaUTKUHEzGNkSPS5W/0u2vRPtqlvcaUePc5OBcwrNde4AlG9j/0c7+49p5Jzewf9+6iC4V9xkwxmdQkl6d2EKcwTfUi6R/9LCSg42bD3ie8DE9hWYNptkGRJO9R3uNKBlucnAecJNz2y8a2L97jbye6PUlzclBgEWAjwkf0xVeIxKRRtxF+PP8Dsmsv+qTkoON6ya6XtGrWJtP0mlFogOizvQaUTLikoM3OP+e2MD+b3f2Fde2S3tycATwFuFjyspMcpE8G4DNGgp+tp/uvz3NlBxs3FDgFcLH9DCaNZhmmxN9nx7mNaJkuMnBqdhsweBtP2lg//9x9hXXzju5gf23gyWxdaWDx3Sh14gk1TYnmjA4x2tEjVNysHG/I3w8vaR/FHLePUA02dvlNaJkxCUHv+bc9i71Hesqzn5mE79mY9qTg2DrTbmf+SyMQhXJozWJtkuu9BpR45QcbNxJRP+Gu3iNSBrlJsxexUpmpl1ccnAX599TgSF17HsxrK0YbO/sGvN6aU8OQrQ93Ads4TUiEUnCjkQ/2z/3GlHjlBxs3B+J9ous4zUiaZTb1/UI2Uj2xiUH3bX0XqO+Y13f2c80YG+in/eTGzmANuH+Pp4HrOA1Ikm1iUQ/KN/2GlFjlBxszP5E/37neo1IGrUp0XO6qdeIkhOXHBwITHFu366Off/a2cc12LUki8nBuJnk13iNSEQacTrRa9WRXiNqjJKDjdmFcLkeXePTb2Wi5zQrpcPikoMLEp0RsU8d+z7e2cc92ICKLCYHAe4kerwikn7uciG92CDWtFJysDE/IJt9FHm2PdHP+Je9RpScuOTgYKw0ZvD2TerY9/nOPi4hftmNkxs5gDbRDTxH9HhF6jI/8DrhN9RM0rv+4GDsB2Rwa3aZBff15m/y6zXLRsAsoiM2hvsMShp2C+FzepvfcBIVlxyE6Bo8V9e4326iHcPbk93kIERHofYAy3iNSETqNR/RkirzgG19BtWA+Yi2tQY2+TVHOK83osmv1yyrEi07MxkY7TMoadilhM/pY16jSVZccnAB4KfObXfWse8XnX3sR7aTg+4I+j40k0QkC0Zi1XGCn+1p2PUsjeL68Lqb/Jru66W1z2sLopMVsrBcVN65a8pd7zecRMUlBwEudm6/uMb9DgI+cfaxKdlNDgLsTvi45gCLeo1IUm0tYDrhN9V7WOkVyYcvEU2GpLmBKWZxwuWT+oCNvUaUrFLJwbWc22dS25pbbgPifewHSpaTgx3YmhXBY0t7iRqRPFuO6A+kT0j/IvZSvVFYqUn3R+NXfAYlDRtB9HdblkrElkoOLkl4tmQPsEQN+93I2ecXwDCynRyE6Dq0WpNGJBs2JDqw+w1gjMeYpLXGAR8Rfg98BqzkMyhp2LJEl/9a22tEySqVHBxPfDutWns6z38d6+PKcnKwk+jAtxO8RiSp9zWiF6CXUIIwD8YCzxM+92kvTSHmx4TP65N+w0lcqeQgwL+d+w6uYb/uOj5n9N+e5eQgwIGEj+1drMEhIum0FdHRxG9hPzol2xbEZpO531nf8xmUJMJdJ/h1svVdXSo5CFYWs94OEHcpjcKI9KwnB91Osc/RjBKRrIhbEuZ5YBGfQUlLLEm03HYP8D8+g5JEnEb4vD7oN5zElUoOguUggvftV8N+b3eee1L/7VlODgIcRvjYXvEbjmTBz4l+aF4HlvYZlDTVEsB/yfbFMs/cjsHD/IaTuHLJwSOc+6pdR2AU0VGYhdF3WU8ODsUaZ8HjS2uJaREx7rWwD5sNvYrPoKSpxgDPED3vv/cZlCTmVuI7P7KiXHJwP+f2Sdio8EoGA586zx3ff1/Wk4PdRMsPqvNYJDvOIXoNexGVlsuypbHlf9zzfozPoCQx7jpyB/oNJ3HlkoMnOPfdXeU+FyNcMa2X4hI5WU8OLgDMIHx8q3uNSFKvk+iMGSUIs2sc8CbR830d1f3Qlva2COHZwD3YLNEsKZccHIOVT4tL8pVzuPOcRwP3ZT05CPBnwsd3mt9wRCQB7hoOfcAH2Hp0ki2LEv3R3YeVFmz2GtzSfEOJDmBa2WtEySuXHByMlUwL3ldNufx9nee8RvG3TtaTg2ADA4LHd4HfcEQkQd3EXzdfxpYYkWxZCVsCyj3fl/lF4a6UAAAfGElEQVQMShKzDOHzOofalsdJg3LJwbgkXzW5CLdi2j2B+7KeHAT4G+Hj+7HfcCQLBhKfIHwHWM9jXJKs9YmOIi0kBtV5lA3u4rSPln94KpVLDgL81bn/F1Xss1w50jwkB90OtPv9hiMiCegE/kD0+vUhsJm/sCRhqxItMdUH3AkM8RiXJGdzwuc2i6WDyiUHITrYYWIV+7zbec6JgfvykBzcjvDxPe83HBFJ2GCis8oLAyHW8BiXJGsTYDLR83w50OUxLknOAYTP7b1+w2mKcslBsN8twft/UsU+3XX39g/cl4fkoLvkwG1+w5Gs6MK+YNwP0Czgux7jkmTsT3TacR9wNTbyTLLhV2Q7iQWVk4Nfde5/l/IN57Wcx88kPFIrD8nBJQkf33R0XRDJgg7gt0SvYfOAY1HFgLTbA5hG9PzeitYXy5LjyHYSCyonBzdx7ptK+eT34oRHofdgyyoU5CE5OJzoSPwFyj5DRNKm1CD/mWSvLGEeHUS0KlIfcBHZWnc4784j20ksqJwc3Me5P1jtIc7GzuO/AIYF7s9DcnAlwsf3KfptLwnpAi4h+iEqjExRR0P6zIeNro07p1eiBEDWuJ0ru/sNpykqJQe7sdJ5wcdsV2Z/v3Eee7Vzfx6Sg2DrkQWPcXm/4YhIQjqAs4hvB9wIzO8vNKlTF3AG4TLihe1mrO0n2XEF4XN8iN9wmqJSchDgJef+fcrs76fOY+9y7s9DchDgWcLHuJHfcESkCbqIfk8UtguxBKKkyyBK98tegBKDWXM/4XO8o99wmqJScjBunehNyuzvIuexf3Duz0NysINo2X2VlZbEdBKdfVTY/gWs6C80qdFKRBMphZGjZ6BGRRa9TPhcZ7GkSKXkIMDZzmPchF/BQGCK89htncfkJTn4AOFj3MZvOCKSsBOw2TPu9ew5bAa1pMNSwH2U7gRUmfjseZjwed7SbzhNUU1y8ETn/jtL7KuDaKldN5GYl+SgO6Nob7/hiEiTdAPnEt82eBAY5y80qdHqwDNEz2MPcBKaGZRFbxM+18v5DacpKiUHwX7HBB9zcYl9VZNIzENyEOAJwsc43m84kkW7YB9Y9wM1B0ssaQRS++rGyoXNInr+viCbs8nEuOXFslg+qJrk4KrOY9xSoQVfdx73DtESpHlJDl5G+Bi/7TccEWmC7YCPiV7T5mKzqIf6C00q6MDKS31O9PxpCYBse4Pw+V7WazTNUU1ysFKp0IJNnf3ElSDNS3LQnTV+rN9wRKTJ9sGWh3CvbzOwz7/WqGtfA7BzNJvo+ZsK7OovNGmiTsKDN3vJZgWQapKDGzmPcUuFFuznPC6uBGlekoPXET7GvfyGI1m1AtEPcWF7BljPX2hSwhpERw8UtklY0kSyqZNwibFesjmyrJrkIMBTzuMOjnnMzc5jTo95TF6Sg2551Ql+wxGRJlmC+KoCfdhsm839hSYlLAv8g/hz9jawvr/QpAU+InzOR/kNpymqSQ6ClQcNPubEmMdc4jxmYsxj8pIcdMurnuo3HBFpgTWxzvK4NsPTwNr+QpMSNgCeJ/6cvYhVBJNsGkr4fE/3G07TVJMcBHu/Bx+3f8xj7qFyWzAvyUG3vKoGi0rTjCB+keM+bKT52cBC3qKTgoWAc7BzEneurkPrCmXdEPLRsKg2OTjBedw/nfsXJrrAd1zZ5LwkB08nfIzH+w1HRJpoCHAp8e2FHuyHxqK+gpP/NwL4OfGVIPqAO4DR3qKTVnFngbiz4LKg2uTgPs5j/kt4INxQorNr49bZy0ty8EjCx/hrv+GISIuMAm4lvu0wGziT+Ko60lpjgfOIL/vfh60lqaoe2bYQ4XP+od9wmqba5ODxzuPuce5fkvDnpVQVibwkB88hfIwa4C9NtzvRtbkK2xdYqVEln1pvKFZ+wF2ItLBNJn60hWTPAKLJ+yyqNjk4imiHanDU3VHOfY+U2E9ekoPuWrM/8huOiLTA9sCbxLcfpmNtuyyWp253A7ESopOJPzef9d+fxeoAEuW28Uf4Dacpqk0Oxq0zE1xf5ZvOfZOI/5zkJTnodrL9wm84ItJiu2PJhri2xCdYP9Jgb9Hl1zDsbx+3jFMf8D62/Ilk3wiqS5qlXbXJwcUIl5DvBZYJ3O9WRLirxH7ykhw8j/AxHuI3HMmLBYkuEuqOcjiWbNZIbjcDsI6h9yh9Pq5Fszrzxq1RP8hvOE1RbXIQ4HrnscFOkf849x1UYh95SQ661/a4Mqwikj3zY2WFS41a/hh1HrVKJ9aRV6ocWB9wC7b2muSH29bP4vmvNjkIViY0+LiLAvfd59z34xL7yEty8OeEjzGu9JaIZNvCwOWUble8g/0O7vYVYI4UBn99QPk+vCyWD5d43UT7trI4+K/a5CDA7c5jf9p/ewe2BEbwvn1L7CMvycErCB+jJgZJS+0IvEHpL7S3sI4klSpI3kjgOKwRV+rv/wawg6f4xK93Cb8XxvkNpylqSQ66jYJ3sYXY13Nun0npTqi8JAfd0jMarSiSL5sCL1G6bfEBcBIwxleAGTYcKwPzKqX//u8De/gKULx6hvB7IYtrTNaSHNzIedxUrNTq0oTX3u4BvlRiH3lJDroJge/5DUdEPPoa5fuQXsdKEasaWPJGAz/B2nKl/v7/Bbb0FaB45VZEWMRvOE1RS3JwT+exr2GJwU1j9lGq1H5ekoP3ET7G7b1GI7lUzaiXmdiPkpU9xZgly2Ij+6dR+u/9EZaUzeJsManOg4TfE1v7DacpakkOdhNthG8HnOvcdlWZfeQlOegu/qzF6kXypxtr27kDTYLbbKxtt7qnGLNkUeyH6ieU/nurdL/8lfB7Yi+/4TRFLclBiLZZ9gVOcW67vczz85IcdH8XbOM3HBHxbAjWX1Su3fE5VlFmRU8xZslyWB+eu3ZwcFP1NXmS8HtiY7/hNEUtycFBRK9RXwEucW6bWGYfeUkOusuD6Lot3owATqN80qoHK4O0O0pc1WIw8A1sRk+pcl+FjqNTUceRWMdG8L1xrN9wmqKW5CDAr53H/xUrk1dtZ0kekoPDCdd270HXE5E8G4KtVeWOZHW3u7FO+WF+wkylgcAuWNnr4HXX3WZi318qDy//S/i98Uu/4TRFrclBdy29e7BZL9UmUfOQHOzCfiMGj3FprxGJSLsYiX23zKB8H96NwK4ocVWLIcDewJ2EZ7O721RsNqHa0PJnwu+NH/gNpylqSQ4CnO88/lqibZpySdQ8JAdHEz6+OSjfIm1gEeB3lE8S9gGfYT++tsDWVpGwLiwZcQmlFygOJgV/g9WRFwFbJ85NhGVNrcnBVSn/OXoH+9yVkofkoHuML/gNR0TaxCjgTKztVu46Og24EitlojVrojqATYALiA5OiUsKTgSW8BKptKPdCL9HHvQbTlPUmhxcjPLJ9c8oXWoK8pEcdI9xst9wRKQNLY61TcolCfuwWTwXYrN3srgeWqO6sOpElxNNYMQlBX+FBn9J0ZGE3yPlqlqlVa3JwfUp/zmaRPlrUR6Sg+4xPuE3HJGwkdhspbcp/2EudMqfhXVM53k00iCs/OM5lC/lVdjeAo6m/I9mySe3I+BzsvfZqjU5CPZFWerzdFqF5+YhOfgrwsf3B7/hiEibKayJ5y4CH7dNxko3b0/5zvmsGwBshiVX36Dy3+0D4KdoTUeJWpzwe2Uu2VvXvdbkIFjZ0FKfp/MrPDcPycHjCB/fTX7DEZE2Nho4EXiPyu2VN7HfwptjbZ28GowlBH9H+aWWCttrwBGoOo9EuWspf0j5wetpVGtyMO45we3HFZ6bh+TgeYSP73d+wxGJ1wXsBDxM5S/KPqwO911YYjEPdXKXxtb1uZbKMwSDIwH2J9+NMCmvg2iCOWuL0taTHPwBpT9Xla43eUgOvkz4+HbzG46ItKlOrG13F9W1W2ZSbNut4yHeVlsYa6ddS+WSrIXt31h7cLCHeCU9niX8vtnPbziJqyc5uFfMcwrbBhWem4fk4COEj+9gv+GISAoMxJYCepTa+vAmAEt6iLfVlqHYh/c5tfXhqbKGlNJNtLLIpl4jSl49ycFjif9M9QBfqvDcrCcHO7FJQ8Hj28FrRCJVWAc4m+pGIhW2l4FLgUOAtUn3l+kA7G9wKHZMk6j+7/AutubMWq0OWlJrIuH30DV+w0lcPcnBkcAsop+vh6p4btaTg+MJH9tsbJaQiEg5qwK/ILoQerntNaz86A+wcjEDWx51crqA1YHvAhcDz1H932EK8Htgw5ZHLWl1OuH30D1+w0lcPcnBQcQn4V+mctm7rCcHVyJ8bL3YDFQRkWqtD/wWqwhRbfvmBexa+j1gDdLfh7ce8H2sXGg11TMK29tY5YjVWh61pNUVhN9Dl/gNJ3H1JAfHYtUy3M/XHVU8N+vJwW0IH9s0tN6gpEgX9ia+jOpH2hS2GdgaG7/GRt58mfYsq7kg1pA6ACuZ+hCVa7i72+dYEnErsjedXJrvK4TfT7PI1rqU9SQHwRb8vtbZdq7ieVlPDl5J+Niu9RuOiKRMJza6dSK2Jk0t7Z1ZwD+x9ZMPxJJlo1obflXmB9YF9sGu//dReY3tuHbsn4D/QRUgpHarEE32LO81omTVkxwEm63itu32ruJ5WU8O/obwsd3rNxwRSbFubEbKVdhMwVraPtOA+4FfYm2o9YARrQ2/KiOxGecHYkv9PIJVv6jlWD/DlubYDGsbi9RiO6KfnSyVkK8nOQhwKtF23jZVPC/rycEbCB/bZX7DEanfEODrwEVUtz5hqW0y1uC4CDgK+0G4OfYjenQT4h7dv+/N+1/r6P7XfgAbCV7vcbyFdaztikpLSeNeJPz+OsNvOImqNzlYrywnB5chOhpra68RiUiazYclv87DZgnW2yb6CCtL/wespMw+wJbYbMWFqTwrqFajgJWxJOeewJHABcA/qK3qhbu9j4383QPNyJbGuWUiL/YbTqLqTQ7WK8vJwVHAF4SPbS+vEYlIVgwDvoFdLxttH/0DuBD4EXaN2hRriyU9SKwDW895VWALrE15DPYd+hC2vlu9x/EGtsbtzmjWjjSmk+ga5Sf5DChh9SYH65Xl5OBKWGnV4LFt7DWiDEjzNPe0mwFc37+BTbnfHhsxMZ7qR1WP6d++UuL+udgX/hRsNM88bBTQLGAONvqpt/+xncBQrMzVICxJNwD7cTq6/3WSes/MwRojt2M/iJ9LaL8iAOcSXpD2UKy072Q/4UibOpHwNe0FsleqTERaZzZwS/8GsALWttsea6dV23EyCtiof4vTg7XrPsTadnOwdt1MrJ1X6Bjvwa5xQ7DE5Xz9/9+Fte3GYO27pMqbzsPW6bkNa9893R+HSBJ+S7gU7f7Y4K9X/IQjbepYrAO/4F1shLmISKOmUZy904GVDi304W1E9X1lY/u3zUrcP5dwO28e1n84u3+bgbXxOrA+vGFYv91grK1Z6MMrtPOS6sObjU0KKPThvZjQfkV6scGVZwZum4D16X3iJSJpVycRnp38JDaoViRzhmEjh47Ffsy8S/2jedplewdLhB7Tf2xDE/triUQNIjqab6LXiJKjmYPJWIvoiKN9vEYkIlk2GBv8dSS2Fm4taxW26/YBcBNwAjazcf7E/loiUV3YenrB92BWkj6aOZiMZYiusT3Ba0QikhfDsZl5xwM3YrMDfbfTGt3eAq7DZjdugg0wE2mWoURnsp7jNaLkaOZgMjbEEsnB49rVa0QiLbY4Vob0F1iS7Tls5I7vBoO7zeqP7XrgdOBrwGJN+HuIVPJDwu/NeVgt/bRTcrBxXdg6X8Fjeh7NpheR1hoL7EJxPYlnqH2Nl1Zsc7DR4X/FRvTuASzZhL+HSCV7E31/7uA1omQoOZiMvxM+prdRZ7aI+LMEVob0DKwN9QLt24f3HywReBrwVWCRJvw9RCo5juhvkDW9RpQMJQcbNwB4ivAxPYnWOE2EOkLT4x2Ks+8KuoClgOWBFbHRkmOx9WjG9P9/0gseT8VGQX2IlWj8AFtX5yVsNO+b2GwcEd/OA76L1dcH+7xcis0Ym+kpJmkPRxJNFP+I5idaRUSCCjPvbgrc1ol1Ji2PlSVdDmvXjcVKQ40FFkw4ji+wtl2hhNV72Lofhbbd6+j6KO3hauAQbBZuwUXY8gwqO5VvBxJNFB+Lld8TEfHhrf7t2sBt3Vgf3gpYH97SWCKuUAJ0UZJfp/lTrO9uCsU+vFexdt4krA+vt+SzRVrnLOBb2O8fsITQVcA6WBJb8uvHWF9u0BHo2pUIJQfTrQf7Un8VG3EaZxDWyBiNNTKCtcgHYlO3C5n2XmwNwuDaNXOxTqMP+zddkCUt5gGHA3dh9fjBGuHnA9/0FJP4txE2SyfoBmztBBER33qxxNwbwJ0lHjOQYifScMJrRXf339aBDYoptU7N58BHWEeRBsxIWvRhbbtHKf6OXRQb/PVV1EGQV6sCv3Fuux9LJouItJN52Fq5r2CzneMMItzOC64fXejD68K+E3uxtRDnYu25WVh/XrAPb3ZzDkUkcXOwQdt/C9y2MpY0PNRLRNIONseWsQi6ElsDVRKg5GD2zcJKqrztOxARD+4BLgQODtx2AFY24ywvETXubmwWR0GzO8ImYyVGgv7T5NdslkWxY5kvcNvHwGF+whERqcscihUlRPLmSaxE24mB23bCSqH92EtEjXsQ68gNmtvE1/uMaNvuiSa+XjONxMr1BWfazAC+g3Wci4ikzSyKsw5F8uZm4HJg/8Bth2B9UBd4iahxd2DLbxU0e2Dme0TbeS80+TWbZSngGmyiU8EHaE1pERGpwTCiNb7nAdv4DEpabgjWoRh8H/Ri66KKiIhIegzEZg+63+l7+gxKWm4AcC/RdXW+6zMoERERacgC2Oxad/3Br/gMSlpuOPAs4fdBD7C9z6BERCSdlsPWogl+qXxCcT1CybZubOSU23l0is+gREREpG6LAu8S/l6fDmzoMyhpmQ5gItG23bk+gxIREZFErIItgxD8jp+CLRUk2TcQm0XqtvOO9RmUiIik21ZYiSY3Qbiez6Ck6QYQnxj8K8X1VkVERCR91sYSgsHv92nAFj6DkqbrAH5PtG33ANaZJCIiIum3HVb1K/hd/wGwms+gpOkGYv11bjvvWqwNKCIiUrcjiH7BfAps4DMoaZpSjYrngfk9xiUiIiLJ2Jfo9/x0YGufQUnTdAGXED3nrwOjPcYlIiIiyTuB6He+Bvln1xBsjUb3nD/Zf5+IiEjDzib6RTMVGO8zKEncEOA2ouf6VWAJj3GJiIhIsuI6jmYAO/gMShI3ALia6Ll+D1jRY1wiIiLSHB3ABUS/+z9GCcKsGUb8WtIvAot4jEtERDLoZ0S/cGYBB/kMShLzJeAxouf4ZWBxj3GJiIhIcxxF9Ht/HrY2iUoQpd9o4juM3sLWFhcREZFs6iB+kP9M4Fse45LkLAc8S/Qcv4CtMy4iIpK4Y4l+8fQBlwODPcYljdkOG0UWN9pIjQoREZHsOhjoJdoGuBEY4TEuaczG2OxA97y+CYzzGJeIiIi0zqnE9+FdiNYcTrMdseWe3PP6NLCQx7hERCQHjie+E+lf2OwzSY8OrKxYD/GNCq1DIyIikn3fI74t8BywvMe4pD7fB+YQPZ+TUJl4ERGRvImrAtYHPACM9RiX1K4TS/iW6pNd0F9oIiKSJ7sAnxH9MvoUKzOqUlTtbxngbuIbiZotICIiki+bA5OJtgmmY5UjuvyFJlVaBLiB+LbdbcBIf6GJiIiIR3tjbTq3ffAJWiooLZYF/kF8O+8qYIi/0EREJI9WAJ4n/ovpfrSWSbvqxBp/X6B1hkRERKSo1PrDfcAjwMr+QpMyOoD9iS8R3wucgZK7IiIiebcG8Crx7by/o+oC7aob66ebSfS8ze2/T0RExIsRwE3ENy6mARNQZ0Q7WZ3SnX5TgC38hSYiIiJtYDBwBfFthVnAiWiNmnZSbhT5VKzah4iIiAhYFYHbKd1uOBgbUC7tYW1syZ+48/UeMN5faCIiIqYDOJz4EgV92OxCdUz49SXgYmxmYNw5uhVY3Ft0IiIi0m6+jZWLj2s3vALshTqPfBoD/BaYTel1hJb1Fp2IiIi0qy5Kz0TrA/4NbO8tOgFYCric+DXBC0sBab1IERFpK0sDdxL/xdUHPAps5iu4nBqJlZKaQfw50RqRIiIiUspY4C+Ubts9B+zuLbp8Gop16E0l/pwU1ohU4lZERETKGQfcS+l23iPAJt6iy6dRWB9eqcTtx2iNSBERaWMdlB9p3gfcDGzgK8CcGIWV/Sp3Hq4FFvYVoIiIiKTGN4DJlG5T3ANs7i26fJgfOBIrA1/qPNyG1gsSERGR6nUChwKfE9+26AWuB9bxFWBOjAZOofR56AOuBBbyFaCIiEgtFgUuoXQZyz7gn1hnU7enGLNoBeB8Spd47QNeBHb2FaCIiIik0ijgXGAOpdsYTwMHAPN5ijGLlgbOovRMwT7gdWBvXwGKiIhI6i0BXEXpMpZ9wP3Arqg6QZJWAS6i9EzBPuBZYDtfAYqIiDRiFeAmSn/J9QFvAkcBC3iKMe06gK2Av2Ojukr9nd/GZnUqGSsiIiL1GgdcTfk2x/vAT7BR0FKf8VhJ13ID7aYAPwQGeopRREREsmVNrBJBuT68V4EJWFUDqV0nluy7g/Lt6TeA/VEyVkREMmAD4B+Ub2DMwkqO7g8M9hNmqqwEnAz8l/J/14+xtWf0NxUREZGkrIqVKC/XBpkH3IW17Yb6CTNVlsDabC9R/u86DVuPZoSfMEVERCTjNgIeoHx7ZCbWh7c7GqhUjZWxPrxXKf93/QhrDw7yEqWIiEgTbQ3cSvnRMYWE1gXY4scdXiJtT4ti6808Sfm/X2FG5tGo40hERESaZzy2Fk25GW59WFnMS7BqB11eIm1PCwGHAY9QuX38PvBTNCNTREREmq8D2AEb6FWpjfIh8HtgQ9SHF7Q41i/3DJX78F4DjgCGe4lURESkhZYFfoONfK70BTkFG5m+PzDSR7AedWILPx+LNcjmUvnv9QT2txrgIV4RERHJp6Wx2WyfULmt8hHWtjsIG/iUN6tQbNuVW8OxsP0b+1upCoSIiIj4sDzWhzeDyu2WyRT78Bb0EaxHXRT78B6i/BqObh+elgASEZHcGQUcD7xC5S/MQnmqh4ETsRFJ87U+5KZbCmsY/AnrPKvm7zIDWzx6g9aHKyIiIvL/5sdGPT9PdW2YHuAx4GfYLMQhrQ+56RYH9gQuBT6gur/LLGxG5uatD1dEREQk1hisisHrVNeemYuVJz0eWJ9slh8dB3wTuIbqBsn1YRMlLgPWbX24IiIi7WkVbMR5tZ0mhYbG88CFWEJtFdJVwmA41hE2ARtZVcux92AjkQ5CC0GLiIhI+1kFW1ul2g6kwkCw54HLsfbROlglhbQYSrFtdzlWIqraYy+MHp+ASoeKiIhIe1sHm004mfz24dVy7PMo9uGpdGjKpelNKyKSNgOAbYB9gB2pPfH1CdbYeBmY1P/fl7HOmbnJhVmTMcCKwApYOYYV+v89jto6vPqw9Qb/BPwZW3tGREREpJ11YTPg9gV2ovYy8VOBF4CXCLftXsHKcvowimJ7bvn+bSVgOWpfU/EZbLT5n7A1o0VERETSYiCwPdaHtz0wrMbnfww8h7XxJmHtvZexwWXzkguzJmMJt/EK/XnjqC0v1ItVyLgaa+tNTjZM8UXJQRGR1hgAbAxshzUyVm9gX3OxDpf3sS/kD7BFkz8I/P8c4AusATIDmN2/zcA6egqJykLd9OFYXfBRWAJwDLBI4P8XBZYAFmgg7k+AO4Hb+zc1JkRERCSturCyUttj7bu1qX9mYA/Ftt0U4D2sPTe5/7YPsXbcNKwdOBMr1zm3/7ZOYET/vkb0/3sY1v5cAOsYGt3/38L/L4K17UbVGTPA58DdwG1Y2+6dBvYlIiIi0i4GAptQ7MNbpYF9zcUShIX+u0K/XaFP78P+x3yOtQmnY316hT68booz9Ap9ePNjbdGFsD670Vi/XbA/b0mK7cN6fIT14d0G3NEfp2SMkoMiIn4shjUytsTWHFzKazTNMQ0rKfUg1ph4DGvoiIiIiGTNGGBbYGts7eTl/IbTFDOxyg8PYcnAR/BXzUJERESkVb6EJQm3xNp5S/gNpym+AB4H7sfaeU9gMwYlw5QcFBFpDyOA9bCa3+tgCcNGRnK3Wg9WLuHJwPYY/kpkiYiIiPg0P1YpYmOsfbc+6Vt/730sEfgw1rZ7HBvFLiIiIpJnY7E+vHX6t42ovdy8T24f3kPA0ygZmDtKDoqItKdObDZhsCZ4YY2/xfyFxXTCa+QEt+ke4xIRERFpZx3YKPNCm66w/ssK2Gh0X7/NZ1Fc3zq4Ps7LWHkrERERESmvC+vDc9dxXhEr8enLNIrtvJcC//8yVrJUck7JQRGR9BmOdS4V6oiPBhbGRi4Vao13UVxHcAgwH1YzfSg2QqjQ2fNp/38Ltc2/oLjezRSK69xMAd7t30REREQkOUOwDqXCWoDBtWNG9/+7m+I6goOAwf3/Pwwb5T21f19T+/9dWJ9wOtG23RSsfVdo22mUuIiIiEhzzE/pPrxCO6+T4jqChT68+fr/fx7WVwfRPrzPKfbbTXb+/73+TaSk/wMARlD88idVlQAAAABJRU5ErkJggg==" width="2399" height="100%" style="display: block; margin: auto;" /> --- ## 2. Initial links formed on exposure-side <img src="data:image/png;base64,iVBORw0KGgoAAAANSUhEUgAAA8sAAAI0CAYAAAAqQrvXAAAABHNCSVQICAgIfAhkiAAAAAlwSFlzAAASdAAAEnQB3mYfeAAAABl0RVh0U29mdHdhcmUAd3d3Lmlua3NjYXBlLm9yZ5vuPBoAACAASURBVHic7N13nFxl9cfxz+6m94QUEggsgZCGtCDFgBSDgHQEFdCACAFFiKKCiiWCJSqWiCiBH4KhSAcNqBg6QZAAoRNKGikkIZDey/7+ODOvuW3K7s7Mc++d7/v1uq9kZnZnzszcmb3nPs9zTh0iIiIiIiK1qTswENg+s/XJ/Nsv8/++QB3QM/PzXYC2QHugE7ANWJm5bWXm8jpgI7AWWAwsAT4A3geWZraFmW1bJZ+ctE6d6wBEREREREQqqAHYFRiS2XbPbEOwxNiVDcDbwFuZf7P/n0kuAReHlCyLiIiIiEia9Af2A0ZmtlHkRoaT4n1gGvA08AIwHRutlipSsiwiIiIiIkm2E3A0MBo4EJtWnTbrgBeBp4B/A/8FtjiNqAYoWRYRERERkSRpD3wSS5CPBoa34r42AXOwdcXvk1tfvCjz71IsKV2e+fk1wGZslHcdUI+teybzbz22lrl95nJ//Oug+2b+vzOwXSviXgk8jCXO/wYWtOK+JA8lyyIiIiIiEnftgc8AZ2IJcudm/v5S4HVya4NnZv6di7sR2u0Ir6Memrnc0Mz7egW4HbgNmFfGGEVERERERCSGRgITsWS3qcRtE/B85vfGACNI1iBhZ+BgYBwwGZhN6c+9CXvu47ARbREREREREUmJfsCPsVHfUpLDzcDjwHeB/bHWTmmzI3A68FesHVUpr8sG4G7gsOqHKyIiIiIiIuWyFzAJWwtcLBFcjI26ngb0cBGsYyOAy4Cp2PrpYq/XDGAs0NFFsCIiIiIiItI8dcDxwCMUT/iWAH8ADnASaXz1Bi7EKmRvo/Br+D7ww8zviIiIiIiISAyNwnoJF0ru1gNTsBHkNE6vLredsBHnmRR+XdcAE4BubsIUERERERGRoH2wdkeFkrl3gYuBro5iTINDgHuwat+FprNfiE5EiIiIiIiIOLMTcCuwlfzJ2+PASVi/YimPXYDfYb2ZC52c+JyrAEVERERERGpRPTZ6uYroRG0bVrV5H1cB1ohuwLeAD8ifNE/BKm+LiIiIiIhIBe0KPEr+5OxprKewVE9nbF1zvpHmNZnbNbovIiIiIiJSZg3A97ECXflaGR3tLDoB62f9R/K3nnoC2M1ZdCIiIiIiIinTG/gP0QnYCuB8NGoZJ7tjiXG+9+sEd6GJiIiIiIikwz7AbKITrweBge5CkwLqgLFET83ehrWZanAWnYiIiIiISIJ9EVhLONn6EEvEJP76A/cSfbLjX0Avd6GJiIiIiIgkz8/Jv+61n8O4pGUuAjYRfj/fQrMDREREREREiqoDfkt0ojwJaOsuNGmlUcAiwu/rXKzKuYiIiIiIiESoA/5AOJlaD5ztLiwpoz5Et/56DxjsMC4REREREZFYqgNuIJxELQNGOoxLyq8tcAfh93oRMNRhXCIiIiIiIrHzE8LJ0xJgT5dBScU0ADcSfs/nYKPPqVTnOgAREREREUmUk4F78OcSi4HRwOtOImqZYUCnPLctBeaX6XGGAx3z3LYEWFCmx6m0OuBq4MLA9U9h7/2mqkckIiIiIiISE3sCq/GPLi4mmQWfXiC6MFkTML1Mj9EL2FDgcX5bpsepljrgesLP4w8ugxIREREREXGpFzbt1pskbcSqJidRoWS5ifJMKb+oyGMkLVkGW8P8GOHnco7LoERERERERFz5G+EE6TynEbVOsWT5qio8RhKTZbATJ+/ify5rgN1cBiUiIiIiIlJtJ5O+qbfFEtkltK5P9B5F7j/JyTJET8l/DNXFEhERERGRGtERmIs/KZoBtHMYUzkEk+XNWILsve74Vtz/bwP3tZB0JcsA5xJ+Tl9wGpGIiIiIiEiVXE54nXIaWkQFk+VNhBPcu1t4322wwmfe+5pA+pLlOuAh/M9pHvmrf4uIiIiIiKRCN+Ajyr+WNw6ikuXg1OmNQO8W3PdJgftZgLVXSluyDFYJPVjx+yKnEZVJvesAREREREQktr4O9PRcXgb81FEs1fAa8KLncjvg9Bbcz9mBy5OBrS2MKe5mAdcErvsOyZ+mLyIiIiIiEqkBeA//iOHlTiMqr6iRZQi3e3q+mffbN3Nf3vsYChxOOkeWAfoB6/A/t885jUhERERERKRCjsWf/KwGejiNqLzyJcu9CE8rbs4a7UsCv/t05vo0J8tgo8ve5zbVbTitp2nYIiIiIiIS5czA5TuAFS4CqbKPgAcC141pxu+fFbj819aFkxjXBi4fAfR3EYiIiIiIiEiltAWW4x8pPNhpROWXb2QZ4LjAbaX2XB4Z+L115Ebj0z6yDDAd//M7z204raORZRERERERCToI/5TrpcB/HcXiwr+B9z2X+wJHl/B7Zwcu30ttjMZn/T1wuZTXLLaULIuIiIiISNCBgcsPAdtcBOLIFuDWwHVnF/mddsAXAtfVyhTsrH8FLh/kJIoyUbIsIiIiIiJB+wcuP+skCrduDFw+DuhT4OdPxN+TeQHwaLmDirmXsannWf2BgY5iaTUlyyIiIiIiEjQ4cHmGkyjcegNbg5tVrOfy2YHLN5He3sr5bAFeDVwX3JcSQ8myiIiIiIgENQYuz3YRRAzcFLgcrHSdtT3w6cB1t5Q9mmQI7iu7OImiDJQsi4iIiIiIV3ugm+fyZqzAVy26FVjvubwvsFfEz50FtPFcnga8VcG44mxB4HKhqeuxpmRZRERERES8Ogcur8XaANWilcCUwHVRPZeDI843VSSaZFgTuBzcnxJDybKIiIiIiHgF+wlvdhJFfNwUuPxF/K/RgcAwz+X1wN0VjinONgUut3MSRRkoWRYREREREa91gcsdnUQRHw/hn1rcFzjGc/nswM/fjY1I16qomQmJpGRZRERERES81uHvqdyJBI8OlsE2wj2Xs9OuOwCfD9x2U6UDirkegcvBadmJoWRZRERERES8tgLvey7XAzs5iiUu/oJ/3Xa25/Ip+JPDecDj1QsrloLVr99zEkUZKFkWEREREZGgOYHLQ51EER9vA//zXM72XD478HN/xT8qX4t2D1ye6yKIclCyLCIiIiIiQa8GLn/cSRTxcmPg8jjgU57LTcDk6oUTS9sBu3kubwHedBRLq7Up/iMiIiIiIpJi2WnWu2MjyEOB5YGf+WS1g4qhO4DfYWu4AQYFbn8SmFXViOLnEKDOc/k1ElzgS8myiIiIiEhtaI+N+g3HEr0Rmf8PAboEfvZXgcsHAz0JJ9G1ZCXwd2z6dZSbqhdKbB0XuPykkyjKRMmyiIiIiEi6DCCXEHuT4l3wj/oVsh3weuZ3wfKGU4Abyhpp8txEdLK8FrinuqHETnvgxMB1U1wEUi5KlkVEREREkqcNsCvwMWAPLBneHRsl7tDM+1oIvIUVsZqZ2V4HzieXLAN8BSXLDwPzgYGB6+8CVlc/nFg5CejtubwcjSyLiIiIiEgFZUeKR3j+3Yfc2tlSbAIWAG9gifDszP9fAVbl+Z2bgMvJjUYfhBX6mt6s6NNlG3As0C9w/RsOYombcYHLN2P7XWIpWRYRERERiYce5EaJs4nxXlg/31Itx58QZ5PimVj/5OaYBTyKv+Lz94GTm3k/afMq4Wrhte5w7GRKVhNwvaNYykbJsoiIiIhIdXXCkuGPef7dA+jfjPtYiCXEr2T+fQ2bSl3uqcC/xp8sn4gV+5pW5seR5KoDrgxc9yC2TyaakmURERERkcrpBuwJjMxsw7HkuF2Jv78SeBcbHX4BS4xfBZaUPdJoDwHPAftnLtcBv8FGEbdVKQaJty8CowLX/dxFIOWmZFlEREREpDx6YiPFIz3bUKyPcTGbgXewZNibGM/BprS6dAnwFLm1y/sDFwO/dxaRxEVv4KrAdfcAzziIpeyULIuIiIiINN8uWJEt7zaghN9rwhLgF7ER4uxU6tk0f01xtTwN3At81nPdz7BR5zedRCRxUAdMAvp6rtsIXOomnPJTsiwiIiIiUthgrAr0vuQS454l/N4WrLDWjMC2sjJhVtTFwBHknncn4H7gQKyomNSe72G9t73GYyd+UqHUpuQiIiIiIrWgO7ameBRWyOoASqtGnZ1G/YJnexFYV5kwnRgD/DVw3UNYK6W4jooXcgDQ1XO5CXikgo/XE5ua7/Ue1t86aY7HTpZ4lxhMBz6BnSQSEREREZEEa4slL+OAydiU6G1Y0lRoW4VVg56U+d2DgfZVjt2V6wi/HsE1q5JuQ4EV+PeBD4FdXQYlIiIiIiItU49Voj4L+BPwPLCJ4onxauAx4JfAacBu1PbszPbYGubg6/Rtl0FJ1ewKzMP/3m/C+iyLiIiIiEgC9MN6Av8MeBhbJ1wsMd4MvISNnn4Fm47dUO3AE6AfNn04+PpNcBmUVNwQYAHh9/2rLoMSEREREZHCBmFraidR+nTqRcAUrCjR8UCPagedYPsCawi/pj9zGZRUzB7AYsLv959cBlVptTyFRERERMSrK5Ys9SyydQC6YcdR2eSqR+Zyd2y6b1es68h6YEORx12JjWiuyvzsemzq7xasyvBGbD1gdvsIWOa5vLo1Tzqh2gB7YWuFRwGHUbwI12qsRVO2+NaTwNyKRVgbPgk8gL9IFsAfgW+iQk9pMQor5tU7cP2NwHkks7hbSZQsi4iISNp1BXYEtgd2KPBvJ1cBttImLGlejE2RnA8szPyb/f8CLAlPql7YAXt22w87aZHPNmx0eRrwP+A54K3M9VJe+2EVsXsFrn8K+By2X0pyjQWuBtoFrr8Om36d6s+UkmURERFJunos2R2EFZ8ZFPh/cDSkUlZhIyxrsJHiUnTHKjJ3q1RQHsuwdabvZLa3M9s72Gh1nAwg17ppFNbXuL7Az6/D+he/gCXIjxC/55Rm+wD/IfxZW4gVRXum6hFJa3UArgHOibjtz8CF2DTsVFOyLCIiIknRGWtZMgKrajwcGAzsQuva9mwGlmDrV5dHbCsCl1dioykrM7+/vBWP7dUeG93ujI3i9MCmG3cHtstsvTz/917uS+sS7g/xJ9CvAi9jVW+rYTg2lfow4BBstL+QJVhF5mnAf7F+xqWeoJDK2BNb/71T4PqNWKXsa6iB5ColhgB/w06CeDUBV2Br/GuCkmURERGJm45YMZkRwDByyXEjzT92WQ7MxqYhL8CSrOC/S8oRdAx0whKVHbBp597/D8z8f7tm3ucKLGn2bq9TfB12MYOA0djI8WGZ+AqZTS45fhp4AyVecbQdcBvw6YjbngbOBWZWNSJpjjbYiPHPCS9LWQ18Gbin2kG5pGRZREREXOqIFWoa6dmGYwdtpdiKrcudndlmef4/G03FDeqBjcYPBnb3/H8wpVeC3oKt/30em177DJZAFyryM4jclOqjCY8+Bu//ZXLJ8ePAByXGJu41AD8EfkQ419iA9av+GZoJEDd7Ajdga9CD3gJOwU5S1RQlyyIiIlIt7bB2My1JjDdj04Rfxw7Ysv++jQ66y6UPlkAPxw6c98r8272E310NTMcS52eB97EZAaOAo4CdC/xuNjl+OLNNo/Uj1+LeV4Brif58vwBcjE2hF7d6AN8DLiH6vboTey/XVDMoERERkbTrBRwH/AKrjLue4n1vm7AR4buAH2PFgUZgRbDEjV2AE7GRwruBdymth3G+bTM2Kj0Bm4rdsXpPRarkEOyESbF9YSp2Ukaqrx1W6XoJ0e/NR5nbRURERKQMBmDJ7UQsGdpK8YPlRVhRoPHA8VihKom/gdhU2/8BayktSV6O9eT9Is1fOy3JMRZrZ5Z93z8ArgSWEr1fbMVGLwe5CLYG1WPf07PJ/1m9k+J9y0VERESkgIFYwZdbKG0UaTFwL3AZNqLYs/ohSws1YNPmL8NGA73JUHDbhrVyKjT6vBU7ofIrrBhUa6qZSzx0AW7H/z4/jxXmA0u+biP/PrEBuB4r7ifl1xkr3vUO+d+D94BjXQUoIiIikmRdsCR3AnYQXGwq7ixgMjbSNALVSkmaYdi60inYmuR87/MW4DmscNMxQNfM73em9P1lbeZxxmJVuyVZBgOv4H9PJ2FTfYMOxIq2FfrumIaNfjZUOO5asD02c2cZ+V/v1djntGv0XYiIiIhIUFvgcKyNyHNYUpTvYGsjti55AjadWtNsk6c3lqBMAuZSOJl5H5uqOYbS3+u+wBeA67B1z4VGnf+HTfPeB51kibvjsCn22fdvPVYQqpgTgNcovJ+9CVyEpgQ3VwPwKaxXcqFZIOuAX2P1JURERESkiD7AWVgitILCU21fAq7CRhM7uwhWWqUNuanV0yi8xnwNNv36sszvlEMjllTdQ+GR6/nAn4Ej0EhjnDRgI5be/WYe8PFm3seXKbx+tglL+B4ATifc/1dy9gF+Ayyk8Ou5EZvyXqy/uYiIiEjNG0FuLepm8h9gLcKSaE2VTa6dga9iiUehwlxbsLY+P8HaP5Xa97ql2mB9mCdgo4n54lqGTe0/vgoxSX7bAQ/hf2/+SctHKBuAz2KzUwoledkpw5OxCu1dWvwM0mMv4HKsnV6x1y5bbK2/k0hFREREEqA9NnVyErCAwutI/w6cD+zqJFJprWxhrvEUXzc8C9snTsN98bXsCZynyD/9fxHwByzJrncTZk3aF5iDf5bJBMr3HuyD7YeltJrbjM2KyM54qIUp+52xk0WTsIJcxV6jJqwf/Tg0Ki8iIiISqSNwEla5utD06jnANdjU6g5OIpXW6otNpb8D/1rS4LYc65t8PvFu29MbOA+b+ZAvcZ4P/BbY21GMtWIMts41+7qvBE6u0GPtAHwXeJXSEsLsNPC/YPv0XqRj9sH22Cj6z4AnKTz7x7utBG7Cli/UwkkEERERkWbpiI1ATMYOnPJNt30eG3mslZGZNMqOxBZbezwL64M9muhKxXHXC0vYppC/cNHr2Guhnt3l0x4rzOZ9nV+iejNO9sIKURWaCRO1rQGeyPzuqViF9zjv9/2xoorfwpa8zKN5z3cT8A/g89j3v4iIiIh4dAXOwIomeUeAvNs6rN/xmagKalJ5p2IWSiDWYSOy47D1ymnSD/ga1oYo6gTBRuA+rOpyWzchpsJArDq597W9BTdTeuuxkdKJ2NTi5iSS3qnb7wAPYsWwzscqRw/HZjFUUkessN2BWEL7Q+BWYDr5T2gW21aTWy6jLgRlprPHIiIiydcWOApLfk8kekRhLVaA5+7Mv2uqFp2Uy2Ds/T0WK7qVLwGcg73HD2KJ5PpqBOfYAGz/PxtLeoKWYknJjdi0XinN4cDt5EbptwA/wHpqx8GuwNHYspHDKU8CvxkrhLUEa5H2AXbiZV3m303Y9+lWYBU26t4Jqw/QLXMf2TX/vbHXri+2j5arINnrwL+Af2Pr+jeV6X5FREREUmMkNsKyhOgRh7XYdNUxqGpsUo0gV5wr38jSFvxFjmrdCKzg1AdEv17PYxXdNU01vzpsf/KuEV8AHOQyqCI6YAnz94D7sQJwLRmpjdu2AXgG+64/A7V7EhEREclrKHAFtvY035S8m7EpuirQlTzZFkoTKTy9eim2pnEM0MNJpPHXAZvq+i+iC4MtBn6Kko+gbtgyDu9r9QRWaCppdsKqu1+FLUeYR+GK8K63VdjJnMnARcABxHuNtYiIiIhzvYCLsXVtUQdYm7Ept2eg1iBJ1BObQnwndrCc70D6ZexEyX5oKV1z7YCNOM4hemT+PuBI9LoOBd7A//pMIl1rvjthVdM/h00pvxWbyjyTlq8bLnXbiJ0E2+y5fD42Ij6gkk9aWqbWvxBERETiqg47gDoXa80SNUr8LHAb1iJoafVCkzLYCVtreTzwaaJHj7Zi7/EUbFrpW1WLLr3qsTXfX8Ne92Bv4LeAP2Frm1dXNzTnTgeux4rHgdU1+Ap2EqeWdAD6YMlr38z/22BJdnvss9qZ3Brl7Hrm7BpmsLZsYMn3Yuz7eSnwYeb6aVjdATKP837Fno2IiIhIivTH1gq+S/TIxFxsPebujuKTlvO2d8o3FXQdlhyPxao9S+XsSv61zSuxqfA7Oouuetpgr4P3+b+F7a9SGdeTe62PcByLiIiISKy1waocTyF6beVq4P+Id3EdCavD3rPfED3917t29nrgOLTO3IWOwJeJXuawAbiB6ArbaTAAO3njfc5/B7q7DKoGXELu9f6641hEREREYmkn4OfYFLyoJOoZbBp2V1cBSrPVYxWpx5N/dkATVqBtIjAaO1ki8XAANu04eNJqG3Yy61B3oZXdIfgrRm/BZj5omWblHUPudb/GcSwiIiIisVGHFRK6j+hR5A+A36EpkElSjyVRVwMLiU6OtwL/xZKRoW7ClGbYFfgj1n4t+F4+B5xKeL1zkozFevN6v3dGO42otjSSe+0fdRuKiIiIiHvdsAPU14hOpKZibYDU+zUZ6sm1eMrXz3UrNsV1HFaNWZKnO/b+RZ0EeR37zDY4i675ugC3438ezwM7uwyqBtVjBdSaUHEvERERqWFDsYRqNeGD7RVYW5ZhzqKT5mgglyDnmzrvTZDVCiY92mOJcbCtUlPmuiQkzYOBV/HHPgn18XXlBXLvQ0/HsYiIiIhUTRvg81jvzqiE6nmsoJBGkePPmyAvJvr93EIuQe7vJkypknrgs8BLhPeDN7F+2XFMmo/H2hllY12PtYUSd24h9358wnEsIiIiIhXXBUuY5hI+kN6IFQ7SusD4q8d6XF+P9UWNSpA3AQ8B5wG93YQpDtVhCWhUBe3Z2JKLOBRua8CKzW0lF9884OMOYxJzObn3RCcuREREJLUGAr/GplUHD5znAt8F+rgKTko2Aus3m69Il3cEua+jGCVe6oCTgBcJ7y8zsUJgrqpL98ZO6Hhj+ifQy1E84ncKufflKsexiIiIiJTdSOBW/FVls9tTwMnEc0qm5IwAriR/m6eNWMugs4AejmKU+KvDeqV716Fmt2epfsupffH39t6GnQhKcgXvtBlG7v150HEsIiIiImVRj02/fIzokcc7gP2dRSelGIiNDk+jeJEuzQiQ5qgDTiB6pPkB4GNViGEMsM7zuCux0W+Jl7bYybjs1H0RERGRxGqLjS5GVcNdhfVGbnQVnBTVC0sipmKjbFFJ8utYH2RVsZbWqgfOAGYRPhFzE7BTBR6zPbbO3vt4M7Ce0RJP2b8nW4HOjmMRERERabYOwNfwT2nMbvOB72C9WCV+umMJ8hSip8pnE+TxwO5uQpSUawdcDCzFv9+tx9aplmv98EDgf4HHuAXoVKb7l8q4h9z7tY/jWERERERK1hVLhKP66c7AWsS0dRad5NMOm3J6L5aQRCXIc4CfU50psSJg3ydXAGvw74sfAl+ndZWzDweWeO5zMzZDQuLvp+TetzMcxyIiIiJSVDfsQDOqZdA0bL2yq+q2kl+2krU3afBuy4BJWM9kvX/iSm9sP91AuHL2Mc28rzrsu2qL534WAAeVK1ipuDPJvXdXOo5FREREJK/+WPun1YQTrYeofjVbKW4HrAjXS0QnyCuAydgJDs0CkDgZDNxHeJ+9B9ilhN/vhs2e8P7uE8D2lQhWKmZfcu/f3Y5jEREREQnpg430eKvHZlutTAEOcBeaROgInIa9N94RNW9F8qnYWmWt15S4Oxx4hXC7sonY1O0ow4A38X9XTUQnhJKoE1bcK1s/QURERCQWstMh1xKuVnsnMNxdaBJQj02fnkT0yL+3knU/RzGKtFQbYCzwAf59emHmem9v5DPwr3tejZ08kuTKFo/chE54iIiIiGPbAb8gnHRtwpKxQe5Ck4BhWKXqqErk2WRiIqoiK+nQG7iW8IyJJ7FidBMC17+FrdWXZPsnufd0qONYREREpEZ1xUYeVxA9krybu9DEoy/wDeBFohPkNcDNwJFAg6MYRSppGFYnIbgsxHv576hlXVr8htz7erLjWERERKTGdAd+TDhJ3gL8FdjVXWiSUQ+Mxk5abCScIG/FKpGPJf86TpG0+TzhqdlNwP+hiu5pci659/b7jmMRERGRGtERuJRwC6itwK3AEHehScZAbLR/LtGjyG9i07A1NV5q0VhseUjUZ+MWtD4/LT5B7n292XEsIiIiknL1WMGb4DrXbHXrvd2FJlj11zHA44SnljYBy4FrgP0dxSfiWhfgdvyfizcIt0hbjrVOq4++G0mIHuTe0+cdxyIiIiIpVYet93qDcJJ8L1YgR9wZiRXjWkY4Qd5Gbpp1Z1cBisTAYOBV/J+PSUA7rGr2JfirYTdhn51hLoKVslmMvZdr0ckPERERKbODsIqxwSTsaeAQh3HVuh5YApyvWNdCrMKv1o2LwPHYaHH287EeOCfi5wYAd+H/LK3HljSo6F0yPUbuvdzZcSwiIiKSEnvhb7uR3V4CjnEYVy3LFuuaDKwj/N5sxKbDn4aNlInUugZsbf5Wcp+TecDHi/zeKcAC/J+vZ1D7oST6E7n38GjHsYiIiEjC7YwVQvEeXDYBs4EvomlsLuwMXAHMJ3oU+RVsfWVvVwGKxFBv4D/4PysPAj1L/P3uwA2ER5m/jUaZk+Qicu/fNx3HIiIiIgnVGRuBCY5YLsOmIHZwFllt8rZ82kw4QV6JjTCPdhWgSIzti78Q4TZsWUJLTvYdBbxHeJRZa5mT4VPk3rfrHMciIiIiCVOPVVB+H//B4Brs4LK7u9BqUn/s5ESw4ri3oquKdYnkNwb/Sb+VwEmtvM9uWDEwb5V5rWVOhgH4C7aJiIiIlGQ08DL+ZGwLdlCoPqPVU4+tpbuP6FHkpcAvgd1cBSiSAB2A6/F/dmZQ3l7iRxNeDvFfVEgv7rLF3T50HYiIiIjE3+7Y9N5gUvYIVthLqqMfNjI1i8KjyB1dBSiSEAOB/+H//NyM9R4vt6hR5lXYZ1Xi6Vly71Vfx7GIiIhITPXC+vFuwn9Q+SZwnMO4as1IbL1x8H1oAlZgB+LqXS1SmsOBJeQ+QxuwgneVdjThitl3UHoBMamev5B7jw51HIuIiIjETBvgYmwKWrB410VAW3eh1YxsX+TXKDyKuA/9+wAAIABJREFUXImRMJE0qsNmZmwh9zlagPWGr5ZewN34P8vvAYdVMQYp7jvk3p8LHMciIiIiMfJJwuuSN2Gjl30cxlUrRgG3YqNdUaPIf0SjyCLN1Q24F//n6Qlge0fxjAFWe2LZhs3iaecoHvE7jtx7M9FxLCIiIhIDOwB/I5yg3QcMdhhXLWiPHTzPoPAochdXAYok2F7Au4QTU9czZAYTXjc9HRjiMigBrABb9j2Z6jgWERERcagdNjXRO8qRXZd8pMO4asEgrN3WB4QT5PXYOuV9nUUnknxnYG3tvIW1TnMakV9b4Gf4p4avAc51GZRQT66d2ALHsYiIiIgjRwCvE+6XPB4b7ZTyqwOOAqYAWwknyW8AFwJdXQUokgJtsBNR3s/WTGCEy6AKOJBwlfub0WwSl14i9150dxyLiIiIVNEuwP34D8y2AbcAAxzGlWbdsKnUwZMTTVjSPBU4HkumRaTlBgBP4/+M3U/8E54ewO2ET57FNcFPO++ypP0dxyIiIiJV0AEbNc5OL8tuLwGHuAsr1YZg6yOD09ybgOWZ2xpdBSeSMocAi8h9xrZgy0ySdBJqLLYMwzvbZ4zTiGrTj8i9B2e7DUVEREQq7VBsHXIwWRuHTVmU8qnHRomnYiP2wST5BdT2SaTcxuLvRf4BMNppRC23N/AO/u+NyUBnl0HVmNPIvfa/dByLiIiIVEgvrO2TN2nbhh149XUYVxr1BX4AzCecIG/A1iAe4Cw6kXTqAtxBuKr0zi6DKoOuhDsUaFp29exB7nX/h+NYREREpMzqsKl7wUrLLwMHOYwrjXbHplOvJZwkL8IKDe3gLDqR9NodeBX/Z24S6epXPBZ/3/XVwOlOI6oN7YDN2Gv+juNYREREpIz2AKYRrnL9bTTlulzqsCmeU4ieav08drLCdS9XkbQ6HltK4m23do7TiCrn48Bs/N8xE9H3eaW9RW7tewfHsYiIiEgrdcQKeG3Ef1A1heRPSYyLzsDXyB1Eebd1wHXYyQoRqYwG7HvO23ptHrCfw5iqoRtwF/7vnKeA7V0GlXLerhF7Oo5FREREWuEzhEce5gMnuwwqRfpjB+jLCCfJizO39XYUm0it6A38B//n70Ggp8ugqqgO+A420uk9UTDSZVAp9gtyr/PnHcciIiIiLdCXcBGYLcDvsAIx0jojsWJo3iq72e1FbD2hpueJVN5IYA7+QoUTsOrzteZI4EP8U9C/5DSidBpD7jUe7zYU0ZoDERFprtOAa4A+nutmABcAzzmJKB3qgWOBiwm3ntkG/BNbL/hwleMStzpgn7We2Fr09lj7rwZsiiyZy9k+5isy163GTmBtAD7Ciu5tqVrU6TAGuBZbagKwCjgLmyZbi6Zi7aXuw04idMBO6h0MfB0rTCWt96bn/8OcRSEiIiLNMhCbeugd5VyNJXcNDuNKum5Y3+m5hEeRV2FVdoe4Ck4qqgO2JvFU4PvA1dj60CeBmVjiG9wnWrMtAV4DHgVuBX4PfAs4DhiMBlGyOgDX43/tZgCDXAYVI52B2/G/Po/hP4EqLdeFXAHHVxzHIiIiIkXUY8WlVuE/OPoXKuDVGrtgI8WrCSc1s4BvkBs5lGRrA+yDzb64GngIm9rrLRYVh20TlqTfD/wKG1kdgq1ZrRUDsRky3tflZmz0XnLqgEsJr2Pe12VQKfIe9ppuQCexREREYmtX4BH8B47LsTWztXQAXU57Y1MXs700vVu29ZMOjpJtANZiaDw2dTWqF3aStpVYW7gJmeeVxBHEw0r4mWPwr8ndgM36kPwOA5aSe83WA2e4DCglHiL3mg52HIuIiIgEtMEOEtfgP2ieAuzgMK6kqsMqhz9K9GjezWhEJsk6Y0nkNdisgHKP9i4EXsJOpkzDEvB/A3dmtpsy/96VuW0q8Gzm51/DKqdH9eVu6bYNKzT3c+CTxP/kzq7AAwVurwMuwz/SvwA4sPKhpcJO2L7m3T/GuwwoBX5P7vU8wXEsIiIi4rE3diDsPTheCJzoMqiEaoeNFL9KOOFYhU3D3slZdNIaQ7H1vlOxEciWJJ1bsdZr/8b2hXHYqNzhwAjKO4LbgLUi2xM4GtsvL8XWxD8GLGrhc2jC1lbfA5yHjarHSTsskbs5z+3dgHvxP58nUB/h5uqCvz9wE3YSp53DmJLsfHKv42WOYxERERFsdOgyYCP+EYLJQC+HcSVRVyzxmU84sXgfG3WplR6taTIAuITwyaRStoVYUvY94BRgD6yydZx0A/bDEvafYlNBm1tkbAt2AuFs4rHm/iosrp9G3LYX8C7+77uJWNVxab467LvNuz88jbUalOb5JP6TDiIiIuLQCGA6/oOc2YRbGElh22MHi8sJJxGvoP7ISdQNaxc0FX8xo0LbeuApLFE7FSsalVR1WPuas4A/YScKSn0d1mFVk4/HTQJ6FLnp5+cFbjsT/1ryVVhbPGm9c/CfdJ2FWiA113bkXr//OY5FRESkZjVgo8neaaRbsdEVVX8t3Z7YCPwmwgnDNCxZUEG0ZNkVK2gVdeIjapuFTWk+DZtZkGbbYc9zEra2t5TXZzF2Iql3lWLcHustnX38T2eub4O9r97YZmInDKV8jsD6e2df44+ATzmNKHmy++9K9PdDRESk6gZha/O8B41zsPWSUpojgf8QTgw2A7ehol1JdATwD4q3ddqCvfcXYol1raoDRgKX4y/yVGi0+Toqm5zWYT3hve/h7lhxwqcD8dwPdK9gLLVsN+BN/N+LFzqNKFmeJPfa7eg4FhERkZpRj/XwXYd/rd61pH9ErBzqsZHiZwknAqux0Ta1+kiWOuALWMXpYsne88A3sWJZEjYMuBJbxlHoddyGrYkeVYEYLo14rCOxegHexO0yNGJXadsBj+N/Pyag170Uk8i9Zkc6jkVERKQmNBJuX7QIOM5hTEnRFqsg/AbhA/8l2BRTFUJLniOBFyic2C3AEkCtuyxdHZYI/xlbD1zo9f0H5RtpHokth/C2ylqNf4nEUjQluJraYOvdve/5zaiQWjHfIPd6Xew4FhERkVSrAy7ADhq9Byw3Aj0cxpUEnbDK1u8RPsh/GziX+FU1luL2Ax6mcBL3IirKVg5dsddxJvlf661Yr+hBrXicLlh160IFyKYDO7fiMaTlvo//JMZDaDZTIUeRe63+7DgWERGR1OqLjdx4DxgXAye5DCoBumFJclT/2ZexUeY2zqKTluoD3IL/oD24Fvlu4GBXAaZYA/BZrFJ4vmR2PdbmqSUnKG4m//vahM0K+QFWCXsUtoa5vsXPRlriS/hH+V/B3gcJG0judXrcbSgiIiLpdDT+dXpN2OhNtSrSJlFf8rd/UmXrZDsTf4Xk4LrW29F682o5GPs85Uts36R565m/VOC+su9v1IjzZqyS+Z1Y72upvE9hFZ6z78EcYIjTiOKpjtwShqWOYxEREUmVLsD1+A8KPwQ+5zKomGvEWmZ5C595k2StcUyuAcB95E+kpmJrXaX6RgMzyJ/gTqL4VN3dgDUUr2AetT0FnI6WUlTbftgMp+z7sBQ4wGlE8TSd3Gukk9wiIiJlsD+2ltZ7QPgIaj2Rzx5Yj+TNhNdQTsEO6iS5ziJ/gakZqMpsHDQAZ5O/Z/M88o8ytwWeo7REOfsza7AkfM9KPBkp2S7417GvRcUmgyaTe320NERERKQV2mDTh71J33rgEjRtOMqBWDIcXOO4AWujVcu9c9OgDdaiJt+62PFAO1fBSaRu2OyOqMQ32+op6KqIn82XJGcLtnWu5JOQZumDnezwvs9fcRpRvHyP3GtznuNYREREEmsXwkVzXgP2dhlUTI3CkuTgAfVq7EBdI/DJNwB4mujEaRow1F1oUoKDsTXLUe/frViFevBXC86XIG/ERuf2rV740kydgQfwT7//jtOI4uNEcq/Lbx3HIiIikkjn4W8JtRX4DVqDF3Qw0UnyB6hHcpqMIlzULjvF8wI0yyIpOmKjxlHVrWdga8yj6gtkk+S3sZHontUOXFqkDfAX/O/lT51GFA+7k3s9/uU4FhERkUTZjnDRovmoEFXQMUSPMr6PTVHXlMz0OIroBGoeWnueVMcSXZl+Q8R1G4G/Agc5iVRaq47wtPqrqe32Xm3I7evzHMciIiKSGAcCs/EfVNyFJdBiRgP/I3xAvRgbceroLjSpgM9ga5GD7/c/0ehi0g0GXiX/lOslwDfR7JC0uIzw1Pu2TiNy6zVy09O7OI5FREQk1hqwKcPefqErsR6jYiMTx+Nvt+EdXRwHdHAWnVTKqcAm/O/3NuAKantUKk26YCcEg5/r5cBeDuOSyrgI/xT8+6jdpUV3knsd1OJOREQkjx2BJ/AfKD6P9RitdfVYkvwi4YPpOViSXKsHWml3EuG2X9uAr7kMSiqiG7CQ8Gf8Q2CYw7ikMs7E/9l+jOI9t9PoJ+Regy86jkVERCSWTgSW4U8GJqLWN/XAaURXzp2FtYlp4yw6qbTh2MyKYIGnc1wGJRXTDZtd83vCn/fZQG93oUmFHI9/ecVz1N5yoy+Qe/4/cxxLzVFFSBGReOuAFTy50HPdUuAs4N9OIoqHtsAY4PvAoMBtr2IHFHdhJxUknbbD1qR7+2FvxXq0/tVJRC3Tk+j+wVm3Yvt0a+1E4dH2a4G5ZXicavkpcHnguoexgn5bqh+OVNCR2DTsbDHGV4BPY2vVa8FewEuZ/98PnOwwFhERkdgYirVI8Y6ePIL1kK1V2ZHktwmPLL2CJdANzqKTammLTckM7gPnugyqhRrJX7iqCWunUw5XFHmcQ8r0ONV0HeHn8TunEUmlfBz/7KqZ1M7fwg7k6pTMdByLiIhILJyL9YXNHhhsAr5L7RYryibJbxE+OJ6RuU2zpWrHrwnvB1c7jajlGimcxK6h9RVw67FR47Qly+2AJwk/l8+7DEoqZm9sNDn7Pr9J7STMs7DnvBnV3xARkRrWBZt26T3wmwuMchiTSw1YkZeoJPlZ4Gh3oYkjn8BfDb4JS5iSun6/kcJJbBO27KI1RpfwGElMlsGm42cTiey2DNjeZVBSMUPxF3l7Gyt+mXZTyD3n4Xl+RieMRUQk1YYS7iV6D7XZIzZb3folwgf1L6OR5FrVjvCJk7kku+BPI+F9fFvg8qOtfIxbitx/kpNlsBHHYI/tO5xGJJU0BH/CPAf7HKXZr8g931Mjbj8FtUUUEZEUOxv/tOv1wHkuA3Ik2yc5uFa7CVuTrCS5tn2HcNL3aacRtV4j4X19Gv7R822EC9mVqhv+75bs/acpWQa4lPBzOtRpRFJJwYR5FlbELq2+TO65/jBw20DgtapHJCIiUgUdgEn4D/DexkZKakk2SX6B8AHvqyhJFpthsRz/vnGt04jKo5HwPn87VtnZe92PW3j/Ywl/nv4Q8ZhJT5YbgOn4n9N09L2RZoOB+eTe73m0/KRSXDQAx0ZcfyC553mb5/o2wDOZTUREJFWGYFOKvQd39wI9XAblwGjgecIH769h1a1rtaiZ+I3Hv398RDr66jYSnSx/MXDdHFqW+P03cD/fJJ3JMsBBhKeYH+M0Iqm0RqzHtjdh3rXQLyTAFcDN2KyQrG7knuMMz/VXomUHIiKSQmcCq8n98duAv5dyLTiO/EnyaShJlpzOWHLs3U8K9SZOkkaik+WOhEfSmzuteHf8yeNmrPBVWpNlsDoP3uf1pNtwpAoG4a/2nvSEuQ+2dGIOcIDn+uy08/XYCPShWG/5JmxNs4iISOK1BybiP5ibh02xqhUHA08QPlh/A/VJlmje9XpNwIdAV6cRlU8j0ckyhPsIN7fn8i8Cv39/5vo0J8t7Ex5dHuE0IqmGnYB3yb3n84FdnEbUOtnP6Gbge9jJY+/SjH2BReT29a+7CVNERKR8diNcuKqWpl0fQHgdZhM2hW4sSpIlv6fw7zMT3IZTVo3kT5ZHBa5vTs/leuC9wO+flLktzckyWPVw73P7ndtwpEp2wt9GbBawg9OIWm5nLFHOPpfHsZNl2cvPkBtVbsJqfoiIiCTWcfinVG7GppHWQvGZvfH3iPQeyGgkWYrpj3+kcBtW2CctGsmfLAO8GbjtrBLv95jA7y0j14s67cnyGfif23vUxnetWGVo7xrmt7HvkCSaTC4h3kq4PZp329NRjCIiIq1ShyXF3jPA7wGfcBlUlQzB/th7W+A0AUuw16S9u9AkQc7Dv/885zacsmukcLL8/cBtj5V4v3cEfs87upr2ZLkj4XZZtdZhoJbtir+t1JtAP6cRtcwehE8URv2/idqZoSYiIinSG/gP/j9o/wR6uQyqCnbC2mF5p5BlR7Yuww5kRUp1O/796Ptuwym7RgonyzsQ7rlcrHhRL8KjUHt5bk97sgy2Ptv7/C5xG45U2VDsxGz2/X+RZCaUUwifcA5ua5xFJyIi0kL74p8Ktg1bZ5nmCs99sOcYPEhfnbm+u7vQJMHm4N+fDnIbTtk1UjhZBngocPv4Ivd5YeDnXwzcXgvJ8jj8z0+tdWrPx7CTtNl94FmSVxgwWLcganvdWXQiIiItMAZYR+4P2UrgZKcRVVYv7OB9Ff4/4Guxyt99nUUmSdcD/z61ifTNTGikeLJ8euD2ORQ+8TY98PMXB26vhWT5QPzP72234Ygje+NvO/c0pRfJi4tp5B9d3oKNPouIiMRee8KtXl4i2f0eC+mCTasO9oLdhE3DHuAuNEmJffDvWzPdhlMRjRRPljsQ7jN9WJ77GxH4uY3YrA+vWkiWuxL+XlIxwdp0IP6TuQ9jn6mkOJb8o8pbgD+6C01ERKQ0OwPP4/8jdjPQyWVQFdIB+Bb+6W1N2BrlG7DXQqQcTsS/j/3LbTgV0UjxZBngz4GfuTHP/V0V+Ll7In6mFpJlgA/wP8cd3YYjDh2Kv+jbA0BbpxE1z0vkH12+1GFcIiIiRR2Ov5BIti1U2tQDp+Ffi92Erce+E6t+LVJOYwifgEqbRkpLloPTitcQXn/ZBng/8HNR/VdrJVkOtt0a7jYccWw0/poat5GcOiJnkn90+fMO4xIREcmrDvgu/rO9C0hnW6gTgNcI/5Gegr/Krkg5XYB/f5vkNpyKaKS0ZBngjcDPnR24/YTA7UuIHj2rlWQ5uHZ7f7fhSAwcj03Jz+4TSZnC3IDVKvC2ocxuaSt6KCIiKdAB6yHs/YP1JNDfZVAVcCDwOOE/zs9g09pEKul8/PvddW7DqYhGSk+WLwv83GOB2+8N3H5VnvuplWT5BfzPcT+34UhMnIE/6RzvNJrSBavcZzfVBxERkVhpBGbg/2P1W2wKZFoMw6ZWB/8ov4FNxa5zF5rUkC/i3/9udRtORTRSerK8Pf7+5d6ey9sBGwL3s2ee+6mVZHkm/uc4zG04EiPB1mJfdxtOSToAS/HHvZnkTCUXEZEacAj+9ckbgC87jai8BmJTXYOFROYDY1E1WamuYBXYh92GUxGNlJ4sAzwY+Nnxmeu/Ebh+eoH7qJVkeQX+59jPbTgSM1fgP/F0ttNoSnM5/n36fbfhiIiI5IzFv9ZpAelZA9cLmIC/+EkT8CE29TNJbTYkPYJtkGa7DaciGmlesvy5wM/OwUaWgrNdLixwH7WQLPfC//zWohkxEvZ7cvvIJuwEXZx1x9rBZWN+s4TfaQP0zGy7AIOAjwEjM/8fBPTN3J7GDh4iIlJh7bGWSN4Dr2nYlMik60R0r+S1WPLcw11oInTGv7ZwK9DNaUTl10jzkuV2hFsiXRK4vBHoXeA+aiFZPhz/83vVbTgSU/XA38jtJ+uI/2fhFnKj4dOwKeQ/Aa4F7geeBt7FKuZHrXEuZdsELAReBP4J3AT8CvgmtjxmFIW/Y0REpEYMwIpZef+ITMIOWJOsLVZpeBHhP5B/Jn2FyiS5Xse/j37KbThl10jzkmWwCr7Bz6338l1Ffr8WkuXv4X9+NzmNRuKsLZYQZveVFcA+TiPK6YG1vPo6cDXwEPAeLU+Cy719iB0j3Yh95j6LjVSLiEgNOAh/MrkRq86bdKOxURbvH7xsr+TBDuMSiRKc1fFLt+GUXSPNT5Y/HvE73u0zRX6/FpLlR/E/vwvchiMx1xn/ifFF2JTlamrAlp6MwU7Kv050q6gkbCuAqVhNheOxAoQiIpIi5+CvLLsYONhpRK33caLbQE1FLVUkvr6Af38tZY1ekjTS/GQZ4OWI38t+VxWrzJ/2ZLkn4dH2XQv+hoitc3+N3D4zk8oneXsBl2Ind9YS/Zlu7bYZ+CizzQZmAa8Az2f+PwsrXPpRBWPYir22vwOOQnVQREQSqwFrA+X9kp8O7OgyqFbaDRs13ob/ec0AjnQYl0gpggVtmrATP2nRSMuS5W9H/F4TpY28pz1ZDvajfc1tOJIgOwDzyO070yhvYtcDa794A7YmuKXJ5xZsTfIT2Cj0FdhU7VOxz/IQWlffoQPWHeMA4DhsAOEHwERsmcdL2Prulsa/Fpv6fhGa0SYikhhdgX/g/0K/jeRWh8xWuA72XlUbKEmaKfj34evchlNWjbQsWe6HFfGZFdiGl/C7aU+WX8L/3H7sNhxJmOHYKGt2//kHrft72Qk4HXiA8IyHUpLil7E1wd8FTgH2wAqP7ofbEdo6YGfspPuF2PfKo8Aqmp88v4oVGx1Y1WcgIiIlG4S/kNBW4DtOI2q5dsA4whWuV2Prhzo6i0ykZU7Cvy+vx4rvpUEjLUuWWyPNyfJRhKd/7uQ0IkmiQ/GfaP59M3+/Hlu6NYnmJY+LsJOD47G1vj1b9zSc8K69noiNzpd6kmBr5ufHoWrbIiKxMQpbr+NNKk9yGlHL1GHTu2bj/+OzCfuD3c9daCKt0gb/1Mgm4DdOIyqfRpQsl0sD8CT+53Wv04gkyT6Hv8DWJSX8Tl/gR4Q7TeTbVmPtni6g+gXFqqkbdlx1LTCX0l6b9cDNWD9oERFx5Bz86yHnA/s6jahlRmP9EIN/bKZga5ZFkqweW4/n3bc3ko61bo0oWS6XXxJ+XqOcRiRJ560NsA3rMRzlY8D/YQlesSTwdaxn8REkvw1lSw3HTj5MxQqQFXvNnsSmoGv5mIhIlTRg63m9X8b/JXmjr8Ow4l3BPyzPko6DX6ldg7C19XcCy7CkJ9hfdIqz6MqnESXL5dAZa1cTfF6bsKq/E7GZN2phI831e/wn6by93o/CEr5iyd4C4NdY9Wvx64OdDPW27sq3zcKmaGs5mYhIBXUB/o7/C/hvJOvLdwfgL4R7ML4FnOwwLpGW6oslM5OAOYQPkr4LnBlx/ZdcBFtGjShZLoeo5xS1bcXa5lyD7U87uwhWEqUeuJvcPrQCK9j1GIX3tZXY3+kjMvchxe2GFeR7h8Kv7XzgK2ikWUSk7HbBqi56D5wucxpR83TC4g0WDFmWub5Wp3RJ8nQHTsBG/LzF9aK2ddhMkDqs+Iv3tuUkezp2I0qWW+szhE8cPo7NSlhK4X2riVxRpcuwokz6HpWgjsDTlHZCZja2LyWxOFdc1GHLy6YQbnvp3WZiJ1nr3IQpIpIuB+M/cFoNnOg0otJli3fNxf+HYi2WRHR3FplIadpghVouw6YtBnsne7ct2NTZCdgBk7c9yhDC/T1nYv1Lk2hHwu2f/lDhx/xxxGMmtXf1EMKV/2djM4iyBmGVeSdhJ2YKHXw3Yf1rp2H73/Ekd9+S8umBtW8qtO9oXW1l7EHx9eDPoEJgIiKtcib+NhDzSM7aoUOA6YSnEv4Vm44tEkf1WPuQ7LrjlRROUGZhycxpFB+RuSji9x9AUx1rTU/gbfz7wWaKj5L3xZLgCVhSHOxFH3Xy5nVs/xyDpm7XmlMoXN363yT3ZFOS9MEKowVPlno/+78iWUvqREScq8PaOHjPBielkNdOwGTCZ7KfBQ5yGJdIPsGiXIUSkPczPzeWlvXB/WvEff6qdeFLgrQD/kN4H7ioBffVFhuVGoftkx9G3G/U1O07M78zEp2oSaN+2N/gfPvAc/iLfUl17ICduMpXRXsWel9ERErSjvAB9V3E/6xjF2A84SlH87BRDa3NkbgoVpTLu63Gpl9fhiUXrd2POxBdPXVCK+9X4q8dcB/h9/7GMj5GcOp2seR5FbZ/j8eWDsT974wUdibwEdHv9VtonWwcDMP6qEe9R9uAP6PPoYhIXj0JV6qcSLzP/rfBRtmWEE4yxuNftyniQmcsEZiArSkutH5vHf7koW0F4tkeq4oafOw/ogPZtGoP/IPwez4tc1ul9Mc/dbvQmvvslFBvy6o+FYxNyqcN4baS2W1T5rZK7mfSfEcRrueS3V7CTnyJiIjHIOBN/ActFziNqLjRWDuT4MHWJGz0TsSFchXlqqT9iO6vq4Q5fToDjxJ+r2dS/e/JzljRyMuwir3BImP5podOxk6KjkD7Z9z0JXr/agJeBPZ1F5oU0Qn727OF8Hv3IXC0u9BEROLlIPwjsx9hPQ7jaihWmCj45T4V+JjDuKQ2VbIoVyWNJHqd6V1AV4dxSfnsDLxA+D1+FZth4FoDuc/OZPKPdHm3xfhbVmnE0p2DiS7itQ74JqpwnRSjsJNnwfdxK3A5OkElIjXuNPxVEmdja1riqA9wLeGzoK8ARzqMS2pPNYtyVdJwog9238KSGEmuwwgvT2kCZhDv6c0DsL9LE7FZF8Fe0MFtLf6WVb2qH3JNOpbotkTzsJkrkixdgDuI/oxdT7yX44mIVMw4/AcizxLPitdtsViDU/aWZa5v4y40qREui3JV2lBgAeHnsQo41WFc0jJ12L4XNbXyBWA7d6G1SFdsicJ47HOVrwVOoanbUl6nYWuRg6/7P9HJiqQbS/R7exs61hKRGtIOuAn/F+Hd2PqVuPkM4elB64FfAN0cxiXpFreiXJXWSPR03W3AH7BRB4m/HYEHid5P7yYd0+uzNQGyLauWUjx5XoR/6nYSP6NxcSbh9kPbgCvQ6GNaHEH05+p29NkRkRrQBfgX/i/AOFa83h07uAl+WU9BVRql/JJQlKvo/4OmAAAgAElEQVTSOgD/R/RznotVT5V4qsNaNkW17dmCnciJ23d8OQVbVhU6uZWd/eGdut2j+iEn0hjC0+K3AOe4DEoqYjDwHuHPzj1oLbqIpFh/bL1a9ktvM/BVpxGF9cAOYILJyhvoYF3KJ6lFuaphLPlPFtwJ9HYXmkQYBDxM9Pu1DPi0u9Cc6Ye/ZdUGCn++t2BJ9iQsIdy5+iHH3icIv45bsNdL0mln4F3Cn5dfF/vFuK89EhGJsge2nmhg5vJqbD3if5xF5FcPfBH7Eva2M/kIm971R+yMtkhLDcJGgkdj08wKrd1cDDyFJSH/xs6w15KDsDWfu0Xc9gFwJZZYbKpmUOLTC5sJcRHQMeL2J7FEZl41g4qpTlgLo1HYNOxPUHxt7ftYov105t8Z2Ih1LdoJeA5/TZPNwOnYSGNS7EbhTh83UZ7vtGHAIQVuvx5LOpNgIPAINtLsdS5wQ/XDERGpjMPwF8daRLx6Hx4OvIT/zOUmbHq4psdJS6W5KFc1dCR/D84mLAkbi6bkVVsnbD+NmnLdhFWHvox0T7suh+DU7WLrnlfhr00QdYIijToRrmewjWQW/zuTwu/xKWV6nLuLPE7SvjMHAPPxP4f1wP4ugxIRKZfP4m/v8AbxmWI2EBu9Cv4hmYoqmErz1VpRrmrZD3iZ/K/ly8Bx6ARDpbUHLgAWkv+9eJDc7CFpnv74p25HVQT2bpux75mJ2Em5OLfjao2bCT/3HzuNqOWKJct/L8Nj9KL4tP+kJctgJ5GDlegXkt79XkRqRLA11DPEY71hZyxJCfZofAs76BYphYpyVU9b4PvY6Fq+1/gNbKS5VkbcqqUv8CNsaUC+134+lghI+XTBpmxfhhWWDLYujNqCLauSfgLpJMLP8V6S+7yKJcubge1b+RgXFXmMpCbLYIMvwZPQdziNSESkhRqAqwn/gXN9EFsHnIVNA/fG9hHwDTS6J4WpKJd722EnHIInurzbCmy0TSOcrbM79jquJf9r/RGWzLn+bq8FDeS+fyZjFeKLJUWL8besal+h2IZidQbKqRfh9kGvkOw2csWS5SbgklY+RlQLvrQky2BtO4PPp1zT10VEqqI9dqbP+0UWh9ZQ+2BT27xxbcUOOvoW+D2pbYPIJcfLKHwA8n7m58ZiBWmkcnYBbiHcRsa7bcRO0p1C4ZH8/tgsAbGTOucBT1B4GcFa7KSFTgK5NQA7GTcRm7lS6POQfd+8LauKFRlrjv9g33/laq0YPOG+AfhYme7blahk+Y3A5Vdacf97Bu5rPXbSNk3Jchus2Jv3+byHrW0XEYm9XlgF1OwX2DZsurNLvbHRveBBxCPYHxYRLxXlSpa9gNsovr5zOdbD+TD8J+5GYAdaN1O7718HbHrjvRRf67gS+C2wg5NIpZiu2DKP8dh3U3CNZ3DbihUX807dbqnB2P6zCfgNrUvEh2BTkr2xXt6K+4uLqGT5h4Tfp5YWQP1t4H5ux06ipClZBuuuEvzO/67TiEREStAIzMR/FvhzDuOpxyqNfoD/C3UB6ssoOSrKlQ47Yu/hhxSfgjgf+DO2BnqF5/pLqx61O32BL2EnGkpdC/sNoJuLYKXFsnUVxmGjvsFpzVHbIvxTt5vzPfcjz/2szNxHS+ozBIt6zaJyU8irKSpZHgf8LXDdH1pw322w2U3e+zmGdCbLAL/H/5yWkewp+iKSciPwl/X/CDjUYTyfJFw9dyM2VU1fprVNRbnSrTPwVeBNiicF3m0J9h6nVQO2pvQKYDrFp+tmtyewaexpOLgWE2xZVegEYRM2g8Y7dbtQO8V25E6aZ+93LnbivNSZG4MIjyqfVuLvxl2+ZPmowHUf0vyTA8FiaAuxz21ak+WehE/0tXa9t4hIRXwS/xfWPGC4o1gGYNPJgn/8p1C+dVSSLCrKVbtGYAf4wdGWQtsK/IlBHKr3t8QALP7x2Pdfvp7IUdtc7PkPrnLM4kY//C2rik3F34Il2ZOwpHvnwP0dgv9vcPbEzPPY8UIxEwKP9wbua56US75kuR5bDuK9/rPNvO/7A7//i8z1aU2WAa4k/Pe7VpfTiEhMnYB/rc3ruKlA2xb7gxNsK/M28BkH8YhbKsolXm2w6Yg3U3xtc3Dbho2U3Qr8ABsh25v4FJPpCRyAVfn/GbbueAHNe45NWNXkicD+1Q1fYqgT/pZVpSxtWIR9j47DZu7cSPik9ZbMvw8Au+V57DaEu1V8uczPz6V8yTLAzwPX/6MZ99uX8HfbsMxtaU6W+xA+uXOE04hERDzOwj9V6n+4GYUZTbia5BpsRCUNa5ykOBXlkmLaYetzvftCKWt1CyXR87B96TpsevPXgVOxkbWhQPdWxtwXGx0/AjgDWzP8CywReYrS1p/m27Zi07GvBD5Beg6epfyys3O8U7eL7V+rseQtaor3FuzYYRLhThSjAz+7inQtnSqULO+O//VqTs/lSwL3+V/PbWlOliG83nuS23BERMxl+L/Up1D9kZbdMo8bPIC9E/VXTTsV5ZLm6Ao8hP97Ynzmtl2BC7HRrmJT9Fu6rcamQS/EpgnOxPbb7PZO5volmZ8rVsG4pdti7ITBl1C7PGmd/vinbjd3xkb2c7gS+Ba5E9sTAz9zY1WeTfUUSpYBng7c9q0S7/elwO+d77kt7cnyMfif20J0ElxEHKoDrsL/xTSZ6iYgnbHRleDUm+exERJJHxXlkpbqD8wgt39sxA5Y88kWP5qIJQGF9rWkbNOwkUEdQEqldME/dXstzdtHFwFfIDxL7KRqPokqKJYsjw3c9loJ9zky8Dvr8RdhS3uy3J7wErw9nEYkIjWrHeHpLhOp7gHY8VjxGW8MH2J/bNL05V/rVJRLymE4NlU6u5+sBo5u5n10xqZVn4/1MP0n8C659Zeutw3Aq8A92JrHs7EerYfiT/QvaubzFmmJXYB/YftcqRXX821bsFkhaVIsWe5G+ETDfkXu8+rAz98auD3tyTLYzCDv8zvXbTgiUos6k/sD2IRNn/pOFR9/V+BB/F+GW7FR7aRWqxU/FeWScjoI/370PpZElks7LBk/EZvG/ROsd/P92FTKd7HaCa1JFpZjLbCexPb3q4EfAudhSf8gCh/0fs1zX5uBw8vwvEWitAW+i41qlutE0ltVfQbVUSxZBrglcPvVBe6vHfBB4OePDPxMLSTLP8D//K51G46I1JpewDP4D7rOqdJjd8DWFq7H/0X4NLBXlWKQylBRLqmUk/Gv+32H/NV3q6ELNuNhAJbgDsH24+y2W+b6vpmf61jGx76W3OvwIXbiUaScRhGePh0cId6c+bc5o81bsQrvnav3VCqulGT5yMDthXounxb42QWEE+FaSJZPwP/8proNR0Rqyc5Y+6XsF9Aa4KgqPfansFGV4B+NbE9CSRYV5ZJquAj/AfmzWHuRWtUWeJzc6/Ey6Uo+xJ1eWCV473f5KqyQ3LvAi8AjWAukv2EnRn+Nfa9/B5sZdDq2vOoI4Ab8fweurNozqZ5SkuV6/MtHmrAq+1GCM+5+GvEztZAsfwz/83vHbTgiUiuGAu/hT1SrUUBrB2x6tfeLbxuacp00Ksol1VSHHYR796v7iU8/ZJf64f8uvxfNzpDWa8RmRPSjPO2drsH/+b2wDPcZN6Uky2AnCrw/MyXiZ/rhb9/ZhB23BdVCstyb8MCKiEhFjcTfw3Me0V/C5dQW+6OxGv+X3ovY+kOJNxXlElfaYUVtvPvXDdgJGzF74y8c9AO34YiE3IT/M/xlp9FURqnJ8i6Eey73D/zMpYH7eSrPY9ZCstwR//Pb4DYcEUm7TwIr8Cc1gyr8mIdiLRK8X3bLUZXruFNRLnGtC+Hig+NdBhRjp5I7AN9G/qmdIi4EZ5Sd5Taciig1WQZLfr0/9+3A7cF14vkqQNdCstwe//Pb5DYcEUmzY/EXxnmV8NnMcuqP/YH0nkHNTrnuV8HHlZZRUS6Jk/7YzBPv6Mt5TiOKv1+Se71WoX6kEh/eYnRNwFfdhlMRzUmWvxL4OW/P5QMCt63D31vZqxaS5V6EB1tERMruC9jZuOyXzbPYF1AltMH+QASn6s7E1qxKPDS3KNc0cuuOVZRLKmkY/p7ra4BjXAaUEPX4e5LORrUgJB68J3KasDZpadOcZLkL4WVpH8/cFjyxMLnAY9ZCsrw7/uc312k0IpJKF+CvIPsw5SnYEWU/YDr+L7Y12NTJdhV6TCmNinJJEhzI/7N33uFyVdX/fu9NJ5BKSAglEErovRNp0ovUoEgRWwREA7agyA9QhKhYUASD0qUYiiIgSFN6Db1DIJQUIJDey/z+WHO+p87Mmbpnznze59nPvTNzyjpz9py9114tXFt0GtZvRTr6AK/gf3/3oPhu4Z5vEx5jLncrTl0oR1mGuGv6xdhYOzPy/l5FjtEOyvJ+hK/vQbfiCCGyxljCFsN/ULimXzUMAC4iXmfxdhTD6gol5RKtxmGEQ0XeBjZwKlFrMoJwbopfuxVHCA4kPN484VaculCusrxXZNtPscRnUStqsXKa7aAsn074+q5yKo0QIjN0EHd7upraWxg6gRMIW4JyWB28/Wt8LlEaJeUSrco3CJdKeZL2rqFcLfthHiLe9/k1t+KINmd1wuPPIuqzcO+ScpXlTuL5QT6LvP5ZiXO2g7I8gfD1fdetOEKILNAFsxAGHy4XUfskTFsDj0fOswBzuc7aINisKCmXaHWSaij/C9VQrgVn4n+nC4Ed3Ioj2pz3CP/O93AqTe0pV1kGODdhH6+tANYrsX/WleVOYDrh69vJqURCiJanG3Aj4YftGTU+Rz9M+Q5aLTyX63VrfC4RRkm5RJboCvyFcL+9AsXY1ooO4Ab873YqsIZTiUQ7cxXh3/pvnUpTeypRlteh8Dj+vxTnzLqyvBPha5uF5jJCiCrojsUkBxXlUg/qcujAXK4/Ivzw+gDV9KwXSsolssrKwL8J9+FxTiXKJr0IT6gfQ54/wg1HEv69TyVbC2OVKMtgSnHSmP7VFPtmXVn+A+Fru8GtOEKIVqY3lvXUe6AsAY6p4fG3wKyUwYfWEszCXK/M2u2IknKJdmAIMJHwQs9opxJlm2GEFzmvciqNaFd6Ey+XdIhTiWpLpcryiQn7zQNWSbFvlpXlXsRjuEc5lUgI0bKsDNyP/zBZDBxRo2P3xuIJoxbNB7BaqKJ6yknKNR0l5RKtzXpYAsDgpPAgpxK1ByMJP8e/7VYc0ab8lfhcIitUqix3wRa7g61PynNmWVmOlhubgbxihBAV0I9wkq35WBbUWnAI8D7hh9UUzBVbVI6Scol2ZUfgY8KTn12cStReBCefS4E93Yoj2pDtiY9zuzqVqHZUqixXQ1aV5e5Y2azgdf3SpUBCiNZkNeB5whaaYsXr07IhYUu153L9a+RyXQlKyiUEfAFbzAuGEGzoVKL2JFgpYQbm2SJEI7mX8Lj3ONlYCJayXDu+T/iaFgJDnUokhGg5hgAv4z9IZgI7V3nMbpgFcyHhh9RDwGZVHrudUFIuIcJ8jXAN5aewxT7ReLoRTij0PLagJ0Sj2I34WHicU4lqg5Tl2jAYm9MGr+kPTiUSQrQcwwjH/H0EbFXlMXcDXiX8cJoOHE82VnzriZJyCZFMUg3le0iXuEbUj8GEQ2xuQc950ViClTs8L4chTiWqHinLteFmwtczE1jVqURCiJZiBOFJzjSqs/p6NZOXB465ArgGPZyKoaRcQhSnK3AZ4d/ClSi0oFnYmrBb/JluxRFtxvrAIsLPhzuwxedWRcpy9XyF+PWc7lQiIURLsTGWYMt7gLyHDTiVMop4zeQ3qU3cc9ZQUi4h0tMbuJPw72Ic+i00G8HJ/XKyVcZHND9jiY+fv3AqUXVIWa6ObQgv4OWAJ2jd6xFCNJjtCFsw36RyK+Vw4G7CD6SFmLuk0vIbSsolRGUMJjyBWwac5FQiUYxf49+rOVhIiRCNoAvwGOHxdAXwRZdCVUF34iWg6j2n6pNwzlZkKGFjkDe3UolSIUQqtgU+xX+AvEplWQG7Yauc8wg/kP4HbFQLQVsYJeUSonqGYwt53m9lPnCwU4lEKToJewG8Awx0KpFoJ9YAphIeY+djVkbRHvQCniS+aPIll0IJIVqHXQknjHqWymKJdwVeIvww+hSLo21X18hg3LGScglRHdsTDuv4lOzUT806fQgneLwHuT6KxrEtZkWMzk+2cymUaAgrYc+bLLnjCyEayEjMLc57eDwDDCjzGErg5aOkXELUh30JP6vewZIRitZhBDAL/x7+yq04os04nvg4PBPY0aVQoq70Bu4nft/vQot1QogU7IYlivIeHo9iq//lcAjwAeGH0FuY63A7oKRcQtSfEwnXUH4ai1sWrcf+WJiJdy+/6lYc0Wb8kvjYPAvYxaVQoi70xea10fv9POXPdYUQbcjuhBXlRyjv4bEutjIXfAAtwWJss5zAS0m5hGgs0Wy296KJTqvzU8KJH7d3K45oM4IJ54KL2Ye7FErUlGFYSGH0Pk9E+RKEECnYj3DszkPAKin37Yol8Aoq2t4xsphRUEm5hHBDF+DPhH9jV6NFpyzQAdyIf1+nUFlCSSEqJamk1Aps/JZ7bmuzB/GSpZWGGQoh2pADsJV87+HxILByyn23IV5v7zOyl8BLSbmEcEtv4HbCv7WLyNZzpt3pRXg8eZRseyWJ5iNJYc4B/8VCrERr0YEZc4IhO5V6Twoh2pSDgEX4D4//YBOWUvTFJqrBOLMcpkwOqoukjUVJuYRoHgYSjjNbBpzsVCJRL4YBH+Pf6yvdiiPakJNI9hh7FwtXE63BEOCfJM/bbsEWYIUQoihHYjHF3sPjLtK5CR8CvE/4wfM2sE99xGwISsolRHOyLvAG/u9vEXCUU4lEvRlJWFk5xa04og3ZDniPZLfsa5DrbrMzCviE5NC4c7A670IIUZRRhBXlOymtKK+BrcYFHzxeAq9Wi8VVUi4hmp+kGsojnUokGsWphMeZPd2KI9qQQSSXGMoBU1Hyr2ZkKIWtyTOwcoNCCFGSLxN2n74V6F5key+BV7CeaQ54GNikrpLWDiXlEqK12IfwM+ddYCOnEolGM57wRHe4W3FEG9IN+C2wnOT5wgRgA2fSCY+VgB8TTzTrtcexEI+q6MAmkPXiMGB+HY8vhEjH0cB1mPIIcDOmPC8tsP3W2IQlWMZjFubG8kfMItusDMeU3b2xbN/FEjm8A9yXb/djScqEEG74CvAXfC+Ol7BEhFOcSSRc0A17Ju+Wf/0CsCuaT4rGswvwV5IrfCzFYuvPxSzOonF0w+qyn01y9vyF2H25EFvwqJpiMXrVtr61EFAIURXHELYo34CvNEfpjVlWkxJ4NWtGSCXlEqL1GUs4LOI+lLG0nRkMfEA4MY/yRAgX9AR+QTiELdjm5T+XzlN/OoEvAW9ReJ53P3XwRpGyLER2OYpw6vwJFFaUDyU8OclhD6RmS+ClpFxCZIcuwKWEf7fXoBwBwjyc5uP3i5+4FUe0OVsBT1F4vjEbc91e15WAGaY3VgnhdQp//zOAb1CneZ6UZSGySVRRvolkRXkwNjkN/naXYCWimiHFvpJyCZFNVgL+Rfg3fBHKWCp8jsPvG8uxqgxCuKIDW6x/k8LzkOVYbfi9HcmYJYZg4X/FvAbnY3O+fvUSoiN/oiALgNNrdPyrsEm3EKKxHAncSDhG2XPH9ujAYgR/Q7gcwqPAt4BX6i9mIl2BLfHjjnejcCKy5cDz+HHHj2AlZoQQzc0ATFHeNf96OZZQ8E/OJBLNyoXA9/P/zwV2xt34JATYQvw3gLOA1Yts9wymC/0dU/hEabph+WaOA46gsNFjKXA58DNgWr2FimroSnAjRGsTraN8M/GHzbrAf4ivzo3F3CIbTTDueDaFVxBzwCTMBXsUqnkoRCuyLmF3ukVYEkIhkujEyhx6/eUN6mhFEqIMemPZmKdRfN6yBLM2fwno5UTS5qYDWwS7mOQ6ycG2GPOGbGg2cinLQmSHIwgryrcQVpQ7MaU0mmb/3zQ24ZWScgnRnmwBfEh4zvE5pxKJVqA/YdfXe3CzsCtEEj0wT73nKB2iOge4Fvgi7b3g3w3YHXOhnkTp720GlkgtKft13ZGyLEQ2KKUobwY8Qfz3ProBsikplxDi84Q9Rz7ElGch0rARVsLQ6z/j3IojRCIjsQX+aFWRpLYMc9Uel98v6/kaBgMnYN/PTEp/PzlskWwMluPCGVKWC9ML2AaLm9wT2IT6JzzqibkW7JI/746YNa2eiYqGAzvkz7dP/u/WWGB91lkLi4ndG1vhGk75yllfbBD/XP44WwFr1FDGNByEuTImKcrdMMVzMeHf+gRgUJ3kUVIuIUSQEwgv5r0ErOlUItGKfAGLb/f60ZfdiiNEQYZhLtovk04pzAEfA//EMr/vRWuXz+uKzYe/BVyB5RlI+z18ilVJGEmTGExcKMvfxybqwXZKhcfaOeFYl2NKZxIDErbfN/B5FywR0n8JZxL22lLgLuBYarcC1A/4HpagKKrQBK1tt2JFuAslO0rLStjE5W7Cq7SFFJnHgd9jAfdpFJkbCH+/X6pQzhMjx7ky5X5XRvb7RuTzLYG/YO69Sdecxi1mW+BXwIsFjpED3gcuwxY+6smBhBXlOzGXIPLnjj6gpgCH1ViGrth3MhazChfqx9GV1L0p/FsVQmSDMYQXzO5H1TJE5ZyF35cWAtu7FUeIkmyFJaoLhqCkacsxZfty4FTMoDWMJlEgAwwAdsJc0X8NPES47FuathCbsx9K9XpOzXGhLA/Lnyc6gd6zzOMMAN4jfg3HFNlnjYTtT85/th7wZMLnhdojwIgyZQ7SBZtElIrZjLa3gMMrPOdBwOQyzxdsz6Q4R7Rw+88rlPV3keOk7ZvRvuVlV+2FuQEXs3LmgIFFjr0B9mMudYxouxXr97XmC4QV039iD5mVMGU06Aa0Arv+VWp0biXlEkIUowv2/A0+C25CC2SiOjqwag9en5qCozhGIVLSB8va3AM/Tvd5Kp+LL8jvPwE4D/gmNh/cGViH2iYR64Jl/N4C2B8zto0F/go8TOmEXMXadOBqzKjW1AuoLpRlMMtWVOGYisU2pqGDeH3GHGbJK0YhZXlj4KOEz0q1mZjLcrn0BP5RwfmC7TzKW106ibD7UiXt1RTnaUZluTdWEinNNa5a4LgHAfNSHiOpTaOyvlKIfbGVOO/4/8YexAcQX0R6E9ijyvMpKZcQIi09sGdA8LmgGsqiVvTCFu+9vvUovkeVEM1EP8xDM4eFyHUNfLYG8HVsETFtDG/aNhfLHP8M9vu4F/OM9bwux+fb3/Kv/5Xf5r/5fV7C9KJyjUPF2jLM0HgmFubabBbygkQvpJExy79POP9/SDeYnp6w74uUXk1JUpZ/RDwT2ydYavKfYQWxL6ew+8QnlGdh7gY8UOBYyzH3hT9jK09XUHz16Q8pz7kTyYryUqzjXgv8FouvODf//+3ErdCtqizfQvzaP8IeYPdi37FnHU1asDmRZLf8HKaYXpeXdxy22lbIPXsOsGnK6yjG3oQV5X9hiRPGR863JC9TJZMIJeUSQiRRzPsGLHPxw/jPhxXAD+stlGg7hmExnl4/u9KtOELEGER4Dl9sDtgl/9kJ2MLiM1Rv4HLdpmG6xDnAIbSwZ2H0whqpLHcHnkqQ4Scl9tuBeEzkPMw6XIokZTn4sJ2LWZqTYnM7MVeBJJeDx0i/Yv7zhP1zmFvRWgX22Yp4JmOvHZninMGJS1CJTGPJXxf4LvaDb0VlOWplnQBsR1yh6w4cBawceT+pv+UwS+6OReTYEluhi+6XZlGnGLsSLv10D5bkJNiPc8CzlGfJVlIuIUQpNscSthRiHeA1/GfFIirPWyFEKUYSHp9PLr65EA1jCGad9frmTMxNuhz6YIm+xmLGs8ex5FeuleBoW4TNbW8GzscMLZnyKnSpLIO5dkZdD5ZSuO5iP+Ad4nJ/JeX5kpRlr83B3AJKsT7JLtvfTrHvdiSnk/9xin27EHdry2EusYVch8EyjkYVnwtTnC+JrVJs02zKcrBfFYtnT6IP8f62AktSl8Z62hV7wEVlObtMOTx2wvqpd5wniYcjzMcerKVqUCoplxCiHDoxd74TCny+OfAB4cnhbo0RTbQx38Hvc0uoPuRIiGpZG8sv5PXLT6ltIrpVMcPJ17Daw1cCd2DztA8oPp+rpH2EJRp7APOk/D2Wc2k/zKCW+fCaQpPialshZTeJIxPk+IBkBTDJnfaqMs5VTFlOY6H1+BxxBfQ9SisowaQUXruijPP2IByn47Vi1viDEravZ1moZlWWK8m4fkbCcc4t8xjdiMdLf0z51uWtCF/b24QtzDngf1gSskIoKZcQolJOxZ4NSdn09yJcXWEK5l0jRCO4jLABYbhbcUQbsy7h0M7p2EJioxmAedxui1+Odj9sTjcKmwuOxqr7jMLcpPfGFpu2xZ7fqxOOsW5barnyEGyHlCnHxQnHuIOw9e7UhG1epbzax4WU5XvLlBcsKD56nIOKbD+UuCI5g9LxX1F2JK6ov0/hDv3VyLbzyzxfuTSjsvwY5cfRdscmfNH+VsmDY6cEmQpZZ5LYgnBSraiiOxN76EWvUUm5hBC1YCj+4twekc+OI2zJeJnCIUVC1INuWL4Xrw8+T3lzQyFqwUaE8xu9R3EDhmgRmkVZ7gFMTDiOlxRkG8K1ZHNY3GS5qzWFlOVDyzwOWCxr9DjXFNn+5ITtf13BecEsiNFjjSyw7bGR7VbQfpblYys4/xEJxzmxguN4PBc5VlqPghGEa0JHF0puxy+boaRcQoh6cBv+syMYrjSGcBKa/9LkJUBEZhlCOAzgFjSuicaxFeHcMe8iD4fM0CzKMlgscNRitgQrkfMW8XN8s4JzJCnLs6m8AHY0i/abRba9OuHcaWKAk/hmwrEKZRvdJWHbq6lfjEGzKcvLqGyF+aLIcZZgWUL4Bk4AACAASURBVF4r5Tek7yseG2Al1ZJ+Y1OxGOy0SbmWEo47VlIuIUQaoqFS62NKyK8j79+M8hkIt2yDec95fTJNPhghqmVbwt57r2P5gkQGSHInnY+57VbLxAr2eRtzAb0x8F43rC5YVLG7EfhLZaLFeA5ThCrhCcIrR+tjcQJJil00C948LHtcJTyS4vgeT+flCcadnoBZ5S8A/okpUlnlNSpzPY/G3b+FuTtXytuR1+tjMe7LC2w/DLgPixkJksPu6QIsXr/QQs9yzBXtvnx7BPPOEEKItPTBwqRW4I/DC4AbgC8GtvsDVtJxRUOlEyLMs1i29mvzr8/DMhLf4UwikXVGAndiz0qwcL29sbJJIgMkKctLsOLYrvg7sCfh0hRRRfktipeuKJdKFVZv3y8HXndgynOSshyNA32JyicWb2J1doNJooYV2HYpVjf5vMj7W2MxqnMx17l7gAexH3qWJjxRJTUtm0VeL8cWcyolWsapA7NUz0jYdi2sREBUUfYmrDsUOMc7+Mrx/TQ+u70QIlv8CqvhHnRnvRF/MdGz3v2ywXIJUYi/YePt97Dx8nrMmPCKS6FEJtkDC4XzSo5OBPYneV4nWpioq2YzTK57Ei7iHWyLSFfeqRBJbthnVXG8aPKsHLaiFGWlhO3+UcV5ASZHjvdOkW27Av9JkCGpzcJ+/KdRWZKWZnPDvrqCc69Cuu+q2rZhwrk3whZC0uz/Yf76TsCPWxZCiFqwE/HQjuDrxZRfjk+IRtAF+Dd+X30DKz0qRK04kPBc7RF867LIGM2oLIPVDktSDn5W5XGTlOXTqjheUtmrUQnbDU3YrhIlLsgLkeOVchHujsXNRpXZYm05lgRq9zLkajZl+ZIKzj2M9N9RNW2LhHNvW2T7eSgplxCi/vTA4u6Wkfwsmo/Vtx2BecAo87BoNvoTznfzH0qX9xQiDV8gnHT4f5iRRWSQZi0iPRSLpU3ieGq/Opir8b5JCkzSe9WcN2n/UorTEuD7WN2135AunqITs5T/D8v03YoTokq+Z5dJaoq5evcGNsUU5ZFYUfoejRBKCNFWnIEpwoWUi5WwOOXXsWSD8zDF+lMsp8IhaDFPuGUm1g9n51/vSzwcTYhyOQbLtO7Nve4CDsBCGkVGaTbLchfgAYpb426p4vhJluWfVnG8ExOOt0/Cdr0Ttru1ivNCvDTQu2Xu34FZNsdgWUyDKe8LtfsonUW5Vpbl30eOU6ll+U8VnHsQ8Wsfj61U17IlTUT/knDuYm0B5v5zIXAUysAohKiOjSjPA8lrn2BVGWRhEc3EofjlzVag0AFROd8kXCrvX8hg0RY0m7J8DukG5W9XePwkZfl3Vcj7w4TjbV9g28WR7R6u4ryd2Ep+8HjPVnE8MOV5a+yaHqBwGaLTSxwnOsmqdCU3qjQ2Uln2slQHj3NbBcephJUI95Vov0nTpmJx5+dgngErNUh2IURr04ElfIw+/5Kat82H2KKrnjOiWfl/+P12IYXnaUIU4hTC8+IbSE6ULDJIMynLexKPj5qBBdEviLy/iHh24TQkKcv3VSFzUu3k1QpsG63JPIvK3dQ2SDhvrZW5EdikKXqeySX2i9bK/m2F5/8HlfXNWijLYBPA4HEmVXicSrg0cN7lwPnEE7p5fWhWwvvRtgzLBHoN5ua9Kc0bhiGEcMdo0ivJ72FKsmori2anA6u24vXh9yg8VxMiyljCz8C/oDlUW9EsyvJqmDUsKMsKLN4EkgfwNynf3StJWf6UyleHXoscq1hG6usSzr1xhec9PuFYP67wWMXoAbyccK71i+zzfmTbKyo892Qq65u1Upb/Rvy6N6rwWOWyGlbyyzvv/zD39xOxrJ5Jv4Vx+fYI6bJpz85vOw77na3agOsSQjQvQ4gvdkYX3XLYuHcCsqqI1mJlwolRH8USnwpRjKiifAlSlNuOZlCWO4G7E2SJWiRvTNjmujLPlaQs54D9KpB7i4Tj3FBk++8kbH9OBecFSygQPdYeFR6rFN8r81zRsl+PV3DODRPO2WhlOSkj+y8qPFYlRL0WDsy/34kptxMT5HsJy8beDbMej8asya9Q2K0+2Cbltx+DJRDTREKI1qI3sB6wCxaCcSj2TDgWex58C5v8jcUqQYwGvpHf5imKK8kvYUqyMgqLVmUY4fwslS7mi+zTgSXDDT4LVU++TWkGZfnHCXI8RXyi3gd4O2Hbr5dxrkLKciU1j/+ccJwji2w/jLib+QeUH+e1acJxplM/xeaLxK/z80W2vzay7UKgb5nnvDjhnI1Wlgdi2Q2Dx5pD41y3RhBWcCcRTiTRCRxBck3yx4nfoyGYkn0OFs8c/Z6S2jzM+nwRNklep5YXKIQoi66YV89B2CLmJVj4zWOYV9N8Sv+mK2lLME+f+zGPmwuArwI7Y89JIVqJzxHOrXKSW3FEE9KBZfsPPgfHOZVIOMW1sjySsLtpDovBHF5g+20J1zbLYROETVOer5CynMNiptOyeYLc0yidKfqfCec9v4zzdpKcLbza+tPF+EnC+Yq5I5+WsP0pZZxvR5IzsTZaWQZ7OEbluI/GuR/eETn39xO26cCU4Gjd7RxWk3mrAsfugv1uTsCU4WdIl9QnmjysV1VXKIRIYihwOKaY/gNzfa4k2V8j2gzMpfUKzCtlJ5QhVjQ3YwgvBu3hVBrRTHTBnmXBZ1w1VXNEBnCpLA8kHt+aw9zBivHdhH1eJp2FtpiyPJ3isbgeQ4gn68phbm2lGEncHXY5cFyKfcEUv+h55wCrF9nn/2FWiErKefTHEmFElaVibnjrEle6PgHWSnG+zYkn1nKpLA8iuaTWrZinQyVshyUaWTvFtruT/l53Yr+dt4j3rwmkswqvgvXRsZhCnKac2FLMzXs8pnhviuqrClEOvTBr1/eBmzCPI9cKcLVtMfAEZp05FnMNF6KZCFbcmIHNXUR704Wwd+QKbGFFtDEdWGcIshA4t0bHfwRbbS507tvwE3h5/Bk4ucRxOzBl5bDI+5djsVfFWANTxoLMwVd8PsKsoIVqIO+JrTitE3n/RUwJWlri/GBWvO9G3luef/9szPU1yprYd3NQwmdfA64scr7LsNpws7B6yhOwLNfLSsi5Pfadbh55fxylk4ndi1kdg7yLue49mLD9Slg5sJ/i34tphBXDmcCAEucFU5b7B15fQuWlxsDu+T3ErcnvYl4B12ITw2IMw1z0j8JcF8G8J95Ncf6nsb7lcQXFQw+6Y7+Dswm7jC/EaldfgLmXp2UosCumRG+bl6WU1Wh2Xu5HsdjqR3Fflk6IZmJD4ABgf2xRrFYeGouwRa6p2FgyH7OcLcF3056V37Y7piAcmn/9IaaoL8aeHYOobdjJu1h+krsxD6mksU6IRtENCy34XP7189g4N9+ZRMIl3bHcSIfnXy/Hcjoorr3NSVKWa8k5FFa8v4cFzgd5AXPfWpTi2P2B5zAlJMixwPVF9ktSls/CFOSgYvYGZll7H1tZWgOb2CS5tM7DJjtp6xz3whYStkn4bD6WvOtVbEKzGqac7EmyNfcG4Mslzucpy1GZn8+3qZgiuiIv2wbYgLFlwrHexr6DUoPJVljceZJb+suYxeFTTDHeAFPGgpPFW7FJW3BFz5WyDJYU508k34N5WIzwC/lzzwP65dsIrMTZ0IT90irLhxNevFmBKdxPldivN3AqcCZhr4JpwBn4q6fl0g1Lbucpz9sCm6TY7x185fkR7Pe7ooLzC9GKdAf2wRL17U/hUKNSTAdexzxI3sSeIR/n2zRs8Tct3bBn14vYIuibCdt0xVeahwKDsWf2hoFWSemoJcDDmOJ8G3Y9QjSaIdjC7pr517dgHlr1nBuL5mMlLNxl3/zr5Zgh6hpnEommop5uWGcXOOcOxGOv5mKKRTnsTDy2dQ42kBciyQ375LxM0WROadp8TFEul74k1zAup40nXVbSy6o8j9cmUZ4r3Q8qPM//sAfX7yLvu3DDDrJ/wrGraWldvjqweOLgvo+R3tV5KNZXoknhHsKU3lqwOn7ysHtJl2hoLn7ysFGo5qXIHh3YotKl2OJgOc+Hhdji0m+xReDtqDz0oxDrEV9wLpdOzNNqX+B0bAH3Xcp/Hj6BVYzQc0A0mm2ABfh9MU1IncgOvbFcNN79X4wlTxXi/6jVxD+pnZ1wvn6YhSm67fEVyv+jhGM9S2E30ULKMtgDMynbdqH2MuaqXCk9gJ9TfgbT6cQtxcUYS3ggKLetAK6isknMdygvKc1f8a0UzaYsg00KryKe3K2c9h5Wgqp3Gec9IuE4x5Yp+xbYQkTwGEsxZbVfmccqRTB52HjSl67ykoeNxZSMSixWQrhmI+A8ylMa38WsGKdi40qpZJHNzmDgC9j3cD/par97z6Q7MY8pJQ8UjeJ4/D64HDjYrTiiQfTDjA/evV+EPbeE+D86sPjVejEBi5ENcgLxh9ArVB4n3QlciO9C43EtNumOkuSGfQq28g9m0TwFiwktlPF5IlY+4xLMsl0ta2ATpAMpbOlbjq28/ysvazkxp2CTjj3zbUdsYaCUsjYJc0u5GlsYqJT1sIWTgwm7R3ssxmKCL8SsnR4n4tcWBltU+GqK811J+NruwZTwWjIciwveC7P4FLPwz8fc3e/DrK6PU777cQfmthx0jZ+C9dFy4/4OwUpzBROMfYZlVP9jBbKlpQ/Wv7345zRlZ5ZiLqKe+/ZE7HkhRLPRgZVrG4Pllijl+bEI86y4L98m1lU69/TCfvt759u2KfaZjY0/F2JhOULUk99i3hFgc6ydsJA4kU36Y2EgO+Rfz8dyId3nTCLRlLRjxtpSynKQtbBYzIHYd/UBtvpfz0F7dcw9dxDmqv0ZFov2Jn5SllqxBqb0DcKUyxzmxv4Z8BI2UaklXTBlbyiwKmbt/hBThlo50UsfbEFgNayvdMEG2lmYF8X7NTrPkcQXn87HYpLLZSXMK+MMwl4YE7GFmycqEbAChhOOfd6e0vXCp2Nu6Z7y/DC1/20IkZbe2CLwGEqHEs3Asl3fjiU6XFBf0Zqa9bDQlqOA3bCF70Isxb6332G/fSHqQRfst3lA/vUbmMJcaHypd94fUTnDsOfG1AKfD8aMF14C29mYceax+osmRPNTzA1biGamA7NQB/vuYorH6JdiBGZ5Dx5zOZZ5PU0ytVrTG1Oex2AuqWncWJdh1uZrsMyVm1J84i1ELeiDeUSVikVegGVYPZjWd62uF2thoRcvUvr3/jBmwReiHvQnXH7xbgp7jh2CQoWalZsoXK50LcwAFQzx26HAtkK0JVKWRStzFPH+W6jMWTkcQlwxnU7pmueNYCgm3zjMbTVN/P2c/Lbj8vuu2nCpRVbpAZyG1Y4vpdR9hdon5co6WwC/xqzwxb7f/5BcUUKIatkIsyZ7fe38AtudBRzXKKFEakZi4WRDEj5bh3BuounULtGpEJlByrJoZTpJtr7sV4Nj98KyWUcT8dyO/W6aha6Y9Xg0Zk0uJ3nYBMxqPZLStaKFCNKJLR4VSwK5BOtjOxc4hkhPD8y9/WUKf98rsO+7Gu8aIZI4FPOy8vrZMQnbXAI82UihREm6YGGEOeKL5COwMMrgnGDThkonRIsgZVm0OknW5ZcxJbIWrE+8rNmn2MS1WemLJQ06B1Pu05TpWYLFP16EXVvaUl6i/dgGi48v1JdmYNanpHrqojo6sBjS/1B4UWwJ5kWi7NmilpxNOJxiu8jnt+U/27HBconCfBv/ngW9ejbBlGPvs8mUVwpViLZCyrJodZLqLueA79X4HCcQL8N1N9XXZW0EwdJVF2Hfl2clKGV9vh1TuvdGk+92pyfWF5aQ3F/mYUqaXK0bw/aE66FG29tYhQIhakEH8Hf8/vUe4RKaz+bf/1vjRRMJDABm4t8vb/zehnDYzOvEK+gIIQJIWRZZYA/i/XgOtXeXXh0rHxY8z3wsGU+rJdFaGT952ATgI0orz0uJJw9rxyoC7chI4DUKWzLHkxwTJ+rP3iQvGHous9fgJkGhyB4rEw59egS/YsPH+OOEngXu+RNh75MuWLm6YPz5q8gDSIiSSFkWWeFu4n35xjqdaxTxhEaPULgWeasQTR4WjddOarOwkhPn5PctVStatBbdgN9T2OX3BuSy3wx0AF/CrH1J9+l9YBdn0okssQ7h8e8SLOwpGNN8tivhBGAL2csIL5rtjhkRvPcmomSfQqRCyrLIClsQHqy9/nxAsZ2qYDXCLmleHNepZMfa2g2r9+yVrnqF0spzDpiU395LHqYSQa3JIOB+ku/xFOAwd6KJAqyELXYlhVksxbxghKiWz2P9yetbZwT+X4F5KnUvuLeoN/cRVpaXEa6c8RTyNhEiNVKWRZa4hnh/fpP6Zno+iHBGyRxWq7mZMmbXktUxC/I5WDxzMCaqUJuLWaovwqzygxsttCibXTCFOMmtdzyKS252dsFcLJN+j3/DlGohquE0wgsx0X72ZXeitTVJSU+D7UFgFWfSiZYnK9agcujEMucGWQAsdiCLENUyDHgDU45z+L/pMylcG7IW9McUweMD780AvkVt6j43M10w9/NtsXiokfnXpWK4p2FuYI8Aj2Ixl4vqJ6Yog28BfyBuGZqM9fFHGi2QqIiewM+AHxCf3zyHlQP6oNFCiZZiIPBz7Nn+AfbcnoIlf5wKfA04PGG/FdgzXZmxG0svbA40FBubo0wHHsM8T+Zj+SbAFjvmYUnabsLmT0Ik0o7KshBZ4zfEM2EvxGJ43q3zuY8C/kw4bvcmLBnWrDqfu5lYBdgSX3neidKxUcswLwBPeZ6IbxkTjeOHwK8S3v8f8EUsiY9oLQ4GrgX6Rd5/H3OnfbvhEolW4yvAxViCr3LYAXi69uKUxUAsbGo1zDNqNSzEZBC20N0B9MYWB3tgXhedWBz2Esx4tABbAJidP+YcrDrGNCx++yNMEf0Ee0auqP9lJXIWtkCWhOcZ5P3fBV/veTO/rxRlIYRoA6LlErx2W4POPwS4M3LuycBuDTp/szIUc8G+CFOIF1PafXsafumqQ7CJjagfY0l2ux5HspVCtA4bAC+R/BvbxKFconVYB3iYdHkrcpj18poGydYf824ahY0XEzDL9vwy5K1lm4olvhyPPVcPAYZT36oZa+Ir9aXk87aZhi3md62jXEIIIZqQH5EcS3VIg87fgQ1AwYF6OaYoKumJ0Ru/dNU1wDuUHuCX4ZeuGoNNjlqtZFezcgHx73seSuKVJVYGbiF5Yi+FWaShA3v2LiGcQKpQW0Lty0gNBA4EzsWqYHyaQo5mafOwxeLfAEcDa9fwe7mR5MR+SYryLEyJ75V4JCGEEJmnOxa3Ex0k3sOUtEaxGfB8RIangQ0bKEMr4ZWuOgdblQ9m7yzU5hBOHjao0UJngJ8T/15nY4sZIlt0Aa4mfr8/wixfQqRhM5I9FZLaWVWea10s8ey1mLuwa4W31m0q8E9skX9rKgsJ3ZXSFuUV2Jg6jnhIhhCpUcyyENnhYMyFF2wF3HMzOo/qB+9y6IYlGPspvivrXOAULCutKExXYAR+7PO2wMaUflZPIxz7/DRKWliII7E4teB3OgsrufaEE4lEvenAEridGnn/NWBn/LhMIYrRE1vY/BGmiCWFauSwON618JNJlaIXVhN4/3wbUaWc8zCF1IsnDv7vWaXnYV5owfjkZcTjmL2EuH0wi7kX/zw0/3cw4ZwllTAds5jfjS0af1Zi+05sjCukaK/Ityuw+zWtSvlEmyNlWYhscQdW2glsQOzABuwtMMtzI9kDs+gE3a4uB76LDc4iHX2B7fGV510oXS9yKfAivvI8EXPnbne2whYVgt4WM4H9cJ+Upxw2Bc4u8vmZwFs1OM/ngO8U+fxkbPLdCnQAv8XK/wS5DTgCdwmKROuxNxYaM4TC8+hjMDfhQvTB+t3R2FhZrnvwAszq7LU38u1NGr/40xPzHvPaiHzbkPLzbizHnsU3Y9/flIRtTgIuLbBvZ37fHwOTyjy3EEKINmA9rByR54LkuSPd40ievljikaBr1GuY8i4qZzhwAn7ysCWkc33zkoftTfvVnV2VeJz4Qlqz1MueFL/XtSobd3OJ86xVo/M0kuuIX0ehbLpCFKIv5imVIx43uwJ4PGGfLviK9jyK/7aCbSl+7orR2GJZq+SuCIYa3U5yMtJCbTk2vo3Gr3PfH3+OE9wuB9yHLYgKIYQQRRmHP4gElahRDmUaTXiAW4glTRG1YWX85GETMLe2UhORYPIwbwKWVW+jDmwiFf0OvuJSqCoopSx/SPXZvAcQn5RmQVnuhVmuosrNAS6FEi3LKMySm5Roavv8NpthZag+SdgmqS0G7scSUo0kW0mpumIK7cnArdh3l+Y7WYBZmpMS9j2EhVMIIYQQqVgZmywnWRb7FNmv3mxDPFnJzSjxRr3wVvTHYavzCyk9IZmd33Zcft9qY9GahVOJX+tvnEpUHaWU5RzmWl4N30lxjlZUlsFqz0afkVNQqTZRGWtjddmjv4+JmDU1TWmjadjC5Sj8OOF2oAsWXnQOVvoqzXcVnNO4NAIIIYRoYb6MP6AErUO/dikUsApxN8h3gZ1cCtUmdMWsx6OxSdkrpJuYTMIvXTWS1isFtgaWYC54TffT2nWUk5TlqGXrhirPMbHE8VtZWQaL/Q+W2cuRHAcpRBo6scRfaUoZee0dLDP/pg7kbVbWwPIKRL0/ktpHmJI92IWgQgghWpsOwivd3gC+FNjSnVj/xzcJl0lahCXtEI1lCOF4ss8oPUHx6mZehMVNr9NgmcvlGuLW81rW+nRBkrIcdTNfSOVeG5slHOvhhHO2srIMFtsdvJ5lKJ+CqIxu2EJkKVfrmdgzaW+yG/ZSK9bBXNFLlc9aDIzHMnULIYQQqdkCP2Y5aEF8mOYYpDfGMjYHB71rab/EU81EF8zK4SUPe4Z0lhIveVizxdhtRVz+k51KVBuSlOVxwAuR975V4fF/FznODcC/Es7Z6spyD+L16e9yKpFoNToxT65JFH9GPgwcRet55jQDHViG/1uIe4NEFyLOQHMIIYQQZXAB/kASTPZ1okOZgqwEXEV4wHsOy+otmoNVMAV4LKYQf0xp5dnL3joeU7xdJQ+LZnN+Gb/+eCtTSFn+fuS9pIy8pehKPEHcfmRTWQarTx+9ru2L7iGEsS/wLMWfg/MxBU7UhmFYONksCn/vH2Lea60caiOEEKJB9MBKNUUHkxk0l8vSCYTdsmdjNShFczIUS6zila4qlTU5h01u7sVcvg+hdK3oatmQuFX5wDqfs1EUUpZXI15GbOMyj30Y8YlnF7KrLAM8SPi6bnYrjmhyBgHXU/hZNxdbqF4DSxp3qhsxM80qwHeBDyh8H57AQkqEEEKIouyO74YdVB6udSlUAtsQroO7AlMAtDrc/HTDMpqOweLxSrkk5giXrhqT37+WtUN/FTnfczRH+EEtKKQsA9wWef+CMo8d3d+r2ZxlZXk/4hbBoU4lEs3KKAp71yzBvGmGOJOu/eiOxYp/ROF7Mg4zHAghhBAFuYKwkuL9f5BLoRIYCNxNeLD7L8p22Yp4pavOwSzK8ymtQM/FTx42isq9H5JciU+o8FjNSDFl+YjI+1NIv+CUZJneKP9ZlpVlsIWb4LWNdSuOaDLWBv5N8nNrBWZpVviQO/phC4OFxpmXUdUNIYQQReiLTZqjA8hkrC5zM9GBTVSDVvAP0EDX6nilq07ArC9pS1dFk4f1THGu3SPHmEW2kr4UU5a7E7d87Z/yuN+L7Pdo4LOsK8unEXfhFALs9/Mpyc+niZhXjGgOhgI3kXyvlmLPOCGEECKRL5JsXf6VS6GKcBDhUkaLMHddkR36EE4eNoPSyvMSLEu3V7oqqUbphZF9rqnnRTigmLIM8PvIZ2lrLkezaY8OfJZ1ZXko4cWb5cidtt3xFm6D46XXFmJeM91cCSeKcjCF45n/iY09QgghRIzbCU8GvdXWbVwKVYS1gacID3QqL5VdOoFNgK9i1ucXSJ6oRtsU4FasNAuYMh38/CiyRSlleXPiE/v+JY65XWSfBYTrNGddWYb4s+Zot+IIh6yClSpKet48jB+eIJqXvtg4kuTB9DrlJz8UQgjRBqwNzCE+cDxN8ybS6gn8lbC8Ki/VPvTGrM9e8rDJFFaaz8FqPAfjblcAqzZY5npTSlkG+40EPy9VX/riyPZ/i3zeDspyNCncb9yKIxyxFqZMJbnx/oDsJApsF6Jeal6bBezmUC4hhBBNyteJW5dzwOkuhUpBUnmpw51KJFzhJQ8bhyUE8/rFAVj8YHBC9I4jGetJGmW5nBjc7sAnke33jmzTDsrykYSv73634ggHDAPeJt7XPyH+mxCtw9qYUSB6X+cD+ziUSwghRJOS5F42H1jXpVAp2AZ4l7DVUOWlRHdgRyxZ3VGE+/WdDuWqF2mU5YHE618XcjscFdnuA+K/qXZQljcl+wstojAbkhznOhFYx51Yokb0JFwZxGuLgEMdyiWEEKIJGUS8tE4OK43R7AwE/kNY7geovMyQyBanE+4bF7sVpy6kUZbB4rhLbQPxkjg/T9imHZTllYi73crltj3YBJhGvI/fiGr0Zo2xxO/zYuAwl0IJIYRoPg4lbKH1/v+iS6FS0gX4BWG5J6MSHgLOJjwJ+plbcepCWmX50Mg207ASXkGGYEphcLsRCcdqB2UZLBla8BqVTDD7DAbeJ96/ryP+exHZ4CTCYWg5LNeFYpiFEEKEuJb4BGE6pTPnNgtJ5aVGF91DZJ1fEu7PZ7gVpy6kVZa7EreWHRDZJmpleajAOdtFWY7GbstjJdt0Ax4k3rcvw7Lzi+zyDeIK83Sy+VwTQghRIX2B90ieKLQK6xOvD3sNlhVZtB/nE+4LZ7oVpy6kVZbBMjpH3UqDvBr5/GsFjtMuyvIswtfYKguHojIuJ96v/0Trud+fQvw6gp5jtaoeMb7IeebU6ByN5ETidzAVZgAAIABJREFUpaWeQfMHIYQQAfYhPlisAPZyKVSZ9CQ+6XmW5k9YJmrPmYT7wfluxakL5SjLm0W2W4zF/QPsFPlsHlZfNol2UZaDZcdyKF41y3yHeJ++i9ZMGFlMWc4B59bgHL2AmUXO0YrKMlioTvRarncqkRBCiKbjj8QHizcwJbSVGI0pA941zAD2dSqRaDQnEe7HVzmVpj6UoyyDZfMNbntK/v2olejqIsdoB2V5EOHrm+tWHFFHNiEen/4G0M+lUFVQSlmeTPVu5ceWOEerKssdwN+JX88JLoUSQgjRXPQEnqc+q9GNZjvCruXLgHNQ/Fm7sD/hPvygW3HqQrnKctSC9iT2m49aifYscox2UJZ3JHx9L7oVR9SJbpjnUfBezyI5sV2rUEpZzlG9t9i9JY7fqsoyWNnBlwhfz2fAUJdCCSGEaC42xAa7qMvmpi6FqpBBwH2Er+V2WtdqINKzPuH7PpPsLZSUqywPIF5z+YLI68kU/57aQVn+FuHru9WtOKJOfJd4Xz7KqUTVk6QsRxfDrqni+GsTToa1HJgdOX4rK8tgc6D5hK/pb04lEkII0XR8lfiA+wStGcPVFVMggvHYbwKbuxRK1J0OwhnSc8DGTiWqPeUqywA3RbaPZoEt5UXSDsrylYSv78duxRF1YADwKeH7fINTiWpDkrL858jrYjkJSvHTyLHuBd6KvNfqyjLAaYSvaQWws1OJhBBCNB3XEB90f+hUouo4lHCG27m0Ri1pUTl3E+6/p7kVp+ZUoiwfnLBPcEJYKltu1pXlDmAK4esr5pYuWpNotvw5WK3xVidJWf4K8FrkvULZ7ovRQVwxPjbhvSwoy51YNuzgdd3nVCIhhBBNx8rEB9hFWEKUVmUE8ArhaxqPxa6J7BG1DvzXrTg1pxJluSswNWG/tN9P1pXl7YlP/JUJO1v0J+46/COnEtWOQsryjyPvVZLDYbfIMWYDK5FNZRlgD+LfpazLQgghQmwOLCA8WDxOa7pje6xC3BX1YWB1l0KJujCc8i2nrUQlyjLArxP28ybVpci6svwnwtd2s1txRB34AeF7/BGm9GWBQsryGliSy2qehVdEjntZ/v2sKssA9xO+tgluxRFCCNGMnEp88D3dqUTV0wGMAZYSnjDJ3TJ7PEm4757nVpyaUqmyvAnmYhhsT2DeJKXIsrK8MuFQjRytn/BJxIl6TGUpJr2QsgxWOzr4fjlVLnoTT/y5S/6zLCvLnyd8bYuB1ZxKJIQQoim5hfCAsQDYwKlEtWF3YDr+dS0FxjqVSNSaaGbjT6k8uU2zUamyXA1ZVpajFsfpKEQja0Td7JeQLeWnmLL8xcj7k0lfIeDEyL5vYIvOkG1luQN4nfD1fdepRMIZWSunIYSoLSdiq/EevYCraf1nx4NYPeYn8q+9zNnXYyvpovW5HsuK7TEAqzcsRJDewPci712GLaCJ7HBo5PUdwMcuBHHAbVgZKY9hWFxuGk6MvL4KUxyzTg5zPw/yBReCCCGEaH5GEHfDysoKaw9sYhy8theweoui9TmbuOUjCzHqsizXjp8RvqZ5WJ12kS2eI3yfj3crTs0pZlkGuCTyWZqay+sQLr24nPBvPsuWZTAvuqg3QpqQFSGEEG3IYYQHzUXA+k4lqi3HA/MJD/rHOJVI1IJ+wAzKnyQ2O1KWa8P6xBMZnu9UIlEPViGc5Go5MNCpRLWnlLK8Q+Sz+UCfEsc8N7LP3ZHPs64sg7mdB69R+U2EEEIU5JeEB40n8WOXssC2wLuEr/FSoKdLoUTVfIf4JPIIpxJVj5Tl6unEwjGiscqlFAjReuxB+D6/VnTr1qSUsgzwYuTzrxc5XgcwKbL9lyLbtIOyfCXha8xSUjiRklaPOxRCNI4fY+UUPHag9bNjB5kIbImVl/I4CXiMbFnR241LsUli9L1hDmQRzcOZWP3YID8imxP+dmejyOtnnEjhnr9FXkeV6SB7YSX4PGZjsc/tRrSvRPuSEEIIEWIgVmrJW2VdSvYUSa+81GLCK+bRVXXROmyFxZsFLQTP07rJ3LYjXgKq3snLfpdwzsF1Pme9OAxzxQ32hzucSiTqSdQr6hyn0tSHNJblwYTLJuawnCRJXBvZ7tKEbdrBsrw/4Wt80K04QgghWoEtCcd/TSJb7tge2wHvEI937eVSKFExZxKfTE4gm31XFGYLYC7hfvARsIZLoURduYrw/f6aU2nqQxplGWxRKLjNzxK26UM4h0cO2DFhu3ZQljchfI1vuhVHCCFEq/BtwgPIZW7FqRt9gZsJX+tEsmdNbwc6iN/LrFqZRDJDiOclWAx8zqVQou5Ef/ej3IpTF9Iqy0dFtvkA6BLZ5huRbV4vcM52UJbXJHyNU9yKI4QQopW4jfAgcphbceqG55YddOOdTfZKj7QDvTD34eikst7JsYR7hgCvEL/3o10KJRrC7YTveRbr5aZVlrsDn0S2+3xkm0cjn/+owDnbQVlelfA1fupWHCGEEK1EB7Yq7Q0iS8i2xXV74lapCUB/l0KJshlGOO7ea790KZSoK2sTn9jngD+4FEo0jJsI3/ej3YpTF9IqywB/jGwXLKe3AeEykcuAoQWO0w7K8lqEr/FDt+IIIYRoNYYRThgyBXNdziqrEi+hMwnYyaVQomx2wSZ20cnlb1EMc9ZYD5hM/F7fBnR1J5ZoIFcSvvffcCtOXShHWd42st0C/HH7/MhndxY5Zzsoy5uRziVdCCGEKEh0kH6I7E9CTwDm4V/zUiz2NRr7JZqXbYEZJCtRWV7waSd2w2onR+/xTUA3h3KJxhJVAJOSWrU65SjLAC9Etv06Vk72/cj7xeK720FZPpDwNT7gVhwhhBCtyn2EB5Q/uhWnIWwMPEv4uh/HLFmiNdiaePxeDngD2NShXKJ6RhMvF5YDbiD7i3kizDcJ94Eb3YpTF8pVlr8f2fZh4mWSPgV6FDlGOyjLYwhf4+VuxRFCCNGqrAp8RnhQqXfd12agB5YcKlizdTZwrEuhRFlsDEwlPtGcg2WOFa3Fylgugej9zAHXIUW5HdmVcD942604daFcZXkw4cWkFdhib3D/i0ucsx2U5b8RvsYfuBVHCCFEK7Mf4cQgy8lmIpUk9ifu7nkVcudtFdYh7iXgTSAvweqOiuZnN6wOavQ+LsPqbCsevT3pRVwxXN2pRLWnXGUZ4hUtom27Evu3g7I8mfA1jnQqjRBCiJbnMsIDyyJgL6cSNY5BxJN/TSWbZUqySE/gryRPGqcCh7sTTZRgJeIeHkFX0v3ciSaahCcJ94usJfmqRFk+PGEfr72c4pxZV5ajyb0WYgsvQgghRMX0Jm7ZmQNs41KoBtKBuZ8vJGzF+AuyTrYKo4HFJE8gJ2CLIqJ52B94j+T79Rww3J1oook4i3DfuMutODWnEmW5G/Bxwn45LKa5FFlXls8mfH13uBVHCCFEVtgFc3sMDjIfAxu6FKrBrAc8SNw6eYhLoURqdsZKhCRNIj8FfogsDK7ZHLid5Hu0HIu31D0SHpsTd81fy6lEtaUSZRng9wn7LQWGpNg3y8pyJ3EX7NEuBRJCCJEtLiA+AE8i3QCcFTqxwXU+cetkf4dyiXT0xMqBFbIyf4jdXyWMaixrA+OJL8h57W1gT2fSiWYmmpfgF27FqSmVKsvDsOdYsB2Z8pxZVpYPJXxt84F+TiUSQgiRKboBTxMfvF+g/Qac9bHa01Er88EuhRKp2Zrk5F9eew2rRaoa2/VlTeAPFF68WIot0vV0JaBoeqIK5Syyk4SxUmW5GrKsLD9B+NquciqNEEKITLIJsID4AH4f0N2hXC7oCpyBJTzzvocVwJ9RLHMr0BUrGRItjxZs7wCno/tZa7YHrie5ZrLXHsIWNYQoRm/iMbrnOZWodkhZrh1fIHxdK4AtnEokhBAis3yf8IDj/f8PzPrcbmwCPEXcynyCS6FEavpjWZeTFoGCE8bxwAhHMmaBTiy+/14Kf8854BXMqi9EWn5CuA8txErHtTpSlmtDd+JJSv/pVCIhhBCZphN4AH/QCZZ2uZH2dF3tCowlbGXOYd/TRg7lEulZEyszVShuNpf/7F7gRGRtTsuWwK+ADyiuJL8LHI89X4Qoh5WBKYT70720fg1uKcu14Vziz/HNnUokhBAi86xJ2H01qGBcTftOeDcFHiY8MC8Afgr0cCiXSM9GWFmwYKmwpLYA+Dvm3tduIQilWBsLUXiJ4t9hDngV+Bb6fYjqOJF43/qmS4FqgJTl6tmGeLjHJU4lEkII0TacSFhxWBp4fTmtv6pfKR2YC3Y0ju4tYF+HconyGIQtckyjtML3KXADdt8HuxDWMZ3ADlgN0ycIe5sktRXAf7B6yu36nBC1pRP4H3F37O0dylQtUparY1XMYyV4PVOBAS6FEkII0V7chD8IvUF4kvwHh3I1A/2Bi4grDrdjlnnRGnTHlOBoXHqhthx4Bvg5sCvZtToPAY4DrgM+Id13Mxe4DPPAEKLWrIspd1HlaA2XQlXBJsRLQG1Y53N+KXK+r9b5fPWiG+FwMa8d7lIoIYQQ7ccgYDr+QPR3wkm/futOtKZhJPAi4QF7JnAy7euu3qpshNVpjlpfirWlWNKq8ZjSvSmtZ03tBmwLjAGuwa4n+Dsv1rw47xOw2FIh6smxxPvg48jNv924lHg/aPcFfCGEEI7YD3/ivAT4DeEB6mx3ojUNXTFFI2r1eBXVZm5FOjCr8SWkt6oG28fA3ZjnwSnA3licr2sluheWkGsU5oJ+LVaPOhhikaatwBSUU7EFNSEayS+J98kJtGe1hnbkDOL3/z5sHBZCCCGc8GfC7thnEx6ofupOtKZiKGaZiw7k92JKimg9ugA7Az/DXLVLxesWa/OB57CyJn/GsrieChyJeSiMwNz7+5M+63zv/ParY33sAMzK+wNsYetabCI5mfTW4qT2GeZZ8lXMRVsIV3Ri4S7RPnoH0NOhXKL+jCV+39/B4peFEEIIZ/QEXsAfnK7ElIfggDXOmXTNx8HAa8TdVS/DlBrRugzCYnn/BrxH5cpn2jYPU1SnAZPy/39G8RJYtWiLMOuxF5vdjiXjRPPSl/gzNgf8GynMWeV84vd7DsqRIIQQoknYFMuK7Q1SxwK/IDxw/dKZdM1HVyyBSjDm27MujgNWcSeaqCFDgEOwWOfbsczZ9Vag69Gm5uUfi1m5pXCIZmct4E3iffk+TJkW2aAL8Hvi93k29qwSQgghmobv4A9Uc4ENiLtFXYL72MxmojemSAUXGnLAFEyZlsUuW3TBstweBfwEq0v+BOG65a7aMsw6/W/gd8BJwF7AwLp8E0LUn8HEEyzmsCR9mzuUS9SGgVgZuuj9nQns5FAu0eRoEiqEcEUH8A/g0PzrpzEXzdOAXwW2uwzLBr2iodI1N2thlvhjCWfJfhFbcLjbhVCioQzCSsOsjlmkB+X/DgZWy7/2FNc+pFtIWYwtxCwGZgAfYd4Mn0T+nwy8nd9OiKzQiY1BNxAvIbUQG4eubrRQoiZsDdyClQwLMhOr4f5UwyUSQgghUrAqZhX1Vnh/kX/fU4699/+KSiclsQ3J9SEfx9x5hQiyEpbAawgwPP//F/D7zUtoEV20D13xy5xNwBaIgm65SR4VvyG79dCzytexxY7ovfwQ2MKhXEIIIUQqdsdPMLQc+Hz+/dGEswVfj8o5FGJvkt0HH8X/PoUoxPP4fWZ/x7IIUS96Y3GpY7GqAtFwlmBbCFxc4LOXgR0bLLson9Ux77Wke/gwSpAphBCihbiA8GqvV7rhG4QV5glADxcCtgBdsRX0d4lPDO4H9nQnmmhyTsTvK/e6FUWImrEacDgWU/80pWuAz8P6/9nY87IrcDzJSvVyYDywcsOuRqSlA1tsL+QdMB7V0RZCCNFidAUewx/M7sR3Bz2G8CTnASwGUyTTHUu29D7xScJjmHu2XG1FkG7AB/j9ZGu34ghRNh3AxsDXgMtJLgUVbZ9glsfvATtQ2HNpewqXdpuEvDGaiU2BBym8GPJld6IJIYQQ1TGc8ErwtwOfHY0lE/I+ewpLYCQK0wM4lXBMuNdeAL6EsmcLn5/g949rHMsiRCl6A3sAZwJ3kK7M2mSsb4/GFOtyFg37YhbJYC6NYHsElR5yyZrY/SnkPfAIsJEz6YQQQogacTT+4LYI2DLw2Z6Elem3gfUaLWAL0h04AXiD+ATiHSx+r58z6USz0B8r4ZYDlmAZ14VoFoZiXjHjMMVnEaWV40n4ynE0C3Kl7EbyszQYxqAyU41jINYnkhJ45bA5wxiUIFQIIUSGuAZ/oHsFy+DrsR3wceDzqSibZVq6AsdhyWmiE4pZwK+BtZ1JJ5qBP+L3iXGOZRHtSzfMLXoM8HfCIQLF4o0fAH4OHIgt/tSL3lgctJeYMtqWAlcSXuwVtWUNLNdJobjkHOZxsKYrAYUQQoh6sTLwOv6Ad2nk840Ix499htXGFOnoBA4jHCMenOTdiL7PdmVdfAVgNsoNIBrDcCw3xe+w51IhK2GwvYdVSPgOVkLPRaWErbGa9sXkvB84GFk2a8V2wHWY90uh7/xlrCSeEEIIkVm2JxyjfETk86GESyXNBw5opIAZYVvMkp8U5/UactFuR27G7wNjHMsisscqWGyvV9v4I0orxksxL6PxWEhJrVyqa8VeWB6NYtfwBpZDYoAjGVuZHsAo4CFKL6CciHJxCCGEaBN+hD8IzsSsD0H6Y3WEgxOqkxopYIYYBlyIuWMnxXxdjGUZFdlne/x7/y6qbS4qpxtmCfw2tij3OoUTZAXbNOCf2GLd54BejRa8AjqAowh7RSW1xVgG7qOAnk4kbQ06sQRuf8HG/2Lf6SdYRnN9n0IIIdqKTsIubhOJD4a9gbsID5wXIZe3SlkFOIXkuOYc5iI5GssMK7JLcBHqaMeyiNagC7AZ8BUs9j2tO/V84GFssW4UrZ83oRM4FPgvpa99Flbmam8sEWO704F5O/2S5NKHSd5PJxHOayKEEEK0FQOwch/e4HhJwjbdgasJD6I3oFXmatkWc3tcQHySshBzoTwEubxlkSPw7/XTjmURzYmXnfocLAP0PEorNzn8DNVjMHfsHg2Wu5FshT1D0y4a3It9L8NcCOuIAdgiyXjgQ9L1oUfy+2jsEUIIIYCdCMcvH19guzGEXfweB1ZrhIAZZzBWg3cSyROXycAvMKuSyAadwFv49/hzbsURDukANgS+DPwWeBCYQzqlZhpwG/b8+DztmzBudeAsSrtoB9sr2Pd9JNnK6Nwf2B84G/NgKZRRPNqmY15jGmeEEEKIBH6AP2jOAzYpsN3RhFfxJ2HZs0X1dAC7Y1b8Qlakl7CJcbMl4BHlcyr+ff2nY1lEY+iGlTv6CpaZ+r8UL80TbJ9gITHnYdn2Vac7me2B32MLCWkV5xwwBbgVi+Peg9ZIvNgLu95TMW+CN0gXsx4c66/FlGvlThBCCCGK0IFNFLxB9GUKxyntgk3cvG0/xZQ8UTtWAb6OucMlTXJWYDGLp9FeLoVZYiVgBv791KJTtugL7AZ8F7gCeJawB0+x9hlwD1bn9kj0G6+ELpgSeDkwlfIUZ699hFn6/wL8ECuXNILGxvB2x+7/Plgit4uxvjGZ8hRjr80GbgGOxXKSCOGcDtcCCCFESvphSb68rNjXYwNqEhsBdwa2XQx8Lb+PqC1ejdRjKJwx+1ksA+w/MPdC0RqcB5yZ//9SLPmbaD3WxizGWwVatLpAIWZjz91nAn/fqYOM7UwHdn/2z7ddqd6SOh+zXn+ELR5PBT7GPK9m5beZjSm087G6xWAhGF4Cx7751ytjCvggLCxn9cD/tSiF9QKWzPNuzDV7aQ2OKYQQQrQlWxJOOPX1ItsOJF6XUZmy68vmwPnYZLqQ5eBNLNvpbsi1rtlZDT+sYQGwqltxRAn6YfHlJ2OLGw9RuuxOsL0P/Av4OWYxHo6MKi7oCxwO/AbLFD6fyizPzdiWAs8Bf8ZqIg+tzVcmhBBCCI9v4w+8C4Gti2zbC7iJ8GB9B+2baKZRdAA7AuOwOLVCE6dZ2P35Gpo0NStX4N+vsxzLIowe2HPveGzh6S7SldoJKiwvYfGgP8BKFw1s6BWIcugKbIMtglyF3bu0LvMu2zIsb8jNWD/bDblWixZEK4ZCiFbkWuC4/P9vA9thLmVJdAA/wiyenlX5LawW5mt1lFH4bIol/TkcK0mVRA7fHe8BzB1vQUOkE8XYDHgR+x19DKyDLVKJ+tMDCynZCNgYS2y4ObA+6b0yZmPK1Qv59hyW82FRrYUVDaUL9lvcEOsfG+bbBph7dKPqNi/H3LwnY+Ppm/n2BjY2L26QHELUDSnLQohWZGXgKWwCCWadPLrEPgcD1+FbledgCvft9RBQFGQYcBBwALAXhZPRLAGexBTnB/L/a+LlhruwWEqAbwJ/dShLFumLKTyb4CvGG2NZ5dPWkl0CvIopwi9jCvLLmMVZtB/9gSFYbPEQTIEehC3AeJm0vZjk3vjK9Qr8hedZ2CLmPOzZ+0n+uAdiizZnARfm9xFCCCFEk7EZ4Viu76TYZ3PC9YKXYaU4hBt6AvtiZWqKuWvnsHt9H3Auprj1TTieqA9749+H11HcfyV0YpbAfbBEaX8E7gU+pDzX1uWYxe4fWGzx0ZiSrfh/0Qj2JBzSJIQQQogm5lj8gXsJsHOKfQZgSldwAno9Ft8s3DIcs1xeT+k6pMsx69l4rC7shshbqp48i//dH+RYlmZmTUyhGA38Cj8D/CLKT4T0Rn7/C4ATsJq1ivkULumKlQ7LYWEy6o9CCCFEkxNMQPQe6RLVdMUsO8HJ6VOoXmizsQmW0O0W/Jq/xdos4L+Ya+CXMZdWWUFrw/H43/MDjmVxSTdgPcza/k1Mkb0JeJ7KshYvwBYibsDcWo/CvGYaFXMqRLncgN9/v+BYFiGEEEKUYCUsAZE3eP+b9ArScfilcXKYQnZAHWQU1dOBKc/fAK7E3IHTKCNzsBI6f8Ssfbvix+yJ9HQjnHF5e7fi1I0OYA1gJLZA8P+w/vY/bDFuGeUrxMuxBEj3ApcAp2Nxn8PRYo5oPY7D79vjHcsiRN2Ry5oQIguMAJ4GVsm//gXw05T77oJZhrzSRSuA87DYWCUuaW5WxVzvdwV2wMqrpI1l/gBLiPQS5ib7BpYlfUbtxcwMY7FyYGDJ8o4rsm2z0g9zlR6W/7tG4H/v/R4VHnsq1oei7W2UfVpkhwHAR5iH1jTsN5RzKpEQdUTKshAiKxyKxfd1YAP3FzElOA2DsMn/PoH3/gscg00KRGvQgbnIboOVqPL+9i/jGLPwFZygwvMeML2WwrYgfbBFhj6YhXU9mifbcndgNWziPhhb/BqCrwSvBayNZdKvlMWYhfjdSJuE9ZF5VRxbiFbiIeBz+f+3AyY6lEWIuiJlWQiRJX6Ob1GeB+yEWQ3T0AWLGTwL3zXyQ0zpfqyGMorGsy4WB7oplhF9E6w0T7kWxMWYcvhB/u97gdcfYgsrn9VG5Kbl98CY/P8XAj+s47l6Yd4DAzEFeCCmEHuK8BBgdfyyONWyDLMOR5Vhr01FFjQhIOxlcg7miSVEJpGyLITIEp3Av/Cz9b6FuefOKuMYBwHX4lsjl2EK+C9rJKNoDrpiltHNsURgGwIb5FuaJHGFWAx8jLknfpRv0/LvzQBmBtpn+b+t5O6/Dva76orFg6+NX5c1iQ7M9bkf5iIfbf0whdhTigfl26oUrsFdCTnMM8Bb2PAWPD7MN89zYHkNzylEVtkEfyH6GbKbw0AIKctCiMzRB3gCsxyC1YI8lPIUkuGYC/c2gff+jiWImlMDGUVz0x9YH1Oc18+3dTDFcCiW7KqWzMZXnudg8a1z820h5iUR/H8hfgzsovzr6P+VsBK+tb0bvsuyp/B6f4/DLPUAT2JKZy+sbna//F9PGe5ThTxp+QR/UWJ6vk3FFig85XgKVl5OCFEb3sKejTkszGGKW3GEqA9SloUQWWQjbBLvTdTPxVzFyqEHVif1u4H33sNKEsktu33pgrn+Dss3LxZ2DcxF2HMPVt3uylmIWeFn4FvkPw289qz2U/KvpQQL0Xguwh8fvwn81aEsQtQNKctCiKxyGHArfsKvUVi93nI5ESv34ik/SzG37AtpLfdZ0VhWxhToQfgKdP98G5Dw/wDMIpsV5mIW86Q2K9+8159h1uFPMIV4gQN5hRDlsQ9wT/7/27AxV4jMIWVZCJFlLgDOyP8/F0v49WoFx9kIuBHYMvDeA1gd1qnVCChEgF751q/E/55S7blGgx9j77lKl0OOcFz/LPxEVnOxuH0wxXZJ/r3NsWR4YL+pQzAX8cUUj2EWQmSD7tji1irAfCzPgEqkCSGEEC1EJ3AnNvHPAa+Tvg5vlJ6Y29mKwPE+xk8mJkQ70QG8hv9b2N2tOEIIB9yM/ww4wLEsQgghhKiA/lgiEm9Av43qvGoOxVbTveOtwJTocssQCdHqnIz/O/iXY1mEEI3nRPxnwJ/ciiKEEEKIStkccxH1BvWfFt+8JGsBDwaOl8PKZ2xY5XGFaCV6YXHG3qLRxsU3F0JkjEFYubUclhVf4Z1CCCFEi3IEvgv1cqp3n+4ExmAxnJ7CvAAYm/9MiHbgZ/j9f7xjWYQQjedx/GfAFo5lEUIIIUQV/Ap/UJ+J1dGtlh2BSYStzPcAa9bg2EI0O6thi0Q5LLnPELfiCCEazJn4Y99PHMsihBBCiCroBO4inPCrf9E90tEfuI6wwvwZcEwNji1Es/MX/H5/jltRhBANZkv83/9jjmURQgghRJUMJGwJvgfoWqNjH4Ufw+m1CflzCpFVRuDHLX6MX5NcCNEeTMYPcRrsVhQhhBBCVMvGmBu2p9D+tYbHHoxl3A4qzNOxOrRCZJU78Pv7txzLIoRoLJfg//6/4lgWIYQQQtSA/YCl+AP8t2t47A5MYZgbOP4K4FJg5RqeR4hmYU/8vv6ehpFfAAAgAElEQVQGSnInRDtxIP7v/ybHsgghhBCiRpyGP8Avo/oM2VHWIV5i6l1g3xqfR4hm4Cn8fi5PCiHahx74i8Nz86+FEEIIkQH+f3v3HS9HVf5x/HPvTSWdNEhoISGU0EMngGJAo3QIRaogCP6UgKBR+KkgoiggAj+QCIIEkCJFDNICiPQSunRCAgkJhPTe7++PZ/e107bd3dmzM/t9v17zys3u3t1ndmbvnmfOOc/5E7kG/gJgWJWfP7vE1HI0l1nS7Vhy5/eTbkMRkRrzTj8a6TgWERERqZL2wGPkvuQ/BvrG8DrbAC/jT5hnAAfH8FoiLrQDPiF3fu/qNhwRqaFTyX32/+g4FhEREamidbF5ltkv+qeADjG8Tjusl3kx4V7mfjG8nkitnUvuvL7dcSwiUjvrY7U5WrEVJ0RERCRFNsfWRs429G+M8bU2BZ7AnzDPBU6L8TVFaqE7MJ9cHYBN3YYjIjU0idx32paOYxEREZEq2w9/hewfxfhazVgF7oX4k+YJwIYxvq5I3C4ndz7/wXEsIlI7F5D77P/EbSgiIiISh1PIfdmvAQ6K+fUGEF6XeQnW6IhjKLhI3DYAVmLn8kKgp9twRKRGdiL3PfYfx7GIiIhITK4m94W/ENi2Bq85GpiNP2l+A9i9Bq8tUm1/I3ce/9hxLCJSG03AdHLTMLTig4iISAq1Ax7BvzZyLQpw9QfGkyuS0pr5eTxqdEiyDCd3Dk9HoyREGsWfyX32j3Uci4iIiMSkO/AWuS/9SUDXGr32PsA7+HuZPwdOwK7ciyTBk+TO3+PyPGaXmkUjIrVwMLnP/d8cxyIiIiIxGgLMwl98q12NXrs9MBZYhj9pfhJVGZVkOBD/lILghZ6uwN21DkpEYrUOVnejFauM395tOCIiIhKnnfGvi3x9jV9/KPAY/oR5GfArrFEiUq+a8I+Q2Ddw/ynAx7UOSkRi9y9yn/uvuA1FRERE4nYA/iWlznMQw4HkCqd454JqaLbUs9PIna//Ctz3LFZxvmOtgxKRWH2f3Of+UsexiIiISA14G/1rgRMdxNAL+BOWYHiT5ieAbRzEI1JMR2Amuc/NsMztm5ErZKdpBSLpsiG5z/d7jmMRERGRGvkduQR1JbC/ozh2AJ7GnzCvwapm93EUk0g+vyR3nt6Que03ntsOdhSXiMTndXKf8c0cxyIiIiI10IQlpN41mLd3GMto4BP8SfMcYAzQ4iguaVybYxeUgp+JvsBS7PxcDmwAzCDX83RuDWMUkdr4NbnvpbMdxyIiIiI10h6YSK4R8BmwkcN4umEJygr8SfMrwAiHcUljOhZLgl8HziS3Pvh1+JeTyf68GluXVUTSZTdyn/PHHcciIiIiNdQdWwon2xB4G5tP7NIQbGmr1sA2ARjsMC5pPL/CP13h79ic/+xc+xVYkpydPvCkkyhFJE7NwOfk/g70dBuOiIiI1NJA4FP86x/XQ1XfkVjy7k2YVwLjsOGwInFrAu7AP58+OwQ7eDGnFSsAJiLpcxO5z/mRjmMRERGRGtsamEeuMXA7djXdtY7AT4D5hOcznwV0cBeaNIjOwEv4K7evJTpZXgt0cROmiMTocHKf8/GOYxEREREHRmI9t9kGwW/dhuOzLnAJ4R69T9D6zBK/9bC1wLNDrgttrgrliUh8upL7/pmDCk+KiIg0pGPw95rVW3XfocBdhBOUF1ARMInXVsAiiifMR7gKUERi9Si5z/mejmMRERERR87HP6z0JKfRRNsHmER4COydWEItEodvYMOx8w3DbgV+5iw6EYnTmdTnyCsRERGpscvwL4kz2m04kbLrM3+MP1lZg/U+D3IXmqTYWeRPlFcDN7oLTURiNIjcZ/0tx7GIiIiIQ03ADeQaBiuArzuNKL/OWG/eAvyJy1LgUnLr44pUyzXkL/D1rMO4RCRe3hUadEFWRESkgbUAd5NrGCwB9nAaUWHZImBL8ScwizK3d3cXmqRMO2Ai0cOxZzuMS0TidQm5z/oPHcciIiIijnUAHsGfCGzlNKLiBmJrMa/Cn8R8CYwFOrkLTVKkO9bLFFXwq6fDuEQkPnuR+5w/7DgWERERqQPdgJfJNRCmk4zhZ0OBOwj3/k0FvoP1DopUYhB2ASl4ju3kMigRiU0LduG1FVtKqpvbcERERKQe9ME/V+sjbO3ZJNia6OWmpgKnoaRZKjOC8CiGE5xGJCJxuoXcZ/0wx7GIiIhInRgITCHXSHgT6OU0ovLsDjxJOGmegpJmqcwJ+M+py9yGIyIxOprcZ/0vjmMRERGROjIEmEmuofA80MVpROUbCbyAkmapruvJnUu3Oo5FROLTA1iJfda/AJrdhiNSmibXAYiINIhtsR7abK/yROBAbHmpJBkJXAzsErh9Clbx9EaseJNIKZqA24BjgEnAzp77umDTFvoDfYH1gX6ZrW/md3tm/u2CFdbrkPl5FbAYa5jPzzzfIuzcXATMwOZQzsIuZGV/VlVukfg8AXw18/NuwIsOYxEREZE6szf+JZruwAqfJNEBWHIT7Gl+DzgO9TRLaTpgleI/BJZhFdknYsls1JrMcW8rgMnABOziz2nYBaKk1BoQqWfnkPus/cpxLCIiIlKH9sMa5NkGw50kN2EGSyReInp49higs7vQpM60A3YAzgD+CrwLrMFNUtyWbSZwP3AesC9ag1ykXEPIfZ5ecxyLiIiI1Klj8K8zexPJn781Ev9SWdntC+ACbL6aNJbOwDeAS4GngSW4T3irua0B/osVKzoBGy4uIoW9T+4ztLHjWESK0pxlERE3jsLmamZ7lW8CvoutPZtUTdg87POAXQP3zQGuzmxzaxyX1M5QYBSWJO9DZSMLVpCbUzwLm1c8w/Pzamw+ciuWiK/MbEuA9kBXcvOawdZ2bQesiyW2/fDPg16fytZ/zfaWPZzZnkfz90WCLgd+lPn5DOA6h7GIiIhIHTsJ/zDUG0jPRcwR2LzPYG/cIuBKbEktSYedgCuwub5t6Z39GEsurwL+B5uqsDFuPgvrYoWHTgJ+A9wNvAUsp/x9m4dNszgEm5stIlbgK/sZecBxLCIiIlLnTsafMF9FehJmsB7m+7Eec28isQy4FhjsLjSpwIbAWGzecTkJ5AzsIsoF2CiEpKw53gIMw4ZbX4kVtytnvvU8YDw2XSFNn2+RcrXDPg+tWMHLddyGIyIiIvXuVPzJ5BVuw4nF1liysIpwz+IEYA93oUmJOmFTBZ4hfPEj3zYF+BMwGkuw06Q78DXgQmwJnFKT5ynARWh0hTSuO8h9Hg50HIuIiIgkwJn4G9SXuw0nNptgPXPeJbSy2ySs5y7J1cHTqD/Wi/wZxRPBZdjST2OB4S6Cdag3dlFgHKW9VyuBu4DdXQQr4tBx5D4H4xzHIiIiIglxFv7G9KVuw4nVAGwd2+xwPO/2Hra+bSdn0QnAjthoAO9SZ1HbCuAfwBFoSGVWM5YE/x9WkKxY4vw0cDi6UCSNYV1yo4xmoKkJIiIiUqJz8Deif+E2nNh1xdZjnko4gZiFzW3t4yi2RrUl1uNZLMGbhB27vm7CTIwWbK7yeKzAXaH39D2sd1rJg6Td0+TO+x0j7u+FXVQVERER8fkF/gb0+W7DqYl22PrTrxBOIJZgxcC2cBZdY9gIW8LMuwZ4VKGq3wODHMWYdN2A7wHvUDhpfh5bekskrcZS+KLwpcBWNY1IREREEuNX+BvP57kNp6ayy05FFZF6But503DV6ukFXIbNN86XvH2ALevU1VGMadOErUf9CIWLpT0IbOMoRpE4DSN3nr8UuG8gVtdC576IiIjkdTH+hvMlbsOpuWHAjUSvcfsB8EOsp07a7nBgJvmTtWexarXNrgJsANnzPF+P/kpszWfN4Ze0+Qg7x9fiH3J9U+b27V0EJSIiIsnxW/wN58tovPmM/bAhe9MIJxILsWqqWzqLLpn6U3he8rto7mytbY4dk3w9zR8BX3UWnUjbbQusF3H7VeTO71Myt21Dbhm2RquoLyIiIm3gndvVClxHY/b0dcCWHHmJcCKRXa95P5TgFXMyMJfohOzTzP0a5u7ObsC/iT4+a4Br0IgKSZZ1gDex0VE9PLfvT+7cvi9z20Oe23auYYwiIiKSYOfi73G6FSuK1aiGY9WFVxI9RHsstu6t5HQB7iA6CVsB/BIN9a0nB2EXL/L1/Gs0hSTJfth32GxsCk2HzLYQO6cX40+eW9Ea5CIiIlKG08kNT2vFEp/2TiNybz1seamotWyXY8Na93QVXB0ZgvXsRCVerwI7uAtNCuiC9cZ5P/fZbRE2VF4kKW4md/5OAY4G7vHc9gH+ufsj3IQpIiIiSXUc/sbEBNQbCPYenEz00lPZaqsnY8MBG803iR52vRTrgdeQ6/q3J9abHDyGa4Er0UUzSYbeWM9ydkpBK/AJ+Wsn7O0mTBEREUmyo4FV5BoUDwKdnUZUX4ZjRb+WEG58Lcjct7Wz6GrrHKILRr2O9TZLcnQG/kJ0UqG/AZIU3yY8Dz9fsqyCdiIiItImB+JfUulJVPQnaF3gR9jQvqgeuceBI4GOrgKMWbAwXHb7Gza8V5LpBGxUQPC4PoX+BkgyPEThJDm7jXQVoIiIiCTfN4Fl5BoWTwPdnUZUv0Zg85ejCoLNw3qbt3MWXfVdQHg/V2EJtCTfjsBUwsf4GfQ3QOrfxtjIn3zLpGW3r7sKUERERNJhP/zDjV8E+jiNqL6tjyWM+ebJTQJOA7q6CrAKLiO8XwvRkMa06Y8VZwse6xeBXg7jEinFWRTvWR7lLDoRERFJjRHYXNxsA+MjYLDTiOpfO+AQ4AH8BdO8yeU4YBdXAbbRTwjvy3y0BEta9QSeJ3pIdgeHcYkU0wy8QPTf3+x2gLPoREREJFV2A+aQa2RMp3GKWFVqA+DnwMdEN9jeAM7E5kDXs/0JNzznkryEvwfWMxq1VbPye61eJ25dgScIn7dXuwxKpATb4C9WGdwOdheaiIiIpM2W+IcXz0PrVJajGXu/xhFdQGk5tlTXaOpvqZ7NsePtjXcOsK3LoNoo30WLVmxN1mrojb9AXnD7ZZVep1a6Yr3Jwf041WVQIiW4mPyfw8McxiUiIiIpNADrCfUmeIc7jSiZemJzl18nuhE3B0uqh7sK0KMH8B7hYl5JrSRbKFleAfStwmsUmy+ZtGQZrFZB8L1bgS6YSX3rCLxP9HDs0Q7jEhERkZTqhVXFzTY4VqMepkrsDlyPzf2NSqxew5Kvfo7i+2tETGc6iqUaCiXLrcCYKrxGvosgSU6WwUaXeOsXtGJVs7WklNSzvYmujH20y6BEREQkvdbBildlGx1rseWEpO06YT0dE4ieZ7camIitg9u5RjEdEBHHjTV67bgUS5ZfrfD5dyjy/ElOlgEOJZx4XOc0IpHi/kz4vP029rd0ALApNt1keGYbmdkO8vy8c+a+LTOP3xCtKS8iIiJ5tAA34G98XIXNzZXKDAR+CrxD/mHafwL2AppiiqET4cTyLWxYY5IF92kJ4fnFlayJfWXgueaSrmQZwsuHrcESCZF60g/7G/ld4LeEa0UsI/rva7nbEuzvynPA/dgUmguB04F9sb/nIiIi0oCasB5lb8PhNuqvQFWSDQMuAWYR3VCbhiVo1Z47+tPA66ykPuZQVyqYLM8C/h647Q9tfO4OhI/TNYSPWdKT5c6E57E/S3wXbkQKWQ+rbH0+MB5bCzxYkND1tgh4Bbgd+84cDWwUw3shIiIidegc/EPcHsYq6Er1dASOwIZpryS6QfYmluRuXOFrdSfcI3pFhc9ZL6KS5W8FbvuCtl3wOTzwPNMibktDsgzWYxbcr285jUgaQXvsot0YLDF+m+j5yEnZZmJ/0y8ADsRqgoiIiEgKHYc/iXsDm9Ml1dcLm7s8kfwNxUlYg7IthcHOCzzXbNLTiItKltsBMwK3t2UN1gmB57gIm+ObxmQZbMipd7+ecxuOpFATsD12EfBJ8l8obOu2ApgOTMZGS0zKbBMz2/2en1/K3PdO5vGfUr2h3NltNdYzfiGwGzbdSURERFJiJLCQ3Bf/Z6Rj6G492wT4GfBfohtfK7FibCdgy0AV0w7r7fA+x/nVDtqhqGQZ4PeB2+8t83n7E27Ib066k+XtCV+s2c1pRJIGPYEjsWKCwYtY5WyLsYJ9d2DrLZ8JHAVcnrn/nCrF2w37rI/ARpL8AEt2b8US7HwrHZSyzcnEfxL2N0ZEREQSbjts+Gn2y34RNrxM4jcMG843meiG13Ks9/MEbKh1lIMDv7MQa7ymRb5keavA7Sspb83lHwd+/+nM7WlOlgH+hX/f/uI2HEmojsBh2EWqYMG9Ytty4HngauAM4GsUHtXUjH0+fxDDfuSzHrAPtszi5cBTWGGwcnudHwGOR9OcREREEm0gdkXf+yWf5LV5k6YZa5iNw3omohpeS4F7sB6cdTy/e3fgceNqFnVt5EuWwYY/eu87q4znfTPwu6dkbk97shxcXmwRVkldpJjs36nrKa8g12SskOSZwK5YYb1ybQ6cVln4FWuHjc44HbgJG95d6vzrJdh78M3M84iIiEjCdCU8h3Mc+mKvtRZseOCV5K+ovRQ7VqdgQxe996VtWG2hZPn7gfveLPE5dyHckM323Kc9WW7Bplt492+U04ik3nXHLkTlGwET3BYC9wHfo/LihV7dqvhc1dIfG/lzO1YropT3Zwbwv5Q3EkZERETqQAs2NM77xf4w+YcAS7w6YkPibwEWULwRNp30LQdUKFnuQXgt1u1LeM5rA78z3nNf2pNlsDW/vft3tdtwpE4Nwqrql/K3531sybyv0LhLEbZgFysvAF6j+Hu2DOul39pBrCIiIlKBMcAa/D12WmPSrWziPB5/UTbv9mdn0cWnULIMVkzHe3+xJbM6EV5ma1/P/Y2QLB+If/9ecRuO1JkdsSkfqymc7H2OjYDZxU2YdW8Y8BtgKoXfx7XAo9iFBhEREUmIUfiTshmoUna96IxVcQ1Wwf6Oy6BiUixZHhW4fzaF50QeHXj8VGwuZlYjJMt98M+1XIV/Hrw0psHYcOJC83CXYVWjR6EpOqVqAvbCpjUV66V/kNJGx4iIiEgd2An/UiCqlF1fgnMIt3UbTiyKJcvN2Pqp3sccUuD5Hg489sLA/Y2QLEP4fd3GbTjiUB9sCHWhqtZfZB4zwFGMadENK1T2HoV7mu8ChjiKUURERMowABum6f0ivwR/b5zUXgvhdYLTuDxJsWQZ4LeBx9yX57kG4h9auhbrTfNqlGT5Mfz7qItgjac9tub7IvInbq9hawZ3dBNiarVgS289Rf73fgVwGRr1ISIiUve6AQ/h/yK/m3QmZ0nRj/Dw4zQqJVkeSnhY8XoRjzsv8FxPRjymUZLlG/Hv46luw5EaG07hIlSvAt9wFl1j2R34N/mPxWRsLWoRERGpYy1Yj7I3KXkP2MJlUA1sEP4G1RS34cSmlGQZ4LnA486OeMy7gcecFPGYRkmW/0jx90vSpzNWrTk4KiW7fYINE25xFF8jG4l/FFdwuwvo7Sw6ERERKclR+Nf2XQAc5DSixrQF/obU+27DiU2pyfL3Ao97K3D/noH7FxO9bmujJMuX4N/Hn7kNR2pgZ+ADohOxWcAPKFwcT+LXDByLXfyMOk4zgK87i05ERERKsh3+JGYt1luRtjV+69kmhHuE0qjUZLkHsCTw2B08918fuO+mPM/TKMnyVfj3cYzbcCRmxxNek9zbY9nXXWgSoTN2QStq+S7VDREREUmA3sBE/F/idwJdXAbVQPrgf+/nuQ0nNqUmywC3BR77x8ztnbH3x3vfPnmeo1GS5Zvx7+MpbsORmHTE1kKOSpI/o3DleHFvB2z+eNTxewDo6S40ERERKSY7j9n7Bf4GsKnLoBpEE7bmqfe9T2PDqZxkef/AY2djycLxgdunkH8URKMky0/i30cVc0qfgcALRCda1xA9DUHqTwdsibtVhI/j+8BW7kITERGRUhyDfwjsHGA/pxE1huBanTu5DScW5STLzdhwdO/jDwUeD9z2iwLP0SjJ8nT8+6hCfemyMfAR4XN5EVZ3QpJnb+Bzwsd0LrCLw7hERESkBMOBT8l9ga8GzkfzquJ0H/5G0xluw4lFOckywK8Dj38ZWOP5/xpsvnc+jZAsD8S/f8tQYac0GQpMI3wefwhs4zAuqdwGwPOEj+18bAkqERERqWN9CPfiPU70mrdSuZ/if69vdRtOLMpNljfDv7xZcHu8yO83QrJ8FP79e9ZtOFJFw4CZhM/hf2JF8CT5OhIuWNgKLAT2chiXiIiIlKA9cAX+hGU6+hKPw174G0tzgHZOI6q+cpNlgGfInywfX+R3GyFZHo9//37vNhypki2wz0fw/L0ZrZucRhcTPtaLsWXyREREpM4dhCVv3mHZF6Bh2dXUghWx8jaW0jZXvC3J8neJTpQXAV2L/G7ak+VO2BxH7/7t7TQiqYbuwNuEz90/o7+5aTaW8DGfjYpsxiZtV6NFRMSdf2LLXtyBzaVqwZKOEcCxwBfuQkuNNdjyISd6bjsZW9Krkd2FLZezTuD2O7Gel0Z2ONDL8//ZwHOOYpHqaAZuJ1wV+Trg+1gClRTnkD8feQqbr1upDsBZ5K+I/wjwehVepxZ+h30PXOq5rTdwL9bDvMRFUCIiIlK6dtjyUt5h2dOwpFkq91X8vQrLgQFOI6qutvQsAxwInBbYhpTwe2nvWX4O/779wW04UgWXET5nb3AaUdstJf8Uiier9BqHFXiNVuB7VXqdWvol4f24k/wXBERERKTOHIx/+OcqNCy7GpqwKrfeRtLlTiOqrrYmy22V5mQ5eGFlLVqjNemOJHy+PkNyq5sXSpbXAoOr8BoTCrxGUpPlJuBuwvtyrsugREREpDybApPwf5k/gFXRlrY7Hf97ugTY0GlE1aNkuTqasKrXwc+eJFdfbDqL95h+RrJHlhRKlluxC6yV6A+sLPIaSUyWAToT/n5djlVIFxERkYToiM0n9X6hfw4c4DKohOuIVRz3vqe3OY2oepQsV8cxhPdL67Im29/xH89l2Hr3SVYsWZ5CZaORzi3y/ElOlgE2Jlz08Tk0HFtERCRxDgHm4R9iNw7o4jKoBIuqAD3KaUTVoWS5cr0Jr717n9OIpFJfIXye/thlQFUSTJY/wP890QrsW8Hzvxl4rpcIv49JTpYBjia8T8c5jUhERETaZGPgP/i/1N8BdnQZVEI1A6/hfy+n4a98nERKlit3G/79WUFpxc6kPjUBr+A/pi+SjrWUg8nym1hVb+9tN7fxuXcOPM9i4CeEP+9JT5YB/kH4u6CT04hERESkTVqwtSJXkPtiX4nNTUtD46+WdiQ8H+9Rkr00pJLlypxJuvZHbA177/Fciy0TlAZRyfJuhJPcbm147msDz/NX4AzCn480JMuDsGH53v36vtOIREREpCI7Ae8Rnmu1qcugEui3hBt/Sa6OfQz+5Z9OiPn1NiG85FRS54F+Das67z0XXie5lZLFPI//mN7pNpyqikqWwUYceW8/uczn7YR/NYZWrDp8WpNlsL/73v2aSrIvnIqIiDS8zljxL++azAuwhEVK0wFbOibYADzRZVBSc0OAOfjPgfnAFi6DkoptS7hXeVunEVVXvmR5bOD2/5T5vEcFfn8KNnUlzcnyeoTfTxXSFBERSYEDCS+J8jegp8ugEqQ/8Cn+92858A2XQUnNDCA8SmM16Sj41uiuwn9cH3EbTtXlS5YHYOew9yJBOWsuPxR43gsyt6c5WQa4Af++3e82HBEREamWvtgXu/eLfiZwmMugEmR7bG6f9/1bgc3JlfTaCPiQcAJwrsugpCqagRn4j+sRTiOqvnzJMsCDgft+VeJzDiR/op32ZHlXwt8BbZnvLSIiInXqBGAR/i/8u7BkWgo7Cv+Q9lasANhol0FJbDbFhpcGG/+3uAxKqmYX/Md1IbbGepoUSpaPDNw3ldLWXP5Z4Pf+7bkv7cky2PJb3v073G04IiIiUm1Dgafxf+HPwpJBKex0YA3hIbnfcRmUVN0wbHmYYMP/HlTQKy2CSd9dbsOJRaFkuQMwO3D/10p4zncDv3Oi575GSJaDhb6udRuOiIiIxKEJK/S1EP8X/wPABg7jSoJvE66K3AqMQ4lUGhyMFcILHt87gfYO45LqCq6dm7akDgonywDXBO4fX+T59gg8fjHQ1XN/IyTLo/Dv36tuwxEREZE4bYKtHez98p+PJdJN7sKqe0cRXoO5FaucPcBhXNJ2LcAlhIfatwK3oWVi0iZYtG8Ht+HEoliyvHPg/qVAjwLPd33g8TcG7m+EZLk3/r8Rq9BFUhERkdQbTXhpnIeBjV0GVecOA5YQbhzOAPZxGJeUb33gCcLHshXrfStlLqckR0f80ynWkL75ylA8WSZzm/cxp+R5rs7AvMBj9w48phGSZQgXhiunkriIiIgk1EBgAv5GwELgh1ivm4RtB0wm3EBciw3LVqXU+jea8NzNVmx5sFMdxiXxGYL/WE9zG05sSkmWfxx4zFN5nuu4wOM+Jjz6qFGS5Wfw72Mpc71FREQkJUYDX+JvDLyOzVeTsO6E5z9mt8+wObBSfzbB1tWNOm7TsGViJJ2G4z/er7gNJzalJMv9CU8p2TzicY8FHvPziMc0SrJ8H/591BKMIiIiDaYfVtDI2yBYg81Z6+0wrnrVDFxE9HzXVqxwzvrOohOvDsDZhJdQy26Po6XU0m5v/Mf8abfhxKaUZBnCI4ouCty/Mf5h62uBQRHP0yjJ8i349/EEt+GIiIiIK/sSXipkLjAGDc2OMoLw+5XdlmAFpHo6i66xNWGjJj4k+vgsBcai87oRKFn2OyLwuGn4Pwe/CNz/WJ7naZRk+Vb8+3i823BERETEpfZYcrwYfwNhErCLw7jqVSfgAqKrZbdihdTGYgVzpDZGYku8RB2PVuA/2Prj0hiCw7DTuvxPqclyB/uyhDQAAA79SURBVMJTb0Zm7msCPgrcd1ye52mUZFnDsEVERCRkU2wd5uDQ7PFoaHaU4dhc73wJ2qfYcODurgJMuWbgEKzXMN8xmAecjJZJazSD8Z8H092GE5tSk2WAqwKPvSVz+z6B2xcA6+R5jkZJlp/Fv4/7ug1HRERE6skhwFT8jYWZwEloiZ2g9sDphJcaCTY+r8AuRkjlumIV3IO9Yd5tBXA1NjdfGk8HYDX+i36dnEYUj3KS5WBv+1JsyshNgduvL/AcjZIsB/+e62+3iIiI+HTGhhovx99o+C8wyl1YdasLcB4wn/wJ3GrgHmwZEl10KN/mwKUUfo/XYPMN1biVqfjPjeFOo4lHOckyhEfCjMGWD/TetmeB32+EZLkv/kKOK7GLoiIiIiIhWxBeUqQVuJ/o5UcaXW/gcsKN2OA2HbgM2N5NmImxHtagf4nC72cr8CC2LrYIwL34z48z3IYTi3KT5bMDjw/+/gcUnrLQCMnyAfj3b5LbcERERCQJRmK9yt5GxCpgHBrqGqUv8L/Y8PViSd5/gZ+hiw9ZvbGlWh7GP5Q2aluODSNVkixBY/GfK/e4DScW5SbL/chfmLAVGx1TSCMky3/Ev3/XuA1HREREkqI9cBrwBf7GxCJsyHYa5wRWqgO2nNELFE+aW4GPsQsQo2mcwmDN2BDZscBECjfms9ssbJmugQ7ilWQIztFdRPr+RpWbLAP8g+jP1BpgwyK/2wjJ8mT8+3eo23BEREQkaXpiicoy/I2KT7EeQVUejvYV4GbCcwQL9Zo+Cvw087tdax1wTNphPcGnYfOLZ1Ha+7EGmxJwIulLeqT6mrD1hL3n0NFOI6q+tiTLhxL9+XqkhN9Ne7K8J+G/wWn5uysiIiI1thG2rJS3GEor8AbWMyrROmON9gmU1oua3VZjjeHrge8C2wIdaxx7uZqBTYDDseJc/yG8nnex7TXgHNSLLOW7Av+59LjbcKquLclye6z3dG5gO6qE3017svxX/Pt2r9NoREREJBV2AZ4h3Ih6Dqv8LPn1Af4H69UJ9tSXmkBPBh7CEoMzsDVBN6J2iXQT0B87D44Hfg3chVXeLVboLGpbiyXIvwaG1WgfJJ22Jnx+7eQ0oupqS7JciTQnyxtgS855900rP1RIw8xERERME9YzcREwJHDf48DPgedrHVTCrIMNtx6V2QZX4TnnAZ8DX2LFxmZltlXYHM7VWJK+HOvlXoLNs+6C9Qr3yDxPT+wY98WKBPUD1vf8v6UKcU7Eino9nIlVpBqeBkZ4/n8fcJijWKptKTZSJestbMRJXM4Arg3cdjpWZyHprgTO9Px/MlZ0cY2bcERERCSNmrEh2B8R7oGYSLp6deK2GdYQvQl4B2u0ldtLW6/bTKzQ0HnYPMFKk22RfL5J+Pzb22lE1aOe5eoYjF0w9O7XqU4jEhERkVTrgA0v/ozwENu70fDatugB7IctSfVPYCrh+eL1uM0GnsLWnz4SGyIuUooWYFPg68APsHnvtwD/Bt4FPsHm267ARklk59++DzyBzUEdDryI/5x8FSs0l3RKlqvjX/j3aSr2HSYiIiISq85YcaZgxeM12LqnO7sLLRU6Y5WlRwPnY4nEi1jPbbE1iqu5zcXWi74Xq5R+MrAHtmaySCl6YvPtf4QVDnyD8BzStmx7AbsTvrD0v7XZrVgpWa7ciYT36UinEaWI5iyLiIiUpiswBkucewXuexxLsB6rdVAp14TNJ+6LFeBaj9wc42agG9a71hlbiik7Vzk7d3ktsCDzXPOxRuR8cnOgv8j8PCvzOyLl2Ay7oLIHNhR/S+y8LMd8LKFekvl/dp59F3I9g72xizl/A47x/O5KYB9sDfSk0pzlygwGXsb/nfQ0dl60OolIREREGlpP4EKs8Rq8mv8ytsRQuQ1mEalvHbCk+FyswNbnlNYr/AlW6+Ba4GzgIGw0ygAKz3NvzjxmN89tfSJedyZWBTmpZuBf/umZmF/vZMJLTp0U82vGpRt2ccF7PiwBhroMSkRERASsoXIu4TnNrcB7WKNMc8ZEkqkJ2B4bSfIgxdfWXovNQb4JG4HyFezCWrUdQng49kv4e2cl/Zqxug/B8/DMQr8kIiIiUmsdsaqjHxJuuEzDGttxNJpFpLoGYZ/lOwjXKAhuy7DhrpcAB2K9vrVycUQ8t6LplY3kEsLnwC1OIxIREREpoAUrqvIq4UbMImwY5lbOohORoB7YesXjsDVpCyXH87Glws7Bim25HDXSDNxPOMbr0RSQRjCW8LF/BVvnXkRERKTujQAmEN3ofgar/JyGZV9EkmYYlmxMpHCl6lXAJKwHbyTQ3kWwBUTNV23FioDpb0t6XUj4mM8g2fPWRUREpEHtjA2Ni2qUT0ZDtEXi1hs4GluvuFBRrjVY79zvgP1JRi/dYCxRCu7LHdRfci+VaQL+QPhYL0DLF4qIiEjC9Qd+QXTDdjFwHVZMSEQq04INk74AW1Kp0FrdM7GCXEeR3PW0NwM+Jbxvj5LcfRK/dbA56cFjPA/Y1WFcIiIiIlXVHhuCPZHoxvvb2BDRWhYLEkm6ftjnajwwh/zJ8WpsaPUFwHDSUxBrY+Ajwvv7KbCLw7ikchthSxIGj+1c1KMsIiIiKbYjVlhoKeGG0HLgLqzKbqG1WEUaUTusLsAlWPIbXEop2Hs8Hkum0zzlYUPgA6Krdp/iMC5pu1FYUhw8pl8A2zqMS0RERKRm+gE/wdZmjmrsT8GGcG/kKkCROjAU+CHwAIXXPF4KPAycDWzpJFJ3+gNPEf2+3Ei6LxakSWds7vwawsfxLWCIu9BERERE3NkTa9QuIroA0aPAyajRK+nXDTgYW3Kt2LJO72DFj76OJRqNrB3Ra/Bme9mPcBealGBP4F2ij9/tQBd3oYmIiIjUh87k5jZHDTFdnbnvBCypEEmDTYEx2Lm9nPzJ8eLMY8Zg83Ul7Bjy98BPAAa6C00irINd5IjqTV6F1bJIyxx7ERERkarZEvg91isU1fBdAtwJHAp0chSjSFv0B74N3Ez+8zs7quIl4CJsrrLWES7NdsB/iX5P52BL1+lvhlst2GihaUQfp0+AfZxFJyIiIpIQLcB+wF+ILvrSCszHEo9D0XA9qT99gMOB/8MqvxcaWj0TO5ePQdXhK9Ee65XM11M/DTgNXYBwYSTwBtHHZS1WAFIjh0RERETK1II1tMYDC4lubC0jN1R1AzdhSoPrip2n2arVUUNMvVML0risU73YBniR/O//28Ah6H2vhX2xNcDzHYv3sBEUIiIiIlKh7Pzme7AEOV8vxcvAz4Ht3YQpDaA78C3gMizxXU3+hGAtNkT4KuAgLLGWeLVgVcLnk/+4vAucjs2hleppDxyLfS7yvffLgIvR0HgRERGRWHQHjgJuI/9Q7ew8uHHAkWiIq7TdYOB4rGL1GxROjrM9Zn/Czrt+DuIVsy7W2x+1xrt3SseV2PrN0nY9sNE9n1J4Tv5dwCBHMYqIiIg0nBZsKN8l2NI6hZKYyVjyPBpLuEWC2mHDo8dgw/+nUvicagVmYEnAaWid8Hq0IbZUXaGLHCuBu7Eh2h3dhJk4LcD+2Jz7JRT+jNxL460HLiIiIlJ3tgJ+CjxL4bmjy4EngPOwNT/VQG5M62GF4i7DzplCSzlltw+BvwInouQ4SYYBt2KJcaHjOxf4M7A30Owk0vq2E3AFdpGo0Pu4Brgf2M1NmCIiIiJSSDf8hZcKNexWZR5zJbau88YO4pV4DQAOxIprTcBGGhRLjIPnhZLj5BsI/AaYTfHj/wl27EdhdRMaUXvgq8DvsCkGxd6zRcDVwGYughURERGRttkE+C5wOzCL4o2+ycAtwPex3hQVpEmGdsDW2DzjPwD/BuZR/Hi3Yks53QP8CNgd6FDj2KV21sGKfBWbvpHdlgIPAWeS/kRwQ2xawb3kX4kguE0Ffgz0rH24UimVhxcRERGvJmA7rMdkT2APYP0iv7Maq6L7WmZ7PfPvgvjClCL6YUPvt8QqoO+ALR9UyoWN5Vil6peA54HngI/jCVPq3DDs4spJQP8Sf2cm8Ipnewa7KJM07YGh2N/BEdic/S0pLX9aAPwT+Dt2IWF1TDFKzJQsi4iISDGDsKR5d6zhuA1WxKaYj7Gk+R0smf4AeB9YHE+YDacdNix+CLnEOLv1LvE5FpC7uJG90PEOatyLXzusaNWxWLGvcpaWWoOt4/wSNlz5PexvwRTq4zxrxj5HQ4HNgS2wETPbYwlzqVZiifFt2HSG5dUNU1xQsiwiIiLl6grsiiXPO2S2cpY9mYYlzR+QS6KnYsurqIHp1wMb+jkIW6ppSGYbjDXwy2nMzySXFGe3KdhQUZFSdQG+hs1X/gY2jaMtVmFTOt7HLqzNBL4AvsSKZM3K/FxJQt0M9MVGWqyH9Y5nfx6EJcdDaXshw8+BhzPboySzB10KULIsIiIi1dCT3HDf7L9bYj1S5ZiJJc3Z7RMskf6MXEN6ZVUidqs91mjvjzXc+2FFsjbIbBtltm5teO4FWO/d25l/38IS4y8qjlokbAtyifNeVLfgVyu25vMy7ELaSmw5pjXYnOGsrthnqjM21aADltR3p7RRMKVaBbyA9SA/jI3E0MWmFFOyLCIiInHphM153BxLnLO9OJtTWVGwBViPzpeZ7QusF2ohVnF2XubfRZ7bFmDDv1dV8LpeHbGhqNnGedfM/3t6tl6Bn7M9XP2APhW+/mrsQsJk4COsdy473P2zCp9bpK3aYZ/v4eTm+m5Bcpeays6/fgZbQu0VLHGXBqFkWURERGotO0cwOz9wSOb/m2T+bUtvarm8ifMSCvdWdyPXQ96O2sQH1lCfjg1b/xRLirPJ8SdUL/EXiVNPYBes1sFQchfMihUOrKXZ2AWn7PSQ7BzrWS6DEveULIuIiEi96YUNQfYm0N75hn0zWzWHV9bSl1gjfBaWEM/Cesc/w5Li6ZlthasARWqgO7nkeSPs892X8Ge90nxlDrkpHNnP25f4ayfMrfA1JKWULIuIiEgSNWFDmft6tl5Yr693897WCevV7pF5jhaswQ65HuPsnMgo2fmTq7Gh3SuwNWaXZn5elLk/3zYHa6SrR1ikNNnPaHAusvdzDLmRIsG5zdWceiEN6P8Bb/xoOzWaROEAAAAASUVORK5CYII=" width="65%" style="display: block; margin: auto;" /> --- ## 3. Direct effects estimated between exposure and network <img src="data:image/png;base64,iVBORw0KGgoAAAANSUhEUgAAA8sAAAI0CAYAAAAqQrvXAAAABHNCSVQICAgIfAhkiAAAAAlwSFlzAAASdAAAEnQB3mYfeAAAABl0RVh0U29mdHdhcmUAd3d3Lmlua3NjYXBlLm9yZ5vuPBoAACAASURBVHic7N13vBxl2f/xz0nvJAESEggcAoFAKEKUFrpRQGlKUUAD+kjkERBFBWyPsYP6oAF5NKDSlS4SqlERCCASmrRQEgiEkAAhvZ6ck98f1+5vZ2Zn++zcM7Pf9+s1r2TLmb1mZ3Z3rrnv+7rbEBERERERaU2bAKOALXLL5rl/h+f+PwxoA4bknj8A6An0BvoBXcCy3GPLcrdXA+uAVcBCYBHwLvA28E5ueSu3dDVz46Qxba4DEBERERERaaLuwHbAjrllh9yyI5YYu7IWeBl4Kfdv/v+zKSTg4pCSZRERERERyZIRwAeB8bllAoWW4bR4G5gJPAw8ATyOtVZLjJQsi4iIiIhImm0NHA5MBPbBulVnzWrgSeAh4F7gEWCD04hagJJlERERERFJk97AgViCfDiwcwPrWg+8ho0rfpvC+OIFuX/fwZLSJbnnrwQ6sFbe1UA3bNwzuX+7YWOZe+duj8A/DnpY7v/bAJs2EPcy4G9Y4nwvML+BdUkJSpZFRERERCTpegMfA07BEuT+Nf79O8DzFMYGz879+zruWmg3pXgc9djc7e41rus/wA3AH4F5EcYoIiIiIiIiCTQemIoluxurXNYDs3J/NwkYR7oaCfsD+wPnANcAc6l+2zdi234O1qItIiIiIiIiGTEc+B7W6ltNctgB/BO4ANgLm9opa7YCTgKuxqajquZ9WQvcAhwcf7giIiIiIiISld2BadhY4EqJ4EKs1fUEYLCLYB0bB5wPzMDGT1d6v54CJgN9XQQrIiIiIiIitWkDjgL+TuWEbxFwCbC3k0iTazPgTKxCdhfl38O3ge/m/kZEREREREQSaAI2l3C55G4NMB1rQc5i9+qobY21OM+m/Pu6ErgQGOQmTBEREREREQnaA5vuqFwy9yrwZWCgoxiz4ADgVqzad7nu7GeiCxEiIiIiIiLObA1cD3RSOnn7J3AsNl+xRGNb4JfY3MzlLk6c6CpAERERERGRVtQNa71cTnii1oVVbd7DVYAtYhDwNeBdSifN07HK2yIiIiIiItJE2wH/oHRy9jA2p7DEpz82rrlUS/PK3ONq3RcREREREYlYd+BbWIGuUlMZHe4sOgGbz/rXlJ566gFge2fRiYiIiIiIZMxmwF8JT8CWAl9ErZZJsgOWGJfaX0e7C01ERERERCQb9gDmEp543QWMchealNEGTCa8a3YXNs1Ud2fRiYiIiIiIpNhngFUUJ1uLsURMkm8EcBvhFzvuAYa6C01ERERERCR9fkLpca/DHcYl9TkbWE/x/nwJ9Q4QERERERGpqA24mPBEeRrQ011o0qAJwAKK9+vrWJVzERERERERCdEGXEJxMrUGOM1dWBKhzQmf+usNYIzDuERERERERBKpDfg9xUnUe8B4h3FJ9HoCN1K8rxcAYx3GJSIiIiIikjjfpzh5WgTs5jIoaZruwJUU7/PXsNbnTGpzHYCIiIiIiKTKJ4Bb8ecSC4GJwPNOIqrPTkC/Eo+9A7wZ0evsDPQt8dgiYH5Er9NsbcClwJmB+x/C9v362CMSERERERFJiN2AFfhbFxeSzoJPTxBemGwj8HhErzEUWFvmdS6O6HXi0gZcQfF2XOIyKBEREREREZeGYt1uvUnSOqxqchqVS5Y3Ek2X8rMrvEbakmWwMcz3U7wtn3cZlIiIiIiIiCt/ojhBOt1pRI2plCz/IobXSGOyDHbh5FX827IS2N5lUCIiIiIiInH7BNnrelspkV1EY/NE71Jh/WlOliG8S/79qC6WiIiIiIi0iL7A6/iToqeAXg5jikIwWe7AEmTvfUc1sP6LA+t6i2wlywBfoHibPu00IhERERERkZh8m+JxylmYIiqYLK+nOMG9pc5198AKn3nXdSHZS5bbgPvwb9M8Slf/FhERERERyYRBwPtEP5Y3CcKS5WDX6XXAZnWs+9jAeuZj0ytlLVkGq4QerPh9ttOIItLNdQAiIiIiIpJYZwFDPLffA37kKJY4PAc86bndCzipjvWcFrh9DdBZZ0xJNwe4LHDfN0h/N30REREREZFQ3YE38LcYfttpRNEKa1mG4umeZtW43mG5dXnXMRY4hGy2LAMMB1bj37YTnUYkIiIiIiLSJB/Hn/ysAAY7jShapZLloRR3K65ljPa5gb99OHd/lpNlsNZl77bNcBtO49QNW0REREREwpwSuH0jsNRFIDF7H7gzcN+kGv7+1MDtqxsLJzV+G7h9KDDCRSAiIiIiIiLN0hNYgr+lcH+nEUWvVMsywJGBx6qdc3l84O9WU2iNz3rLMsDj+LfvdLfhNEYtyyIiIiIiErQv/i7X7wCPOIrFhXuBtz23hwGHV/F3pwVu30ZrtMbn/SVwu5r3LLGULIuIiIiISNA+gdv3AV0uAnFkA3B94L7TKvxNL+DTgftapQt23j2B2/s6iSIiSpZFRERERCRor8DtfzmJwq0rA7ePBDYv8/xj8M/JPB/4R9RBJdwzWNfzvBHAKEexNEzJsoiIiIhINPq5DiBCYwK3n3IShVsvYGNw8yrNuXxa4PZVZHdu5VI2AM8G7gseS6mhZFlEREREJBp9gVuBOdi0OdOA84ETsMJPaUqm2wO357oIIgGuCtwOVrrO2wL4aOC+6yKPJh2Cx8q2TqKIQA/XAYiIiIiIZMRi4Hjg68CPgYmBxzuxrrlzcsvcwP+TUgiqNzDIc7sDK/DViq4HfoFdCAHYE9gd627sdSr+3Gom8FLTo0um+YHb5bquJ5qSZRERERGR6GwEfg48DPwJ2NrzWHdgm9xyaMjfLqY4gc7/Oz+37jj0D9xeFeNrJ80yYDpwoue+ScDXAs8Ltjhf1cSYkm5l4HbweEoNJcsiIiIiItF7BNgN+B3W2lyNTXPLh0IeW48lzHNDlpcoTlAaEZxPuCPCdafRVfiT5c8AF1B4X/YBdvI8vga4JZbIkml94HYvJ1FEQMmyiIiIiEhzLMOSrK8CF1KchNaiFzA6twRtBN4ivEV6DtZiXYvVgdt9Q5/VOu7DLlRslbs9DDgCuCN3+7TA82/B9n2rCuuZkEpKlkVEREREmmcjcDHWLftGrAt21NqwRG4r4KCQx5dSnEDn//8mxRWbV2NzKueLAffDkvVgi2Gr6MLGLp/vue9ULFnuA3wq8Pyr4gkrsQYHbkfZ60FERERERDJoEHATlkAnZVlPePXuRYHnbd+E98O1Jyh+L0rZAUua889dhxWuOjmwjtcpPePQIRS//xc3uA1JdCf+bax2GELiqGVZRERERCQey7FWyEeAi0jGWM6elO7e7TUNeAB/63QrVch+GXgMG58MhTmXjww872osqW5lOwRuv+4iCBERERERSacPYUmn65blRpYVwNPY3NI/Bz5HusY319KyDDA58Pw5WBf2/O0uYLsyf98KLcub4m+B7yDF1bBLdREQEREREZHmeRybs/c214E0YAA25/BBWLfke7BK0Fl1I/7iZ6Px51MPYgl0KzsAG0Of9xwpLvClZFlERERExI2lwHHAl7FkM21eAb6EzSX9HWCh23CabhnwlzKPXxVTHEkW7Jb+oJMoREREREQkM8YDr+K+a3U1ywPAMaS/4a3WbtgAHyX8PVkJDKzwt1nvht0beBf/9k10GpGIiIiIiKTeIOBA4FHcJ8OllveBfZv1BjhQT7LcDXiD4vfmyir+NuvJ8qcoPl6SUMSubqqGLSIiIiISn57AOGBXYJfcv+OwrsxJNwTY4DoIx7qAjwPDA/e/4CCWpDkncPtaWndubhERERERKaMf1hL7JeAKYBY2Ntl1C3E1yyLgXOCfgfv/HOUb5Fg9LcuNyHLLcnDburCLQSIiIiIi0uIGAftjrWvXkK7E2Ls8hnWnzfdAPSzweFduO7NAyXI02oCZ+LdrutOIRERERETEiSH4E+Pn8c8tm7alE5vCqlQS/Fjg+Y+R/uJeoGQ5Kp+leLuyNLZdRERERERCjASOBX4E3AW8jfvkNqplLZbsj63wHkyg+GLAV6p8/5JMyXLjNsO67Hu36RanEYmIiIiISOQGUNxi7DqhbcayEJgCbFrDe3NLYB2rgJ1q+PskUrLcmDbgVoovwIx2GZSIiIiIiDSmO1aFehIwFRtzuR63Sew64J0mrv8ZYDLQp473ayQ2FZB3fS9hXdLTSslyY75F8fZc4DSiiLW5DkBEREREJAbbAHsDe+WW8Vi1ahc6sUTzP7nleeC5XIwzsEQ+Sg8DFwF3YglNvSYBVwfuuw+bSqmzgfW6sjcw0HN7I/D3Jr7eEOy483oDeLmJr9ksRwG34x+7/jiwH5peTEREREQksQYCE4HvAHdg3Y5dtRavxSpjXw78N7AP4Un6cGBBhK+7DutKHvX0PZeHvNYvIn4NSbaxwFL8x8BiYDuXQYmIiIiISLERWEvXhVh3aldTNq3Ivf40bNzz/lTX5bkb8NeIYng39z5sWdU7V7veWEt18HW/3qTXk2TZDphHcff1Q1wGJSIiIiIiZjTWJXga7opwLcES46m5WMZR/3RKUyKI5xUsQY+ja/lwrPtwMIYLY3htcWdHYD7F+/2/XQYlIiIiItKq+gEHYV2q76a4+2ccy7vYeN/vAUcDoyLcvoOxMZ71xjYTOIHoxzlXsiewMiSeH8cch8RjF8KHM/yfy6CaTQW+RERERMxAYDBWhKfc0gcYhJ1HDc797eDc7U2w1sWBQA9gDTZmtZxlQAewPPfcNVh33g1Y6+U6bDxgfnkfeM9ze0UjG51Aw7ECXBOwbswfxLr+xqUDa6WdiXU3fgJ4AUsMojYMeBrrRl6LTuAeLDH9V9RB1eBA7CLCwMD9vwa+igo9ZcUErJjXZoH7rwROJ53F3aqiZFlERESybiCwFbAFNo6z1L+uKiM3aj2WNC/Euki+CbyV+zf///lYEp5EI7HxjgdjLchjYnztLqwq9b9zy2NYdeqOGF47P075wzX8zVKswNal2D5Ngg9iFbGHBu5/CDgROy4lvSZjx1uvwP35gnVdsUcUIyXLIiIiknbdsGR3NFZ8ZnTg/8HWkGZZjrWwrKT6ZGsToCfWUt1s72HjTF/JLS/nllew1uq4DMdaJPfHWqz2JL5z0qVYZep8i/Ej2IUGF76HjVWuxmvY+Oxp2DYkzR5Y4h/8rL2FdRF/NPaIpFF9gMuAz4c89hvgTJrT2yJRlCyLiIhIWvTHpiwZB+ycW8YA29JYN90OYBE2bc+SkGVp4PYyrDVlWe7vlzTw2l69sdbt/lgrzmCsK/cmwKa5Zajn/97bw2gs4V6MP4F+FngGq3rbqOFYq3F+GRvBOquxDkuM/4W1GD+GXSxIgoOw+XwrjTN+ArgE+CPJ79K8GzAd2Dpw/zqsUvZltEBylRE7An/CLoJ4bQR+QPUXeVJPybKIiIgkTV+smMw4YCcKyXE7tZ+7LAHmYl1W52NJcfDfRVEEnQD9sERlS6zbuff/o3L/37TGdS7Fkmbv8jzlx2Fvjs0lPAGb6ziuluPlWFfqhymMN05i1/PhwFOUHqfciY0PvRhr+U6TTbHE/qMhjz0MfAGYHWtEUoseWIvxTygelrIC+Bxwa9xBuaRkWURERFzqC+wOjPcsO2MnbdXoxMblzs0tczz/n0u83YvTYDDWGj8G2MHz/zEUipVVsgEb5zsL6177H2xc+KFYcrwH9U+hVIu3KSTFM7EENOnjJ7th43snhjy2EvgDNhXV3DiDilh34LvA/1Cca6wFLsIKk8UxLlyqtxvwe2wMetBLwCexQnciIiIi0gS9sBbHM7Gk4BnshLma6XHWYy2aN2FdAE/AWpx7xrkBGbc51hp8OlbQ50HcTNMUtmzA9v80bE7jYFfftPguxds2Hzif6i9WpMV/UfrzPQvYz11o4jEYu4BRal/dCAxwFp2IiIhIRg0FjgR+ilXGXUN1idFc4GasAJKSYve2xSri3oG14nfS/OR4BdZafCFwFDZuO+0Owj+f8tPY+9rHZVBNcgDW8l9pP8/AepZI/Hphx98iwvfN+7nHRURERCQCI7HkdirWclRNUrUAKwo0BUuKhsUdtITaAvgMcA22j5qdHC/BkvGvEl837jgNwypDd2EJ4lFkdzjkZKwnSH7fvgv8EHiH8H3fifUYGe0i2BbUDfuenkvpz+NNWE8TEREREanTKKzgy3VU14q0ELgN63I6ERgSf8hSQl9sn1yIXejoovkJ8jLgAeBcst3Nsxt2IeAarJdEVg0AbsC/j2dhhfnAkq8/Uvp4WAtcgRX3k+j1x4bAvELpffAG8HFXAYqIiIik2QBqS6jmYAnCZCxJyGpLWlqNxvbNdKrvIl/vUqmXwapcHJOxqt1ZMpzst9KNwQq9effpNKyrb9A+wD8pfzzMxFo/K02tJZVtgfXceY/S7/cK7Ht9oJsQRURERNKnJ3AINo3Iv/GPtwwu67BxyfkxprVOUSTNNxg4EbiK0uMUo1pWAPdgvQj2xqqbDwM+DVwOvFrmbzuxOZG/i3XJ1kWWZDsS60af339rsOJelRwNPEf54+hF4Gyyf7Ehat2BD2NzJXu7xAeX1cDPsfoSIiIiIlLB5sCp2Ji1cpWQu7ACRb8AjsC6+Eny7AKch7XkVVt5vJ5lFfBX4FtYleNqpv5qx5KqW7HkutS63wR+g01NpZbG5OiOtVh6ew3MAz5U4zo+R/nxsxuxhO9O4CSK5/+Vgj2A/8XGx5d7P9dhXd5HuQlTREREJD3GYS2AMyifUC3AkugsdpXNCu/Y49k0t/V4DlbMbSKNV3TuAeyfi/vFMq/5Hta1/yiqn4tborcpNl+0d9/cTf0tlN2B47DeKdX0WrgGOIZsj3Wv1u7At7Gp1Sq9d/liayOcRCoiIiKSAr2xrpPTsDley7UW/gX4IrCdk0ilGu3YBYybKN9C2+iykPguluQv4DxE6e7/C4BLsCQ7a9Wzk2xP4DUK+6ELu8gR1T7YA/tuqmYcfQc2vvl8YDyt0WW/P3axaBpWkKuaz+7LwDmoVV5EREQkVF/gWKxydbnu1a8Bl2Fdq7M4/2sWdKfQCjuL5iXHq7DeBq4Tkc2A03OxlEqc3wR+CXzAUYytYhI2zjX/vi8DPtGk19oSuAB4luqP2XnAH7ALfLuTjd4HW2Ct6D8GHqT64RTLsPoEh9IaFxFEREREatIXa4G4BjtxCjuh2oAlXFNonZaZNNoUG0t+M6X3ZaNLJ1bI7cfAwYRXMnZtKJawTad04aLnsQRfc3ZHpzdWmM37Pj9NfD1OdscKUZXrCRO2rMSmJ/s5cDywE8k8rvNGYEUVv4b14phHbdu7Hpum7FPY97+IiIiIeAwETsaKJnlbgLzLamy+41NQFdQk2xb4CnA/5SuRN7K8jhX6OZH0VTAfDnwJe3/CpqZaD/wZq7rc01GMWTAKq07ufW+vw02X3m5YS+lUrGtxPcd8BzbX8F1YMawvYpWjd8Z6MTRTX2zYxD5YQvtd4Hrgceq/CLaCwnCZtH2GE09Xj0VERNKvJ3AYlvweQ3iLwiqsAM8tuX9Xxhad1GIcNp78KKyqdNTnap1Yi+CdWOvsk9gJd9qNAD4DnIYlPUHvYEnJlVi3XqnOIcANFFrpNwDfAS5yFpHfdsDh2LCRQ4gmge/ACmEtAt7O/X8ddpFxHXYRZhX2WVqOtbr3w4ZHDMqtY0ju382w924YMJLoCpI9j03Ndi82rn99ROsVERERyYzxWAtLqTlzV2EJ0SRUNTap8uOPp1J9sZ5al3exLp2TKJzEZ9k4bDz3u4S/H7OwImXqplpaG9aV3dujYT6wr8ugKuiDJczfBG7HCsA14/MU97IWeBT7jjgZTfckIiIiUtJY4AfYtD2luuRdi7VMqkBXMvWjMJZ8Cc05wX4eSxhbuVJ0H6yr692Ed2NfhI3P3tpVgAk1CBvG4X2vHsAKTaXN1sAJ2HzwM7AxwF24T4BLLcuxiznXAGcDe5PsMdYiIiIizg0FvoyNaws7werAxt+djKYGSarNKRSmWkv0J9krc+vWHNjhtsRaHOdS/N5twMY2fwQNURwLvID//ZlGtsZ898Oqpp+IdSm/HuvKPJvmFc/LL+uwFvoOz+0vYi3iI5u50VKfVv9CEBERSao27ATqC9jULGGtxP8C/gjciI3JlGTZFqvAezzwIaI/73oeazW9G5trdkPE68+ibsDHsMJgh1Hc6v4y8H/Y9EMr4g3NuZOwYm/9c7dXAv+FdeFvJX2wi1sjsbHGm2NTUPXDxif3wt6j/Bjl/Hjm/BhmsB4jYMn3Quz7+R1gce7+mcCE3P9HYmOjRURERKSCEdhYwVcJb5l4Heteu4Oj+KS87bG5YZsx//F6rCvpWVhFXWnMdpQe27wcGyO6lbPo4tMDex+82/8SNvZbmuMKCu/1oY5jEREREUm0HlgV6+mEj61cAfyOZBfXaWXbAudgrUVRj4f0FmkbHNcGtZi+wOcIH+awDmtlDquwnQUjsePWu81/ATZxGVQLOJfC+32W41hEREREEmlr4CdYF7ywROlRrBv2QFcBSkmjKSTIUbcgv4MV+DkK6/Yp8dkb63YcvGjVhU23dbCzyKJ3AP6K0RuwXi0aptl8R1B43y9zHIuIiIhIYrRhhYT+THgr8rvAL1EXyCQaB0yhOV2s52Ddflu5enWSjAZ+jbXsB/fVv7EKy2neT5Oxbv3e752JTiNqLe0U3vt/uA1FRERExL1B2AnqcxSffHdiY1EnoblfkyafIAcrBDe6dGJJ9xRgp5i2RWq3CdaD4C2K9+Hz2Ge2u7PoajcAuAH/dswCtnEZVAvqhhVQ24iKe4mIiEgLG4u1GK6g+GR7KTYti5KlZNkJ+D5WGTnKBHk1cDs2Pnbz2LZGotAbS4zDLprMwS6E9XAWXXXGAM/ij30amsfXlSco7IchjmMRERERiU0P4FPY3J1hSdMsLGFSK3JybEVzxiCvplCgS2PP068bcBzwNMX7+kXgMySzpfkobDqjfKxrsGmhxJ3rKOyP/RzHIiIiItJ0A7CE63WKT6TXYYWDNC4wOYZiSewMoq1i7U2QB8S2NRKnNiwBDaugPZfktDR3x7r6d1KIbx4277e49W0K+0QXLkRERCSzRgE/x7pVB0+cX8fm3VW322TohxVnmo6/wFGji3eKp/6xbY241oZN+/YkxcfES8DxuKsuvRlwXyCmu7GLROLeJynsl184jkVEREQkcuOB6wlPuh4CPkEyu2S2mt5YK+A1FIrqRLEsya3zBCwJl9bVBhyNfxxqfnmM+Kec2hN4zRNDF3Ah6a7gnTU7Udg/dzmORURERCQS3bDE636KT4o3ADcCezmLTvK6Y9MwTQOWEV2C/D6FOZBVGEmC8t2zw1qa7wJ2iyGGSdhQgPzrLgOOjeF1pTY9seE5+a77IiIiIqnVEziV8Gq4y7G5kdtdBSf/317ApcA7RJcgvwdcARyGHQcilbQBJ2GVsr3HUidwNbB1E16zN3acel/vKWC7JryWRCP/e9KJeqeIiIhICvUBvoS/S2N+eRP4BjYXq7izJVZYLaxCcb3Laqwgm1qQpRG9gLMpvnizBhunGtX44VFYd2/va1yHErCku5XC/trDcSwiIiIiVRuIJcJvU5xIPQWcgloZXepLoVDXBqJJkNeiKtbSHAOxubuD860vxpLpRipnHwIs8qyzAzi/kWAlNj+isN9OdhyLiIiISEWDsBPNxRQnUzOxlkZX1W1bXTcK45CDSUe9Sye2X8/BqgeLNNNmWKGttfiPw9nAETWuqw37rvJeLJoP7BtVsNJ0p1DYdz90HIuIiIhISSOw6Z/CkrD7gIPchdbydsLmig3rCl/v8jyWaIyIbzNE/r8xwG0UH5e3AaOr+PtBIX//ALBFM4KVptmTwv67xXEsIiIiIkU2x1p6vNVjN2JTrUwH9nYXWksbCkzGWn2jTJCnYImKSBIcAvwH/3G6DpiKJcRhdgJexP9dNRUNC0mjfljvlvz3k4iIiEgi5LtDrqK4W+5NwM7uQmtZvSmMQ+4gmgR5DjYucFyM2yFSix7YhaF38R+7b+Xu986NfDL+ucJXYJ8ZSa98j5n16IKHiIiIOLYp8FOKu1uvx8bCVtMFUqK1M3bhIqrpnpZicyFPROPLJT02A35DccG6B4Fdsc+I9/6X0EWgLLibwj4d6zgWERERaVEDsTGqSwlvSd7eXWgtaROi7Wa9AZiBVbLWdDmSZjthdRKCw0K8t/+CpqzLiv+lsF8/4TgWERERaTGbAN+jOEneAFwNbOcutJbTRqGatbcraaPjkM8Hhse4HSJxOJHw3ha/Qz0msuQLFPbttxzHIiIiIi2iL3AexVNAdQLXAzu6C63ljMQS2leJJkF+CytotEecGyESs8nY8JCwz8D16AJRVkygsF+vdRyLiIiIZFw3rOBNvmiKtxvjdOAD7kJrKb2wOalvIppiXWty6zoKK4gkklUDgBvwH/8vAE8H7luCzQ/eLXw1khKDKezTWY5jERERkYxqw8Z7vUBxknwbViBHmm8nrBDRIhpPkDuxMc2TsTHnIlk3BngW/+dgGnbxqQdwLsXFCWdinztJr4XYvlyFLn6IiIhIxPbFKsYGk62HgQMcxtUqBgJfBP5NNN2sX8HG7o2KcyNEHDsKay329qb4fMjzRgI34//MrMGGOnSPJVKJ2v0U9uU2jmMRERGRjNgd/7Qb+eVp4AiHcbWK/JRP7xNdN2tN9yStpjswBetJkf88zAM+VOHvPgHMx/85ehS1MqfR/1HYh4c7jkVERERSbhusEIr35HIjMBf4DOrG1kx9sDHhM4imFXkWNu5yaJwbIZIQmwF/xf+ZuAsYUuXfb4JVx/ZOLbUG+AZqZU6Tsynsv686jkVERERSqj/WArMa/8nle1gXxD7OIsu+sVgr8ns0niAvwcZhqtiatLI98Rci7MI+Y/Vc7Pso1hrt/Zz9C7Uyp8WHKey3yx3HIiIiIinTDZgEvI3/ZHAldnK5ibvQMq036eF94QAAIABJREFUhVZkb8tVPUtnbj2TsGm9RFrZJPwX/ZYBxza4zkHYRahgK7PGMiffSAr77CHHsYiIiEiKTASewZ94bcBOCjXPaHPsgF2EeJfGW5HfzK2rPc4NEEmoPsAV+D8jTwGjI3yNw4E3Aq/xCLBdhK8h0VuK7avFrgMRERGR5NsBK/gUTL7+jhX2kmj1IrpW5LWoWJdI0CjgMfyflWuBfk14rbBW5lVYfQBJpn9R2FfDHMciIiIiCTUUmAqsx39S+SJwpMO4smpb4CLgHRpvRX4OK1QzONYtEEm+Q/DPPb6WeBLXw7HeHd7P6U1UX0BM4vMHCvvoIMexiIiISML0AL6MdUHznti9hyVgPd2Flkn7YyfNHagVWaRZ2rAxwxsofGbmY3PDx2UoxfMyv4El8JIc36Cwf85wHIuIiIgkyIEUj0tej3Uj3NxhXFkzEJiMtQA32or8MpYEaP+IhBsE3Ib/c/MAsIWjeCYBKzyxdGG9eHo5ikf8jqSwb6Y6jkVEREQSYEvgTxQnYn8GxjiMK2vyBbuW0FiCvA61IotUY3fgVYoTU9c9ZLaneNz0LGBHl0EJYAXY8vtkhuNYRERExKFeWKukt5UjPy75Iw7jypJuWFI7ncYLdr2JzW+tojMilZ2MTWuX//wsx4rnJUVP4Ef4u4avBE53GZTQjcJ0YvMdxyIiIiKOHAo8jz8ZW4klY73dhZUZg7HCQa/TWIKcnxf5BDRHq0g1emA9OLyfo9nAOJdBlbEP/tbvjcB1wACXQbU473CkTRzHIiIiIjHaFrgd/4lZF3ZyNtJhXFkxHhvjnW+ZqHd5Czvh3zre8EVSbSTwMP7P0u0kP+EZTPFQmBdIboKfdd59sZfjWERERCQGfbBW42AS9zRwgLuwMqE3VrQnOAax1qUL+CtwNGpFFqnVAcACCp+nDdgwkzSN6z8dWENhG1YBpzqNqDX9D4V9cJrbUERERKTZDsLGIXsTsyVYN+EeDuNKu82xk/Hg/Km1Lsux1mi1IonUZzL+OeHfxWoFpNHOFA+RuQbo7zKoFnMChff+IsexiIiISJMMxZIwb2GpLuzES0Wi6rcDVlF3FY0lyS9hyfbgeMMXyYwBwI34P1ePA9u4DCoCA1G3bJd2ofC+3+E4FhEREYlYG9Yt+F38J1vPAPs6jCvN2oimqnW+YNdRpKt7qEjS7AA8i//zNY1szVc8Cf/QmRVYlW9prl5AB/aev+I4FhEREYnQLsBM/CeQK4Gvoy7X9eiDnbAGu0XWuizCCnalvcVLJAmOwj9f+Rrg804jap4PAnPxf59cgvu5orPuJQpj3/s4jkVEREQa1Bcr4LUO/0nVdJSg1WME9n6+R2NJ8ixsPGXfWKMXyabu2Oeyk8JnbB6WUGbZIOAm/N8tDwFbuAwq47yzRuzqOBYRERFpwMcobnl4E/iEy6BSajw2pttbLKjWZS12Yjsh5thFsmwzrFq897N2FzDEZVAxagO+gbV05rf/DbJ/ocCVn1J4nz/lOBYRERGpwzCKi8BsAH6JFYiR6vQATgQepbFW5AXAt7Eq2SISnfHAaxQ+a13YsIZuLoNy5CP4e7yswYaKSLQmUXiPp7gNRTSGTEREanUCcBn+xOwp4Azg304iSp+B2Lym5wBbN7CeJ4BfYa3J6yOIS5KnD/ZZG4KNFe0N9MO6BQ/KPacfhWJMS3P3rcAuYK0F3seK7m2ILepsmAT8lsJQhuXY3MO3O4vIrRnAHsBtWKtyH+BqrCfLWVhhKmnci57/7+QsChEREanJKKzrobdFcwXwZezEXSobjrUULKaxqtbTSe9crlLQB9gNOB74FnApcDPwIDAbS3wb6XEQVuztOeAfwPXYhZavAUcCY1AjSl4f4Ar8791TwGiXQSVIf+AG/O/P/ahnS1QGUJj54D+OYxEREZEKugFfwlpVvCdH96ACXtUag82PvIb6E53l2PQ0O8YcuzSuB9YidwaWEN+Hde31FotKwrIeS9JvB36GtazuSGtNNTYK6yHjfV+uxVrvpaANOA//OOZ5WLd1adwb2Hu6Fl3EEhERSaztgL/jP3FcglVZbqUT6HrtT+PzI88Fzqd1igllwUhsiqEpWNfVVbhPhBtZlmHTwl2Y2640tiAeXMVzjsDf62MtNlRCSjsI67GQf8/WoPmYo3Afhfd0jONYREREJKAHdpK4Ev9J83RgS4dxpUE3LKFotGjXLKxlT60Kydcf2+eXAXOIvrX3LeBp7JiYiSXg92Jj1W8Crsr9e3PusRnAv3LPfw5YSGMXbIJLF/Ak8BPgQJJ/jG4H3Fnm8TbsgpS3pX8+sE/zQ8uEUdix5j0+pqALqo34FYX382jHsYiIiIjHB7ATYe/J8VvAMS6DSoH+WIv7S9SfhKzDkh6dpCffWGy87wysBbKe/d2J9Ry4F+umfw7WKncIMI5oW3C7Y3N47wYcjl2IOQ/r2n8/VlG93uN2KXArVrRuZIQxR6EXlshdW+LxQVjBKu/2PIDmEa5Vf+DP+N/Hq7GCdFK7L1J4H893HIuIiIhgrUPnYwmbt4XgGmCow7iSbjjwIxor2vVebh0jYo5dajMSOJfii0nVLG9hSdk3gU8Cu5C8RGIQVuX4ZOx4vI/ai4xtwC4gnEahWrdLv8Di+lHIY7sDr+L/vpuKVR2X2rVhLcre4+FhbKpBqc2BFN7Dq9yGIiIiIuOAx/Gf5MxFFZfLGQtcTv2tihuxLrtnYa0ykkyDsOmCZuAvZlRuWQM8hCVqx2PdVNOqDZu+5lTg/7ALBdW+D6uxqslH4SYBPYxC9/PTA4+dgn8s+XJsWjxp3OfxX3Sdg6ZAqtVmFN6/xxzHIiIi0rK6Y63J3oSvE2tdUfXXcLtjre3VJgxhy5NoPHLSbYcVtFpC9Rc+pmEJ10AH8cZpU2w7p2Fje6t5fxZirY6bxRTjFtjc0vnX/2ju/h7YfvXGNhu7YCjROQR/b5v3gQ87jSh98sfvMjT+W0REJHajsbF53pPG17CTHCnWaGXrLqx18qi4A5eaHArcQeVpnTYAfwXOxBLrVtWGTRf0bfxFnsq1Nl9Oc5PTNmxOeO8+3AErTvhwIJ7bgU2aGEsr2w54kcJ73YF9XqQ6D1J477ZyHIuIiEjL6AZ8BTtp9SZyvyX7LWK1asOS20eovxV5HdYSrZar5GoDPo1VnK60P2cBX0Xjy0vZCfghNoyj0sWj+4AJTYjhvJDX+gjwtue+DqxXjVrsmmsoVkDOuz8uQu97NaZReM8+4jgWERGRltAO/AP/icsC4EiHMSVRfvqnJ6g/SV6GdWdXi0CyfYTK+3k+lgBq3GX12rBE+DfYeOBy7+8dRHcxaTw23Za3B8iK3H352++gLsFx6oGNd/fu82uxSuVS2lcovF9fdhyLiIhIprUBZ2Anjd4TliuBwQ7jSpre2Fjil6k/SV6Ajc3U+5psHwT+Rvl9+SQ2HVgfRzFmxUDsfZxN6fe6E5s2bXQDrzMAq25drp7A48A2DbyG1O+b+C9i/BX1ZirnMArv1W8cxyIiIpJZw7CWG+8J40LgWJdBJcxAbI7bt6g/SX4aS7Q17UyybQ5cR+mx5xuAW7Ax6hKt7sBxWKXwUp+jNdg0T/VcoLiW8jUFXgC+g1XCnoCNYe5W99ZIPT6Dv5X/P9h+kGKjKLxP/3QbioiISDYdjn+c3kas9SauirRJNwz4CbXPJetd/o6m2EqLU/BXSPYuXdg0R2OcRdda9gdmUvpz9SK1jWf+bJl15fdvWItzB1bJ/CZs7mtpvg9jw1Ty++A1YEenESVTG4UhDIscxyIiIpIpA4Ar8J8ULgZOdBlUgmwNXIq/yFktSyfwZ2CvuAOXuozE9lep/TkDG+sq8ZsIPEXpBHcalbvqbg+spHIF87DlIeAkbAiGxGc81sMpvx/eBfZxGlEyPU7hPdJFbhERkQjsRfGY27+jQlNgBc6mYl09602Sb0KVrdPkVEoXmHoKVZlNgu7AaZSes3kepVuZewL/prpEOf+clVgSvlszNkaqti3+ceyr0NR6QddQeH80NERERKQBPbDCUh0UflzXAOeiaTq2xU6Ove9NLcta7KRFXXTTowdwIeH7cw32WVE13mQZhF3MCkt881M9Bf0i5LmlkuR8wbb+zdwIqcnmwGP49/MXnEaULN+k8N6c7jgWERGR1NqW4qI5zwEfcBlUAuyMJbn1JsnLsZP3kXEHLg0ZCTxM+D6dCYx1F5pUYX9szHLY/rse6Jd73mElnuNNkPPznO8ZX/hSo/7AdPzd789zGlFyHEvhfbnYcSwiIiKpdDr+KaE6gf+ltcfg7YKdIJebQqbc8g7W8jgk5rilcRMoLmqX7+J5BuplkRZ9sVbjsOrWT2FjXsNqDuST5Jexlmh9htOhB/B7/Pvyx04jSoYdKLwf9ziORUREJFU2pbho0ZtYpdFWtTs2prjc9DHlltexKaT6IWl0GOEJ1DxsXmVJn48DSwgfGhG8bx1wNbCvk0ilUW3Az/Hv01/T2tN79aBwrM9zHIuIiEhq7APMxX9ScTOWQLeiPWgsSX4VG8uoOZLT62OEF267G7Uupt0Y4FlKf34XAV8FhroKUCJ1PsVd71v5u/k5Ct3TBziORUREJNG6Y92Dvd2Ll2FzjLaiA7Fpf+pJkPPdOY+jtVsusuB4YD3+fdsF/ADt26wYgF0QDH6Gl2A9SiRbzsJ/8fN2Wndokfe41xR3IiIiJWwFPID/RHEWNsdoq/kw8E/qT5L/jU1RovGr6XcsxQXcuoAvuQxKmmIQ8BbFn+fFwE4O45LmOAX/Z/t+Ks+5nUXfp/AefMZxLCIiIol0DPAe/mRgKq039c3+2JzR9SbJD6MkOUt2xnpWePdxJ/B5l0FJ0wzCetf8iuLP9lxgM3ehSZMchX94xeO03n7+NIXtV9GzmOlkQUQk2fpgFWHP9Nz3DnAqcK+TiNzYD7u6PrHOv38YuAibnkSyYVNsftbtPPd1Av+FFXlKiyGEzx+cdz02XrdRW1O+tf23WIG7tPgR8O3AfX8DjsCGqUh2fAQrZpmfH/vZ3H2LnEUUr92Bp3P//zPwSYexiIiIJMZYbEytt/Xk77TWnL/74p9/s9ZlJq1dHTyremJdMoP7+wsug6pTO+WP4T9E9Do/qPA6B0T0OnG6nOLt+KXTiKRZPoS/d9VsWue3sA+FOiWzHcciIiKSCF/A5oXNnxisBy6gdYoV7UNjSfKM3Dokm4LTy2wELnUaUf3aKX8sr6TxCrjdsFbjrCXLvYAHKd6WT7kMSppmd6w1uRUT5jnYNnfQuoXOREREGIB1u/Se+L0OTHAYU5wamSe5C0uwPxR71BKn/fBXg9+IJUxpHb/fTuVj+9QGX2NiFa+RxmQZrDt+PpHIL+8BW7gMSppmR/xF3l7Gil9mnffi8c4lnqPhtSIikmljKZ5L9FZaY47YXak/Se7ETiQ0pUb29QJeovhiUprnF28n/MKP9/Y/GnyN6yqsP83JMsAHKJ5j+0anEUkz7QjMp7CvXwO2dRpR8/2MwvYeF/L4cajFWUREMuw0/N2u1wCnuwwoJh/A5s+sN0m+gdJX2SV7vkFx0vdRpxE1rp3iY3sm/tbzLmB0nesfhP+7Jb/+LCXLAOdRvE0HOY1ImmkH/AnzHGAbpxE11+cobOt3A4+NAp6LPSIREZEY9AGm4T/BexlLIrNsHHANlvDW2916j9ijFpeGAEvwHwu/dRpRNNopPsZvwCo7e+/7Xp3rnxxYz7PAJSGvmfZkuTs2rZB3mx5HXVOzrB1rVc7v73n4q+OnUXfg4yH370NhO//oub8H8GhuERERyZQdgWfwn9zdBgx2GVSTbYtdHAiOOa12mQHsGXvUkgRT8B8L75ON+VbbCU+WPxO47zXqS/weCaznq2QzWQarnh/spXKE04ik2bbB5tjO7+83SH/C/APgWqxXSN4gCtv4lOf+H6JhByIikkGnACso/PitxT+XctaMwpLkDupPkj8Ye9SSFP2x5Nh7TJSbmzhN2glPlvtS3JJea7fiHfAnjx1Y4ausJstgdR682/Wg23AkBtvir/b+BrC9y4AatDk2dOJ1YG/P/Quw7VuNVbg/iELvrJ/FG6KIiEhz9Aam4j+Zm0d2pznaHLiQ4uI7tSTJqm4t3vF6G4HFwECnEUWnnfBkGYrnEa51zuWfBv7+9tz9WU6WP0Bx6/I4pxFJHLYGXqWwz98k3UW/8p/RDuCbWHLsHZqxB5Y854/1s9yEKSIiEp3tse5TrdDteijWbXY59SfJe8UdtCTWQ/iPjwvdhhOpdkonyxMC99cy53I3rIXN+/fH5h7LcrIMVj3cu22/dBuOxCSYMM8hvdNKbYO/J9Y/sYtl+duP4K/5cbSTKEVERCJyJP4ulR1YN9KsFZ/pj21XsPtotctM4OC4g5ZEG4G/pbALGOM0omi1UzpZBngx8NipVa73iMDfvUdhLuqsJ8sn49+2N8jed62EG4V/DPPL2HdIGnmLYHZiw7VK/Xbu5ihGERGRhrRhyaP3CvAbwH4ug2qCXljV3bepP0k+JPaoJQ1Ox3+s/NttOJFrp3yy/K3AY/dXud4bA3/nbV3NerLcl+LpsrI+w4AUbIN/DPNsYLjLgOq0C8UXCsP+vxGbLUBERCRVNgP+iv8H7W6si3JW9ATOwD/fZS3LQ2guVCnvBvzHzLfchhO5dsony1tSPOdypWq/QymuE7C75/GsJ8tg47O923eu23AkZmOBhRT2/5OkM6GcTuXZI1Y6i05ERKROe+LvCtaFjbPs5jKoCHUDTgBeob4k+T+5vxep5DX8x86+bsOJXDvlk2WA+wKPT6mwzjMDz38y8HgrJMvn4N8+Ta3TenbFhh94PwdpqxESrFsQtrzgLDoREZE6TMKmdsj/kC0DPuE0oui0YdvyPPUlyc8Dx6Hxg1KdwfiPn/VYF9ssaadysnxS4PHXKH/h7fHA878ceLwVkuV98G/fy27DEUc+gFXPzx8HD1N9kbykmEnp1uUNwJ3uQhMREaleb4qnenmayl0m02JfbM7SepLkediY5h6xRy1ptgf+42i223Caop3KyXIfiueZPrjE+sYFnrcOm8LNqxWS5YEUX2jp7jQicWUf/DMz/I10XXT7OKV/WzcAl7kLTUREpDrbALPw/4hdC/RzGVREdgZuor4keRFW4Kx37FFLFhyD/3i6x204TdFO5WQZ4DeB51xZYn2/CDzv1pDntEKyDPAu/m1M6zRC0rgD8Rd9u4tCdfg0eJrSrcvnO4xLRESkokOwpDD/w5WfFirttgamUbm4SNjyHvYepOnqvSTPJIovQGVNO9Uly8FuxSux1lOvHhRXpD8qZF2tkiwHp93a2W044thE/IXvbiU9vQ1OofTv7acdxiUiIlJSG3AB/mRyPumfFmpTrBhZsJpuNcuK3N9uEnvUkkVn4D++prkNpynaqS5ZBivk433eaYHHjw48vgirWB/UKslycOz2Xm7DkQQ4EuuSnz8mfu02nKp1x2oVeKehzC9ZK3ooIiIZ0Ae4Bv8P1oPACJdBNag/1hq8lNqT5FXAVGBY7FFLln0R/3F2udtwmqKd6pPl8wPPuz/w+G2Bx39RYj2tkiw/gX8bP+g2HEmIk/EnnVOcRlO9YJX7/LKly6BERESC2oGn8P9YXUx6i1f1As7CPydltcs67MR7i9ijllbwGfzH2/Vuw2mKdqpPlrfAhnnkn+edc3lTYG1gPbuVWE+rJMuz8W/jTm7DkQT5Mv5j42y34VSlD/AO/rg7SE9XchERaQEH4B+fvBb4nNOI6tdG/XMld2FFv7JS6VuSKVgF9m9uw2mKdqpPlsEKE3mfOyV3/1cC9z9eZh2tkiwHe8kMdxuOJMz38f+mpeG3/Nv4j+m33YYjIiJSMBn/WKf5pHcM3ESKuyhWu8zApvQRabbgNEhz3YbTFO3UliyfGHjua9icy8HeLmeWWUcrJMtD8W/fKjS/uxT7JYVjZD12gS7JNsF6dOVjrmY6vR7AkNyyLTAa2BUYn/v/aGwI1RCyMYOHiIjErDfwe/wnXjNJZ9fjccCd1JckPwYcGn/I0sL64x9b2AkMchpR9NqpLVnuRfGUSOcGbq8DNiuzjlZIlg/Bv33Pug1HEqob8CcKx8lqkv9ZuJZCa/jD2DCq7wO/BW7P3fcqVjG/nt/6/IWDt4AngbuBq4CfAV/FhsdMoPx3jIiItIiRwKP4f0Smka75GcHmF613GqjZWHdttcqIC8/jPx4/7DacyLVTW7IMVsE3eGLrvX1zhb9vhWT5m/i37yqn0UiS9cQSwvyxspTk9J4ajPUEOwu4FLgPeIP6k+Col8XYOdKV2GfuOKylWkREWsC+wAL8rTVfdBpR7QZgYxpXU/uP4Hys63laC5dJNgR7dVzkNpzItVN7svyhkL/xLh+r8PetkCz/A//2neE2HEm4/sAjFI6XBViX5Th1x3p/TcIubj9P+FRRaViWYkO2pmBzvW8a3dskIiJJ8Hn8lWUXAvs7jag2PbFE11uMrNrlfWyKmr6xRy1S7NP4j88X3YYTuXZqT5YBngn5u/x3VaULXFlPlodQ3NquYoRSyVCsu37+mJlN85O83YHzsIs7q6jtt7rapQP7XX8fq/swB/gPMCv3/znYucL7TYyhE3gOGyN+GFbZW0REUqg7Ng2U90v+cawbc1ocRX0VrtdhV7M3jz9kkZKCBW02Yi2rWdFOfcny10P+biPVtbxnPVkOzkf7nNtwJEW2BF6ncOzMJNoLx4OxYU2/x8YE15t8bsDGJD+I/W7/AOuqfTz2Wd6Rxuo79AFGAXsDR2INCN8BpmLDPJ6mvh5r+WUV1vX9bGBMA3GKiEiMBgJ34P9C/yPpqQ65D/AQ9V3xvQk7aRdJoun4j9nL3YYTqXbqS5aHY0V85gSWnav426wny0/j37bvuQ1HUmYnbCxu/vi5g8bmMu4HnIQV1wz2eKgmKX4GGxN8AfBJYBes8Oh43PYAawO2AT6CXaC6BGshX07t5yHPYj3aRsW6BSIiUrXR+AsJdQLfcBpR9cZiFTDrubr7F+zEQCTJjsV/3K7Biu9lQTv1JcuNyHKyfBjFFwO3dhqRpNGB+IdiTa3x77thQ7emUVvyuAC7ODgF6yU2pLHNcMI79noq1jpf7UWCztzzz0HVtkVEEmMC/rG9K7CT86TbDPsh6qD2JPlx4ODYIxapTw9gHv5j+H+dRhSddpQsR6U71i3Vu123OY1I0uwE/AW2vlbF3wwD/gd/cdByywrsYvcZxF9QLE6DsPOq3+Lv5l5uWYNNkzU+/nBFRCTv8/jHQ74J7Ok0osr6Yz/GK6g9SZ4DfApNAyXp0g0bj+c9lteRjbFu7ShZjspFFG/XBKcRSdp9jcKx1IXNMRxmV+B3WIJX6Xf4eWzO4kNJ3zSUUdkZmx9+BtVd8H8Q64LeSHd4ERGpQXfgQvxfxo9g4wCTqg270v06tSfJi7HxQKpAKWkxGqvofhPwHpb0BOcXne4suui0o2Q5Cv2x6WqC27Ueq/o7Ffv+1BQ2Uqtf4b9I553r/TAs4av0Gzwf+DlW/Vr8Nscuhj5KdRf8z0GzdYiINNUAbKyu9wv4TyT7y3dv/HNAVruowrWkxTAsmZkGvEbxsXwBcErI/Z91EWyE2lGyHIWwbQpbOrFpcy7DjqdtXAQrqdINuIXCMbQUOBm4n/LH2jLgD1gLcrfYo06n7bGCfJVm9HgT+C/U0iwiErlt8c+j2Im1uCbV1sA1WPevWpLkLqxFbnT8IYtUZRPgaKzFz1tcL2xZjfUEacOKv3gfW0K6u2O3o2S5UR/DP7Z0I/BP7DvwHSp/X+aLKp2PFWVq1a6xUlpfir97Si1zsWMpjcW5kqINmIh9Lsud/8zGLrJqaJmISAT2x3/itAI4xmlEpQ3AqmFWM/4puDwM7Bd7xCLl9cAKtZyPdVsMzp3sXTZgXWcvxE6YvMMHdqR4fs/Z2PylabQVxdM/XdLk1/xeyGumde7qHbELJsFkZYDnOaOxyrzTsAszlS4+rsQSowuxisRpPbYkOoOx6ZvKHTsaV9scu1B5PPijqBCYiEhDTsE/DcQ8kjl2qAc2TtNbnbva5UXsCqtIEnTDpg/JjzteRvnjdw6WzJxA5RaZs0P+/k7U1bHVDAFexn8cdFC5lXwYlgRfiCXF3t+GUhdvnseOz0mo63ar+STlq1vfS3ovNqXJ5lhhtODFUu9n/2cke0idiEjitGGVo71Xg5NayOtjwAvUniQvAL6ArmaLe8GiXOWO27dzz5tMffPgXh2yzp81Fr6kSC/grxQfA2fXsa6eWKvUOdgxuThkvWHfuzfl/mY8ulCTRcOxYVCljoF/4y/2JfHYErtwVaqK9hy0X0REqtKL4hPqm0neVcexWKtYrUnyKqxlZFD8IYsAlYtyeZcVWPfr87HkotExZn0Ir556YYPrleTrBfyZ4n1/ZYSvEey6Xen7eDl2fE/Bhg4k7XdGanMK8D7h+/olNE42CXbC5lEP20ddwG/Q51BEpKQhFFeqnEqyrv5visVUzRyDwR+Bm7DCQCJx6o8lAhdiY4rLjd9bjT956NmEeLbAqqIGX/vX6EQ2q3oDd1C8z2fmHmuWEfi7bpcbc78R+173TlmlGQnSoQfF00rml/W5x5p5nEntDqP0lJpPo0KnIiJFRmPjd70nLWc4jcivN9a6VmkMZ9jyEBobJfGJqihXM32Q8Pl1lTBnT3/gHxTv69lYL4e4Y9kf+2xMp7jIWNgyB+vWOxkbz6/jM1mGEX58bQSeBPZ0F5pU0A/77dlA8b5bDBzuLjQRkWTZF39xrPexOQ6T4ijgVWpPkt/AugTq5EqaqZlFuZppPOHjTG8GBjqMS6KzDfAExfv4WayHgWvdKXx2rqF0S5d3WYh/yiq1WLqzP+FFvFYDX0U1QdJiAnbxLLgfO4Fvo3MoEWlxJ+CvkjgXG9OSBGOBe6h9s3nuAAAgAElEQVQ9SV6BdWONq5VOWk+cRbmaaWfCT3ZfwpIYSa+DCZ8h4CmS3b15JPa7NBXrdRGcCzq4rMI/ZdXQ+ENuSR8nfFqieVjPFUmXAcCNhH/GriBZw/FERGJzDv4TkX+RjIrXQ6lvXHIn1jqRhG2QbHFZlKvZxgLzKd6O5cDxDuOS+rRhx15Y18onsLoPaTIQG6IwBftclZoCJ9h7w9t1W6J1AjYWOfi+340uVqTdZML37R+xIUYiIi2hF3AV/i/CW7DxKy7l50t+l9pbk2cAu8YfsmRU0opyNVs74d11u4BLsFYHSb6tgLsIP05vIRvd6/M1AfJTVr1D5d+HBfi7bqfxM5oUp1B8IbsL+AFqfcyKQwn/XN2APjsi0gIGUNy1OQkVrydi4+hqTZJfBI50EK9kSxqKcjVbH+B3hG/z61j1VEmmNqw+Q9i0PRuwCzmuv+ObKThlVbmLW/neH96u24PjDzmVJlHcLX4D8HmXQUlTjMHqvgQ/O7eisegikmEjsPFq+S+9DuC/nUYEO1LffMmLsGrd+tKWeqS1KFccJlP6YsFNwGbuQpMQo4G/Eb6/3gM+6i40Z4bjn7JqLeU/3xuwJHsalhBuE3/Iibcfxe/jBuz9kmzahvDiqj+v9IdJH3skIhJmF2w80ajc7RXYeMS/OopnAPB14AJqq2baAfwG+B8swRGp1misJXgi1s2s3NjNhdiUY38D7sWusLeSfbExn9uHPPYu8EMssVgfZ1DiMxTrCXE20Dfk8QexRGZenEElVD9sCqMJWDfs/ag8tvZtLNF+OPfvU1iLdSvaGvg3/nogHcBJWEtjWmxP+Zk+riKa77SdgAPKPH4FlnSmwSjg71hLs9cXgN/HH46ISHMcjH8+ywW4m/uwG3YCt5DyV/rDlhmoWItUL8tFueLQl9JzcG7EkrDJqHdH3Pphx2lYl+uNWHXo88l2t+soBLtuV/r9WY6/NkHYBYos6kdxPYMu0ln87xTK7+NPRvQ6N1d4nbR9Z44E3sS/DWuAvVwGJSISlePwT+/wAu66mB2Evxt4tcsLaLykVNZqRbni8kHgGUq/l89gdQN0gaG5emNDT96i9L64i0LvIanNCPxdt8MqAnuXDux7Zip2US7J03E14lqKt/17TiOqX6Vk+S8RvMZQKnf7T1uyDHYROViJ/i2ye9yLSIsITg31KG7GG47EunRWKroSXN7HWkh6xR+ypICKcsWnJ/AtrHWt3EWtybROi1tchmHDTsr1xnkTSwQkOgOwLtvnY5W0vb2zSi3BKavSfgHpWIq38TbSu12VkuUOYIsGX+OsCq+R1mQZrPEleB53o9OIRETq1B24lOIfuLhPYntiCXu5E+ywJT9f8rCY45VkU1Eu9zbFLjh4e6sEl6VYa5taOBuzA/Y+rqLyBUVdoGi+7hS+f67BKsRX+i1biH/Kqlrqc9RiR2wcdpSGUjx90H9I9zRylZLljcC5Db7GrCpeI63JMsBPKd6eqLqvi4jEojd2pc/7ReZiaqiJWEtTLUnyRuAfwG4xxyrJNZpCcvwe5Y+dt3PPm4wVpJHm2Ra4juJpZLzLOuwi3Scp35I/AuslIHZR53TgAcr3xFmFXbTQRSC3RmIX46ZiSVK5z0N+v3mnrKpUZKwW92FjZbeLaH3BC+5rgV0jWrcrYcly8DzlPw2sf9fAutZgF22zlCz3wIq9ebfnDWxsu4hI4g3FKqDmv8C6sDGZcdoeu5Jea5L8BpqCQlSUK212B/5I5fGdS7A5nA/Gf+FuHPbZv5bW3X99sO6Nt1F5rOMy4GJgSyeRSiUDsQvFU7DvpuAYz+DSiRUX83bdrtf2WHK2HjtGGknEd8S6JHtj/XYD60uKsGT5uxTvp3oLoF4cWM8NhLc0pzlZBptdJfidf4HTiEREqtAOzMZ/FfjEGF+/H3aCUOlkL7iszP2dxpC2JhXlyoatsH24mMqf+Tex6d++hXXZzt9/XuxRuzMM+Cx2oaHasbBfAQa5CFbqlq+rcA7W6yXYrTlsWYC/63Yt33Pf8axnWW4d9fy2Bot6zaF5XcjjFJYsnwP8KXDfJXWsuwfWu8m7niPIZrIM8Cv82/Qe6e6iLyIZNw5/Wf/3scrTcWgDTgbmU/kkIHhF/UqsC6a0DhXlyrb+wH8DL1Lb98EibB9nVXds7uofAI9TubtufnkA68aehZNrMcEpqyoVvlyBv+v24DLr7kXhonl+va9jF86r7bkxmuJW5ROq/NukK5UsHxa4bzG1Xxw4JrCOt7DPbVaT5SEUX+hrdLy3iEhTHIj/C2sesHNMr/0B/N2+q12eIPpiJJJMKsrVusZhJ/jB1pZyy1L8iYGL6v1RGInFPwVrJSw1J3LY8jq2/WNijlncGI5/yqpKvbM2YEn2NCzp3iawvgPwJ+D5CzOzsPOFSi4MvN4LZGfO7lLJcjdsOIj3/uNqXPftgb//ae7+rCbLAD+k+Pe7VYfTiEhCHY1/rM3zxFOBdghW0GQD5X/Ug8ti7IcpKz8UEk5FucSrB9Yd8Voqj20OLl1YS9n1WBfTE7GLdEkpJjME2Bs4FfgxNu641l42G7GqyVOBveINXxKoH/4pq6oZ2rAA+x49B+u5cyXFLdb53+s7sfHNYXrk1uX9u89FvH0ulUqWAX4SuP+OGtY7jOLvtp1yj2U5Wd6c4os7hzqNSETE41T8XaUeo/mtMN2BM6icAAWXDuxEsFz3MUkvFeWSSnph43O9x0I1Y3XLJdHzsGPpcqx781nA8VjL2lhgkwZjHoa1jh+KDTX5CtZadCXwENWNPy21dGLdsX+I9bLJysmzRC/fO8fbdbvS8bUCS97CunhvwH6Tp1E8PePEwHOXk61xqOWS5R3wv1+1zLn81cA6H/E8luVkGYrHe09zG46IiDkf/5f6dJrf0rIn8Cjlf6DDln+S/ukmxE9FuaQWA7FpbbyJ7pTcY9sBZ2KtXZW66Ne7rMC6Qb+FdROcjR23+eWV3P2Lcs+rVMG43mUhdsHgs2gOeWnMCPxdt2vtsZH/HC4Dvk5hfO7UwHOujGVr4lMuWQZ4OPDY16pc79OBv/ui57GsJ8tH4N+2t9BFcBFxqA34Bf4vpmtobgIymPq6XM/HroTrSzP9VJRL6jUCeIrC8bEOO2EtJV/8aCqWBJQ71tKyzMRaBvVdKM0yAH/X7VXUdowuAD5N8ZzDx8a5ETGolCxPDjz2XBXr3DPwN2vw96LLerLcG+uB4N2+XZxGJCItqxfF3V2m0rwTsDbspHUR1f3Yek+Gp2KtSZJOKsolUdgZ6yqdP05WAIfXuI7+WLfqL2JzmN4NvErtF++atawFngVuxcY8noadPB+EP9E/u8btFqnHtsA92DFXbcX1UssGsvc7XilZHkTxhYYPVljnpYHnXx94POvJMljPIO/2fcFtOCLSivpT+AHciHWf+kYTX283rDWk1h/X6VjLkKSPinJJlPbFfxy9jSWRUemFJePHYN24v4/N3Xw71pXyVWwO90aShSXYFFgPYsf7pcB3gdOxpH805U96v+RZVwdwSATbLRKmJ3AB1qoZ1YWkl2LdgnhUSpYBrgs8fmmZ9fUC3g08/yOB57RCsuyd33sj8Fu34YhIqxmKf6xwB/D5Jr1Wf2wsYa1dH18GPt6kmKQ5VJRLmuUT+Mf9vkLp6rtxGID1eBiJJbg7Ysdxftk+d/+w3PP6Rvjav6XwPizGxmiLRGkCxd2ngy3EHbl/a2lt7sQqvPePb1Oarppk+SOBx8vNuXx84LnzKU6EWyFZPhr/9s1wG46ItJJtsEQ0/wW0EjisSa91EsVTRlRalgPnYVdXJdlqLcqVn+tWRbmkFmfjPyH/Fza9SKvqiRU5zL8fz5Ct5EPcGYpVgvd+ly/HCsm9CjwJ/B2bAulP2IXRn2MXxL+B9Qw6CSsUdijwe/y/Az+MbUviU02y3A3/8JGNWFIcJtj9+Echz2mFZHlX/Nv3ittwRKRVjAXewH91c78mvM4Y4F6Kv8wrLdNRF9wkU1EuiVMbdhLuPa5uJznzIbs0HP93+W2od4Y0rh3rETGcaKZ3ugz/5/fMCNaZNNUky2AXCoLnO0HD8U/fuRE7bwtqhWR5M4pb40VEmmo8/jk85xH+JdyIftjJbXBC+UrLK9RepEeaT0W5xJVeWFEb7/H1e+yCjZgP4C8c9B234YgUuQr/Z/hzTqNpjmqT5W0pnnN5ROA55wXW81CJ12yFZLkv/u1b6zYcEcm6A4Gl+JOaqItmHUXlcarBZTWWXJcauyPxU1EucW0AxcUHp7gMKMGOp3AC3kXprp0iLlyD/zfjVLfhNEW1yTJY8ut93tcDjwfHiZeqAN0KyXJv/Nu33m04IpJlH8dfGOdZiq9mNmI0cBflk6qwZTp2pVXcUlEuSZIR2LhIb+vL6U4jSr6LKLxfy9F8pJIc3mJ0G4H//n/snXe8HFX5/9/3phMgjZAQSiB0AlIC0iJFUbqAFKkRUREQidhQkS9FhaBYUERDbwKGIghKR5qAQOi9hpICJKSQ3u7vj2fnN3V3Z7ad3dnP+/U6r3t3d8ozO2fnnOc8za04dSGLsvyNyHbBmsvbRj6bT7i2cpB2UJYHEr6+mW7FEULklUOx1TjvYfM49gCqBT0xpWkBxZWrpPY+sn64REm5RLOyMTAJv//NBfZ0KVCL0Ek4KdDbWLyfEK4JLuR0YWXS8kYWZXlFbNE5uO02hc+iCwtXlThnOyjLGxC+vklOpRFC5JLjCGeQvZfaJOwAy3L5CsWVrKS2GDi/hjKIdCgpl2gFtiNcW3Qq1m9FOlYGXsL//u5G8d3CPd8hPMZc6lacupBFWYa4a/oF2Fg7M/L+50scox2U5d0JX9+DbsURQuSNUwhbDP9BbeKChxF/0Kdp92NWI1F/lJRLtBr7Ew4VeRPLqC+ysSHh3BS/cSuOEOxFeLx53K04dSGrsvz5yLYzsMRnUStqZ4ljtIOyfDLh67vCqTRCiNzQQdzt6UqqtzD0wB7+c0hWuIq1ycCYKs8typMlKdc0lJRLNA/fJFwq5X+0dw3latkd8xDxvs9j3Ioj2pzVCI8/C8lfQs+synIn8fwgn0Ren1XmnO2gLE8gfH0nuRVHCJEHumEWwuDD5XyqT8K0M5aEIouSvKRw7pWqPLdIRkm5RKuTVEP5n6iGci04Ff87XQB81q04os15l/DvfBen0tSerMoywJkJ+3htObBumf3zrix3Ygv7wevbzqlEQoiWpwdwPeGH7U+qPOZQzOW6VAKopPYgysZaa5SUS+SJ7sDFhPvtZSjGtlZ0ANfhf7dTgNWdSiTamSsI/9Z/51Sa2lOJsrw2xcfxB1KcM+/K8naEr20WmssIIaqgJxaTHFSUyz2oS9GJuegGY9/StKmYy7Usl9WjpFwir6wI/JtwHx7nVKJ80ofwhPpR8uf+KlqDAwn/3qeQr4WxSpRlMKU4aUz/eop9864s/5HwtV3nVhwhRCvTF8t66j1QFgOHVXG8rYEnKK6YJbVlmAV6UBXnFeG4YyXlEnlkKDCR8ELPsU4lyjfDgQ/xv+8rnEoj2pW+xMsl7etUotpSqbJ8dMJ+c0kXvpZnZbkP8Rjug51KJIRoWVYE7sN/mCwCvlLhsQZhJR2yulw/BmxZ8RW0N0rKJdqJdYE3CE8K93YqUXswmrBnynfciiPalEsIj2n3uxWnplSqLHfDFruDbeWU58yzshwtNzYdecUIISqgP6aoeg+TeVgW1Eo4mLD1IU37BBsMSpU2EGGUlEu0K9sCHxGe/OzgVKL2Ijj5XALs6lYc0YZsQ3yc29GpRLWjUmW5GvKqLPfEymYFr+tclwIJIVqTVYFnCVtoShWvL8a6wF2UVtqibTnmcq3SLuVRUi4h4MvYYl4whGADpxK1J8FKCdMxzxYhGsk9xD3T8rAQLGW5dvyA8DUtAIY5lUgI0XIMJVzGaSawfcZj9MAslgsorrwltWeQNagUSsolRJhjCNdQfgJb7BONpwfhhELPYgt6QjSKnYiPhUc6lag2SFmuDUOwOW3wmv7oVCIhRMsxnHDM34fAFhmPMZrsNZNnYQ/+Vn8Q1wMl5RIiTlIN5btR3XXXDAHew78nN5EPy55oHYKVOzwvh6FOJaoeKcu14UbC1zMTWMWpREKIlmJDwpOcqWSrZdwfOB/LXJ1FUZ6ATbCEoaRcQpSmO3AR4d/C5Si0oFnYkrBb/KluxRFtxnrAQsLPh9tp7fwnUpar52vEr+dkpxIJIVqKjYHJ+A+Qd7EBJy2HY4pbFiX5Jcxlqt1RUi4h0tMX+Bfh38U49FtoNoKT+2Xkq4yPaH5OIT5+/sqpRNUhZbk6tiK8gNcFPE7rXo8QosFsTdiC+TrprZQjgDspreBF2wLMfbJd0/QrKZcQlTGE8ARuKXCcU4lEKX6Df6/mACPdiiPaiG7Ao4TH0+XAV10KVQU9iZeAqvccauWEc7Yiwwgbg7y51cYuhRJCtA6jgBn4D5CXSZcVsDu2qjmXbIryA8BGtbyAFkBJuYSonhHYQp73W5kH7ONUIlGOTsJeAG8Dg5xKJNqJ1YEphMfYeZiVUbQHfYD/EV80OdSlUEKI1mFHwgmjniZdooMdgBfIpiTPwOJq28VVstKkXANdCCtEk7MN4TrtM8hP/dS8szK2COvdu7uR66NoHKMwK2J0PrK1S6FEQ1gBe97kyR1fCNFARmNucd7D4ynKK2r9yJ7Aq11qJisplxD14UuEn1VvY8kIReuwIVbxwLuHv3YrjmgzjiI+Ds8EtnUplKgrfYH7iN/3O9BinRAiBTthiaK8h8d/sdX/UuwLvE96JbkLK0G1W+3FbwqUlEuI+nM04RrKT6LM+a3KHliYiXcvv+5WHNFmnEt8bJ6FecqJfNEPm9dG7/ezlJ/rCiEEOxNWlB+h9MNjHeDfZFOSF2Mxt3lK4KWkXEI0lmg223vQRKfV+Tn+/VyAudcL0SiCCeeCi9kHuBRK1JThWEhh9D5PRPkShBAp2J1w7M5DwEpFtvUSeAUV6zTtIfKRYVBJuYRwQzfgr4R/Y1eiRac80AFcj39fJ5MuoaQQtSKppNRybPyWe25rswvh3BZZwgyFEII9sZV87+HxILBikW23wtwdsyjJn9D6CbyUlEsIt/QFbiP8Wzuf1n6uiDB9CJf/+i/58kISzU+SwtwF/AcLsRKtRQdm3AmG7KT1nhRCCAD2BhbiPzzuwiYsUbwEXsG4sjRtAq2ZwEtJuYRoHgYRjjNbChzvVCJRL4YDH+Hf68vdiiPakONI9hh7BwtXE63BUOAWkudtN2ELsEIIUZIDsRhi7+FxB8luwvsC75FeQe4C3sQy1bYKSsolRHOyDvAa/u9vIXCQU4lEvRlNWFk5wa04og3ZGniXZLfsq5DXWLNzMPAxyaFxZ2B13oUQoiQHE1aU/0VcUV4duJFsSrKXwKvZY3OVlEuI5iephvJopxKJRnEi4XFlV7fiiDZkMMklhrqAKSj5VzMyjOLW5Om0lhFHCOGQwwm7U98M9Ax87iXwCtYvTdMeBkY25Aqyo6RcQrQWXyT8DHoH2MipRKLRjCc80R3hVhzRhvQAfgcso3io2frOpBMeKwA/pXji2cewEI+q6MAmkPVif2BeHY8vhEjHIcDfMOURzHJ8OJb8AMz16CJgywzH/ARTQi/FHkrNwghM2d0Ny/ZdKpHD28C9hXYfdk1CCDd8DbgY34vjBSwR4WRnEgkX9MCeyTsVXj8H7Ijmk6Lx7ABcQnJFjyVYbP2ZmMVZNI4eWF3200nOnr8Auy/nYQseVZPFgpS19auFgEKIqjiMsEX5OnyluQ9mSa0kgVezZIhUUi4hWp9TCIdF3IsylrYzQ4D38fvDTShPhHBDb+BXhEPYgm1u4XPpPPWnEzgUeIPi87z7qIM3ipRlIfLLQYRT50/AV5Q/B7xKtt/0W5i11iVKyiVEfugG/IXw7/YqlCNAmKfTPPx+8TO34og2ZwvgCYrPN2ZjrtvruBIwx/TFKiGUmrNOB75JneZ5UpaFyCdRRfkGTFHujymapZJbRdsSrISUi5T7SsolRD5ZAfgn4d/w+ShjqfA5Er9vLMOqNAjhig5ssf51is9DlmG14XdzJGOeGIplsS7lNTgPm/P1r5cQHYUTBZkPnFyj41+BuS0IIRrLgcD1hGOUD8Pi/y4E1shwrEeBbwMv1lLAEnQHNsePO96JcCKyIMuAZ/Hjjh/BSswIIZqbgZiivGPh9TIsweCfnUkkmpXzgB8U/v8U2B54yZ04QtADs2KeBqxWYrunMF3o75jCJ8rTA/NgPBL4CsWNHkuwnDlnAVPrLVRUQ1eCGyFam2gd5RuBNTHLchbPkFnY5LURVp5g3PHsMnK9hVnGD0Y1D4VoRdYh7E63EEtCKEQSnViZQ6+/vEYdrUhCZKAvlo15KqXnLYsxa/OhWK4YEaYDWwS7gOQ6ycG2CAvVaWg2cinLQuSHrxBWlG/CrMIzKf3wibbrMfeXeqGkXEK0J58BPiA85/icU4lEKzCAsOvr3Vi8uxDNQC8sm/8zlJ9fzQGuBr5Key/49wB2xlyo36L89zYdS6SWlP267khZFiIfRBXlOzHX5CxK8pTCcWqNknIJIb5A2HPkA0x5FiING2EeT17/GedWHCESGY0t8KepMrIUc9UeV9gv7/kahgBjsO8nrRHndczLcQUH8gLJMcszae+VjiB9sNpqA7F4qg+Bd6lvrb/emMvsYKxjfIq5d0zFr4lba0YAq2BlOrw+MaNwzml1OmezsCbmEtgT+37fx1fm0tIPi1kZjK0uTsfcSBpZG3RvzIrcq/D6eWBd0ifk6gKuAb5HbRbM+mIuNV7c8VYUV3qXYnU0vbjjB6lfXxdCuGEMVq/Uiz97Ecuh8IEziUQr8mXgH/hKxRHAte7EEaIow4HDsT46MuU+H2N5Yp4AHscU6Tl1ka7+dAc2BbYNtE1S7vsJplD/Dfgv2ebkNceVsvwD7EsL8gCWeCgr2xNPSPYp8B2SE/0MBP4aee8SzKUHzK3nEMzlczR+giSPpdiE/hqsXu3yCmSO0h84BtgLc0dLSmY0F7O23YZ1nmoSp62AZUo+HNiO0lnLF2CKzP+AO4D7Ka/IXEfYPepmzK03K0dj34nHPKwIeTkuJ6wk3o3dY4/NgROxrJpDEvYfRHmFcRTmQrMHsFmRbd7HrLtXYA+/erEX9h17ivJssmWifwvr7/dXIYOScgkhijEW+D3+gtn9mAfLbGcSiVbmNCypD9jYsRPwpDtxhCjLFljCqkOB1TPstxx4BZuDP4PF678OvIdjBTLCQGADYEN8BXkU2azBC/F1nDtosgTRLtywhxfOE3VF2DXjcQZilt7oNRxWYp/VE7Y/vvDZuliHTOuy+gjWMSqlGzaJKBezGW1vAAdUeM69gUkZzxdsT6U4R7Rw+y8qlPX3keOk7ZvRvuVlV+1DupJJg0oce31stStL2aUuTJkdnlL+LHwZS3bgnWdZBpmqLQelpFxCiFJ0w56/wWfBDZgHlRCV0oEtwHt9ajKO4hiFSMnK2AJPL/w43WepfC4+v7D/BOCXwLew+eD2wNrUNolYN8x78jOYgWgMFiZ3CfAw5RNylWrTgCuxRYSmLjccFbxRMcv7E1c4pmCxjWnoIF6fsQu4qMx+xZTljTE366w3eiawZUqZg/TGXIkq7WBd2A8kSzzncWRTppLayynO04zKcl98V45ybZUix90bs/BX+t1NpbK+UowvYZb/SmR5Ftg64/mUlEsIkZZe2DMg+FxQDWVRK/pgi/de3/ovvneVEM1Ef+AxrJ/eRNhjdXXgG9giYtZErOXap5gl+ins93EPZrGdUGjjC+2awut/Frb5T2GfFzC9KKtxqFRbihkaT6V0eF7TEb2QRib4+kPC+e8i3WB6csK+z1N+NSVJWf4x8UxsH2Opyc/CCmJfSjiDZ3TbLBbmHpgbWtKxlgEPYa7i44DLKL369MeU59yOZEV5CdZxrwZ+h6XAP7Pw/23ErdCtqizfRPzaP8QeYPdg37FnHU1asDka+66S7sG7mNvI77F7dgnWF5O2nUP6mI1S7EZlivJ8rD8Xq1sXREm5hBBJlPK+Actc/DD+82E58KN6CyXajuHAR/j97HK34ggRYzDhOfwciscvdyt8NgZbWHyK6g1crttUTJc4Awt9bFnPwuiFNVJZ7okFsUdl+FmZ/T5L2PW0C7P4bZzinEnKcvBh+ylmaU5SJjoxV4Ekl4NHSb9i/ouE/bswt6I1i+yzBRbsn7TfgSnOGZy4BJXINJb8dYCTsB98KyrLUVf9CZhVNarQ9cRiuVeMvJ/U37qAfxOPvQ+yObZCF90vzaJOKXakMkX5IUov6vTFlPBx2EO61GriEvwMjruRTvkWQrQ+m2Hl6IqxNhZj5z0rFmLjphD1YDTh8fn40psL0TCGYtZZr2/OxNyks7Ay8HnMEHEZZuCZQfb5X73bQmxueyNwNmZoyZVXoUtlGcy1M+p6sITidRf7A28Tl/trKc+XpCx7bQ7mFlCO9Uh22f5Oin23Jjmd/E9T7NuNuFtbF+YSW8x1GGAN4orPeSnOl8QWKbZpNmU52K9KxbMnsTLx/rYcS1KXxnraHXvARWX5v4xyeOxF/Pst12Zh7tBRebtjVuBTMKtw0oKA14LlDXZDMYdCtCOdmDvfmCKfb4YlNgxODndqjGiijfkufp9bDOziVBohTFF8A79fzgC2qeHxV8EMJ8dgtYcvB27H5mnvU3o+V0n7EKtgcD/mSfkHLOfS7phBLffhNcUmxdW2YspuEgcmyPE+yQpgkjvtFRnOVUpZTmOh9fgccQX0XcJZoJMIJqXw2mUZztuLcJyO10pZ4/dO2H5ohnNmpVmV5RMqkOEnCcc5M+MxehCPl/6Q7NblH5E9duQ2bLHEQ0m5hBCVciL2bNg/4bPPE66BOxnzrhGiEVyE3/emY2OdEC5Yh8GDX0oAACAASURBVHBo5zSKV02pJwMxj9tRwA6YoWN3bE53MDYXPBYrbXUw5ia9G7bYNAp7fq9GvCpQW1LLlYdg2zejHBckHON2wtawExO2eZls2XyLKcv3ZJQXLCg+epy9S2w/jLgiOZ3y8V9RtiWuNL1H8Q799ci29awTDc2pLD9K9jjantiEL9rfKnlwbJcg01Ep910bkz/L728q9vBTUi4hRC0YhoUpdRG33B1J2JLxIsVDioSoBz2wUCOvDz5L5ZUehKiUjQjnN3oXq6IiWpxmUZZ7ARMTjuMlBdkK84kPfjaf7Ks1xZTl/TIeByyWNXqcq0psf3zC9r+p4LxgdamjxxpdZNsjItstp/0sy0dUcP6vJBzn6AqO4/FM5FjlPAo6McV1XoIcxdpy4EEsFX9SuEKwKSmXECItt+I/O4LhSmMJJ6H5D01eAkTklqGEwwBuQuOaaBxbEM6B9A7ycMgNzaIsg8UCR11DF2Mlct4gfo5vVXCOJGV5NmZFrIRoFu3XS2x7ZcK508QAJ/GthGMVyza6Q8K2V1K/GINmU5aXUtkK8/mR4yzGsrxWym9J31c2pXhCt2JtIUrKJYSoPdFQqfUwJeQ3kfdvRPkMhFu2IrzAnCYfjBDVMoqw996rhEPgRAuT5E46D3PbrZaJFezzJmZJuz7wXg+sLlhUsbseuLgy0WI8gylClfA44ZWj9bA4gSTFLpoFby6WPa4SHklxfI8nC/IE407HYFb5c4BbMEUqr7xCZa7n0bj7N7CENZXyZuT1eliM+7LAez2A72Nx0VlrRka3X4a5ot1baI9gCrUQQqRlZSxMajn+ODwfuA74amC7P2IlHZc3VDohwjyNZWu/uvD6l1hG4tudSSTyzmjgX9izEixcbzcsHE7kgCRleTFWHNsVfwd2JVyaIqoov0Hp0hVZqVRh9fY9PPC6A1Oek5TlaBzoC1Q+sXgdKx8UTBI1vMi2S7C6yb+MvL8lFqP6KeY6dzfmwvtyFXI1I1ElNS2bRl4vwxZzKmXLyOsOzFI9PfDeSOw+VZpQ4W185fg+Gp/dXgiRL34NDCHszno9/mKiZ707t8FyCVGMa7Dx9vvY/PFazJjwkkuhRC7ZBUuk6pUcnQjsQXheJ3JA1FWzGSbXvQkX8Y66mqYp71SMJDfs06o4XjR5Vhe2ohRlhYTt/lHFeQEmRY73doltuwN3JciQ1GZhP/7vUVmSlmZzw76ygnOvRLrvqtq2QcK5/5hh/8mF6xuD9W0hhKgV2xEP7Qi+XkT2cnxCNIJuwL/x++prWOlRIWrFXpjRyutjj+Bbl0XOaEZlGax2WJJycFaVx01Slr9XxfGSyl4dnLDdsITtKlHigjwXOV45F+GeWNxsljq9y7AkUDtnkKvZlOULKzj3cNJ/R9W0zySce1SZfWZidbKVlEsIUS96YXF3S0l+Ds3D6ttuiJUXUeZh0WwMIJzv5i7Kl/cUIg1fJpx0+AHMyCJySLMWkR6GxdImcRS1Xx3sqvG+SQpM0nvVnDdp/3KK02LgB1jdtd+SLp6iE7OUP4Bl+m7FCVEl37PLJDXlXL0XYC79o7Gi9Fljm4UQohw/wRThYsrFCpgXzKvAFCwHx1JgBpYnY1+0mCfcMhPrh7MLr79EPBxNiKwchmVa9+ZedwB7YiGNIqc0m2W5G3A/pS1rN1Vx/CTL8s+rON7RCcf7YsJ2fRO2u7mK84KlpQ8e752M+3dgls2xWBbTYMr7Yu1eymdRrpVl+Q+R41RqWf5zBeceTPzax2Mr1bVsSRPRixLOXarNx9x/zgMOQvVNhRDVsRHZPJC89jFWlUEWFtFM7Idf3mw5Ch0QlfMtwqXy/okMFm1BsynLZ5BuUP5OhcdPUpZ/X4W8P0o43jZFtl0U2e7hKs7bia3kB4/3dBXHA1Oet8Su6X6KlyE6ucxxopOsSldyL44cp5HKspelOnicWys4TiWsQLivLCH5PpRqU7C48zMwz4AVGiS7EKK16cASPkaff0nN2+YDbNFVzxnRrPwffr9dQPF5mhDFOIHwvPg6Kk/GKlqMZlKWdyUeHzUdC6KfH3l/IfHswmlIUpbvrULmpNrJqxbZNlqTeRaVu6mtn3DeWitzG2KTpuh5JpXZL1or+3cVnv8fVNY3a6Esg00Ag8d5q8LjVMKfA+ddBpxNPKGb14dmJbwfbUuxTKBXYW7eI2neMAwhhDuOJb2S/C6mJKu2smh2OrBqK14ffpficzUhopxC+Bl4MZpDtRXNoiyvilnDgrIsx+JNIHkAf53s7l5JyvIMKl8deiVyrFIZqf+WcO6NKzzvUQnH+mmFxypFL+DFhHOtV2Kf9yLbXlbhuSdRWd+slbJ8DfHr3qjCY2VlVcIW5Qcw9/ejsayeUbnewEq3jMNcshckbBNtswvbjsN+Z6s04LqEEM3LUOKLndFFty5s3BuDrCqitViRcGLU/2KJT4UoRVRRvhApym1HMyjLncCdCbJELZLXJ2zzt4znSlKWu4DdK5D7MwnHua7E9t9N2P6MCs4LllAgeqxdKjxWOb6f8VzRsl+PVXDODRLO2WhlOSkj+68qPFYlXBE5996F9zsx5XZignwvYtnYe2DW42Mxa/JLFHerD7a3CtuPxRKIaSIhRGvRF1gX2AELwdgPeyYcgT0Pvo1N/k7BKkEcC3yzsM0TlFaSX8CUZGUUFq3KcML5WSpdzBf5pwNLhht8FqqefJvSDMryTxPkeIL4RH1l4M2Ebb+R4VzFlOVKah7/NeE4B5bYfjhxN/P3yR7nNTLhONOon2LzVeLX+YUS218d2XYB0C/jOS9IOGejleVBWHbD4LHm0DjXrQ0IK7hvE04k0QEcADxD/Lt6jPg9Goop2Wdg8czR7ympzcWsz+djk+S1a3mBQohMdMe8evbGFjEvxMJvHsWeD/Mo/5uupC3GPH3uwzxuzgG+DmyPPSeFaCU+Rzi3ynFuxRFNSAeW7T/4HBznVCLhFNfK8mjiCYxmASOKbD+KcG2zLmyCMDLl+Yopy11YzHRaNkuQeyrlM0XfknDeszOct5PkbOHV1p8uxc8SzlfKHfl7CdufkOF825KcibXRyjLYwzEqx700zv3wtsi5f5iwTQemBEfrbndhNbKLxfZ3w343YzBl+CnSJfWJJg/rU9UVCiGSGIYthp2DLea+QjxJZLO06ZhL62WYV8p2KEOsaG7GEl4M2sWpNKKZ6IY9y4LPuGqq5ogc4FJZHkQ8vrULcwcrxUkJ+7xIOgttKWV5GqVjcT2GEk/W1YW5tZVjNHF32GXAkSn2hXDiJ6/NAVYrsc//YVaISsp5DMASYUSVpVJueOsQV7o+Jl1Jo82IJ9ZyqSwPJrmk1s2Yp0MlbI0lGlkrxbY7kf5ed2K/nTeI968JpLMKr4T10bGFfdKUE1uCuXmPxxTvkai+qhBZ6INZu34A3IB5HLlWgKtti4DHMevMEZhruBDNRLDixnRs7iLam26EvSOXY/Mh0cZ0YJ0hyALgzBod/xFstbnYuW/FT+Dl8Vfg+DLH7cCUlf0j71+KxV6VYnVMGQsyB1/x+RCzghargbwrtuK0duT95zElaEmZ84NZ8U6KvLes8P7pmOtrlDWw72bvhM+OAS4vcb6LsNpws7B6yhOwLNdLy8i5DfadbhZ5fxzlk4ndg1kdg7yDue49mLD9Clg5sJ/j34uphBXDmcDAMucFU5YHBF5fSOWlxsDu+d3ErcnvYF4BV2MTw1IMx1z0D8JcF8G8J95Jcf4nCJe5uBy758Xogd3v/wOGBN5fgNWuPgdzL0/LMGBHTIkehfXzclaj2cCT2O9/YuGv67J0QjQTGwB7AnsAO1M7D42F2CLXFGwsmYdZzhbju2nPKmzbE1MQ9iu8/gBT1Bdh4SaDqW3YyTtYfpI7MQ+ppLFOiEbRAwst+Fzh9bPYODfPmUTCJT2x3EgHFF4vw3I6KK69zUlSlmvJGRRXvL+PBc4HeQ5z31qY4tgDsFjN4ZH3jwCuLbFfkrJ8GqYgBxWz1zBX0/ewlaXVsYnNFgnHnItNdtLWOe6DLSRslfDZPCx518vYhGZVTDnZlWRr7nXA4WXO5ynLUZmfLbQpmCK6vCDb+tiAsXnCsd7EvoNyg8kWmJKX5Jb+ImZxmIEpxutjylhwsngzNmkLrui5UpbBkuL8meR7MBeLEX6ucO65QP9C2xBzgx6WsF9aZXl/wjH1yzGF+4ky+/UFTgROJexVMBX4Cf7qaVZ6YMntPOV5FLBJiv3exleeH8F+v8srOL8QrUhP4ItYKcQ9KB5qVI5pwKuYB8nr2DPko0Kbii3+pqUH9ux6HlsEfT1hm+74SvMwbAFufUzZ91olpaMWAw9jivOt2PUI0WiGYgu7axRe34R5aNVzbiyajxWwedaXCq+XYUaJq5xJJJqKerphnV7knJ8lHnv1KaZYZGF74rGtc7CBvBhJbtjHF2SKJnNK0+ZhinJW+pFcwzhLG0+6rKQXVXker71FNle6H1Z4ngewB9fvI++7cMMOskfCsatpaV2+OrDBPLjvY6R3dR6G9ZVojP1DmNJbC1bDTx52D+kSDX2KnzzsYFTzUuSPDmxR6S/Y4mCW58MCbHHpd9gi8NZUHvpRjHWJLzhnpRPztPoScDK2gPsO2Z+Hj2MVI/QcEI1mK2A+fl9ME1In8kNfLBeNd/8XAV9xKpFoOmo18U9qpyecrz9mYYpue1SF8v844VhPU9xNtJiyDPbATMq2Xay9SNg9Niu9gF+QPYPpNOKW4lKcQnggyNqWY2WMKpnEfJdsSWkuwbdSNJuyDDYpvIK44pmlvYuVoOqb4bwHJBwnbZy7x2bEF2iWYDGF/TMeqxzB5GHjSV+6yksedgqmZFRisRLCNRsBvySb0vgOZsU4ERtXyiWLbHaGAF/Gvof7SFf73Xsm/QvzmFLyQNEojsLvg8uAfdyKIxpEf6yagHfvF2LPLSH+Px1Y/Gq9mIDFyAYZQ/wh9BKVx0l3Aufhu9B4XI1NuqMkuWGfgK38g1k0T8DKURXL+DwRK59xIWbZrpbVsQnSXhS39C3DVt7/WZA1S8wp2KRj10LbFlsYKKesvYW5pVyJLQxUyrrYwsk+hN2jPRZhMcHnYdZOj6Ox78RjHhbzXI7LCV/b3ZgSXktGYPHxn8csPqUs/PMwd/d7MavrY2R3P/asy6MC703DvDGyuF2CWYAvIJxg7BMso/qfKpAtLStj/duLf05TdmYJ5iLquW9PxJ4XQjQbHVi5trFYbolynh8LMc+KewttYl2lc08f7Le/W6GNKr05YLkPrsTGhvfrJ5oQgHlxnFz4/1MsLPBld+KIOjMACwP5bOH1PCzs7V5nEommpB0z1pZTloOsicViDsK+q/ex1f96DtqrYe65gzFX7U+wWLTX8ZOy1IrVMaVvMKZcdmGK1yfAC9hEpZZ0w+KghwGrYNbuDzBlqJUTvayMLQisivWVbthAOwvzonivRuf5ChZPFeQcrLRXVlbAvDJ+QtgLYyK2cPN4JQJWwAjCsc/bUL5e+DSszJWnPD9M7X8bQqSlL7YIPJbyoUTTsWzXt2GJDufXV7SmZl0stOUgLOt/Z4ltl2Df2++x374Q9aAb9tvcs/D6NUxhLja+1Dvvj6ic4dhzY0qRz4dgxgsvge1szDjzaP1FE6L5KeWGLUQz04FZqIN9dxGlY/TLsSFwV+SYy7DM62mSqdWavvilq64inRvrUszafBWWuXIkpSfeQtSClTGPqHKxyPOxDKv70Pqu1fViTSz04nnK/94fxiz4QtSDAYTLL95Jcc+xfVGoULNyA8XLla6JGaCCIX6fLbKtEG2JlGXRyhxIvP/+o+Qe6diXeC6BaZjFzDXDMPnGYW6raeLv5xS2HVfYd3DDpRZ5pRfwPax2fDml7mvUPilX3vkM8BvMCl/q+72L5IoSQlTLRpg12etrZxfZ7jSy5w4R9Wc0Fk42NOGztQnnJppG7RKdCpEbpCyLVqYDK1EV7cO71+DYfbBs1tFEPLdhv5tmoTtmPT4WsyZnSR42AbNaj6Z8rWghgnRiWdtLJYFcjPWx7YscQ6SnF7ZY9yLFv+/l2PddjXeNEEnsh3lZef3ssIRtLgT+10ihRFm6YWGEXVi4X5ANsTDK4JxgZEOlE6JFkLIsWp0k6/JL1M7Fcz3g/sjxZ2BWsmalH5Y06AxMuU9TpmcxFv94PjYpT1vKS7QfW2Hx8cX60nTM+pRUT11URwcWQ3oXxRfFFmNeJMqeLWrJ6YTDKbaOfH5r4bNtGyyXKM538O9Z0KtnE0w59j6bRLZSqEK0FVKWRavTgWXUjvbjH9T4HGOIK513Un1d1kYQLF11PqYUe1aCctbn2zClezcsEZpoX3pjfWExyf1lLqakydW6MWxDuB5qtL2JVSgQohZ0AH/H71/vEi6h+XTh/WsaL5pIYCAwE/9+eYtnWxEOm3mVeAUdIUQAKcsiD+xMvB/PofYDwGrAzZHzzMOS8bRaEq0V8ZOHTQA+pLzyvIR48rB2rCLQjowGXqG4JXM8yTFxov7shi2AFXPNvgo3CQpF/liRcOK5R/ArNnyEP07oWeCePxP2PumGlasLxp+/jDyAhCiLlGWRF+4g3pf/XqdzHYw/MfDafylei7xViCYPi8ZrJ7XZWMmJMwr7lqsVLVqLHsAfKO7yex1y2W8GOoBDMWtf0n16D9jBmXQiT6xN2DJ5IZY7IxjTfLor4QRgC9lLCS+a7YwZEbz3JhKPYxZCJCBlWeSFzQgP1l5/3rPUTlWwKlYCJ/jbmY/VZc6LtbUHVu/ZK131EuWV5y7grcL2XvIwlQhqTQYD95F8jycD+7sTTRRhBWyxKynMYgnmBSNEtXwB609e3/pJ4P/lmKdSz6J7i3pzL2FleSnhyhlPIG8TIVIjZVnkiSuJ9+c3qG/tx70JZ5TswiytzZQxu5ashlmQz8DimYMxUcXap5il+nzMKj+k0UKLzOyAKcRJbr3jUVxys7MD5mKZ9Hu8BuUfENXzPcILMdF+drg70dqagyg9Hj8IrORMOtHy5MUalIVOLHNukPnAIgeyCFEtq2PKcR9sUPB+0z8HflXH8/YH/ggcFXhvBvBt4KY6nrcZ6Ia5n4/C4qFGF16Xi+GeirmBPYK5sD8FLKyfmCID38b6c9QyNAnr4480WiBREb2Bs4AfEp/fPIOVA3q/0UKJlmIQ8Avs2f4+9tyejCV/nAIcAxyQsN9y7JmuzNiNpQ/wGhZS1S3h82nAo5jnyTws3wTYYsdcLEnbDdj8SYhE2lFZFiJvnEc8E/YCYFPg7Tqf+0DM6haM270BS4Y1q87nbiZWAjbHV563o3xs1FLgdXzleSK+ZUw0jh8Bv054/wHgq1isvmgt9gGuxhb1gryHudO+2XCJRKvxNeACLMFXFj4LPFl7cTIxCAubWhXzjFoVCzEZDAzA5v59scXBXpjXRScWh70YMx7NxxYAZheOOQf4BFs8+BhzO59W+P+jwrYuOA1bIEvC8wzy/u+Gr/e8XthXirIQQrQBA7FBLOp6dEeDzj8E+Ffk3JOAnRp0/mZlGOaCfT6mEC+ivPv2VPzSVftiExtRP04h2e16HMlWCtE6rA+8QPJvbBOHconWYW3gYdLlrejCrJdXNUi2AZh308HYeDEBs2zPyyBvLdsULBxrPPZc3RcYQX2rZqyBr9SXk8/bZiq2mN+9jnIJIYRoQn5EcizVvg06fwc2AAUH6mWYoqikJ0Zf/NJVV2FW/3ID/FL80lVjsclRq5XsalbOIf59z0VJvPLEilhYSNLEXgqzSEMH9uxdTDiBVLG2mNqXkRoE7AWcCdyJhTy5UIgraXOxxeLfAocAa9Xwe7me5MR+SYryLEyJ75N4JCGEELmnJxa3Ex0k3sWUtEYxEng2IsNTwAYNlKGV8EpXnYGtygezdxZrcwgnDxvcaKFzwC+If6+zscUMkS+6kZwI8UPM8iVEGjYl2VMhqZ1W5bnWwRLPXo25C7tWeGvdpgC3AD8GtqSykNAdKW9RXo6NqeOIh2QIkRrFLAuRH74E3FX4fym+m9GvsIRfjaIHcGrhnJ4r61xs8L+mgXK0It2BDfFjn0cBG1P+WT2VcOzzkyhpYTEOxOLUgt/pLKzk2uNOJBL1pgNL4HZi5P1XgO3x4zKFKEVvbGHzx5gilhSq0YXF8a6Jn0yqHH2wmsB7FNqGVco5F1NIvXji4P+eVXou5oUWjE9eSjyO2UuIuzJmMffin4cV/g4hnLOkEqZhFvM7sUXjT8ps34mNccUU7eWFdhl2v6ZWKZ9oc6QsC5EvbsdKO4ENiB3YgL058GqDZdkFs+gE3a4uA76LDc4iHf2AbfCV5x0oXy9yCfA8vvI8EXPnbne2wBYVgt4WM4HdcZ+UJwsjgdNLfH4qliW/Wj6H/V6LcTw2+W4FOoDfYeV/gtwKfAV3CYpE67EbFhozlOLz6MMwN+FirIz1u0OwsTKre/B8zOrstdcK7XUav/jTG/Me89qGhbYB2fNuLMOexTdi39/khG2OA/5SZN/Owr4/Bd7KeG4hhBBtwLpYOSLPBclzR7rHkTz9gL8Tdo16BfiMI3nywghgDH7ysMWkc33zkoftRvvVnV2FeJz4Alqz1MuulL7XZ9foPDeWOc+aNTpPI/kb8esolk1XiGL0wzyluojHzS4HHkvYpxu+oj2X0r+tYFuCn7viWGyxrFVyVwRDjW7DFifTXvcybHw7Fr/O/QD8OU5wuy7gXmxBVAghhChJMHFRMAPzIQ5lGkM4HncBljRF1IYV8ZOHTcDc2spNRILJw1ptApaVDmwiFf0OvuZSqCoopyx/QPXZvAcSn5TmQVnug1muosrNni6FEi3LwZglNynR1DaFbTbFylB9nLBNUlsE3IclpBpNvpJSdccU2uOBm7HvLs13Mh+zNCcl7HsIC6cQQgghUrECltgrOqBMxY8/csFWxJOV3IQSb9QLb0V/HLY6v4DyE5LZhW3HFfatNhatWTiR+LX+1qlE1VFOWe7CXMur4bspztGKyjJY7dkPCF/LZFSqTVTGWlhd9ujvYyJmTU1T2mgqtnB5MG7H6UbTDQsvOgNLBprmu/LaFOz7EkIIITJzGP6AErQOnedSKGAlfNc1r70DbOdSqDahO2Y9PhablL1EuonJW/ilq0bTeqXAVgc+JXxN99HadZSTlOWoZeu6Ks8xsczxW1lZBov9D5bZ6yI5DlKINHRiib/SlDLy2ttYZv6RDuRtVlbH8gpEvT+S2oeYkj3EhaBCCCFanweIT3SXYMm+XPNNwm7ZC7GkHaKxDCUcT/YJ5ScoXt3M8zH3+rUbLHNWriJuPa9lrU8XJCnLUTfzBVTutbFpwrEeTjhnKyvLYLHdwetZivIpiMrogS1ElnO1nok9k3ZDSXbLsTbmil6ufNYiYDyWqVsIIYRIzWb4iZ+CFsSHaY5BeiMsY3Nw0LuG9ks81Ux0w6wcXvKwp0hnKfGShzVbjN0WxOU/3qlEtSFJWR4HPBd579sVHv/3keNcB/wz4Zytriz3Il6f/g6nEolWoxM4HPPAKfWMfBg4iNbzzGkGOrAM/zcR9waJLkT8BM0hhBBCZCBoOQlmTP66S6ECrABcQXjAexbL6i2ag5UwBfgUTCH+iPLKs5e9dTymeI/EzQJNNJvzi/j1x1uZYsryDyLvJWXkLUd34gni9iCfyjLAPsSva5uSewhhfAl4mtLPwXmYAidqw3DgN8Asin/vHwDforVDbYQQQjSIXlippuhgMoPmclmKZsuejdWgFM3JMCyxile6qlzW5C5scnMP5vK9L+VrRVfLBsStynvV+ZyNopiyvCrxMmIbZzz2/sQnnt3Ir7IM8CDh67rRrTiiyRkMXEvxZ92nWFWK1bGkcSe6ETPXrAScBLxP8fvwOBZSIoQQQpRkZ3w37KDycI1LoRLYinAd3OWYAqDV4eanB5bRdCwWj1fOJdFrweRho6ht6apfR871DM0RflALiinLALdG3j8n47Gj+3s1m/OsLO9O3CI4zKlEolk5mOLeNYsxb5qhzqRrP3piseIfUvyejMMMB0IIIURRLsUfPJYG/t/HpVAJDALuJDzY/Qdlu2xFvNJVZ2AW5XmUV54/xU8edjCVez8kuRKPqfBYzUgpZfkrkfcnk37BKckyvVHhszwry2BhA8FrO8WtOKLJWAv4N8nPreWYpVnhQ+7ojy0MFhtnXkRVN4QQQpSgHzZpjg4gk4AV3YmVSAc2UQ1awd9HA12r45WuGoNZX9KWroomD+ud4lw7R44xi3wlfSmlLPckbvnaI+Vxvx/Z79HAZ3lXlr9H3IVTCLDfzwySn08TMa8Y0RwMA24g+V4twZ5xQgghRCKHkGxd/o1LoUqwN+FSRgsxd12RH1YmnDxsOuWV58VYlm6vdFVSjdLzIvtcVc+LcEApZRngD5HP0tZcjmbTPjbwWd6V5WGEF2+WIXfadsdbuA2Ol15bgHnN9HAlnCjJPhSPZ74FG3uEEEKIGLcRngx6inOzroyvBTxBeKC7mnxZCYVPJ7AJlq19PKa8JU1Uo20ycDNWmgVMmQ5+fhD5opyyvBnxif2AMsfcOrLPfMJ1mvOuLEP8WXOIW3GEQ1bCShUlPW8exg9PEM1LP2wcSfJgepXsyQ+FEEK0AWtimaajA8ezNG9Jnd7AxYTlfQbFh7ULfTHrs5c8bBLFleYzsBrPwbjb5cAqDZa53pRTlsF+I8HPy9WXviCyfTQBYDsoy9GkcL91K45wxJqYMpXkxvtD8pMosF2Ieql5bRawk0O5hBBCNCnHELcud9H8sTxJ5aUOcCqRcIWXPGwclhDM6xd7Yl4SwQnR245krCdplOUsMbg9gY8j238xsk07KMsHEr6++9yKIxwwHHiTeF//GNjNoVyiOtYCniR+X+cRf9YJIYQQ3EjyoDHCpVApUHkpkURPYFssWd1BhPv1vxzKVS/SKMuDiNe/nTw5EQAAIABJREFULuZ2eHBku/eJ/6baQVkeSf4XWkRxNiA5znUisLY7sUSN6A1cRvz+LgT2cyiXEEKIJmQw8dI6XcAdLoVKSVJ5qftReSlhnEy4b1zgVpy6kEZZBovjLrcNxEvi/CJhm3ZQllcg7nYrl9v2YBNgKvE+fj2q0Zs3TiF+nxcB+7sUSgghRPOxH2ELrff/oS6FSkk34FeE5Z6EJSkS7c3phCdBZ7kVpy6kVZb3i2wzlXhugqGYUhjcbsOEY7WDsgyWDC14jUommH+GAO8R799/o3lzeYjqOI5wGFoXlutCMcxCCCFCXEVybFarJERSeSkR5VzC/fknbsWpC2mV5e7ErWV7RraJWlkeLnLOdlGWo7Hbq7oVR9SZHsCDxPv2RVh2fpFfvklcYZ5GPp9rQgghKqQf8C7xicLFLoXKSFJ5qauwrMii/TibcF841a04dSGtsgyW0TnqVhrk5cjn3yhynHZRlmcRvsZyJbdEa3Mp8X79Z1rP/f4E4tcR9ByrVfWI8SXOM6dG52gkRxMvLfUUmj8IIYQIsBvxwWI58HmXQmWkN/FJz9PAOi6FEk44lXA/ONutOHUhi7K8aWS7RVjcP8B2kc/mYwtoSbSLshwsO9aF4lXzzHeJ9+k7aM2EkaWU5S7gzBqcow8ws8Q5WlFZBgvViV7LtU4lEkII0XT8kfhg8RqmhLYSx2LKgHcN04HdnUokGs1xhPvxFU6lqQ9ZlGWwbL7BbU8ovB+1El1V4hjtoCwPJnx9n7oVR9SRTYjHp78G9HcpVBWUU5YnUb1b+RFlztGqynIH8Hfi1zPGpVBCCCGai97AM8QHi1ZMjrQ1NjEIWsnHofizdmEPwn34Qbfi1IWsynLUgvY/7DcftRKV8iZpB2V5W8LX97xbcUSd6IF5HgXv9SySE9u1CuWU5XK/7zTcU+b4raosg5UdfIHw9XwCDHMplBBCiOZifWywi7psjnQpVIUMBu4lfC230bpWA5Ge9Qjf95nkb6Ekq7I8kHjN5XMirydR+ntqB2X524Sv72a34og6cRLxvnyQU4mqJ0lZji6GlfIcKcdahJNhLQNmR47fysoyWJ3teYSv6RqnEgkhhGg6jiY+4E7EVuJbje6YAhGMx34d2MylUKLudBDOkN4FbOxUotqTVVkGuCGyfTQLbDkvknZQli8nfH0/dSuOqAMDgRmE7/N1TiWqDUnK8l8jr+cCK1V4/J9HjnUP8EbkvVZXlgG+R/ialgPbO5VICCFE05FUTurHTiWqjv0IZ7idjy0KiPxyJ+H++z234tScSpTlfRL2CU4Iy2XLzbuy3AFMJnx9uzqVSNSDaLb8OVit8VYnSVn+GvBK5L1jKjh2B3HF+IiE9/KgLHdi2bCD13WvU4mEEEI0HX2JD7ALaU13bI8NgZcIX9N4WtNiLsoTtQ78x604NacSZbk7MCVhvy7ggRTnzLuyvA3xib8yYeeLAcRdh1t5IThIMWX5p5H3KsnhsFPkGLOBFcinsgywC/HvUtZlIYQQITbDLLDBweJxWrOkhsdKxF1RHwZWcymUqAsjyG45bSUqUZYBfpOwXxfpPC3yriz/mfC13ehWHFEHfkj4Hn+IKX15oJiyvDqwlOqehZdFjntR4f28KssA9xG+tgluxRFCCNGMfIf44Pt9pxJVTwcwlnAt1Q+Ru2Ue+R/hvvtLt+LUlEqV5U0wF8NgexzLBFuOPCvLKxIO1eii9RM+iThRj6k8xaQXU5bBakcH389Sc7kv8cSfOxQ+y7Oy/AXC17YIWNWpREIIIZqSmwgPGAvIR7KknYFp+Ne1BDjFqUSi1kQzG8+g8uQ2zUalynI15FlZjlocp6EQjbwRdbNfTL6Un1LK8lcj708ifYWAoyP7voYtOkO+leUO4FXC13eSU4mEM/JWTkMIUVuOBl4OvO4NXEpru2ODxW1tjVnVwM+cfS22ki5an2uxrNgeA7F6w0IE6UvcY+YibAFN5If9Iq9vBz5yIYgDbsXKSHkMx+Jy03B05PUVmOKYd7ow9/MgX3YhiBBCiOZnQ+JuWGOdSlQ7emGJvoLX9hxWb1G0PqcTt3zkIUZdluXacRbha5qL1WkX+eIZwvf5KLfi1JxSlmWACyOfpam5vDbh0ovLCP/m82xZBlifuDdCmpAVIYQQbch+hAfNhcB6TiWqLUcB8wgP+oc5lUjUgv7AdLJPEpsdKcu1YT3iiQzPdiqRqAcrEU5ytQwY5FSi2lNOWf5s5LN5wMpljnlmZJ87I5/nXVkGczsPXqPymwghhCjKOMKDxkTyFcoxCnib8DX+BXM9F63Ld4lPIr/iVKLqkbJcPZ1YOEY0VrmcAiFaj10I3+dXnEpTH8opywDPRz7/RonjdQBvRbY/NLJNOyjLlxO+xjwlhRMpydNEVwhRX34G3Bt4vRVwsiNZ6sFEYAusvJTHccCj5MuK3m78BZskRt8b7kAW0TycitWPDfJj8jnhb3c2irx+yokU7rkm8jqqTAf5PFaCz2M2FvvcbkT7SrQvCSGEECEGYaWWvFXWJeRPkfTKSy0ivGIeXVUXrcMWhMuFdQHP0rrJ3LYmXgKq3snLfp9wziF1Pme92B9zxQ32h9udSiTqybmE7/UZTqWpD2ksy0OwMTu4zYZFjnd1ZLu/JGzTDpblPQhf44NuxRFCCNEKbE44/utt/FISeWJr4m7ZVwF9XAolKuZU4pPJCeSz74rifAb4lHA/+BBY3aVQoq5cQfh+H+NUmvqQRlkGWxQKbnNWwjYrE87h0QVsm7BdOyjLmxC+xtfdiiOEEKJViA7MF7sVp270A24kHqudN2t6O9BB/F7m1cokkhkKvEP4/i8CPudSKFF3or/7g92KUxfSKssHRbZ5n3gpyG9Gtnm1yDnbQVleg/A1TnYrjhBCiFbiFsKDyIFuxakbSW7Zs8lf6ZF2oA/mPhydVNY7OZZwz1DgJeL3/liXQomGcBvhe57HerlpleWewMeR7b4Q2ea/kc9/XOSc7aAsr0L4Gme4FUcIIUQr0QG8hz+ILCbfFtdtiLtlTwAGuBRKZGY44bh7r53rUihRV9YiPrHvAv7oUijRMG4gfN8PcStOXUirLAP8KbJdsJze+oTLRC4FhhU5Tjsoy2sSvsYP3IojhBCi1RhOOGHIZKy2bV5ZBcsIGhw83wa2dymUyMwO2MQuOrn8HYphzhvrApOI3+tbge7uxBIN5HLC9/6bbsWpC1mU5VGR7eZjIUdgdcaDn/2rxDnbQVnelHQu6UIIIURRjic8mDxE/iehY4C5+Ne8BHPl7eFSKJGJUcB0kpWofiX2E63DTljt5Og9vgH9VtuJqAKYlNSq1cmiLAM8F9n2G1g52fci75eK724HZXkvwtd4v1txhBBCtCp3Ex5Q8prwK8hGwNOEr/txzJIlWoMticfvdQGvASMdyiWq51ji5cK6gOvI/2KeCPMtwn3gerfi1IWsyvIPIts+TLxM0gygV4ljtIOyPJbwNV7qVhwhhBCtyirAJ4QHlZOcStQYemEW5WDN1tnAkS6FEpnYGJhCfKI5B8scK1qLFbFcAtH72QX8DSnK7ciOhPvBm27FqQtZleUhhBeTlgOPRfa/oMw520FZvobwNf7QrThCCCFamS8RTgyyjHwmUkliD2Aq4UH1SuTO2yqsTdxLwJtAXojVHRXNz05YHdTofVyK1dlWPHp70oe4YriaU4lqT1ZlGeL5N6Jt6zL7t4OyPInwNY52Ko0QQoiWZzzhgWUh8bIUeWUw8E/C1z+FfJYpySO9gUtInjROAQ5wJ5oowwrEPTyCrqS7uxNNNAn/I9wv8pbkqxJl+YCEfbz2Yopz5l1Zjib3WoAtvAghhBAV05e4ZWcOsJVLoRpIB3Aill00aMW4BFmZW4VjCdfUDrYJ2KKIaB72AN4l+X49A4xwJ5poIk4j3DfucCtOzalEWe4BfJSwXxcW01yOvCvLpxO+vtvdiiOEECIvbI+5PQYHmY+ADVwK1WDWBR4kbp3c16VQIjXbYyVCkiaRM4AfIQuDazYDbiP5Hi3D4i11j4THZsRd89d0KlFtqURZBvhDwn5LgKEp9s2zstxJ3AX7WJcCCSGEyBfRUh1dwFukG4DzQic2uM4jbp0c4FAukY7ewBkUtzJ/gN1fJYxqLGth4R7RBTmvvQns6kw60cxE8xL8yq04NaVSZXk49hwLtgNTnjPPyvJ+hK9tHtDfqURCCCFyRQ/gCeKD93O034CzLlZ7Ompl3selUCI1W5Kc/Mtrr2C1SLu5ErBNWAP4I8UXL5YA52CLHEIkEVUoZ5Gf8JhKleVqyLOy/Djha7vCqTRCCCFyycaEY3e9di/Q06FcLugGnIIlPPO+h+WYhUyZlpuf7ljJkGh5tGB7GzgZ3c9asw1wLck1k732ELaoIUQp+hKP0f2lU4lqh5Tl2vFlwte1HPiMU4mEEELklu8THnC8//+BWZ/bjU2IZ2WdAoxxKZRIzQAs63LSIlBwwjge2NCRjHmgE4vvv4fi33MX8BJm1RciLT8j3IcWYKXjWh0py7WhJ/Ekpbc4lUgIIUSu6QTuxx90gqVdrqc9XVe7E7cyd2Hf00YO5RLpWQPLcF4sbrar8Nk9wNHI2pyWzYFfA+9TWkl+BzgKe74IkYUVgcmE+9M9tH4NbinLteFM4s/xzZxKJIQQIvesQdh9NahgXEn7TnhHEo9lno+VOOnlUC6Rno2AizHrVCnlbj7wd8y9r91CEMqxFvAT4AVKf4ddwMvAt9HvQ1TH0cT71rdcClQDpCxXz1bEwz0udCqREEKItuFrhN3eggrzpbT+qn6ldGAu2B8SHqDfAHZ3KJfIxmDg58BUyit8M4DrsPs+xIWwjukEPovVMH2csLdJUlsO3IXVU27X54SoLZ3AA8TdsbdxKFO1SFmujlUwj5Xg9UwBBroUSgghRHsxAX8Qep3wJPmPDuVqBgYA5xNXHG7DLPOiNeiJKcFJmeCT2jLgKeAXwI7k1+o8FDgS+BvwMem+m0+BizAPDCFqzTqYchdVjlZ3KVQVbEK8BNQGdT7noZHzfb3O56sXPQiHi3ntAJdCCSGEaD8GE7a8TSCc9Ot37kRrGnYEnic8YM/ErAbt6q7eqmyE1WmOWl9KtSVY0qrxmNI9ktazpvYARgFjgauw6wn+zks1L857DBZbKkQ9OYJ4H3wMufm3G38h3g/afQFfCCGEI3bHnzgvBn5LeIA63Z1oTUN3TNGIWj1eRrWZW5EObBHkQtJbVYPtI+BOzPPgBGA3LM7XtRLdB0vIdTDmgn41Vo96CdmubzmmoJyILagJ0UjOJd4nJ9Ce1RrakZ8Qv//3YuOwEEII4YTgKu7rmIIcHKh+7k60pmIYZpmLDuT3YEqKaD26AdsDZ2Gu2uXidUu1ecAzWFmTv2JZXE8EDgRGY6WrBhRa2qzzfQvbr4b1sT0xK+8PsYWtq7GJ5CTSW4uT2idYwrOvYy7aQriiEwt3ifbR24HeDuUS9ecU4vf9bSx+WQghhHBGb+A5/MHpCkx5CA5Y41wJ14TsDbxC3F31YkypEa3LYCyW9xrgXSpXPtO2uZiiOhV4q/D/J5QugVWLthCzHnux2e1YMk40L/2IP2O7gH8jhTmvnE38fs9BORKEEEI0CSOxcjreIHUE8CvCA9e5zqRrPrpjCVSi2ZbnYQsLK7kTTdSQocC+WKzzbVjm7Hor0PVoUwryn4JZuaVwiGZnTczTKdqX78WUaZEPugF/IH6fZ2PPKiGEEKJpOBF/oPoUWJ+4W9SFuI/NbCb6YopUcKGhC5iMKdOy2OWLbliW24OAn2F1yR8nXLfcVVuKWaf/DfweOA74PDCoLt+EEPVnCPEEi11Ykr7NHMolasMgrAxd9P7OBLZzKJdocjQJFUK45BZgv8L/T2Iru2OBXwe2uQg4HouRFMaamCX+CMJZsp/HEpbc4UIo0VAGY6VhVsMs0oMLf4cAqxZee4rryqRbSFmELcQsAqZj9b+nYYnJgv9PAt4sbCdEXujEwgSuI15CagE2Dl3ZaKFETdgSuAkrGRZkJlbD/YmGSySEEEKkYADhWM2zC+97yrH3/iWodFISW5FcH/IxzJ1XiCArYL+5ocCIwv9fxu83L6BFdNE+dMcvczYBWyAKuuUmeVT8lvzWQ88r38AWO6L38gPgMw7lEkIIIVKxM36CoWXAFwrvH0s4W/C1qJxDMXYj2X3wv/jfpxDFeBa/z+zhWBYh6kVfzHvpFKyqQDScJdgWABcU+exFYNsGyy6ysxrwD5Lv4cMoQaYQQogWIpiZ8gP80g3fJKwwTwB6uRCwBeiOraC/Q3xicB+wqzvRRJNzNH5fucetKELUjFWBA7CY+icpXwN8Ltb/T8eel92Bo0hWqpcB44EVG3Y1Ii0d2GJ7Me+A8aiOthBCiBajO/Ao/mD2b3x30MMIT3Lux2IwRTI9gW8D71HcPVuutiJID+B9/H6ypVtxhMhMB7AxcAxwKcmloKLtY8zy+H3gsxT3XNqG4qXd3kLeGM3ESOBBii+GHO5ONCGEEKI6RgCz8Ae2EwOfHYIlE/I+ewJLYCSK0wv7DicTnzQ8hy1CKHu28PgZfv+4yrEsQpSjL7ALcCpwO+nKrE3C+vaxmGKdZdGwH2aRDObSCLZHUOkhl6yB3Z9i3gOPABs5k04IIYSoEQfjD24Lgc0Dn+1K2K3qTWDdRgvYgvQExgCvEZ9AvI3F7/V3Jp1oFgZgJdy6gMVYxnUhmoVhmFfMOEzxWUh55fgtfOU4mgW5UnYi+VkaDGNQmanGMQjrE0kJvLqwOcNYlCBUCCFEjrgSf6B7Ccvg67E18FHg8ykom2VaumOlpl4kPqGYBZwHrOVMOtEM/Am/T4xzLItoX3pgbtFjgb8TDhEoFW98P/ALYC9s8ade9MXioL3ElNG2BLic8GKvqC2rA+dQPC65C/M4WMOVgEIIIUS96Au8ij/g/TXy+UaE48c+wWpjinR0YLWt/0vyJO/vyJ2wXVkHXwGYjXIDiMYwAgsL+T2Wu6KYlTDY3sUqJHwXK6HnolLClsCdZeS8D9gHWTZrxdbA3zDvl2Lf+YtYSTwhhBAit2xNOEb5wMjnwwiXSpoH7NlIAXPCKMxNMSnO6xXMRbueFhrRfNyI3wfGOpZF5I+VsMU4r7bxh5RXjJdgXkbjsZCSWrlU14rPY3k0Sl3Da1gOiYGOZGxlemEhWg9RfgHlaJSLQwghRJvwI/xBcA6wYeTzAYSto0uA4xopYI4YjrlhBxOsBWO+/gxs6kw60Ui2wb/376Da5qJyemALn9/BFuVepXiCrGCbCtyCLdZ9DujTaMEroAM4iLBXVFJbhGXgPgjo7UTS1qATS+B2MTCT0t/px1hGc32fQggh2opO4A78AXEi8cGwb2SbLuB85PJWKSsBJwAvkDwpeQxLltPPlYCiIQQXoQ5xLItoDbphC2pfw2Lf07pTzwMexhbrDqb18yZ0YmEu/6H8tc/CylzthiVibHc6MG+nc0kufZjk/XQc4bwmQgghRFsxECv34Q2Of0nYpifhpGBdwHVolblaRmFuj/OJT1IWYC6U+yKXtzzyFfx7/aRjWURz4mWnPgPLAD2X8spNF36G6rGYO3avBsvdSLbAnqFpFw3uwb6X4S6EdcRAbJFkPPAB6frQI4V9NPYIIYQQwLaE45fHFNluLGEXv8eAVRshYM4ZgtXgfZPicWK/QuVS8kQn8Ab+Pf6cW3GEQzqADYDDgd8BD2JhMWmUmqnArdjz4wu0b8K41YDTKO+iHWwvYd/3geQro/MAYA/gdMyDpVhG8WibhnmNKRxICCGESOAH+IPmXGCTItsdQngV/y0se7aong5gZ8yKX8yK9AI2MW62BDwiOyfi39dbHMsiGkMPrNzR17DM1P+hdGmeYPsYC4n5JbA/qtNdjG2AP2ALCWkV5y5gMnAzFse9C9C/wXJXQh/sek/EvAleI13MenCsvxpTrpU7QQghhChBB3AT/iD6KhZfm8QO2MTN23YGpuSJ2rES8A3MHS5pkrMcs+x/D1jbjYiiSlYApuPfTy065Yt+wE7AScBlwNOEPXhKtU+Au7E6twfSXm7DtaIbpgReCkwhm+LstQ8xS//FWELML2OJMBsZw9sTu/9fxBK5XYD1jUlkU4y9Nhsb64/AcpII4ZwO1wIIIURK+mNJvkYUXl+HuQYmsRHwr8C2i4BjsLqcorZ4NVIPpbiL3NNYBthbsBqYojX4JXBq4f+/YMnfROuxFmYx3iLQRpTcw2c29tx9KvD37TrI2M50YPdnj0LbkeotqfMw6/WH2OLxFOAjzPNqVmGb2ZhCOw+rWwwWguElcOxXeL0ipoAPxsJyVgv8X4tSWM9h9arvxFyzl9TgmEIIIURbsjnhhFPfLLHtIOJ1GZUpu75shsUvv01xy8HrWLbTnZFrXbOzKn5Yw3xgFbfiiDL0x+LLj8cWNx6ifNmdYHsP+CfwC8xiPAIZVVzQDzgA+C2WKXwelVmem7EtAZ4B/orVRB5Wm69MCCGEEB4n4A+8C4CtSmzbB7iB8GB9O+2baKZRdGCJ2cZhcWrFJk6zsPvzDTRpalYuw79fpzmWRRi9gC2Bo7CFpztIV2onqLC8gMWD/hArXTSooVcgstAdG+eOB67A7l1al3mXbSmWN+RGrJ/thFyrRQuiFUMhRCtyFTZRBMvSvDXmUpZEB/Bj4Gx8q/IbWC3MV+ooo/DZBLOUHICVpEqiC3PHuwu4D3PHm98Q6UQpNgWex35HH2Ex6AtcCtRG9MJCSjYCNsZ+R5sB65HeK2M2plw9V2jPYKEQC2strGgo3bDf4gZY/9ig0NbH3KMbVbd5GebmPQkbT18vtNewsXlRg+QQom5IWRZCtCIrAk9gE0iw8iQHYApXMfYB/oZvVZ4DHAncVicZRTLDgb2BPYHPUzwZzWLgf8D9hfY/NPFyxR1YLCXAt4BLHMqSR/phCs8m+IrxxlhW+bS1ZBcDL2OK8IuYgvwiZnEW7ccAYCgWWzwUU6AHYwswXiZtLya5L75yvRx/4XkWNqbOxZ69HxeOuxe2aHMacF5hHyGEEEI0GZsSjuU6KcU+m2FuYUE3sVPqJaAoS2/gS1iZmnI1SOdhFuczMUW7X8LxRH3YDf8+vIri/iuhE7MEfhELJfkTcA/wAdlcW5dhFrt/YLHFh2BKtuL/RSPYlXBIkxBCCCGamMPxB+7FWNmocgwE7iU8Ab0Wi28WbhmBWS6vpXwd0mWY9Ww8Vhd2A+QtVU+exv/u93YsSzOzBqZQHAv8GlNqX8LcnrMmQnqtsP85wBisZq1iPoVLumOlw7qwMBn1RyGEEKLJuRR/gvke6TL2dscsO8HJ6ROoXmizsQlWu/NG/Jq/pdos4D+Ya+DhmEurrKC14Sj87/l+x7K4pAewLmZt/xamyN4APEtlWYvnYwsR12FurQdhXjONijkVIivX4fffLzuWRQghhBBl6IMlrvEG7ztIryAdiV8apwtTyPasg4yiejow5fmbwOWUd9v22hyshM6fMGvfjvgxeyI9PQhnXN7GrTh1owNYHRiNLRD8H9bfHgDexUI3sirEy7AESPcAFwInY3GfI9Bijmg9jsTv2+MdyyJE3ZHLmhAiD2wAPImfvOts4NSU++6AWYa80kXLgV9isbFKXNLcrAJsjynAn8XKq6SNZX4fS4j0AuYm+xqWJX167cXMDadg5cDAkuUd6VCWSumPuUoPL/xdPfC/936vCo89BetD0fYmyj4t8sNA4EPMQ2sq9hsqlVxTiJZGyrIQIi/sB9yMWWq6gEOBCSn3HYxN/r8YeO8/wGHYpEC0Bh2Yi+xWWIkq7++ADMeYha/gBBWed4FptRS2BVkZW2RYGbOwrkvzZFvuCayKTdyHYItfQ/GV4DWBtbBM+pWyCLMQvxNpb2F9ZG4VxxailXgI+Fzh/62BiQ5lEaKuSFkWQuSJs7C4P7CJ6/ZY+ZQ0dCvsexq+a+QHwFeBR2soo2g862BxoCOxjOibYKV5sloQF2HK4fuFv+8GXn+ALax8UhuRm5Y/AGML/58H/KiO5+qDeQ8MwhTgQZhC7CnCQ4HV8MviVMtSzDocVYa9NgVZ0ISAsJfJGZgnlhC5RMqyECJPdGI1l/cpvH4Dc8+dleEYewNX41sjlwI/B86tkYyiOeiOWUY3wxKBbQCsX2iDqjjuIuAjzD3xw0KbWnhvOjAz0D4p/G0ld/+1sd9VdywefC38uqxJdGCuz/0xF/lo648pxJ5SPLjQVqF4De5K6MI8A7yFDW/B44NC8zwHltXwnELklU2w8BWAp8hvDgMhpCwLIXLHSsD/MMshwN1YMp0sk+ARWBzzVoH3/o4liJpTAxlFczMAWA9TnNcrtLUxxXAYluyqlszGV57nYPGtnxbaAsxLIvj/AvwY2IWF19H/K2EFfGt7D3yXZU/h9f4eiVnqwX5r72FW4N6Fz3vjK8NeHoF68jH+osS0QpuCLVB4yvFkrLycEKI2vIE9G7uwMIfJbsURoj5IWRZC5JGNsEm8N1E/Czg94zF6YXVSTwq89y5Wkkhu2e1LN8z1d3ihebGwq2Muwp57sOp2V84CzAo/Hd8iPyPw2rPaTy68lhIsROM5H398/BZwiUNZhKgbUpaFEHllfyzhVwe28n0IVq83K0dj5V485WcJ5pZ9Hq3lPisay4qYAj0YX4EeUGgDE/4fiFlk88KnmMU8qc0qNO/1J5h1+GNMIZ7vQF4hRDa+iHlugYU/7e9QFiHqhpRlIUSeORv4aeH/T7GEXy8V37woGwHXA5sH3rsfq8M6pRoBhQjQp9D6l/nfU6o912jwY+w9V+ksdBGO65+Fn8jqUyxuH0yxXVx4bzP8ZHovA/tiLuKLKB3DLITIBz2xxa2VsAWuVaguDEQIIYQQDaYT+Bc28e/CaummrcMbpTfmdrY8cLyPsIRgQrQbHcAr+L+Fnd2KI4Q+xH0zAAAgAElEQVRwwI34z4A9HcsihBBCiAoYgCUi8Qb0W/FLQ1XCfthqune85ZgSnbUMkRCtzvH4v4N/OpZFCNF4jsZ/BvzZrShCCCGEqJSNMddQb1A/rfTmZVkTeDBwvC6sfMYGVR5XiFaiDxZn7C0abVx6cyFEzhiMVZrowrLiK7xTCCGEaFEOwHehXoZfi7lSOoGxWAynpzDPB06hOsu1EK3EWfj9f7xjWYQQjecx/GfAZxzLIoQQQogqOBd/UJ9JbSzB2wJvEbYy3w2sUYNjC9HsrMr/a+++4+Wo6v+Pv25JI52QAgECBEIJPXQCKAQQlU4oUgWlWIgCGgRLEAsKiMAXNIIgQaqAYJAOIl167yGBQAIhPaSX+/vjs/ubuuXe3dmzM/t+Ph7zyM3u3tnP7s7cPZ8553yOXSRqw9Z5HuQ2HBGpsXPxvvvOdRyLiIiIVKAZuAfvi/1tvCrClegL3EAwYZ4NHFWFfYvUu6vwjvtxbkMRkRrbCu/8f8pxLCIiIlKhfgR7gh8EWqu078Pw5nDmt1tzzymSVRvjzVucgbcmuYg0hsl4U5wGOo5FREREKrQJNgw7n9BeXcV9D8QqbvsT5k+xdWhFsupuvOP9FMexiEhtXYF3/p/gNhQRERGphn2B5Xhf8N+r4r6bsIRhgW//q4A/AT2q+Dwi9eLLBNczV5E7kcbxVbzz/x+OYxEREZEqGYP3Bb8C+FqV978e0SWmJgP7VPl5ROrBs3jHuUZSiDSOLngXhxfk/i8iIiIZcCVeA38esHmV959fYmoJmsss2XY03vH9qNtQRKTG7sQ7//d2HIuIiIhUSSfgIbwv+Q+A/gk8zxbAcwQT5mnAgQk8l4gLrcCHeMf3jm7DEZEa+jbeuf9Hx7GIiIhIFa2OzbPMf9E/TjLDyFqxXuYviPYyD0jg+URq7Sy84/omx7GISO2sidXmaMNWnBAREZEM2RhbGznf0L82wefaAHiEYMI8Gzg5wecUqYVewFy8OgAbuA1HRGrIP3pqM8exiIiISJXtDizF+7I/M8Hnaga+C8wnmDRPBNZJ8HlFknYx3vH8B8exiEjtjMM793/sNhQRERFJwkl4X/YrgQMSfr61iK7LvBBrdHRO+LlFkrA2sAw7lucDfdyGIyI1MgLve+wxx7GIiIhIQi7D+8JfAGxZg+ccDcwkmDS/Auxcg+cWqbYb8Y7jHzmORURqown4GG8ahlZ8EBERyaBW4D68xv4UYGANnncgMAGvSEpb7ucJqNEh6eLvYfoYjZIQaRR/wTv3j3Yci4iIiCSkF/Aa3pf+80CPGj33HsCbBHuZPwWOw67ci6TBo3jH7zEFHrNDzaIRkVo4AFXEFxERaQhDgRl4X/x3Y73OtdAJGAssJpg0PwpsWqMYRCqxP8EpBeELPT2A22odlIgkqhtWd6MNq4zfyW04IiIikqTtCa6LfFWNn38Y8BDBhHkx8EtgtRrHItIeTQRHSOwZuv8k4INaByUiifs33nn/JbehiIiISNK+BizH+/I/10EM++MVTvHPBdXQbKlnJ+Mdr/8O3fckVnG+S62DEpFEnYZ33l/kOBYRERGpAX+jfxVwgoMY+gJ/whIMf9L8CLCFg3hESukCTMc7b4bnbt8Ir5CdphWIZMs6eOf3245jERERkRq5AC9BXQbs4yiObYDHCSbMK7Gq2Ws4ikmkkF/gHadX5277je+2Ax3FJSLJeRnvHB/mOBYRERGpgSbgOrwGwHxga4exjAY+JJg0zwLGAC2O4pLGtTHwO6LnRH9gEXZ8LgHWBqbh9TydVcMYRaQ2zsf7XjrDcSwiIiJSI52AB/AaAZ8AQxzG0xNLUJYSTJpfAEY6jEsa09FYEvwycDre+uB/xjs2b/T9vAJbl1VEsmUnvPP8YcexiIiISA31IjjE7A1sPrFLGwITCSbMbbnbhjqMSxrPLwlOV/gHNuc/P9d+KZYk56cPPOokShFJUjNevYJlQB+34YiIiEgtDQY+Irj+cT1U9R2FJe/+hHkZMB4bDiuStCbgZoLz6fNDsMMXc9qwBrWIZM81eOf5EY5jERERkRobDszBawzcjF1Nd60L8GNgLtH5zD8AOrsLTRpEN+BZgpXbVxGfLK8CursJU0QSdCjeeT7BcSwiIiLiwF4E5wtf4DacgNWxeMI9eh+i9ZkleYOwtcDzQ66Lba4K5YlIcnrgff/MAlrdhiMiIiIuHEmw16zeqvsOA24lmqA8g4qASbI2AxZQOmE+zFWAIpKo+/HOc33fiIiINKhzCA4rPcFpNPH2AJ4nOgT2FrQOpiTnK9hw7ELDsNuAnziLTkSSdDreef5bx7GIiIiIQxcRXBJntNtwYuXXZ/6AYLKyEut9Xt9daJJhP6BworwCKwQkItkzBO9cf91xLCIiIuJQE3A1XsNgKbCv04gK64b15s0jmLgsAi7EWx9XpFquoHCBrycdxiUiyXod73zXUoYiIiINrAVbVzbfMFgI7OI0ouLyRcAWEUxgFuRu7+UuNMmYVuBB4odjz3QYl4gk6wK8c/37jmMRERERxzoTLGoyEyt0VM8GY2sxLyeYxHwOjAW6ugtNMqQXtg54XMGvPg7jEpHkjMQ7z+93HIuIiIjUgZ7Ac3gNhE9Ix3zgYdh60eHevynAN9HSH1K59bELSOFjbDuXQYlIYlqwC6/56Uk93YYjIiIi9WANrBctnwy8D6zpNKLybU78clNTgJNR0iyVGUl0FMNxTiMSkSRdj3euH+I4FhEREakTg4HJeI2EV7F5wmmxM/Ao0aR5MkqapTLHETymLnIbjogk6Ai8c13V70VEROT/2xCYjtdQeBro7jSi9hsFPIOSZqmuq/COpb87jkVEktMbWIad658BzW7DESlPk+sAREQaxJZYD23f3P8fAr6Ozd9Kk1HAr4EdQrdPxiqeXoMVbxIpRxNwA3AU8Dywve++7sAgYCDQH5vCMCC39c/9bp/cv92xwnqdcz8vB77AGuZzc/tbgB2bC4Bp2BzKGdiFrPzPqsotkpxHgC/nft4J+J/DWERERKTO7EZwiaZbsMInafR1LLkJ9zS/DRyDepqlPJ2xSvHvAYuxiuwPYsls3JrMSW9LgUnAROziz8nYBaJBSb0BIg3kDLxz7XzHsYiIiEgd2htrkPsT5jQnlqOAZ4kfnj0G6OYuNKkzrcA2wGnA34C3gJW4SYo7sk0H7gLOAfZEa5CLtNdQvPPpJcexiIiISJ06kuA6s38j/fO3RhFcKiu/fQaMw+arSWPpBnwFuBB4HFiI+4S3mttK4HXgr1ixsoHVedtEMu0dvHNoiONYRErSnGURETcOB27EG4Z9LfAtbO3ZtGoC9sd63nYM3TcLuDy3za5xXFI7w4D9sCR5DyobWbAUb07xDGxe8TTfzyuw+chtWCK+LLctBDoBPfDmNYOt7dqKVaMfiM199s+DXpPK1n/N95bdl9ueRvP3RcIuAs7M/fwd4E8OYxEREZE6dgLBYahXk52LmCOxeZ/h3rgFwKXYklqSDdsBl2BzfTvSO/sBllxeBnwXm6owBDfnwupY4aETgN8AtwGvAUto/2ubg02zOAibmy0iVuArf47c7TgWERERqXMnEkyY/0J2EmawHua7sB5zfyKxGLgSm8Mm6bMOMBabd9yeBHIadhFlHDYKoS/p0AIMx4ZbX4oVt2vPfOs5wARsukKWzm+R9mrFRhe1YQUv07aMooiIiNRYfvh1vmH9R7fhJGJzLFlYTrRncSKwi7vQpExdsWP1CaIXPwptk7FhlqOxBDtLegF7AedhS+CUmzxPxioBa3SFNKqb8M6HAxzHIiIiIilwOsEG9R/chpOY9bCeOf8SWvnteaznLq3LaWXVQKwX+RNKJ4KLsaWfxgIjXATrUD/sosB4ynuvlgG3Aju7CFbEoWPwzoPxjmMRERGRlPgBwcb0hW7DSdRa2Dq2c4gmEW9j69t2dRadAGyLjQbwL3UWty0F7gQOA1ZzEmn9acaS4P/DCpKVSpwfBw5FF4qkMayON8poGpqaICIiImU6g2Aj+hduw0lcD2w95ilEE4gZ2NzWNRzF1qg2xXo8SyV4z2OfXX83YaZGCzZXeQJW4K7Ye/o21jut5EGy7jG84z5uFEpf7KKqiIiISMCPCDagf+o2nJpoBY4CXiCaQCzEioFt4iy6xrAutoSZfw3wuEJVvwfWdxRj2vUETgHepHjS/DS29JZIVo2l+EXhC4HNahqRiIiIpMZ5BBvP57oNp6byy07FFZF6Aut503DV6umLrX26mMLJ27vYsk49HMWYNU3YetT3U7xY2j3AFo5iFEnSZnjH+XOh+wZjdS107IuIiEhBvyLYcL7AbTg1Nxy4hvg1bt8Fvo/11EnHHQpMp3Cy9iS2xFOzqwAbQP44L9Sjvwxb81lz+CVr3sOO8VUEq8Nfm7t9axdBiYiISHr8hmDD+WIabz7jAGzI3lSiicR8rJrqps6iS6eBFJ+X/BaaO1trG2OfSaGe5veBLzuLTqTjtgQGxdx+Kd7x/a3cbVvgLcPWaBX1RUREpAP8c7vasOSwEXv6OmNLjjxLNJHIr9e8N0rwSjkRmE18QvZR7n4Nc3dnJ+A/xH8+K4Er0IgKSZfVgFex0VG9fbfvjXds35m77V7fbdvXMEYRERFJsbMI9jjdgBXFalQjsOrCy4gfoj0WW/dWPN2Bm4lPwpZiRXY01Ld+HIBdvCjU86/RFJIme2PfYTOxKTSdc9t8vEKO+xI8zrUGuYiIiJTtFLzhaW3ALUAnpxG5NwhbXipuLdsl2LDWXV0FV0c2xHp24hKvF4Ft3IUmRXTHeuP8531+W4ANlRdJi+vwjt/JwJHAbb7b3iU4d3+kmzBFREQkrY4GluM1Jiai3kCw9+BE4peeasOGbp+IDQdsNF8lftj1IqwHXkOu69+uWG9y+DNchc37bPSLZpIO/bCe5fyUgjbgQwrXTtjdTZgiIiKSZkcSHH58L9DNaUT1ZQQ2r3sh0cbXvNx9mzuLrrbOJL5g1MtYb7OkRzfgr8QnFfegvwGSDt8gOg+/ULKsgnYiIiLSIfsTXFLpUVT0J2x14AxsaF9cj9zDwOFAF1cBJixcGC6/3YgN75V0Og4bFRD+XB9DfwMkHe6leJKc30a5ClBERETSbz+CjeYngF5OI6pfI7H5y3EFweZgvc1bOYuu+sYRfZ3LsQRa0m9bYArRz1h/AyQNhmAjfwotk5bf9nUVoIiIiGTDKILDjZ8F1nAaUX1bE0sYC82Tex44GejhKsAquIjo65qPhjRmzUCsOFv4s/4f0NdhXCLl+AGle5b3cxadiIiIZMZIbC5uvoHxPjDUaUT1rxU4CLibYPVVf3I5HtjBVYAd9GOir2UuWoIlq/oATxM/JLuzw7hESmkGniH+729++7qz6ERERCRTdsSrMtoGfEzjFLGq1NrAz4APiG+wvQKcjs2Brmf7EG14ziZ9CX9vrGc0bqtm5fdaPU/SegCPED1uL3cZlEgZtiC4ukN4O9BdaCIiIpI1mxIcXjwH2M1pROnSjPXSjye+gNISbKmu0dTfUj0bY5+3P95ZwJYug+qgQhct2oA7qvQc/QgWyAtvv6jS89RKD6w3Ofw6vu0yKJEy/JrC5+EhDuMSERGRDFoL6wn1J3iHOo0onfpgc5dfJr4RNwtLqke4CtCnN/A20WJeaa0kWyxZXgb0r8JzlJovmbZkGaxWQfi9W4pdABKpV12Ad4gfjj3aYVwiIiKSUX2Bx/EaHCtQD1Mldgauwub+xiVWL2HJ1wBH8f0tJqbTHcVSDcWS5TZgTBWeo9BFkDQny2CjS/z1C9qwqtlaUkrq2e7EV8Y+0mVQIiIikl2rYcWr8o2OVdhyQtJxXbGejonEz7NbATyIrYPbrUYxfT0mjmtq9NxJKZUsv1rh/rcpsf80J8sABxNNPP7sNCKR0v5C9Lj9Bva3dC1gA2y6yYjcNiq3HeD7efvcfZvmHr8OWlNeRERECmjBekT9jY/LsLm5UpnBwNnAmxQepv0nbM54U0IxdCWaWL6GDWtMs/BrWkh0fvHWFez/0tC+ZpOtZBmiy4etxBIJkXoyAPsb+S3gt0RrRSwm/u9re7eF2N+Vp4C7sCk05wGnAntif89FRESkATVhPcr+hsMN1F+BqjQbDlwAzCC+oTYVS9CqPXf07NDzLKM+5lBXKpwszwD+Ebrtkg7uuzPRz+kKop9Z2pPlbkTnsT9JchduRIoZhFW2PheYgK0FHi5I6HpbALwA3IR9Z44G1k3gvRAREZE6dAbBIW73YRV0pXq6AIdhw7SXEd8gexVLcodU+Fy9iPaIdjSBrDdxyfLXQrfNpGPrCB8a2s/UmNuykCyD9ZiFX9fXnEYkjaATdtFuDJYYv0H8fOS0bNOxv+njgP2xmiAiIiKSQUcTTOJeQVfOk9IXm7v8IIUbis9jDcqOFAY7J7SvmWSnEReXLLcC00K3H9SBfU8M7eN8bI5vFpNlsCGn/tf1lNtwJIOasGkRZwOPUvhCYUe3pcDHwCRstMTzue3B3HaX7+dnc/e9mXv8R1RvKHd+W4H1jJ8H7IRNdxIREZGMGAXMx/vi/4RsDN2tZ+sBPwFeJ77xtQwrxnYctgxUKa1Yb4d/H+dWO2iH4pJlgN+Hbv9nO/c7kGhDfmOynSxvTfRizU5OI5Is6AMcjhUTDF/Eas/2BfAicDO23vLpwBHAxbn7z6xSvD2xc30kNpLke1iy+3cswS600kE526xc/Cdgf2NEREQk5bbChp/6GywHOI2ocQzHhvNNIr7htQTr/TwOG2od58DQ78zHGq9ZUShZ3ix0+3La1zj9Uej3H8/dnuVkGeDfBF/bX92GIynVBTgEuINowb1S2xLgaeBy4DRgL6xSdSHN2Pn5vQReRyGDgD2wZRYvBh7DCoO1t9f5fuBYNM1JREQk1QZjV/T9X/JjnUbUWJqxhtl4rGciruG1CLgd68FZzfe7t4UeN75mUddGoWQZbPij/74ftGO/r4Z+96Tc7VlPlsPLiy3AKqmLlJL/O3UV7SvINQkrJHk6sCMdqy+wMXByZeFXrBUbnXEqcC02vLvc+dcLsffgq7n9iIiISMr0IDqHczz6Yq+1Fmx44KUUrqi9CPusTsJGAvjvy9qw2mLJ8ndC95W75vIORBuy+Z77rCfLLdh0C//r289pRFLvemEXogqNgAlv87FpEadQefFCv55V3Fe1DMRG/tyE1Yoo5/2ZBvwU6O8gXhEREalACzY0zv/Ffh+FhwBLsrpgFVevB+ZRuhH2MdlbDqhYstyb6Fqs5ay5fGXodyb47st6sgy25rf/9V3uNhypU+tjVfXL+dvzDrZk3pdo3KUIW7CLleOAlyj9ni3Geuk3dxCriIiIVGAMNhTb32OnStlu5RPnCQSLsvm3vziLLjnFkmWwYjr++/9YYn9diS6ztafv/kZIlvcn+PpecBuO1JltsSkf/u+AuO1TbATMDm7CrHvDgd8AUyj+Pq4CHsAuNIiIiEhKfIVgUjYN2M5pRJLXDaviGq6C/U2XQSWkVLK8X+j+mdiFhUKODD1+CjYXM68RkuU1CM61XE5wHrw0pqHYcOJi83AXY1Wj90NTdMrVBOyGTWsq1Ut/D+WNjhEREZE6MILgUiALUKXsehKeQ7il23ASUSpZbsbWT/U/5uAi+7sv9NjzQvc3QrIM0fd1C7fhiENrYEOoi1W1/iz3mLUcxZgVPbFCZW9TvKf5VmBDRzGKiIhIO6yFDdP0f5FfQLA3Tmqvheg6wVlcnqRUsgzw29Bj7iywr8EEh5auwnrT/BolWX6I4Gvc32044kAnbM33BRRO3F7C1gwuNlpD2q8FW3rrMQq/90uBi9CoDxERkbrXExse5v8iv51sJmdpMYDo8OMsKidZHkZ0WPGgmMedE9rXozGPaZRk+RqCr/HbbsORGhtB8SJUL2JTcSR5OwP/ofBnMQlbi1pERETqWAvWo+xPSt4GNnEZVANbn2CDarLbcBJTTrIM8FTocT+MecxbocecEPOYRkmW/0jp90uypxtWrTk8KiW/fYgNE25xFF8jG0VwFFd4uxXo5yw6ERERKcsRBNf2nYfmMbuwCcGG1Dtuw0lMucnyKaHHvRa6f9fQ/V8Qv25royTLFxB8jT9xG47UwPbAu8QnYjOA7wGdnUUnYNObjsYufsZ9TtOAfZ1FJyIiImXZimASk5/HnLU1fuvZekR7hLKo3GS5N7Aw9NhtfPdfFbrv2gL7aZRk+TKCr3GM23AkYccSXZPc32PZ311oEqMb9p0at3yX6oaIiIikQD/gQYJf4rcA3V0G1UDWIPjez3EbTmLKTZYBbgg9Nr/mcjfs/fHft0eBfTRKsnwdwdd4kttwJCFdsLWQ45LkT4CD3IUmZdgGmz8e9/ndDfRxF5qIiIiUkp/H7P8CfwXYwGVQDaIJW/PU/95nseHUnmR5n9Bj82suHxu6fTKFR0E0SrL8KMHXqGJO2TMYeIb4ROsK4qchSP3pjC1xt5zo5/gOsJm70ERERKQcRxEcAjsL2NtpRI0hvFbndm7DSUR7kuVmbDi6//EHAw+Hbvt5kX00SrL8McHXqEJ92TIEeJ/osbwAqzsh6bM78CnRz3Q2sIPDuERERKQMI4CP8L7AVwDnonlVSfonwUbTaW7DSUR7kmWAX4Ue/xyw0vf/ldh870IaIVkeTPD1LUaFnbJkGDCV6HH8HrCFw7ikcmsDTxP9bOdiS1CJiIhIHVuDaC/ew8SveSuVO5vge/13t+Ekor3J8kYElzcLbw+X+P1GSJaPIPj6nnQbjlTRcGA60WP4X1gRPEm/LkQLFrYB84HdHMYlIiIiZegEXEIwYfkYfYknYTeCjaVZQKvTiKqvvckywBMUTpaPLfG7jZAsTyD4+n7vNhypkk2w8yN8/F6H1k3Ool8T/ay/wJbJExERkTp3AJa8+Ydlj0PDsqupBSti5W8sZW2ueEeS5W8RnygvAHqU+N2sJ8tdsTmO/te3u9OIpBp6AW8QPXb/gv7mZtlYop/5TFRkMzFZuxotIiLu/Atb9uJmbC5VC5Z0jASOBj5zF1pmrMSWDzned9uJ2JJejexWbLmc1UK334L1vDSyQ4G+vv/PBJ5yFItURzNwE9GqyH8GvoMlUGlxJoXzkcew+bqV6gz8gMIV8e8HXq7C89TC77DvgQt9t/UD7sB6mBe6CEpERETK14otL+Uflj0VS5qlcl8m2KuwBFjLaUTV1ZGeZYD9gZND24Zl/F7We5afIvja/uA2HKmCi4ges1c7jajjFlF4CsWjVXqOQ4o8RxtwSpWep5Z+QfR13ELhCwIiIiJSZw4kOPxzORqWXQ1NWJVbfyPpYqcRVVdHk+WOynKyHL6wsgqt0Zp2hxM9Xp8gvdXNiyXLq4ChVXiOiUWeI63JchNwG9HXcpbLoERERKR9NgCeJ/hlfjdWRVs67lSC7+lCYB2nEVWPkuXqaMKqXofPPUmv/th0Fv9n+gnpHllSLFluwy6wVmIgsKzEc6QxWQboRvT7dQlWIV1ERERSogs2n9T/hf4p8HWXQaVcF6ziuP89vcFpRNWjZLk6jiL6urQua7r9g+DnuRhb7z7NSiXLk6lsNNJZJfaf5mQZYAjRoo9PoeHYIiIiqXMQMIfgELvxQHeXQaVYXAXo/ZxGVB1KlivXj+jau/90GpFU6ktEj9MfuQyoSsLJ8rsEvyfagD0r2P+roX09S/R9THOyDHAk0dd0jNOIREREpEOGAP8l+KX+JrCty6BSqhl4ieB7OZVg5eM0UrJcuRsIvp6llFfsTOpTE/ACwc/0f2RjLeVwsvwqVtXbf9t1Hdz39qH9fAH8mOj5nvZkGeBOot8FXZ1GJCIiIh3Sgq0VuRTvi30ZNjctC42/WtqW6Hy8B0j30pBKlitzOtl6PWJr2Ps/z1XYMkFZEJcs70Q0ye3ZgX1fGdrP34DTiJ4fWUiW18eG5ftf13ecRiQiIiIV2Q54m+hcqw1cBpVCvyXa+EtzdeyjCC7/dFzCz7ce0SWn0joPdC+s6rz/WHiZ9FZKFvM0wc/0FrfhVFVcsgw24sh/+4nt3G9XgqsxtGHV4bOaLIP93fe/rimk+8KpiIhIw+uGFf/yr8k8D0tYpDydsaVjwg3A410GJTW3ITCL4DEwF9jEZVBSsS2J9ipv6TSi6iqULI8N3f7fdu73iNDvT8amrmQ5WR5E9P1UIU0REZEM2J/okig3An1cBpUiA4GPCL5/S4CvuAxKamYtoqM0VpCNgm+N7jKCn+v9bsOpukLJ8lrYMey/SNCeNZfvDe13XO72LCfLAFcTfG13uQ1HREREqqU/9sXu/6KfDhziMqgU2Rqb2+d//5Zic3Ilu9YF3iOaAJzlMiipimZgGsHP9TCnEVVfoWQZ4J7Qfb8sc5+DKZxoZz1Z3pHod0BH5nuLiIhInToOWEDwC/9WLJmW4o4gOKS9DSsANtplUJKYDbDhpeHG//Uug5Kq2YHg5zofW2M9S4oly4eH7ptCeWsu/yT0e//x3Zf1ZBls+S3/6zvUbTgiIiJSbcOAxwl+4c/AkkEp7lRgJdEhud90GZRU3XBseZhww/92VNArK8JJ361uw0lEsWS5MzAzdP9eZezzrdDvHO+7rxGS5XChryvdhiMiIiJJaMIKfc0n+MV/N7C2w7jS4BtEqyK3AeNRIpUFB2KF8MKf7y1AJ4dxSXWF187NWlIHxZNlgCtC908osb9dQo//Aujhu78RkuX9CL6+F92GIyIiIklaD1s72P/lPxdLpJvchVX3jiC6BnMbVjl7LYdxSce1ABcQHWrfBtyAlonJmnDRvm3chpOIUsny9qH7FwG9i+zvqtDjrwnd3wjJcj+CfyOWo4ukIiIimTea6NI49wFDXAZV5w4BFhJtHE4D9nAYl7TfmsAjRD/LNqz3rZy5nJIeXQhOp1hJ9uYrQ+lkmdxt/secVGBf3T6ymUwAABCZSURBVIA5ocfuHnpMIyTLEC0M155K4iIiIpJSg4GJBBsB84HvY71uErUVMIloA3EVNixblVLr32iiczfbsOXBvu0wLknOhgQ/66luw0lMOcnyj0KPeazAvo4JPe4DoqOPGiVZfoLgayxnrreIiIhkxGjgc4KNgZex+WoS1Yvo/Mf89gk2B1bqz3rYurpxn9tUbJkYyaYRBD/vF9yGk5hykuWBRKeUbBzzuIdCj/lZzGMaJVn+J8HXqCUYRUREGswArKCRv0GwEpuz1s9hXPWqGTif+PmubVjhnDWdRSd+nYEfEl1CLb89jJZSy7rdCX7mj7sNJzHlJMsQHVF0fuj+IQSHra8C1o/ZT6Mky9cTfI3HuQ1HREREXNmT6FIhs4ExaGh2nJFE36/8thArINXHWXSNrQkbNfEe8Z/PImAsOq4bgZLloMNCj5tK8Dz4eej+hwrsp1GS5b8TfI3Hug1HREREXOqEJcdfEGwgPA/s4DCuetUVGEd8tew2rJDaWKxgjtTGKGyJl7jPow34L7b+uDSG8DDsrC7/U26y3Jno1JtRufuagPdD9x1TYD+NkixrGLaIiIhEbICtwxwemj0BDc2OMwKb610oQfsIGw7cy1WAGdcMHIT1Ghb6DOYAJ6Jl0hrNUILHwcduw0lMuckywGWhx16fu32P0O3zgNUK7KNRkuUnCb7GPd2GIyIiIvXkIGAKwcbCdOAEtMROWCfgVKJLjYQbn5dgFyOkcj2wCu7h3jD/thS4HJubL42nM7CC4EW/rk4jSkZ7kuVwb/sibMrItaHbryqyj0ZJlsN/z/W3W0RERAK6YUONlxBsNLwO7OcurLrVHTgHmEvhBG4FcDu2DIkuOrTfxsCFFH+PV2LzDdW4lSkEj40RTqNJRnuSZYiOhBmDLR/ov23XIr/fCMlyf4KFHJdhF0VFREREIjYhuqRIG3AX8cuPNLp+wMVEG7Hh7WPgImBrN2GmxiCsQf8sxd/PNuAebF1sEYA7CB4fp7kNJxHtTZZ/GHp8+PffpfiUhUZIlr9O8PU97zYcERERSYNRWK+yvxGxHBiPhrrG6Q/8FBu+XirJex34Cbr4kNcPW6rlPoJDaeO2JdgwUiXJEjaW4LFyu9twEtHeZHkAhQsTtmGjY4pphGT5jwRf3xVuwxEREZG06AScDHxGsDGxABuyncU5gZXqjC1n9Aylk+Y24APsAsRoGqcwWDM2RHYs8CDFG/P5bQa2TNdgB/FKOoTn6C4ge3+j2pssA9xJ/Dm1ElinxO82QrI8ieDrO9htOCIiIpI2fbBEZTHBRsVHWI+gKg/H+xJwHdE5gsV6TR8Azs79bo9aB5yQVqwn+GRsfvEMyns/VmJTAo4ne0mPVF8Ttp6w/xg60mlE1deRZPlg4s+v+8v43awny7sS/Ruclb+7IiIiUmPrYstK+YuhtAGvYD2jEq8b1mifSHm9qPltBdYYvgr4FrAl0KXGsbdXM7AecChWnOu/RNfzLrW9BJyJepGl/S4heCw97DacqutIstwJ6z2dHdqOKON3s54s/43ga7vDaTQiIiKSCTsATxBtRD2FVX6WwtYAvov16oR76stNoCcB92KJwWnYmqDrUrtEugkYiB0HxwK/Am7FKu+WKnQWt63CEuRfAcNr9BokmzYnenxt5zSi6upIslyJLCfLa2NLzvlfm1Z+qJCGmYmIiJgmrGfifGDD0H0PAz8Dnq51UCmzGjbcer/cNrQK+5wDfAp8jhUbm5HblmNzOFdgSfoSrJd7ITbPujvWK9w7t58+2GfcHysSNABY0/f/lirE+SBW1Ou+XKwi1fA4MNL3/38ChziKpdoWYSNV8l7DRpwk5TTgytBtp2J1FtLuUuB03/8nYUUXV7oJR0RERLKoGRuC/T7RHogHyVavTtI2whqi1wJvYo229vbS1us2HSs0dA42T7DSZFukkK8SPf52dxpR9ahnuTqGYhcM/a/r204jEhERkUzrjA0v/oToENvb0PDajugN7I0tSfUvYArR+eL1uM0EHsPWnz4cGyIuUo4WYANgX+B72Lz364H/AG8BH2LzbZdioyTy82/fAR7B5qCOAP5H8Jh8ESs0l3ZKlqvj3wRf0xTsO0xEREQkUd2w4kzhiscrsXVPt3cXWiZ0wypLjwbOxRKJ/2E9t6XWKK7mNhtbL/oOrFL6icAu2JrJIuXog823PwMrHPgK0TmkHdl2A3YmemHpp7V5WYlSsly544m+psOdRpQhmrMsIiJSnh7AGCxx7hu672EswXqo1kFlXBM2n7g/VoBrEN4c42agJ9a71g1biik/Vzk/d3kVMC+3r7lYI3Iu3hzoz3I/z8j9jkh7bIRdUNkFG4q/KXZctsdcLKFemPt/fp59d7yewX7YxZwbgaN8v7sM2ANbAz2tNGe5MkOB5wh+Jz2OHRdtTiISERGRhtYHOA9rvIav5j+HLTHU3gaziNS3zlhSfBZWYOtTyusV/hCrdXAl8EPgAGw0yloUn+fenHvMTr7b1oh53ulYFeS0mkZw+acnEn6+E4kuOXVCws+ZlJ7YxQX/8bAQGOYyKBERERGwhspZROc0twFvY40yzRkTSacmYGtsJMk9lF5bexU2B/labATKl7ALa9V2ENHh2M8S7J2V7GvG6j6Ej8PTi/2SiIiISK11waqOvke04TIVa2wn0WgWkepaHzuXbyZaoyC8LcaGu14A7I/1+tbKr2Pi+TuaXtlILiB6DFzvNCIRERGRIlqwoiovEm3ELMCGYW7mLDoRCeuNrVc8HluTtlhyPBdbKuxMrNiWy1EjzcBdRGO8Ck0BaQRjiX72L2Dr3IuIiIjUvZHAROIb3U9glZ+zsOyLSNoMx5KNByleqXo58DzWgzcK6OQi2CLi5qu2YUXA9Lclu84j+plPI93z1kVERKRBbY8NjYtrlE9CQ7RFktYPOBJbr7hYUa6VWO/c74B9SEcv3VAsUQq/lpupv+ReKtME/IHoZz0PLV8oIiIiKTcQ+DnxDdsvgD9jxYREpDIt2DDpcdiSSsXW6p6OFeQ6gvSup70R8BHR1/YA6X1NErQaNic9/BnPAXZ0GJeIiIhIVXXChmA/SHzj/Q1siGgtiwWJpN0A7LyaAMyicHK8AhtaPQ4YQXYKYg0B3if6ej8CdnAYl1RuXWxJwvBnOxv1KIuIiEiGbYsVFlpEtCG0BLgVq7JbbC1WkUbUitUFuABLfsNLKYV7jydgyXSWpzysA7xLfNXukxzGJR23H5YUhz/Tz4AtHcYlIiIiUjMDgB9jazPHNfYnY0O413UVoEgdGAZ8H7ib4mseLwLuA34IbOokUncGAo8R/75cQ7YvFmRJN2zu/Eqin+NrwIbuQhMRERFxZ1esUbuA+AJEDwAnokavZF9P4EBsybVSyzq9iRU/2hdLNBpZK/Fr8OZ72Q9zF5qUYVfgLeI/v5uA7u5CExEREakP3fDmNscNMV2Ru+84LKkQyYINgDHYsb2EwsnxF7nHjMHm60rUURTugZ8IDHYXmsRYDbvIEdebvByrZZGVOfYiIiIiVbMp8HusVyiu4bsQuAU4GOjqKEaRjhgIfAO4jsLHd35UxbPA+dhcZa0jXJ6tgNeJf09nYUvX6W+GWy3YaKGpxH9OHwJ7OItOREREJCVagL2BvxJf9KUNmIslHgej4XpSf9YADgX+D6v8Xmxo9XTsWD4KVYevRCesV7JQT/1U4GR0AcKFUcArxH8uq7ACkBo5JCIiItJOLVhDawIwn/jG1mK8oapruwlTGlwP7DjNV62OG2Lqn1qQxWWd6sUWwP8o/P6/ARyE3vda2BNbA7zQZ/E2NoJCRERERCqUn998O5YgF+qleA74GbC1mzClAfQCvgZchCW+KyicEKzChghfBhyAJdaSrBasSvhcCn8ubwGnYnNopXo6AUdj50Wh934x8Gs0NF5EREQkEb2AI4AbKDxUOz8PbjxwOBriKh03FDgWq1j9CsWT43yP2Z+w426Ag3jFrI719set8e6f0nEptn6zdFxvbHTPRxSfk38rsL6jGEVEREQaTgs2lO8CbGmdYknMJCx5Ho0l3CJhrdjw6DHY8P8pFD+m2oBpWBJwMlonvB6tgy1VV+wixzLgNmyIdhc3YaZOC7APNud+IcXPkTtovPXARUREROrOZsDZwJMUnzu6BHgEOAdb81MN5MY0CCsUdxF2zBRbyim/vQf8DTgeJcdpMhz4O5YYF/t8ZwN/AXYHmp1EWt+2Ay7BLhIVex9XAncBO7kJU0RERESK6Umw8FKxht3y3GMuxdZ1HuIgXknWWsD+WHGtidhIg1KJcfi4UHKcfoOB3wAzKf35f4h99vthdRMaUSfgy8DvsCkGpd6zBcDlwEYughURERGRjlkP+BZwEzCD0o2+ScD1wHew3hQVpEmHVmBzbJ7xH4D/AHMo/Xm3YUs53Q6cAewMdK5x7FI7q2FFvkpN38hvi4B7gdPJfiK4Djat4A4Kr0QQ3qYAPwL61D5cqZTKw4uIiIhfE7AV1mOyK7ALsGaJ31mBVdF9Kbe9nPt3XnJhSgkDsKH3m2IV0LfBlg8q58LGEqxS9bPA08BTwAfJhCl1bjh2ceUEYGCZvzMdeMG3PYFdlEmbTsAw7O/gSGzO/qaUlz/NA/4F/AO7kLAioRglYUqWRUREpJT1saR5Z6zhuAVWxKaUD7Ck+U0smX4XeAf4IpkwG04rNix+Q7zEOL/1K3Mf8/AubuQvdLyJGvcS1IoVrToaK/bVnqWlVmLrOD+LDVd+G/tbMJn6OM6asfNoGLAxsAk2YmZrLGEu1zIsMb4Bm86wpLphigtKlkVERKS9egA7YsnzNrmtPcueTMWS5nfxkugp2PIqamAG9caGfq6PLdW0YW4bijXw29OYn46XFOe3ydhQUZFydQf2wuYrfwWbxtERy7EpHe9gF9amA58Bn2NFsmbkfq4koW4G+mMjLQZhveP5n9fHkuNhdLyQ4afAfbntAdLZgy5FKFkWERGRauiDN9w3/++mWI9Ue0zHkub89iGWSH+C15BeVpWI3eqENdoHYg33AViRrLVz27q5rWcH9j0P6717I/fva1hi/FnFUYtEbYKXOO9GdQt+tWFrPi/GLqQtw5ZjWonNGc7rgZ1T3bCpBp2xpL4X5Y2CKddy4BmsB/k+bCSGLjZlmJJlERERSUpXbM7jxljinO/F2ZjKioLNw3p0Ps9tn2G9UPOxirNzcv8u8N02Dxv+vbyC5/Xrgg1FzTfOe+T+38e39Q39nO/hGgCsUeHzr8AuJEwC3sd65/LD3T+pcN8iHdWKnd8j8Ob6bkJ6l5rKz79+AltC7QUscZcGoWRZREREai0/RzA/P3DD3P/Xy/3bkd7U9vInzgsp3lvdE6+HvJXaxAfWUP8YG7b+EZYU55PjD6le4i+SpD7ADlitg2F4F8xKFQ6spZnYBaf89JD8HOsZLoMS95Qsi4iISL3piw1B9ifQ/vmG/XNbNYdX1tLnWCN8BpYQz8B6xz/BkuKPc9tSVwGK1EAvvOR5Xez87k/0XK80X5mFN4Ujf759TrB2wuwKn0MySsmyiIiIpFETNpS5v2/ri/X6+jf/bV2xXu3euX20YA128HqM83Mi4+TnT67AhnYvxdaYXZT7eUHu/kLbLKyRrh5hkfLkz9HwXGT/eQzeSJHw3OZqTr2QBvT/AGnP+UHEeDGeAAAAAElFTkSuQmCC" width="65%" style="display: block; margin: auto;" /> --- ## 4. Repeat identifying links, find ambiguous, delete indirect links <img src="data:image/png;base64,iVBORw0KGgoAAAANSUhEUgAAA8sAAAI0CAYAAAAqQrvXAAAABHNCSVQICAgIfAhkiAAAAAlwSFlzAAASdAAAEnQB3mYfeAAAABl0RVh0U29mdHdhcmUAd3d3Lmlua3NjYXBlLm9yZ5vuPBoAACAASURBVHic7N13vBxl9cfxzy3pjXQS2gUhhI4JVUKJhA6CQEQUI4qiAoII/hAEXaQFVKRZsECoUlWKIOTSQ5UmPQiEmpCEhPR+d39/nF12drbvzs6zs/t9v17zSmbr2buz9+6Z53nOaUFERERERKQ59QfWB4YDawNDk/+m/j8UaAUGJm/fF+gG9AB6A3FgYfK6hcn9ZcBKYCnwMTAbmAPMSv47F/gQ+AhI1PLFSXVaXAcgIiIiIiJSQ23AhsCmnm1U8t8RDuNaDkwH3kxubyT/nQ4schiXJClZFhERERGRRrI2sD0wNrntQnpkOCreAR4Hnktu/8FGqyVESpZFRERERCTK1gP2BSYAO2HTqhvNMixpfgy4F3gKWOM0oiagZFlERERERKKkO7ArliDvC2xZxWOtBmZg64o/Tm5zSa81ngusApYkb7sSS1y7sKnSqbXLrcCA5GP2x6Z+D8LWQg/FpnsPS24jsIR+UBVxLwCmAv9ObjOreCzJQ8myiIiIiIjUu+5YYvx1YH+s0FY5PgFeIb0+eHpym4G7EdohwGjSa6hHJfc3wZLtUiWAF4GbgBux4mEiIiIiIiLSwMYCl2KjvIkSt9XAq8CVwCRgC6I1SNgXGAecBFwLvEvpr70LmJa875CQ4xYREREREZEaGgqciRW5KiVBXIMliD/D1iz3CD/kmtsA+AZwA9Z+qpSfy3LgFmzKuoiIiIiIiETUJtgo8lKKJ4KzsURwEtGrdF2tVmzE/TRs3fIqiv+8ngeOBXo6iFdEREREREQqsC9wHxCncML3CfA74AtEa1p1rQ0HTgSepnjS/BFwOtUVFxMREREREZEa2hl4hOJTie8CJmJFvqSwDbAR5+kU/rkuBiYD/dyEKSIiIiIiIn5bAXdTOJmbAZxMuh2TlG88cAdW9Cvfz3km8D2gm6MYRUREREREmt46wBQKJ2+PAYdRXuskKWxj4HJsNDnfz306cKirAEVERERERJpRC1ZcagH5k7V/ANu7CrBJrIVN0Z5H4fdhpKsARUREREREmsVGQCf5k7Mngd2cRdec+mJJ8yJyvycLsD7Nra4CFBERERERaVStwE+AZeROyF4GDnQWnYCNIF8JrCb3e/QAsKGz6ERERERERBrMQOAe8ldhPgGtSa4nWwBPkPv9mg/s5y40ERERERGRxrAN8Da5E697sdZGUn9S68pzTc2OY22mNC1bRERERESkAkcCS8hOtj7FErEWd6FJiTqAf5P7ZMfdWJEwERERERERKVGM3AnW46i6chT9mNxrmV9F76eIiIiIiEhJfknuRPlKoLvDuKQ6uwKzyH5fZ6DCXyIiIiIiInm1AL8lO5laARzjMC4JzjpYey//e/we8DmHcYmIiIiIiNSlFuAP5F6fvJPDuCR4PYB/kP1evw9s7DAuERERERGRunMmudsM7eAyKKmZNuA6st/z6VirsIakinQiIiIiIlKOfbHKyN5eyXOAvYCXnERUmXWwUdNcFgGfBPQ865J/7fZCYF5Az1NrbcBfgW/6Lp8K7A+sCT0iERERERGROrEZluB5RxfnAqNdBlWh58hdmCwBPBXQcwwAlhV4nosDep6wtJJ7hPnXLoMSERERERFxaQDwJplJ0ipgvMugqlAoWU4AWwbwHMcVeY6oJctgo/GPk/1ajnIZlIiIiIiIiCtTyE6QjnMZUJWKJcuTA3iOZ4o8RxSTZYDhWIEv72tZCKzvMigREREREZGwHUh2ovcnpxFVr1iyPAtor+LxNy/y+FFOlgG2I3uK+X1OIxIREREREQlRD+BtMpOil8lfHCsq/MnyGmz9tfey/at4/F/5HutjGitZBjie7Nd0qNOIREREREREQvJ/ZCZDq4GxTiMKhj9ZXgVc6rvspgofux2Y6Xssf/LcCMlyC/Agma/pbaJ/IkVERERERKSgPmSPtl7mNKLg5EqWx/guWwkMruCx/dPWP8ZaazVasgxWCX0Vma/re04jEhERERERqbGfkJkEzaey5LEe5UqWAV70Xf6DCh77Nt9jXIRVDW/EZBngcjJf1ztUt95bRERERESkbrUC75KZBMUcxhO0fMnyyb7Ly+25PBhY4XuMrWjsZHltYDmZr+3LTiMSERERERGpkX3ITH6W0jijypA/WR5G9rTizcp43BN99306eXkjJ8sAV5L52u51G071Wl0HICIiIiIidenrvv3bgHkuAgnZHOAe32XfLOP+R/v2r6kqmui40re/N3biQUREREREpGG0Y4mxd6RwvNOIgpdvZBngEN91HwFtJTzmlr77eQuENfrIMmSv9/6W23Cqo5FlERERERHx2xEY5NmfBzzqKBYX7gZme/ZHYiOlxRzj2/8nzTEan/JP3/5+TqIIiJJlERERERHx29m3fx/Q5SIQR9YAf/NdVmwqdjtwpO+yKUEFFBH+6ev+4yhSlCyLiIiIiIjf9r79citCN4KrfPsHAwML3P5AYLhn/2NgatBB1bkXsUrgKetio/KRpGRZRERERCQYpaxpjYpRvv3nnUTh1stY8pfSE/hqgdsf7du/BhuhbiarsJ+b16YuAgmCkmURERERkWAMAW4Bnk3+Oxk4FpgAbES0vntv6Nt/20kU7k3x7eebij0M2N93WbNUwfZ7x7ff4SKIILS7DkBEREREpEHMBr4GxIDTyU6Ol2FJ59tYQuH9911gdUhxFtMdGODZX0Nmsatmcj1wIdAjub8j1nP5dd/tjgK6efafznGbZvGBbz+y7aOULIuIiIiIBGcNcCZWOfo6MhOF3sBWyc2vC0sy/Ml06v8Laxdylj6+/aVYG6BmNA8rWvVlz2XfBH7qu90k3/6UGsZU75b49v3HU2QoWRYRERERCd79WFJ8PbBXCbdvw6ardgB75rj+UyxpzrW9C8SrjNeru29/Vc5bNY8pZCbL3wB+Rro6+FhgG8/1K4CbQ4msPq307ffIeasIULIsIiIiIlIbc7A+s7/Akqtq1iwPxJKysTmuW0H2SHTq3xlkJy/FLPft9yrz/o3mHqyy9drJ/VTP5XuT+0f7bv9P7ORGs+rt21/mJIoAKFkWEREREamdLuDn2LTs68lsLRSUnsDmyc0vDnxI7mT6bXIndalp1y3J/d7Yetx6WVMdtjXAjcCPPZd9E0uWu5NdIXtKOGHVrbV8+/5p2SIiIiIiIhmGAfdhiWi9bPPJXb17tu92G9Xg5+Hac2S+xkLTzbf03XY5Ntp/uO/yD8nfQmw82T//i6t9EXXoTjJf4xFuwxERERERkShoAU7DRitdJ8rlbPck456ITQXvG/QPxoFykuVct/8BcLfvsvML3L9ZkuXXyXyNO7gNR0REREREouSLwCzcJ8GVbqnp3Y8AV2MVwI8gWuuby02WT/Dd/nVsarr3stEF7t8MyfJa2NKD1OvrAvo7jUhERERERCJnbaAT94lvtdsS4Apg42B/PDVXbrI8CCumlu/n8ESR+zdDsnwAma/vJbfhVKeainwiIiIiIlK5j7Gqyr8g3YYoSmZhVb7Xx0Zd33IbTs3Nx6Zd5zMlpDjq2YG+/cecRCEiIiIiIg1jPDAT96PEpWwvYe2S/P2Yo6bckWWwZDDXz2Q52VWg/Rp9ZLmd7KUF+zmNSEREREREIq8d2I7sBK6etgXAQaRbSkVdJclyO7lPatxQwn0bPVn+MpmvbSHW1iyy1GdZRERERCRcGwBbYe2IUv+Opv5HavsD72KJULNaAxwGrOu7/DkHsdSbH/r2/4at8Y4sJcsiIiIiIrXRCmwCjAE+n9zGYIWi6t1i4FJgH2D75GUtwBnAka6CqhNPug6gDu2MjZx7/dlFICIiIiIiUl/agS2ASViSORWYh/up0+Vu07H+wb2Tr+sQ3/VdWJ/lRlDJNOxqNPI07Klkvq5Ot+GIiIiIiIgL3chMjKcBy3Cf6FazPQIcTHa3nBbgv77bPkxjrFtWshyMQ8l+XV90GpGIiIiIiNRcH2A34MfAdcAr2LpV18ltEFsXcBc2hbaQ/XPc95gSf371TMly9QYA75P5mqY6jUhERERERALXRvaI8UrcJ7VBb4uSr2+DMn429/oeYwHQUcb965GS5epNIfP1rAY2dxmQiIiIiIhUbyTWCimGjbDOx30iu7CGjz0z+VoHVvCz2hAr+uV9vBexkfeoUrJcnePIfj3nOo0oYI2w1kBEREREpJh+WB/jHZPbDliy7Mps4KXk9irwMtALK4wUdAupF4BLsFY+q6t4nBOAy32X/R04HEuUomYCmScO4sDtNXy+YcDuvsvexNaER8044AEyj9XXsWrvkW4XJSIiIiLSyNqAbYHvA1dh64y7cDdaPANLKs8CDiB3kr4W8E6Az5lajzyhgp9fPi3AzTme62cBPofUvw2AOWQeA4uxnuEiIiIiIlJH+mAjXafhfjr1zGQMMWyK97AS4m/Bkukgnn8FcC2wWSk/uAr0Jbs6dhfwrRo9n9SXEcBrZL//B7sMSkREREREzNpYIjoZd0W4VmPTp68FTsKS9b4Vvp6TA4hnNpagD6kwhnLkGlmMAz8M4bnFnfWwaeOaWSAiIiIiUgdasArVx2KJ6duEnxivBJ4GrsBaJo0huHXF21Ndsj8dS9Z7BRRPqfbACmL5E+aTQo5DwtFB7s/ezagOloiIiIhIKLoBY7Gk6xZgLuEnx6mp1KdhI8a1SkSrWac8DRtdd5moHIxN+/bHdqbjuCRYWwDvkf0+3wH0cBiXiIiIiEhD6wPsDZwPPE74U6rnAv8CfgHsBwyu7cv9TCXrlFcAV1NfhZT2BZaRHeudwACHcUkwDsZ6avvf31uxE1siIiIiIhKQXqSLcU0l98hkrbZV2DrjK4FJ2IiZqxHQk/LEmGtbAFwKrOsk0uJ2BxaRHfd07Gcs0dOG1QSIk/2+3gS0uwtNRERERKQx9MTWt8aARwk3OX4TuAY4HlsbXC8jYdtR2s/hbSyp7uMmzLLsCnxC9mtYCEx0GJeUbwTwILmPyT9iibSIiIiIiJSpHVtznBo5Xk44iXGqOnVq1Hj9Wr/QCg2geJGyx4BDgFZHMVZqPawQWq7XdBewjrvQpEQTyX3SYwUq3iYiIiIiUhZvcnwXNpIYRnK8GEvGY1ihq6isj72N/Mn+TdgIeJT1AP5C7tf4KVbVXMW/6k8HcD+537cPgZ2cRSYiIiIiEiFbAD/CCmMtIZzk+F3geuA4rMBV1EZdAU4g9zTli7HexY3kx0AXud/L+4BN3YUmHt2BU8j/OX4IGOYsOhERERGROjcEm555JblbyAS9rSFzSvWGtX+JNbc1mVPSZ2Kj4gMdxlQrW2LrxQu9x11Ya7AONyE2vVbsM/0Wud+fZdhsEa1PFhERERHx6AnsCVwIPE/uirhBbquBp7Dqu/sC/Wr/EkPVD3gDe60vYCcA6qXYWNC+SuYo5TKs3dVicr/3S7H3fS0XwTapCdjnOt/n8VFglLPoRERERETqzEbYetJbyN0GqFYjxxNpzNFVr+ux9dUHuQ6khtqxpNf7Pr9Juif0xtiU3nzHxDzgHGDtUKNuHm3AoVgv83zvwafAMWhNuYiIiIg0uaGkp1a/T+2nVr9NOjkeFMLrqxcjafxRuqHAA2S+33eRe7T4IKz/cr7jZCV2wmbHmkfdHPphJ8FSMxtybauwz6ZOVIiIiIhIU2oDxgHnAy9S26nVceC/wCVYC6RmSo6bzS7AR2S+95MpXICtG/B9bN12oePoIeBwrMK2lGcLrHBcoer0XcB1NEZNABERERGRsgwBvg7ciE1zrWVy/ApwOXBY8nml8R2LjQSnjoNPgL3LuH9v4AxgPoWPr0+BPwO7E80q6GEZiVW2foHin9e7saJzIiIiIiJNYwusiu1UbHplrRLkOdh02WOB9UJ5ZVIvegJXkXk8PE/lI5R9geMpXkE7gS0ZmAxshxJnsLZO3wI6yd+qK7Utx3pgb+kkUhERERGRkPXGqtteSm3XHi/DEvDTgLGoCFCz2hibYu89Nq4FegXw2K3YsXwXpS0TmEv6hM2IAJ4/Ctqwz99pwDSKJ8gJYDZ2gmGkg3hFREREREKVqlx9F7CC2iXIqaJcB2GjidLcDiBzyvQK4KQaPdc22AmgjyntWO0CngbOA74EDK9RXGHrDuwA/BC4meJT1lPbGuA+4Ci05ltEREREGlg3bMTtEuB/1C45nglMwdY5N0qyIdVrwUYyvaOYHxBOxep2YD+s/Za3f3Mp2wzgJuBkrBBZvfdybscqpx+JfdafpPyTYc8CP0KVrQOh6TMiIiIi9ak3sCdwIFZRelgNnmM51ne1M7k9j33hFkkZBNwA7Ou57BHgCGx6b5j6Yp+Fg4G9gAEVPMYcrI3SdGyN9BvAW1hF78XBhFlQd6zVVgcwGkuORyX//znsxFg54tjn9h7gb9jrkYAoWRYRERGpH0OA/bFexHtRm+mTM7C1x3cn/11Rg+eQxrAtcDs27R/sRMrlWMXlNa6CSmoHvoAl8fth07arzW1WYMn0rOS/c7AK3wuxpHQpVjRvBXaiKaWVdOI+ILnfF+iPneQagSXIw4HBVcYIVtn+fuBebKr1nAAeU0RERESk7myErfsstVhPudua5GOnCnOJlOIoLDlMHUeLsJ7H9WoE8A3gCuAZalsJPuztfeA24FRgJ6zIl4iIiIhIw2nFktYY8Cq1+XI9B6tQPJHKpqpK8+qBFdbyHk9vAJu7DKoCPYGdsfW7fwNeI7MndL1uM7ElERdgU85VwdohTcMWERERqb2ewDisqvThBP8FOA68gH3Jvhtbh5wI+Dmk8a2DjWDu5LnsDmASNrIcde3YWuFNyVwv3IFNkQ6i/VUxcSwHa8FGvy8icw11I/ycG4aSZREREZHa6Iu12jkcW1PZJ+DHn4+tW/wX8G9sbaVIpXbDWhOlqiivAc7EkrlmOfHSFzuRNRRbazwSW3ecaz1yN+wkWGrN/yJsGYV3XfMirBf0bGwd9Nzk9gTWDork88yt4WsSEREREakL/YCvAX8HlhH8FM3pwIXYKLXWLUoQWrA18941vnOwSuxSG1NI/6x3cxuKiIiIiEjt9MamV19L+X1gS9lexdY3qziXBK0vcAuZx9t/gPVdBtUETiP98/6e41hERERERAK1FraO8y5sumWQyfEKrKXTSdgaUpFaGAW8QuaxdyXWB1hq60ukf+aXOI5FRERERKRqA0knyEFX1Z2PjfBNwtZIitTSl4AFpI+/5cC3nUbUXDYh/bO/z3EsIiIiIiIVGUTtEuT3sJG8g9BonoSjDZiMVWT2HofbuQyqCbVhJygSWA9lEREREZFIWAv4FlZpejXBJsgvAmcDY0J7NSJmCDa933s8/gubMSHhewl7D+JYYUARERERkbrUk3SRrqXUpkDXqLBejIjPWOBd0sdkHBthbnUYU7O7mfT7oZF9EREREakrbVgLpiuBhdQmQd4krBcjksexZC4hWAgc4jQiAfv9kHpPJrkNRURERETERtLGAZcCs6lNgrxxWC9GpICewJ/JPEZfADZyGZR85gjS78sFjmMRERERkSa2BZbIvk1wyXEXMA21eJL6sx7wDJnH6/VYT3CpD1uTfm/+6TgWEREREWkyHcBpwBvUJkEeGdorESndfsA80sfsauxzIPWlJ7AGe4+mO45FRERERJrACOAU4HmCS5DXAA8APwCGh/dSRMrSgiXFXaSP3Q+BnV0GJQX9j/QJjR6OYxERERGRBpSqZH0LsIpg1yCfhiXgIvWsP/B3Mo/fR4C1XQYlRd1J+v3awnEsIiIiItJAxmKFuj5BRbqkeW0NvEXmcXwl0M1lUFKSC0m/Z4c7jkVEREREIm59bLTXnxxUs72HJd1jQnwdIkH4GrCE9LG8GPiK04ikHEeTfu9+7jYUEREREYmiAVgf0qlAnGAS5E+w0bdx2FpPkShpByaTeUxPB7Z0GZSUbUfS79/fHMciIiIiIhHRBkwArgWWEkyC/Gny8Q7Ckg2RKBoGPEjmsX0nsJbLoKQi/UmfAHzRcSwiIiIiUuc+D1wCzCaYBHkpcBNwMKo2K9E3DphJ+vheg62xb3UYk1TnI+y9XI6dJBQRERER+cwg4ERsZCWIBLkL6MSmbvcN8XWI1NKxZFZ7nwvs5TQiCUIn6fd0I8exiIiIiEgdaMVGya4kuGnWr2OjbBuG9zJEaq4XMIXMY/05oMNdSBKgy0i/rwc4jkVEREREHFoHq2b9NsEkyPNRoS5pXJsAL5F5zF+LJdDSGH5A+r091XEsIiIiIhKyHsBE4C5sjWW1CfLK5GNNBLqH+DpEwnQgVpQuddyvAL7jNCKphT1Iv8d/dRuKiIiIiIRlc6y9zVyCGUV+FjgJGBrmixAJWQs2+6KL9LH/PrCDy6CkZoaTfp+fdByLiIiIiNTQAKwQ0TSCSZA/BC4Ftg7zRYg4Mhi4j8zPwINYuyhpXJ9g7/WnrgMRERERkWC1AOOB67H2J9UmyEuwgkbjUUscaR6fB94h/TmIYzMz1E6o8T1O+n0f4TgWEREREQnAWtgo8ssEO816UJgvQqQOTAKWkf4sLAQOdRqRhOkvpN/7LzqORURERESqMJbgWj6lqllvE+orEKkPPbBlBt7PxOvAZi6DktCdQvr9P95xLCIiIiJSpv7YKPKLVJ8gdwFTUTVraW7rAk+R+dm4EejjMihxYj/Sx8AVjmMRERERkRKlRpGXUH2S/D62BrMjzBcgUod2Bz4m/dlYjVXAlubUQfpYeMBtKCIiIiJSSD9sFPl5qk+QlwO3ABOwQmAizawFW5e/mvRnZA5ap9rsWkmfkJzpOBYRERERyWFzbP3kIqpPkl/FRsoGh/oKROpXP+BWMj8nj6Hqx2K8JycHOo5FRERERIBewLeBp6k+QV6ArbdTT2SRTJtiJ5C8n5cr0Zp9SbuB9LGxs+NYRERERJraOkAMmEv1SfKz2LTtvmG+AJGIOBg7keRdmnC0y4CkLp1J+hj5tuNYRERERJrSOGwNsXfNZKVrka8FxoQbvkhktGMF7eKkPzf/A7ZyGZTUrcNIHye/chyLiIiISNNIFex6hepHkV/D1iIPCvUViETLUKCTzM/O3WgtquS3GZnHioiIiIjU0ChsZOtTqkuQV5CuaC0ihW0HvEf68xPHPoetLoOSutcNWIUdM287jkVERESkIbViSe1dZE7/rGSbjipai5TjWGAl6c/QJ8A+TiOSKHkdO266gN6OYxERERFpGGth/VtnUF2CvBL1RRYpV0/gr2R+lp4HNnQZlETO30kfP9s6jkVEREQk8rYHpmBTpatJkt/FRpGHhBm8SANYH/gPmZ+na9HIoJTvPNLH0JGOYxERERGJpFbgIGAq1RfsmgZMxCr3ikh59gfmk7m+/ySnEUmUHUX6WPql41hEREREIqUfwUy1XgRcCWwZbvgiDaMFm4nRRfpz9QGwk8ugJPLGkj6ebnUci4iIiEgkbEQwVa3fxL7gq32NSOX6A/8g87P1MDDcYUzSGHqTPgHziuNYREREROraOKzY1hoqT5C7sOnaB6GCXSLV2hZr65P6fMWBS7G2PyJBeJd0sUUtjxERERHx6I6tIX6K6kaRP8W+xHeEGr1I4/o6sJTM5QyHO41IGtG9pI+xTR3HIiIiIlIXhmJTpD+guiT5OazXqyrxigSjHVsG4f2cvQFs4TIoaVgXkz7ODnEci4iIiIhTWwJ/BpZTeYK8CrgR2DHk2EUa3TrAE2R+3v4JDHAZlDS075I+1k53HIuIiIhI6FqAvYH7sTWPlSbJ84ALgHXDDV8c6YEVZxuMFX3bCFtDOxbYKrm/YfI2A4E2N2E2jN2AWaQ/b2uw2R9a+y+1NI70MXet41ianhaNi4iIhKcbNq3uJ8D2VTzOW8AVwF+wNZQSXd2wBHcUdtJjODYlfwQwLLmNBPpW8RzzgNnAXGBm8t85yf9/iFVJfx/7ci7mWOwzlirc9QlwJNDpLCJpFq97/r+ZsyhEREREQtIX64/8HtWtR56GFf/SiGH09AK+gE2x/BVwJzAdm0JfzTER1LYMeBGrvn4ucBSwDc03sNIXuJnMn82zwAYug5KmMxs79hajmQwiIiLSoIYDMWA+lScxK7GpeFuFG7pUaSR2YuNS7CTHCtwnxJVsS5LxXwpMIrpFrTYv4TajgJfJfP1XYhXqRcL0MOljcD23oYiIiIgEaxMsuaimaNdsrALvyJBjl8psAHwfK/5UzcmRQtua5GPPB2Zg/X5fxkY+X0vuf5C8/tMaxZDApm5fg01LHhzED6/GBmKjxYUcRObPbDlwTI3jEsnnD6SPxb0dx9LUmm1qjYiISC2NwwoAHUDlU+f+C/weuA77wi71qQdWAGpfYD+qW1v4ETYl+z2soNSc5Ob9/ycVPnYbtgZ6GOl10EOxkzBDsdHUUcCgMh5zHWyUeRLQBfwH6w37byxxj1cYa61cjY3s59IGnJXcWpOXvY/1T/5P7UMTycm7bnlzrBikiIiISOS0AUdgX6wrHanrAu4Axoccu5SnDZgATAEWUv77/BZwEzY1/0isinW/EOMvZAi2pvrb2IyGO8msBF3q9hHwG+Dz4Yaf13FYXBfluG4IloR447+H8k4ciNTCXmQuBRARERGJlB7YyNp0Kk+SU+uRS1lPKe5sgSW4Myj9vV1Eeq3vRGxEN4pGYlOUY8BUrBBYqT+D15L3+1zIMadsQXqt+HG+68aQ+X7GsZMErYi4ty7pY/NRx7GIiIiIlKw/Vtm6klG31LYQS6LWCTl2KV1P4DvAS5T2nq4GHgFOx0ZVGzXp6g7sgSWWL1Jan/A41nLpIML7ufTCprKmYjjQc90kMpP+hcCXQ4pLpBQtpGevVLoEQ0RERCQ0w7BRsgVUniTPwNY0Dwg3dCnD2sA52DrhYu/nTODPwGE073s6Epu6fQs2ml7sZ/YmcDzQp8Zx/dH3vFtis0H+7Lv8RdyNfIsU8gzp43SI41hEREREZPuR3QAAIABJREFUctoQGwUuZ/qpf3sBG81SYc36tQVW4XklxadXT8HWLqvfdaZewFeBuyjeP3o+cCHWXi1oh+Z4vs2Ap32X3UDtk3aRSk0hfazu6jYUERERkUxbYeuJV1N5kjwNm3oq9Ws94CqsLVOhKdZ3YYlgLzdhRs4QbAT5CQp/RhYDv8SWNwRhXawFlPf9XIK1YvO+n6cF9HwitXIa6WP2WMexiIiIiACwC5YYlbIWM9eWKtq1ZdiBS1kGYetuC80YSK0t38BRjI1iW6yib6Gf9SdYctCziudpBR7Gqst7H9v7Wf4Iq/otUu++RPq4/a3jWERERKTJjcOS5EpHkRdhidV6YQcuZWkHTsFGH/O9l6l1tX0dxdioRlB8PfgMrMdxJX5R4HETwP+Ar2PV53tX+iJEQrIJ6WP3PsexiIiISBNqBb4CPE/lSfJHwKkEN41Uamcb4Fnyv5evYlWRG7WSdb3oBZwAfEz+9+LvWHJdql3JHlEuts3DPvu3Y72ht6juZYkEqo1067P3HcciIiIiTaQ7cAw2glhpkvwWto6sR8ixS/m6YVN88xXv+gB7L1WALVx9sPclX4X5Bdj70lLkcQYCH1J43Xm+bT5wEbBxcC9LJDAvk15K0M9xLCIiItLgumNVqd+i8iT5JVTZOkq2J7Pfrj9R+gkq2uXaUGwJQ76TGZ1Y4a58bqb0GgOp0efnsURc773Us1tIH7vbOY5FREREGlQP7Ivxh1SeJKcqWxcb5ZL6MYn8RaXuwnoES/3YBCvQlev9mgvsmeM+389ze++WGnFeghUa27qGr0EkSDHSx/E33IYiIiIijaYPcBK2rrjaJFmiowc2Upnr/ZyJ9eGV+tSCndhaTPZ752/5tDmF+2L7R5HVU1mi5gjSx/P5jmMRERGRBtEXS5JnUVmC3IWNPO4QduBStXWBp8h+T+PYqOIAd6FJGTYEppL783kDtk4510mw1HTsldgU1l3CDlwkQFuTPrb/4TgWERERibh+2MjTPCpLkldhPZJHhx24BGIUVqzL/74uxqqeS7S0YJ/nXFWuZ5M7Sf5f8j6DHMQrErSepJcRvOE4FhEREYmowdjarkK9cwttS1CP5KgbTe6Rxv8BWzmMS6q3H1aMrdBMkJuwXukijSZVkHI16r4gIiIiZRgGXEju9Y2lbAuBc4EhYQcugdocW4vsf3//hU3XlejbGPgv2e/xR8CmDuMSqbW7SB/v6gUuIiIiRY0ALgaWUlmSPB8biVYiFX3bkHva/V+AVodxSfAGknv2yCvY7BKRRnQR6WP9cMexiIiISB0bCkwmfzugYtsnWJK8VshxS20MBt4h+33+I0qUG9EI4HPkLvz1KNZHXaTRfIv0cX6W41iaTrvrAEREREowBDgB+DFWxKtcc4HfA7/Fpl5L9HUDbsOqJnv9HjtWEqFHVJk+wP4Frp+GVXWv1gBg7wLXP4SdTKpnqZ/Dl4B/kvl6dgV+A/ww7KBEauw1z/83cxaFiIiI1J1U4a6FVDaSPBurjNs75Lil9v5A9vt9udOIKtNB4WP4ooCe56Qiz7NrQM8Tlp7Aw2S/ju84jEmkFvqTrvb+guNYREREpA4MwpLkBVSWJL+PJQe9Qo5bwnEM2e/5VKI5Y66DwsfyxwTzul4o8jxRS5bBlmXMIPN1rAR2dBmUSA2kKv0vB9ocxyIiIiKOpPokV9oC6l0sSe4ZctwSng5gEZnv+wyiW9G8g+LH9QFVPsdWJTxHFJNlgK2x1m/e1/IGOlEmjaWT9PG9keNYREREJGR9sSS5UC/VQts7WJKsHpSN70Ey3/vFRLudSgfFj+9bqnyOS0p4jqgmywCHkZ6mmtomO41IJFiXU/jk2Q7oJLGIiEjD6Qf8jNytf0rZ3gS+gaalNYuJZB8D33UaUfU6yH5Nb/j2V2BLEyrRHZhT5PGjniwD/Ins6dijnEYkEpzjSB/bp/qu6ws8EXpEIiIiUjN9sJHgj6l8uvWxRHONqlSmO9ltou4HWlwGFYAOso/vm4AXfZcdX+HjH+p7nA+B3+V4zqgny/2xWgXe1/QPpxGJlK8NGJ3j8vGkj+u/+q67Fni+xnGJiIhICHpi7Z/8I12lbjOw4k7dwg5cnPsumcfCaqI9/Tqlg9zJ8o98lz1T4ePf6Xuc84DLcjxn1JNlgCPJfl1jnEYkUr7LgRPJPBG4Nulj2juKfBQ6MSQiIhJ53YBJZI8MlrqlqltrTXJzagPeIvOYuMJpRMHpIHeyPBibSuy9fKsyH3sYsMr3GKNp3GS5BXiKzNd1s9OIRMq3CbAG+BdW8T0ltVzp0+T+50gXt7s0zABFREQkGK3YOtM3qSxJ/gBVtxbYn8zjYhWwvtOIgtNB7mQZbLTIe3m5PZdP8d1/WvLyRk2WAQ4kewbCSKcRiZTvFuz4nQ1MSF72OOnjen1stkmqsN0pDmIUERGRKkygeG/XfNtsrDq22r8IwO1kHh9Xuw0nUB3kT5YP8V1ebs/l//runyqG1sjJcgvwCpmv7XSnEYmUb1vSiXAcGzm+ivQxfSOZx/jhbsIUERGRck3AznhXkiTPQUmyZOoLLCfzOPmC04iC1UH+ZLkdmOW7rtSey9v57rcMWCt5XSMny5C93vu/bsMRqUgn0EXmTKvU//2t0nZwFKOIiIiUaCfgASpLkudiSXLv0KOWencYmcfKW27DCVwH+ZNlgIt915Xac/kK3/2u91zX6MnyEGzNp/f1beA0IpHyTSDzGPYmzv5kebijGEVERKSIz2OFSCpJkj/BkuQ+oUctUXElmcfMxW7DCVwHhZPlLX3XrcSKfxXSHTsB5b3fBM/1jZ4sAzxK7inoIlHyHNknfvzbSqLfQq9utboOQEREIms01t/xWawAUzmWAhcCGyf/XRpsaNJAdvTt3+ckCndewdb+p3QHjihyn4Ox0dWUD4GHAo6r3t3v29/JSRQi1bkA6wZQSGp6toiIiNSBDYApFD/bnWtbBvyazC/yIvn0IvM4iwMDnUYUvA4KjyyD9Vz1Xv90kcf0z/Q4x3d9M4ws+6ewat2yRFErMJ3MKdj+qdn+E0MiIiLiwCBgMtnFlkrZVmHTadcJPWqJstFkHkfvuQ2nJjooniwPBlb4bpOv5/LaWLsk72039d2mGZLloWS+viVoqqpE03fJ/7d1DfAXd6E1Pk3DFhGRYnpj64rfTv5bTs/jOHArsDnwPeCjwKOTRrahb3+Gkyjcm4eNFnsdlee23ySzvdQ0bGSq2cwFFnv2+2AJtEjUXIu1U4znuK4NeD/ccJqLkmUREcmnG3AsVn14Mum2M6XqBMYAX6HxKhhLOPxTrmc5iaI+XOPb9yfF3su9ptQkmmjwHy+DnEQhUp2VwG/In7c14oybuqFkWURE/FqAicCr2NTpEWXevxPYHtgLrROU6virpDdzIbh7gI89+8OBvX232QnYzLO/HLitxnHVM//xoqr7ElV/xGZK5CrkpWS5hpQsi4iI1wTgGayX6yZl3vdJ4ItYkvxswHFJc/JXge1yEkV9WAPc4LvMP4p8tG//NmBhrQKKgDW+/Vwj8SJRsBi4PM91moYtIiJSY2OwipqV9Ep+Ftg3/JClCUwi81i7zm04NdFB8QJfKVv4buftudwT+NR3/Z55HqcZCnwBvEbma9zCbTgiVRlGdqG/ONZOTmpEI8siIs1tA2yq9X+wEeFyvIcV7doB+HfAcYkALPLtD855q+bxKvCcZ9/bc/nLZNYVeI/m663s5z9e/MeTSJTMAf7qu2w+1m1CREREAjQEK9rlP0tdyjYXq4rdI/SopdmMIfPYe81tODXRQekjywAn+G6b6rl8n+/ysws8RjOMLPfBRt287ev80/pFomZDMo/r/7kNp/Fp7YaISHPpC5wKnJL8fzkWAb8CfktzF1qS8PhbRX0OO0mz0kEs9eJG4NekT1btgC2D8E65TmDtZprZZmT2Vf6A5l7zLo1hBvAY6ZNbH2I919fGiv4Nw4pyDsNapfVJbi2kZ570IzMHXIgl4Muw362rsL/x87CWVXOBmcl/5yT/P4vsmgANScmyiEhzaMX6sk6m/OrWq7D2Mz/H/nCKhOVTrHjN+sn97sC2pEdTm9F84C7gcM9l15I5ajoN64vezHb27T/vJAqR6nUH1sXW3G+OJaypE0F7AC9V+fj+Fn2lWI2dgHoHm/HzavL/qa1hKFkWEWl8E7AejVuXeb84cDvwUxrsj59EytOkk2WA8TR3sgzWc9mbLA/1XX91iLHUq919+884iUKkPK1YQrwD1gpuRyxJrrclBN2AjZLbBN91H2Kft6ew39XPodloIiJSh7YDHqSyCtdTsRE8EdeOI/PYfNxtOIHroLw1y2CDHTNz3C8BLMGmWRbS6GuWe2DLRryvb3unEYnk1obNgjgbeIDs47YRttXAi8AfgEOB/oH85EKikWURkcazHnAm8B3K73rwNDaS/HDAMYlU6g7gCtLTDnfG1i438zTjNcD1wE9yXHcb1pO1mR1A5gmDWaj3u9SPtYF9gP2wLhSDqnisZVjl+znAx6TXGM9KXrYMWJC87SJs3f5yrLgn2HeEAcn/D0ju905uQ8leAz2S9ProUrUD2yS372PJ8xNYF417sWnkiTIeT0REpCIDsTXJyyn/zO/rwEQyC+KI1IunyDxef+k2nEB1UP7IMthUzVyf5fEl3LfRR5bvJfO1/c5tOCKsi53cepbMatalbHHs5OA9WIHN7wNfBI4hneiGbQA2W+PrwDnAzcALVPb94yPshKi/zoCIiEggegA/xipXlvtH6kPsD269rYcS8foemcftbGzkoxF0UFmynLrvRr6tlBNejZwsb4aNnnlf2xinEUmzGgBMwgryrab0v8sLsSJ9k4GDsFaPudTjye12bI31JOBKrPBXOScH3sNe96ZhBy4iIo2nBRsNfovyk+Ql2B+kYmsbRepBf+wLpPcYPtFpRMHpoPJkuVKNnCxfR+brUmEvCdvuwC2UPso6D/gbcDSwSfjh1twgYH/gYmwWW6nfU54Gvgv0Cj9kERGJut2xL4HlJsmrgd9T3nojkXowmcxjeQ7pvqFR1oGS5aBMInsU61CnEUmz6I6dvH6S4n+Hu7Dp2JOxStLdHMTrUgdwLHZCYQHFf15zsJ/Vug5iFRGRiNkE+wNTbpKcwCpcbxV+yCKVicVim5911lknnnXWWXccdthhO2IzIrzH9G+cBhiMDpQsB6EFeJfspMQ7nbWSfrIihQwBfkb+6vT+WQ4noZPVXj2AQ4BbKT4SvxJrmxdqp456nOsuIiLZBgKnAT/C/riU41ngVOCRoIMSCVIsFls7Ho/v2tLSMiGRSOyLp79yIpH4wTnnnDMMa7GSEsdmWUwLOdQgdQAzfJfdDHy1hs95GfBD32W7AY/V8Dlr7TiKF/LqwirvTsNakE3DiguJlKsPcAJwOoWLbL2PTbG+GpgeQlxRNgA4GBuh35fCXZs6sVotL9c6KLWOEhGpb+3At7Fqk8PKvO/7wHnAX7CkQqSu/PSnPx3Ys2fP8YlEYs9EIrFnPB7fFCCRSGTdtrW1dXfss3AM6SS6FfgzsANql9TMNsFGj71mYVNjB3suawM+n9x+6LmdN3l+Af2+lPy6YVOIzyL/CHEcuBO4BHgUGxWV4hYC1ya39YHjsTXLuWaETACex05CnE0NT3opWRYRqV8TsDYRW5Z5v0+BC4FLSfdSFHHu5JNP7jVgwICx8Xh8l0QiMaGlpWX3eDyeb71eF/Ai0Nna2to5b968x7BpeMcA95OeHTcaK+p0KEpymlE/4B9kFitcCowD3sEqhI8Ddkn+u7nv/iOwkayJyf3FWGGhVPL8ODY9VOQrwPlYn/dcFmPJ22U0dx/4ILyPzab7JVb47CSyi5+1Y8n017Gf+fnU4KSppmGLiNSfzYBfY9Ujy7EKmAKcCcwNOCaRsk2cOLFt9OjR2yanVU9oaWkZB/QscJd3WlpaOhOJRGdra+vUWCy2IM/t/oR9SfI6FxvtiZp2PNPNk5ZgRW1qZQhWYdxrJtE7udaKjeAd4Lv8+1jbmlxGANuRTp63x0ag81kD/Jd08vwItX1vpP6sD/wR2C/P9XOBi7BZLgvDCqrJtGKf8zOxmUS5vA/8AOtHHRglyyIi9WMw8HNs6lE5fY8TwG3AT7GRFBFnYrHYRl1dXRNaWlomAHtibUPymQVMSyQSnW1tbffEYrEPS3yanljS4v3SlAC+Ru2LY0n9uBD4P99l12FVsUvVB5uWnUqex1G8yvo7ZI48v4am2jaiFuyk3K/J3WZxKXAFcAFKksM0AWtDla9g6a3Y96hABg2ULIuIuNcTK9x1OtmjPcU8hhW5eDbooERKccYZZwxvb2/fLVdRrhw+AR4CHk8kEtPOOeec56p46hHAf4B1PJetBI4A7qjicSUafoKN5nk9h1X1rmbadBs2tT+VPO8GbFDkPguxYzGVQKeWDEh0bQr8FTsO/FZjo8jnAB+HGZR8ph1bkvNzYGSO6+cAJ2LFEquiZFlExK2DsCIgG5V5vw+w6UjXoRENCVEsFusL7BSPxydgZ/jHkP/7xLJEIvFEamp1W1vbC7FYLMh1xTtjybe3Qvwq4Ejg7wE+j9SX08hd0Gt7alPoZyTp5HkXbCS6tcDtV2NVt1PJ84PAvBrEJbVxOHAVuUeTH8EKfL0ZakSST2/gF9igQa5aXH/E1juvqvQJlCyLiLixIzaN6Atl3m8hVuH6MjRyISGIxWLtXV1d23jWHe+OVYTNJaso1+WXX17r4/Rb2AiQ9zvNauAorCe5NJbzgDN8ly0Fvoj1sQ1DP+x3eCp53gXoVeQ+/qnbr9YyQKlIG1b34Odk50iLkpdfjgoJ1qNtsM4f2+W47jngMOC9Sh5YybKISLhGYmdBv0PhkQm/OHADNvVwdg3iEgEgFou1dnV1fT6ZHI9LJse5RlhSSi3KVUvfA35P5meqCxtRKNZ7V6KhG1bh/we+yxcDB2Itelxpx76sp5Ln8VgRtUJmYV/iU8nz09hJHnFjMNYPea8c192DFY37INSIpFztwClAjOxCknOx3vUPlvugSpZFRMLRHfuSdw6FE49cOrE/AC8FHZQIlF2U62PgsUQi0RmPx+8977zz6uUL5NeAa8ieincDNm1yWegRSVCGYYXbxvsuX4hVKH4y9IiK87es2ozC37uXYFW3U8nzY4CLE0/NaCPs7+yGvstXYifc8lVWl/q0NbYMx9/iazXwbeD6ch5MybKISO1Vui55OnAqcHfgEUlTi8Viw+Lx+O7J0eN9KFzAaHEikXg6NXpcZVGuWjsC+yLkT5hfxPowzwg9IqnWLlh12xG+yxcA+2IjslEwHKvenkqetyNzrb1fF/Y3IJU8P0KF00iloFHAA8C6vss/wnpv1+OJGCmuP3by9BDf5QngBGwmUkmULIuI1M5mwG+Bfcq833zgl9j00TVBByXN59RTT+3Tt2/fnUssyrU8kUg83tLS0tna2vo48FQsFovScfglLGH2z+CYhy1/+GfoEUkl2oCTgfPJXiM/A/sSHOXZNr2xz6G3cNjAIveZRTp5nga8gNbPVmM0lij7qylPwxJlVbqOthastdz5ZC7RSWAdSC4r9UFERCRYg7B1yeX2S14N/CF5X02/k4rlKMq1G7YUIJeMolzAtFgstiK0YGtjFDYNb4sc192NrXGeGWpEUo4tsKJtO+a47iFs7eGcUCOqPX/LqnFkTwv2W4QVNUslz9OAqH92w7INMBUY6rv8OqwlkdaPN46J2HIc70m3BFZB+5Jid1ayLCISnG7AcVhxibXKvO/d2JnOtwOOSZpAsijXZsAuyXXH+1C4Z/dnRblWrVrVOXny5E/DiTRUfbH2LxNzXLcAaz/0Z9R6rZ50w77Ank32FOUE1lf5Z9gJnmZQbsuqNdi651Ty/BDW21wyrY31xfZPvf4LdiJNo/WNZ3/gdjILfyWwWhc3FbqjkmURkWBMwM5Q5hrJKuQFLEl2WclVIshXlOuLWDXXfOq1KFetpabhnUPudlf3Y3UBXg4zKMlpb+A3wJY5rvsUK8zT7FPo+wI7kU6ev4BN5y7E37LqNZr7BFEP4GHs5+j1R2w2WFQS5b5Y4bt83onY84RhH+AfZLZ5Ww7sBjyb705KlkVEqjMK+4J3YJn3m4d9gb+C5hklkSr4inLtDXQUuPmSRCLxlKco1/M09xfkrbBRox1yXBfHRhx+SrS++DWK7YELsCrsudyNdRL4MLSIosPbsmossAewXpH7fIwlBqnk+RlgVe1CrDtXA0f7LrsMO2kdpd+RX6dwVeedgacCeJ5LgRMLXN9OtL7D7Im1AvMuS3oP+z00N9cdlCyLiFRmLdLrknONWOWzCrgcS5QX1iAuaRDlFuUCngemtba2ds6cOfORP/3pT1pzl6kd+7yeB/TJcf0qYArwc9TLPAyjgHOBw8l9XM/GZgVcG2ZQDcA/dbvQ7w2ApVjNAm/hsEZclgGW9F3qu+werChglBI+KJ4s/5HsnuTl6o6dpPKv6/aKWrIMtib9L77LHsUS6axilkqWRUTK0wJ8A1s7N7zM+3ZiZ69fDTooiT4V5QrNJsCfsFG4XJZgo0+XAW+FFFMz2Rmrcv1lslt8gY3uXYutXZ4fYlyNqj82o8Lb87lngdv7W1Y9RmO0XNsceI7M1z4dm44dxYKaxZLlhVi7teVVPMeh2KybQqKYLEPuEfMzsZOpGZQsi4iUbiw2Krxzmfd7E/vi96/AI5JI8607LrkoV2tr6wOxWEyJRHUmAL/GprHmEgcexJLmu4nWFM160wocgBVV26XA7Tqx6fD13Ms76roBW5NOnovVO4BwW1atj50kWRLgY7Zj/bjHeC5bgCXK0wN8njAVS5YBjqRI8aoi7gQOKnKbqCbL7cC/yVz+sQr7nveK94ZKlkVEihuMTc08gcKVSP0WAJOxXsvNtCZM8vAkx+OwP9L+/p5es4FHE4lEZ1tb279jsdj74UTZVNqwmSJnY1/S83kBq5x9C1ZvIJ8WlFR7bYh9qT+WwmtpnwVOx5JlCd9GZI48b17k9oux5NNbOKyaEUyvHljv41uB35FjWmwFjsfqg3gdgX2eoypXsjyXzCnT9wH7Vvj4w7Ep2KnZHwnsd98Q3+2imiyD/axeJfNn9jAw3nsjJcsiIvml1jieDQwo435xrKffqTReL1Apw+mnnz60W7dueySnVu9F4b6pKsrlTk+s7dspFD6BsQobjbgBuIvMBKEPcDPW3/mq2oQZCYOBr2Bf5r9A4e+aL2P1G25Dx3o9GQFsRzp53p78S0Igu2XVI1T3t29vLNF7A/tM3lPFYw3EZnd5k7xbsGQ5ynIly5cAJ5H+zMWxQpCVdD84FfiVZ/8xrPL6WN/topwsg9VMuNV32WHY73FAybKISD57YNMvtyrzfg9h65JfCjogqX+xWKw38AVPUa5CfVFTXzA7VZSrbnTHEr0fkf2l0G8R9oXqTmzk+UZsiUYX9mXrjtqFWXfWxkawvpz8t1BilQDuxWbcPICS5Cjog/0uSyXP47Ail4VU27LqRmwaMdhyiFOwOg3lOhubGZayBCsuN6uCx6onuZLlH2HrjHfzXHY6NsOtXP/FpuunfAcrGNZoyTLY7yPvCPzrWAu7OChZFhHxWwdrY/KNMu/3AVYcQpVbm8jEiRPbRo8eva2nKNeu2DTCXOJYUtUJPL5ixYqHL7roosWhBSvlGouN0hxJ7kJU+azAEu67ahFUnWgDtsVOCB1E8RFkgJXYiN6FqMhh1LUBo0knz7sBGxS5zxysTdVzWAL9GHZM5LM2tp64P5aMtQLXYH9nPyoxzv7Au9jockrOIk4RlC9ZXgz81XPZm9h7Vc6Jiu2x9yplGTbb4EEaM1neHBvgaPNcdijWk1nJsohIUk/gJ1hxmd5l3G8pllz/BvuSLA3OV5RrbwpP0VdRrujbADgK+3K6WYn3mYlNRX0GW9v5AtH+/TAA+wK9Y3LbjdKWpiSwUcUbsCnqjdqSSLJbVhWaVQOwGktQUqPPD5Bd/fwHwO89+3FsKcSlwPnY7I5CfojNEEuZj01LboSTlPmS5b9io+Z9PZeX23P5d9iylJTrgElYbYFGTJbBfkd9zbP/CMmOCUqWRURsZOQSrMhJOe7G/hi/G3RAUj9isdjIeDy+S3L0+ABs9kE+c4BHEolEZzwev++88857L6QwJRxjsMT5q9hIS6lWY1NIn8ZG1l7HRnzqMXlcGxuJ2gxrObRDcr+c4oavY18+b0C/H5tVP+zESqktqyB76vbrwBPY+uk2320/xaZYFyoC9iKZ1e7PBmKlvoA6ly9ZvhTrF/9Nz+Xl9Fzujp3s81ZI3xMbVW7kZHlL7ORNKjdOYL8DpytZFpFmtgmWJO9f5v2mY9Mz7ws8InEuFosNicfj45PJcbHKsEsTicSTKsrVdL6F9WouZ3p2LnOwIkbTseT5HawK+hxsdCjI9jkpg7BKt8Ow0cCNgE2xhHgU5RUzTIljX6TvxdZwPx9IpNJI2rHENZU8jye7srLfLOxzsSvZJ2sSWGLzP+BnZBdp2orM2iFd2CyRUqdw17tCyfJ4LLlNKafn8lewWSAp72G/I1Kf8UZNlgEexY61lHOBs6r9JS8iEkV9sHVLP6ZwIRq/BdhZ6aDaWUgd8BflisfjnwdaE4mcOW9GUS7g0VgsprZgzeU0bOlFasDhAeBkLAnYD+tb26fExxqW3HbLc/1yLHn+GJs6uiB5+ULsy+tSMtvS9cJG77onY2jFkt9UgjyU8n7nFTIXuB+rVHw/8ElAjyuNaQ02q+I5LKGD7JZVm5E563UE+WdwpG73OWwtvL8I2Jd8t59K4yTKxTyMnXhLzZYbABxMaT2Xj/btX0PtemrXm6vJTJYPQsmyiDShg7A1TB1l3CeBncH9CfbFVSLMX5QrHo8XK8r1RktLy7TkuuP7YrFYsXVy0pgN1ZQHAAAgAElEQVTasF6t3/dcdh1wDDbN+mXgD9ixtB02fXlHYCeKFz/Kpxf2u6qjwvsHbRr2mp/GXm+zfImW2ngnuaUKYw7HPjep5Hk78v9uTkmNOO+BzWiYApxF9oyxf1QdbXQksM/pLzyXfZPiyfJIrA6H93GaqWjpXdgoeWrK/zbAOpqGLSLNYiMsST6gzPs9h61LfjLwiCQ0vqJce1G47clnRblWr1794AUXXDAvpDClfvXEvnwe7rnsMmxEuZSEcTiWOO+ATevfFNiY4EZ5g7SE9LTwl7BE/XvJ61Zj6xcfcxKZNJNW4HisenpPyquzlJoW7F3nvC6NNbJcaBo22Of2bdInE0rpufxTbNZMymdFrpIafRo22Fr5L3j2J2pkWUQaXTesquN5lD41Eqxq5i+xkaRG+kPQFM4444wR7e3t45Kjx/vH4/F1W1ryftf6rChXW1vb/bFY7N3wIpUIGIStw90lud+F1Sz4XRmPMTv5GHd6LmvHvryOIr1eeCTptcRDKV4QqRJLsLWgc7Cp1O9iiXEqQf4wx326Y+u0u2FTXrfPczuRIIwF/oxV1I5TfkFifzGwD2isRLkU72JrcPdI7rdiCXahnsuTfPvXBB5V/XuSzGR5RyXLItLIxmNfaEtt9wK2ruoq4AxAI4oRkSzKtTOW0EwgefY7z7pjFeWSUnUA/8ZGgsH6wk7CEsZqrQHeSm735LlNf2zN5tDk/73rkMFmSLRgifcabOR3CXY8e9c3L8bWPc/GeqaW6zisWuz2WLXsO7BpsqUUDBIpVT/gHGw2V2pEtFgV9gTp398t5E6s18LqCdwbQIxRcg2ZI8Pfwkbqc/2925nM70pLgdtqFln9es63v6mSZRFpRCOxs6ffKPN+j2B/pF8OPCIJVL6iXHlurqJcUomtsC/XqVZhn2JFcsKcgrwouU0P8TlzWQEcAvwH+/06BriS7JEokUp9GVvasG6B23RhSdwi7KRQ6vOxMHn5UuyE1o5Y8pdyP3bCqDeVnSyKqluxn2m/5P4o7GeTq+fy0Tnu2wj9qMv1tm+/Q8myiDSSbtianZ8Dfcu430dYFc2bi91Q3CizKBd41h2rKJdUYE/g79hoLljf0f3IbEXTbGYCE7Gqwz2wk5EvAL91GZQ0hK2xk1I/x5LgBaST38We/ZUlPt5lZCbL05Jbs1kK3E5mIvxNspPlnljLKK9mnIIN2Wu6hytZFpFGsSvwe2yqYKnWJO9zFnZ2WupIvqJcedYev9PS0tIZj8cfb2tr64zFYjPDjFUaylHYUoxuyf1XsMq6hQrjNIsnsGrgVyf3f4X1iW626a0SrJcI9kSUvz5JLfqVR8UUMpPlI7G2md4lFIeSWfTyXWy9czPyj6b3VbIsIlE3GDgf+C7lFQGZhlXabOaRoroSi8XWjsfjuyZHj/eLx+PrFSjKNRd4OJFIdCYSiannnnvujBBDlcZ1EnAx6Sn9D2HTQxc6i6j+TMGmcn4fK6R0fXL/LYcxiXj5Zx2VOiLdiB7FphZ/Lrmfq+fy0b77TKF528L5j5UeSpZFJKpasRGgi7GEuVSpKteX07x/DOrC//3f//Xr3bv3jp51xyrKJa60YKOkp3guux37HbPCSUT17USsgvcepKuF74Rm6Eh98Bee6+UkivqQ6rkc81zm7bm8LvDFHLdvVr19+8uULItIFI0B/oD1LC1VHLgBm370SS2CksJOPvnkXgMGDNglmRyPA3aMx+P5/g6pKJeEpQdwLZlr9srpodyMVmM9p/8DbIhV0b0Wm86pn5m45p92PSDnrZrHFGw9eGrGzN7AetjSkqPJbLX1MPBOeKHVHf+xskTJsohEyUDs7OjxZPdRLOQFrPVJrgqQUiP+olwtLS3j4vF4ob6xnxXlWrly5f0XXnihpr5KrQ0E/gnsltxPAD8FLnIWUXTMw5Ljx7HRmIOx+g9nuwxKBCtG57W+kyjqx3tYt4/xyX1vz+WjfLdt1sJeKRv49j9SsiwiUdCCVV79FTCsjPstwJLrK7CWE1JjvqJcE7BkJF9RrlnAtEQi0RmPx/913nnnfRRiqCIjscJUWyf3V2GjLH9zFVAEvYi1j7oV+z39C+C15L6IK/4aFqOcRFFfppBOlsF6Lj9Ouoc82Ij87SHGVI828e3PULIsIvVuG+B3wC5l3CeBFZ05FZhTi6DE+Ipy7RuPx9cvUJTrE+ChRCLRmaxY3cxTvcStLbBEeb3k/mJsWvH9ziKKrtuxkfjTsIT5KuB1rIq4iAuv+fa3w47NZq5zcTs2cODtuXyJ7za30tyVwyF7ed+rSpZFpF71wab0nQKU87tqOjZN+4FaBNXsYrFYX2AnT1GuMUBLnqJcyxKJxBOpqdVtbW0vxGIxrWcU18YD/yC9Nm0W1hrqRWcRRd8ZWNu+A7Ae93cB26P6EOLGG1gF+9RnfBCwOfCqs4jcW4olw9/2XDbGd5spoUVTv3b17T+tZFlE6tFB2GjyesVu6LEU+DXWRkqFoAKSLMo1Nh6P75JIJCbE4/HdSfef9evCEo7O1tbWznnz5j12+eWXN3PLDqk/h2GzTlJr518D9gPedxZRY4gDXwOexJKSDmw6+35YsT6RMMWxnuD7eS47kOZOlsHWI387z3UzgMdCjKUebYj9/kpZAzylZFlE6skmWEunfcq8393YaLK+8FapUFGuPNOrPyvK1draOjUWiy0IN2KRkvl7KD+FnZjT6GcwFmEFv54C1sJqFlxIZjsukbDcTWayfDh2PDazx7B+6BvnuO5qmnuaOtgx4jUNWKBkWUTqQW/g/7AqtD3KuN9bwA+Bf9ciqGbhK8q1JzZlrWhRrra2tntisdiHIYYqUokW4AJsTW3KP7GRUH8/VqnOdOCrwL+wjgU/xkbzrnIZlDSlO7GT76mTY9thxfxechaRewmsxdsvc1x+ffjh1JUW4BjfZXdAeesARURq4RDgUspr7bAMm279a0DTfMt0xhlnDG9vb98tOXq8Tzwe36BYUS7g8UQiMe2cc855LrxIRarWHVuHd6TnsiuwUWatn6+N+7Cq2Ocm93+HFft6xllE0ow+xGqX7OW57ATgWDfh1I0/YGu6vZaSXUG82exFZmXwVcCNoGRZRNwZiSXJ/mkvxdwNnIh+sZfMX5QLK+qholzS6Pr9P3vnGeZGebXhe3bdOzZgDARj00zoELqB0GsICZhAaE4BkhCKvVo7pIAgFJtdY4gT+AIJAQIhlBBK6KYb0wkEQg2YatONjbt3d74fzww7Go20klbSqJz7uubafV/NaI52pdGc95zzHOAmYB9v7KKISjIug+qI85Dg1xGoPvwWJPhl7eGMcvInUp3lY9E1oJ4zoj7FWrtF8cvQ+Fa8birmLBuGUW4agB+jqPDALvYN8j5K6bOLfBckk8ke7e3tW/h1xybKZdQhI4A7gS29cRvwE+DPsVlUX7hISGgDYBv0/7gRKZHb9cUoFzcDbwGjvXFv4Awsumyksg+wW2iu1f/FnGXDMMrJFsBlpPexy8ZKlDb0K6z/XyTJZLKhvb19K885HtvR0bGb4zgDwUS5jLrk66iHsl/asQgYh2kblJulSH38aWA1YEfgj8D4GG0y6os2YAq67/D5IXAJ1irOED2Q8GOQ+wiUjZizbBhGOeiHVnMTSPQlVx5ENUYvl8KoaiYoytXR0bGn4zjZRLk+BB51XXdmR0fHXeeee+57ZTXWMMrHDqjH76re+EPU+/e52Cyqb95BCtn3o/rx45Dz/Ic4jTLqir8gjYJNvHEjEpzbHi3GG/XNZDrfGyAti18FdzBn2TCMUnMgujEamccx85Ay9tUlsagKSSaTq3sR41xEub50XfdJP3psolxGnXAIEmTp643fBPZDqvlGfMxC7aNmeOOL0ALog7FZZNQTbajbxh2Bua2Qk3RO5BFGvbAp8JvQ3FVoQe8rzFk2DKNUrAn8DqXh5UobEv06C/iyFEZVC4lEov+AAQN29EW5Ojo6solyLXVd9zHHcWY2NDQ8BjyRTCbbymqwYcTLz9G1w28T8xRwEPBJbBYZQX6P2vYcj+49b0TlOG/FaZRRN9yJ3nPjAnNnoYyTO2OxyIibVVBNe7Bd6afA6eEdM4YlDMMwCsRBN0QtwKA8jnsOOBF4phRGVTphUS7HcXZFaYtRpIhyAbOSyeSyshlrGJWDg9oUnRmYuw21iloSi0VGJnoCM4FdvfELwM6obY1hlJpVUQuz4YG5haiWvhpLvdYH9gjNzUavsVR8l84SF5/LkaBfNdGIynX2D81/D7ghvLM5y4ZhFJPNkYDLDnkcsxj4LVIebC+FUZWIJ8q1MbCz4zh7AfuSfXHhK1GuFStWzJwyZcr88lhqGBVLD+D/gB8F5q5Ai26WWVGZDEcLomt745tR+8Bqu9k2qpN9UTp2UDvlFeQwL4jFIiMOLkJ17EEuJ4NKujnLhmEUg76o/ud0MkdDo/gX8DOgLgSngqJcaEV4WJbdTZTLMDIzAEUA/MiA9VCuHrYGHkXCjyAxnfPiM8eoM5oItAXyuB84GMtGqQcmAtNCc7OAPYEVUQeYs2wYRnfZHUV3NszjmLnAKcA/SmJRhRAS5doHWDfL7otc130iIMr1HBZtMYwo1kDRoa29cTtadLss4xFGpXE08Ffv9w7g22jx1DDKwZ9IzUgBOUwHotRsozaZBEwNzb0LbAt8nOkgc5YNwyiU4agu+Zg8julAX1IJalDAKyzKhW7mM11nl6I67VkNDQ0z586d+/Bll11mbSwMIzvroX7J63vjxajO7I6MRxiVyoXABO/3L1Eq7H/jM8eoI/oAD6H2UUEeR9kqlpJdeyRJ1bYAZRKMBf6d7UBzlg3DyBcHOcgXkj2NOMzzqJbwqa52rBZMlMswysp2KPq4mjf+DKVOzo7NIqM7hEV2XkfOyxexWWTUE8OAe+nMUPF5BvgWKoUyqp8GFE1OhOYXo++PB7p6AnOWDcPIhw1QynVYgTEbS1AtYU0IeIXqjnMW5WpoaLg/mUx+XiYzDaPWOBi4js4617eQk/V6bBYZxWAVtIDqZwrcCxxADXxXGFXBYOAulNUQZC5qM2ULcdXNIOBqVOYRZDFaEMmp17s5y4Zh5EIf4Bfe1ruLfYP8C/U/facURpUDzzke29DQsLPrugeh/tGZ+Ah4xHXdmY2NjXcnk8l3y2SmYdQyP0D1yD288dOoh3LGGjOjqhgDPEnnwuMUInqdGkaJ6I8yHHYPzbcBvya9xtWoDrZAavujQ/ML0ELr47k+kTnLhmF0xW4omjwmj2PmIcf66pJYVEJOP/301Xr27PlNL7V6b2BUlt1NlMswSstk5Dz53AccSg1qHtQ530Y3tg3oGnoUyiQwjHLQH7gJ2C/isSvRor/1A68exgOXoE4tQT5EC63P5vNk5iwbhpGJocD5wPHkfq1wUa+6ZqpEUTKZTPYDdgqIcm2FbtiiaANewKs7NlEuwygZjehmJ9j38ip0PbLPXG2SpFOAZxmwC6ofNYxy4CC15PNIvwd4G2mu3Ftmm4z8GAH8AfhOxGPPoIXWvDP+zFk2DCOML+DVSqeQTi78B32ZPFEKo4rFuHHjGseMGbNlQJRrFzKnlncglcSZwGPLli176IILLrCIlmGUlv7A9aiNi89UlJprmRu1i4P+7+O8cZctXQyjBByE2poNiXjsRuCnSFzQqBz8+9bpKNAT5jLgZDL0Uc7lyQ3DMHzWBy5FEdZcWQpcgFZjC7oQlZqQKNc+SNQjEybKZRjxMQy4DdjJG7ejm5xLY7PIKCcDkKjSZt74MSQoWZHfLUbNMgaVBWwc8diHqOXZ9djiXSXwdeD3pNecg+5Pf4qykgrGITXFqdhciV3gDKMa6I1qjE8nPwGvu4CTgDmlMKpQksnkmh0dHTt70eMDgbWy7P4x8LDrujM7OjruOffcc6tWjMwwqpxRqIfyht54GYoW3BSbRUYcrItE3Fb1xpcCP4vNGqNe6YvKAhKoLCTMM+i+6f5yGmV8xdrAb4Af0in+GGQ28GPgle6eyKG0qyJDsMbehlHp7Aj8Ca3O5cpHqLanIgS8ksnkqh0dHbt7zvFYsr+Wxa7rPm6iXIZRUXwDuANY3Rt/jkSfZsVmkREne6KFE/8m+CfAH+Mzx6hjtgL+7P2MYiZymvMSjTIKZii6/zyFdAEvKEG7UnOWDaN+6QecQeZV0yhc4BqUghRbzU53RLmAR5LJpGW8GEblsBdKeRzojd9GrT1ejcsgoyKYCEzzfl8J7A08HJ85Rh3TC/glyr7rFfG4i65hF2K9mUvFmiiT8WdE15MD3IO0c4qaIWjOsmHUJ/uhdlAj8zjmdbS6n1MT92JSgCjXq47jzPLqju9JJpNVocxtGHXIsSizpac3fhE5yh/EZpFRSfwJ+JH3+0dI8Ou9+Mwx6pyNgHOB75JZ9+kpJDR1E1qsN7rH1ihAczjRCxWgVOtfAf8shQFRzvJKirdydwjWl8wwKolVUM/SfLQKliEl2vOB5aUwKoqQKNfeZF5JhIAo18qVKx84//zzTanSMCqfU9FNpX/T+QBq+WGLW4ZPH3RPup03/jcwFqVaGkZcbIfupaJEpXzeQ22MrgbmlcOoGqI38C0k7rhrlv3eRy3nrqRIKddRRDnL84mW3TYMo7oZhy7c+bSDehhFk0ueDvnLX/5yRI8ePcZ60eMDkHhDJr4S5WpsbLw3mUy+XWr7DMMoGo3ADKRS6nMNiiBaiYQRZgQS/PKFGq8Fjo7PHMP4in1RJ5Cts+zTjkTArkWRT2s/GU0D6q1+NHAY2QMkn6Egzu+R4nVJMWfZMGqfryE10QO72jHAJ0AT6jVYEjxRrh2BnVHN4jZZdjdRLsOoDXojx/iwwNzvUJpdRywWGdXAjqgEyC+/aUYCPoZRCewFnIZKSDLpp4AyIm4FrkMOdL1nSDhIc+Zw4EhgnS72fw24GEXry5a5bM6yYdQuDnA80AIMyuO4G5GIwifFNMZEuQyj7hmKbhTHemMXqZqa02PkwrF09kvtQGmad8ZnjmGksT5KHf4R0L+LfZchtf+Z3lYvatpDkdr9XiiIk621p89jyEm+mRKmW2fCnGXDqE3WBy4jez1NmLlIZfDWYhiQpygXBOqOTZTLMGqOdVFf9jHeeDlwHHB9XAYZVckM4Ofe7/OB7YE34jPHMCIZCoxHKcWZWk6FmYPUnB8DnqR23tfDUI339sA+3u+5dGD5GPg7Evl7sWTW5UBcznKUWM9/gZcLeK5V0OpEmNvRqk2YvsBBobnngDcD4yEoHWBvYFNgVRTp+sjb71ZUd/BFAfZm4msofWMfYBTqNdkfCZ18iJSI70IfpM+LcL4hSM1vR2BL9BoHe4+53jnmog/vC0jd70lyU/Y7lNSI4SvASwXYuCWwQWC8gtwcuW+Tqpj3Jvof+zjoPfMt1NtzFHLiVgLvAnvQdU1JA6pR2Q/VWIxAf8M+KCL7Cfp73QU8Qvnq8Hqg9OmkZ0suuMDlqIVUt2ppChHl6ujoeKyxsXFmMpmc251zG4ZRsWyGIoC+DsF8JAD6SGwWGdVKD+BeOheCX0M34dZ5xahUvo6c5u+TXweSz+i8934K3cd+VHTriks/5DdtT6eDvEHWI1JZDNyC6rvvo0LUxONylg9BofSg7Po85Bx9nMfzOOiPenBo/jLUZyuKtZB6WpCTgEvQSsdE4Dd09nvMxGIkUz6D7tVZjUTiAEeSWYY+yHJU33UehTnrQ4BzUIpIrs6Uz2eoMfvkLvZbQWcbELzz/SbPc4FUUk8LjHN9b36OFlF8LkH/Y1Dqx8XAJlmOH0bmBQkHLTKci1oI5MIc1J/vekpbZ7sF+v9kq/0N8z+Uqv1QISdMJpNrdHR07OJFj/dHiz6Z+AR4yHXdma7r3nfOOefMKeSchmFUFXug73t/MXYucABahDWMQhiKBL9Ge+PbkIq61bwblcwWqCTuCBQcG1XAc3yBFoiC2+uoN325hMN6A2sAG6L74DGB379Gbr5MkIWofvtm5NMtKpqlRaJHTOe9BTl8pwbmRiAxof3J/YJ3GumO8oukOli50ge4AUUbc6E/cJF3/m9RWJH+USi9IB+ntTcSthjvnfuJPI4dg1ZqsqkMZ2MYeq1dOcuVymTU/qirD3KmOtrVUN+8bDL2UYxCYg7fQyuLxVbu6wuciSLDuaS2gKLoF3rH5dwOatKkSQP79eu3vV933NHRsQ2A60auASxxXXe2iXIZRt1yNFrA8zN9/ou+461PrtEdPkeL1o+he7GDUTbVGTHaZBjZmIyCXMfRGbzZCF0P90f3lbn4AkNQtHb7iMeWooDjPBSc+Ahlpi5CAb4V6H5vCYrY+s51bxQR7oEChY53nl7ovncNYDjKeB1OajCqUP4D3I2yLx9D96QVS5w1y73QH+gboflfoTdUV2yLCuOD6baLvflXshyXKbK8C1rtCbLE27cdrZYMyPCcM5ETGZX2nYlTkLOdyXFbiCK5w9GbOIrFSFH07hzOtypaSFgjw+MrgE/RB6wPWv1aI8K+V1BKSTYqMbL8kvczF1YnXdxqffShXj/DMSvQ/2s5+ltneq88hlLti6WAuAtKoc41yg3qU/ljUlPTI5kwYULfwYMH7+w5x2PRBTrTIluKKNdnn3326IwZM8rWl9kwjLLRQNeL2qeiBTl/8fEhlFVm6bJGsfguWsD272WPQEEPw6gUHOACFMwA3Sdtg5zFIP2Ab6LuIDsg3ygfYdZKpg3dgz+JAnz3AR/EalGexBVZBjkX30M37IMD82cBj3pbJoaglNZeofmTyO4oZ+JIOtU5QV/qU9E/1Fdda/D2mYhqYoPs5e1/KrmxD9GO8ieoyfnfUaqazxgUST6NVIGk/uiLYStSa66jOI90R3kOUkq+A9XqhhmAag++iVZud+jiHJXKGOCHgfF8FO24BwkoLER/m41QtD98EzgEvRfWDc0vQpkBNwDPkLoytgmKqpxC6mLHzigN/PhCX4zHYHQBPp7cU16Wos9XKxnUBCNEucZ2dHRkW+38SpRr+fLl906dOtVuhA2jthmItBpuzPB4+OYQlF53FPktKBtGV9yMssV+id53V6LSoi4Xgg2jDDgo6BP0DaaR7iiDAih30qnu3gBsTGcUeVt0j5opeFYptCPf4nnkHD+JPo9la/NUCipBDfsw0r9030cO4KcZjrkJiUgFuRqlN3RFVGQ5yJnAb8meLvpT4A+kOintyBF6sovzD0MflDVD848A48hes70J+iCF+5A9jiKMmeTU+yJHPChj/wRqpp6P4vDXUWSgq8h/pUWWg9wMnICiwLnyD7SCHWQWWsXuanVsAxSRXi80fwiFq05/C0XJ80mnfwQ51q+HHwiJcu1F9hSbecAs13VndnR03HHuuedW1eqgYRjdZga6EfpzxGO9UWuf7wXmrIeyUUoaUGmfX0L3DnIsitr60DDypBHpJwUDNWcCZ3fjOR2U5boRqhH2a4XXR6Wsfbvx3PnQjj5fb6MA5eve9iparKq5Np9RzrIffeou/0B/tFz4A2pZE+QOdPEL23cS8PvQ3KsoZSGXlYtszvIf6GxJ0BW/Rk51kIdRFDYbv/WODfISilrnEpUbA8wm3aE5nMwr/bsDD4TmtkXR0FJQqc7yzejvlE+PtkOQ8nmQWUjpOdcoyUi0sha0/Rn0P8iH4SgT4Jg8jvkC1cpcjvdZColy7Uf2JvCfAg+6rjvTU6x+K0+bDcOoHXZApSRHkP59MwQtAPqaDi66MUyWyzijbhmIgga+cOejSMyzousgjZqlEbgC9QUHXQub0D1tKRmInObVvW0Eqjnug7IRG1DQrJc35zvX7XQGzuZ7P79E9/KfoCCeX//sj+tKfybKWS4WB6P2TbnQBzmA4V5kzShl1GcrdEEMpiIvRSkKufbgyuQsf4BWa3JNFeiBaj83Dcy5KPr6aoZjeqF05+GBuQ5gJ7qOSAf5GXLsgzwC7JZh/6OAawLjFehvXqr/fSU6yx+hL9J8Isogx3jnwHgRihZ/mOfz/AilbAfZlezlBkHGAZeizIRc+Rfwk2QyuQDYwRflQi2vMqVup4hyNTY2/juZTFpEyDCMnug7bxOUlXRv4LE1UdbTFt54BfAD4G/lNNCoazZE91F+u8KLKUzs1TC6Qy903fOzX11UjhcO8hlVRJw1y0GWobStZ0lt2XQeclaeQIXu15PqKIMuhsVoVt1Cfjn1bcgJ/HtgzkE3CJnUog8k1VEGpQ/l4ygD/BGYRGq/tl2RExfVxDxc293gzdWT+NIfyN9R/gapjjLoCzhfRxmk9H4OqXXjB9G1szwK/b/3zvVEPXr0+GinnXaa8c1vfrPNdd0rOzo6diN18SJIO0qpNFEuwzCy0Uxn5C6YBbUJKjXx28YtQjeKQWfaMErN6+g+8k4U2TsVZe2FF6kNo1T0Rn6Kr2vUjsRUr4zLIKM4VIqzDHLyTiR1Jboncka3QjWa4cbW16OagO7S7j1XvtyGUheCinXZ2grtEjF3VQHnbUfOVzideyzRznK4DroHijZfUcC5q5W/FnDMfhFz10TM5cIKpJp+dGBubIZ9QQsaP0ZiEJmUtQFwHIc11liD0aNHu5tsssmHw4cPH+o4zjn+YxF8JcrV0NBwXzKZLKRft2EY9cMGpLbl8VP2vonKVPxo3jzUQ/n5sllmGJ3cizqqTPHGl6A+tLlmcBlGofRDZSh7eeOVqFXoTbFZZBSNKGd5BYULDwWZ2/UuaVyH6muDSsEjUeQ17Cj/Dwk1FYP/Uli0cCmq39o/MLclitpGFbiH1aTbkQNVCHeR7izvCPwlYt/HUbp3sH/wH1CLo0spXyPzuJiLhAjyJby48QGZU+xz4TlSnWU/HTqcDr8ZWg3fLtMTrbLKKowaNYrRo0czatQo+vbti/dcIyJ2/0qUq7Gx8c5kMplN4M4wDCOIg74ngvcLXyDRw2vp7A36ClpgjOqsYK9wkDkAACAASURBVBjlYioqBzgSBVxuQPog9r1nlIoBKHi2uzf2u/3cEptFRlGJcpYXIxGkuDgVOZWbBebCjvJyJDCSj5JzNl7o5rFBZ7kPqp15KWLfTUPj1ym83+5/SHeAw8/v8zkSXBsXmOuDvlQmow/0vcD9ZFYgr2aiZPpzIeysvkn3mrGH0/z7oLKD4Pt4O1R6kJI23b9/f0aOHMno0aNZb731GDw42G0tjU9d133CcZxZruvO/O1vf/tsN2w2DKO+GY/EkoIchb4//O+fJ5AgZy1+fxjVx4+QQvC2qPTpVpTJtTROo4yaZAhwN9JOAt3TfwcrQ6kpKikN22cpctafIbXVUZBmVN9cLN7pxrFzIuaiRKh6klqPDdAdZeFFSLQqGEnMJn7VjFLmVgvND0XS9j9Ezvd/UbT8YdRvupCIe6XxeQHH9KYztdBn1wKfKxtDSXWWnwYe6dWr155rr732V9HjESOiAsZfYaJchmGUglWRyGZwYbYDaXz43ILSDc0RMSqFpagt6dNIFXhrpP1xbLaDDCNPVkdOsS9suBiJG4e7zxhVTiU6y6BU10mkKz6D6qOKrSrXnQh11LFR0ceoUGB3I+MLSHWWs0U930EiUTcDozPs04Ai+psBP0E3RY8icYJrqd42DIWkmZer13h4AcUFjj/yyCP/N3LkyIaoAzo6Ovjggw+YM2cOc+bM6fjggw/eaWtrm4fUwnNtZ2UYhtEVF6HvlaD4QfC69HsksplPOz7DKAfvogjfg6g07hgUZLk4TqOMmmEN4D46Mzq/QHoNj8dmkVEyKtVZbqBTdj1MO8VvedQdByNqNb1fjnPdXYkPp3BnisT7vIAc4VNRP+k1u9i/AbWj2g2Ju5yEaqWrjUJu5LLmOReRNAWu5ubmHgsWLFhEQDhu/vz5zJkzh7feeou33nqLZcu+ess2ABt7m99/eSFSiJ+FsgQex9IjDcPIj31QunUmlqLU651Rxs1Cb1vg/XwSta8zjLiYjRZzLvHG05Dg192xWWTUAuugssX1vfF81E7v6dgsMkpKpTrLvwb2yPDYYajP8CUZHi+ErpzMbEQpFUdFjKOim1lVjnMgHJVcELlXKkuA84ELkBjBPig9eyuyvx9GAXegL57f5WtoFbIoYu5R9DcoJvPCEx0dHScPGTJkUFtbG2+88QZ33XUXixZFmZORQegGNtj2ah6dzvOzwFNEi9AZhmH0Q50m2lEbnij6IgHOkaH551HbxztLZp1h5M6lKE32RPRevhZpg7wZp1FG1TIKOcqjvPFHKGuzGC1sjQqlEp1lP4qZjWnopr87wlxBBnW9S0aiIpBRrXgWkC7I1d3oZbimdn4ex/pK3L4a9wAkgLGXt21OetTTAaajm6FH8jW2AML9octJVE/m95GoTUlZunTp7/r06XNyR0cHI0aMYOXKomS/j0ACb77I22L0f3zW2x6hMMVwwzBqjyTpTnAmfIf6BeBc1Cql2NlfhtEdTgbGoPvLocDtSEi2WCKxRn0wBt0zr+WN30Pih1EtW40aww1txRYwyofVkEMStKcDrQSG7XyVwiKza0U8V3f6oF0c8Xxh9W6fT0P7vd2N864acd6HuvF8YdYCzkIR1vB5Huzi2CWh/Qt1MK+jsPfm56Hjomrfc2Fh6HmeKvB58mXMwQcf3JFIJNxEIuFus8024b9/qbZ3UF/z01Abst7leLGGYVQUmyN9iq6uF23ez9koHdswKpnhqI7Zf//+k4gyKMPIwJbAx6Tev68Xp0FG+YgUEIoJB7iKzhUbnxmoburG0PxGFC8Ve8siHruAzOk9z4XG61C4kNRWOTx/d/gAOBM5TeEc4F3Ibnd4tTacLp4rcV+Iws7xVnSvdVSuvDps2DC/Xx877LDD0oaGho/LcN51UG/A6egGeBFSR/dVRDfBbi4Mo5ZpAC4n+72BrwHxJIqq7IQidYZRyXyElIp9rZdDUMmfYXTFNiii7HeTeQ1lYloqfx1RKZHlSRG2PENndGswemOG9xmf53miIssucr7zZRjq+Rx8nplZ9j8r4rzHFXBeULQ0/Fzjsh5ROK0R59oxy/6vhfYtpHZtIOnRjXJHln9J+usuW+uJRCLxqB9dTiQSRyBBtnEom2EWEqYrV9TZ375ACpBJFE0qx+KBYRjFoSf6zA5F34WrkFoONIHskeQOJNq1bflMNoyichh6H7vez8PiNceocMaiIJh/HXyZrsVxjRqjUmqWdwTOCc0tBI5AzijozXoEchKCtay/Ryvcr3TThmPIf5XxSNLrau/Lsv8DpNdj/wBF1POhP4oABllJ6eqIo3paZ6u3/h+wYWC8NYpU5NP79xjif3/eiWrwgkxGZQElb5Xiuu40x3HGesMESpG+kc4si54oZXIsWvncBvh6ic0aTGddO+jv8Bqdtc+zgH+T3//aMIz8WRVdZ4ejm7fVUN/PEd7P1ZBT3IAEuwotq3C9n6+j0phnkGBlL0wk0Kg+bkIBgGaUKfUXVNb3UpxGGRXJbmhx0C/5fA6pXlt3kTok7sjyKij3P2zHERn2j1r5/g9S5syFTJHlhahvWq4MBOaGnmNFF8/hoFWp8Lnzrfc6L+I5rs/zOfLh+IjzbZ1l/99G7L97lv3DDEb1s4W+N4sVWQa1ygrb0dyN58uZZDLZkEgkXglEl7+Zw2Ej0PspiRZuFlP+6PNC5DRfjCLhfuqSYRj50YBEZb4L/AK4ApVIhPUv4thWImGbfyHn4wTUWcGyTYxKpwG9b/338lto8ckwfPYnVX/nKQovmzRqgDidZQeJLIRtuKyLY26LOOaPOZ4zk7PsorZAuUQz/frq8PF/z+HYn0Uc9z65K4/uTXrqt4uii9mO6U7ayL2hcy0nu4L4HhH2zSK3GvmeRL8n4nKWd6AzZcvf2oDDu/GcDnAQOVx4m5ubf+o7y01NTYXUBTaiWuNj0Wfkv6S/nnJsc4EbUI/vsZh4mGFE4WduJFEdcCU4xcX4vPcp4t/IMIrBIPR96L9n7yX+bDajMvgWqWVuD1O49o5RI4S/5OajleFibF19QZ4Scf4X6TpKPIxUVUN/C6cmR5HNWXbRF3y2FOO+qG9f+Lgvyc3h7YnS2MLHv4HSaLNxONHq1Fd3cdxlyMH9CxLnylXYrQGlIYfPd0sOx70VcdyfyN4OaiSKhvr7t4eOj8NZBphC+mvpQEJY+aw0DkeLJf4X9Kjsu8OECRP6JhKJTzyHuWPixIlb5Gt8BIPQDexk4rshX4E+BxfTKR5mGPXGMFTOcxXR18xibx3o+vg5+tx/Tnr3glJsS9GCaRLYnsy9mw2jnGyEdDj892lLvOYYFcARpOrl3EXumatGjeKgN0OpSCJRqyi2Qb2SgxGmxahZ/Ms5PPdY1MIouBK4EKUHZ1OoWwtFcoP8DX1AfCfyfeQQ34bSgduRUvB+wE9Jrcf1ORX4XQ52A2yM6jvDH8A2VBN2A6qhmY8crG2QQ7EX6byL6lYXZDnfZSiV2ucDVI/7jGfHh+imaTly/tYFdgV+SLoTsxyJu3TVgP1E4P8i5t9ATvPjqJfxEOQ07oeENvwFlvdQK6xjAsfOJzfn9HNSUwEvAU7K4bhMNKKUrf0iHluE/l8Poj6jn6Ebw8HotW2IVLR3AnYm9SZxNDCnq5M3NTWd4TiO/zn6R2traykESUaTWvu8LeXvcz2P1Nrn2XQqlxpGLdAAfAOl+O2HPmeFOo5LUR3xu6ilyTzvp//7J+hauMLbFmd4nsuBQ5H+x5XoOjsIpaUOR/XPa3i/r4a+Q9cDvlag3aDr5L3oRvQez2bDiIP90Pe7/zn8IQosGPXHj1EGnu8L/AuVki2LzSKjYijlavKZGc45CDlN4f1/kKftUWrFT5P9Jj8qsvwzJO5V6Ou8OE+7QS03wn18893mkJuK92XdPI+/taOLSS44wP0FnmcBWvSYHpqPK7IMWti4psDXk2nrMrIMMHny5MGJROJzP7qcSCQ2L8Lr6Yr+yHk+FS0GfEhprxVR20oUhb8a1UNa6yqjWtkBtUH8iPw/B58ix/L3wM+BfVAWTjE+CxujDJNsZTWZ6I8WAr+HhCuvBZ4ntx7N4e+VWWgheljhL8UwCiZ4/7cUBW2M+uInpGYzXo8yQQ0DKO3N7pkZzvn3iH3/WoDtDaTX07rARVmOyeQsO0iRO596znZgKoXftGyDlKML+dvOIvc65GI4y5+Sf2uqwcCjeZ5nLp1tSSrJWfY5hdS0rUK3d1HEJicSiUQyIPR1Q9FeTX6siWp5pqD331JKe/2I2haQ2rrKbq6NSmUkckTDrfSybbWwQBRcaLua/NLL29Dn+1jveQyjHDik3pfOxdoD1RPNpF6HrsXq140QpbyxPTPifCdE7PcandLs+TIcpZwFn68DNZ+PIpOz7LMvUtfu6rU9T3ZRrVzpBZyGUqNz+Zv+F9Uu53MDtTrqR309+UcI5wEXULgKYF+0avtlF+dZjtKzg4qUlegsg9KrzyD3/5m/zUELF3uQe904kB5dnjBhwmZFei3doQe6mT8B3RTHJR72pnd+X0zIVoONuOgJfB+VEOTy3u1A3yVTkJJ0rdbGfQ1ljl1P+vU507YAlUONicFeo/7oS6qezGxMiLIemEzqdSeYhm0YgByuUrZ5WEp6rv8g0mu0lnn7FkpUD8mVqJ40TFTN8kmortXHQdHNvVHf2tXRivdHKBJ8K13X7OaLg1La9kHpucPRyvoC5OC+juqM/1eEc41CdXMboNqzoXQuVixENzMvoZ5ysylOT+FBqOZ6N/Q/GIz+pu+i1Pnb0d83SPj/6qKoblcMIXUxYTmlq33dGDm/GyJHfxh6f3+J3n+vo8Wg58heS98lzc3NZ7muewaA67rXT5s2LVN7tThZA312/NrnnSl/K5lFqH7cr31+hPT3lmEUk6FIp+EkdH3Lxhcoenq3t80trWkVRyMS+fLrtrch++Kvi2qbLwJmemPDKAUj0f2I3+7wSvIvDzQqg15IA2dhln3OBn4TGP8BOBm7xhghqi29qxjk4iwbRsUxefLkwe3t7W+jxYCO9vb2LadPn17sRZti04giQ0HneSvKv3Lri4fNQsKCz2CiHUb32QCYiNKG+2XZbzkSi7kWLXouL71pVcO6wFHetnEX+76InOa/ogVxwyg2Y5Heiq99k+3+cD26uQhulIwEcCMS6Q3jABeirE6fqaiXvWEYdJ2GbRgVSyKRODtQu3xd3PYUyEBSW1d9TPlTt/3a0D9irauM/FkNpU5H9bz3N1+46lRSy0uMzGyC/q5zyf75fRuVf1i6pFEKgm1NV6ASiSia6NRYMSqH1VEGT1TdeQMq+QteT84sn2mGUR2Ys2xULRMmTBiaSCQWeM5y+6RJk7qKxFQLayIBuYuRg7GM8jvQ85DznkTiYUNK+YKrhM1RSYghBqBFnmydDL5EizC18tmMgwb0GZxF9s/si+QvPGkYuRAURv0UtVcMMw2lahuVhe8Mh0VUG9H/K6gZMaGslhlGlWDOslHVJBKJcwLR5ZvitqdE9KNwRd1ibW2kKxPXWySrHxI8nIJ0BuqVRhRt+oTM75d3kKqqLbIUlx2RMFi2llQzgUoQPTRqh55I78J/jz1PukL7dWhhdzWMSmFrOoVGg99ZvYCbSHWUTy67dYZRJZizbFQ1XnR5vq+MPXHixB3itqlM+K2rkkggaQnld6AXomjXFM+Wekiv3R3dWHyKoqp94jWn7GwGPEnm98TrwJFYq5FSsw4S4FlB9P9hOXAWnbWmhtFd1gDeo/M99g9StX781pi/Kr9pRgQOqdkovo5Eb+CWwHwbJtxmGFkxZ9moehKJxORAdPmRuO2JiUppXTUXuIHO1lW1eLP+F1IjqCeQ3tWg1uiJFgcylQR84j1u7WXKy0iU5t5O5sWL3WKzzqg1tiZ1YTYoAvW2N/ch1rKwEjiK1GtBL+Qw3xuYWwkcHZeBhlEtmLNsVD3JZLJPIpF4x3eYm5ubD4rbpgphMGpRlkT1x59Rfud5EVrdvhiJh40q5QsuE0NJ79H+b9T6pxbZFi2+RP1/F6De8eGUTKO8bIMyTKL+R+3AdOovC8IoDceQ+t7yv2+DC2lWOx8v/VCnm+Ai2kDggcB4OfCduAw0jGrCnGWjJkgkEj8IRJdfSSaTlgYazWjktF6MWkZlikiVOvrsi4ftBfQt5QsuEd8j+rU9AGwXo13F5lgyp/jfTtd9lI3ychCpqbLB7TnUmsowust0Ot9XC1EtvT9uQwukRnycR+pnvw14PDBeDOwTm3WGUWWYs2zUBOPGjWtMJBIvBhxmq8HJjQF0iofdAHxE+Z3nqNZV1dD3Plj3Fd7uAzaNz7Ru05tUBdzgNg84LD7TjC4YjD5LUWUYnwJ7x2eaUSM0oh7p/vtqDunvtW1is66+GU16G7/gtWARsGds1hlVTzXcnBWbgaQ7x/eilELDqCqam5sPcl33dm/4wYABAzZMJpNLYjWqOlkT3ejsjBzpbSh/CucC4GngMeBZ7+fnZbahK74GvAQMyvB4G/BnJLQ0r1xGFYGvIaXUcITcBa4AEqhvp1HZ7IkWPMJtftqA01GrH7fcRhlVxTDgJ8jB+gBlBb2PylAGILG/9SOO60BtiX5UFiuNIP8EDia6W8US9Nl/2RsvRNllIAd7MZ2154YRST06y4ZRUzQ1NT3gOM7u3vAXra2tU2M1qDboiXoM+47zNsDXY7DjLTqd51loUa8jBjuCnAT8vot9lgAzkGp4pTuZX0eth0aE5hcBPwRuLLtFRncYhAT/vh3x2B/RYnncnyGjshkJXEW6UNxnKFNhPaLV75ej7MXPSmpd/vQDhiN17/7IqfTbKg1CUfN+3s8vvfkv6ExfXoFe2ydoEXRBuQzPgT3R9bsQ7gV+ib5fDSMj5iwbRpUzceLEHRoaGmajz/P8nj17bnD++edX2pd1LTAC+AadEeid6GxNUS4WAS/Q6Tw/DHxcZhsavPOOzWHfz4ELUL34slIaVSCZHOX/ISGYl8pukVEMHGAScC7pqu1/A45D0WbDyEQDyig5h3Sl6w6io5ggtexyLlj3QHX5G3nbeihTajXkII+g+GKEy+h0nD9GpUxvISX614HXkHNdanoA/wE2JL/uDE+jTgYPlsIoo/YwZ9kwaoBEInETcKg3vKS1tfWkOO2pExqBMaSmb48h801UqZiHHGc/Av00pb9R2RhFuXNtm/QOcAZwDZUT1dsauIf0Xtl3oNYilR4RN7pmd+DvwOqh+euRwvHKsltkVBtbogWWjXPY1xdzHElnqm8x+RqwPVq03Qh934ym8toVdqBrvu84Pwc8BbxKcdOdT0ELsbnY0wC8gXpi31RkO4wax5xlw6gBJkyYsH5jY+NLyHlpdxznGy0tLc/HbVcdMgilb/vO8w6kO2OlZiVabfed52eRmFixOQPVJufDy0gRPO7U5u1QCt7g0PylKM3cbqRqh42A+0lXMf8HUngvhVNj1BZ90HVrEnK8uopiHgrc3M1z9kdO8Q7IQd4eRYyrmQWo5vtJ5Dw/gdLaC2Eo8Ca6hmfyZVzvsbnou+oKLKPEKABzlg2jRkgkEuchIQuAx1pbW3fBbvorgdGk1j5vS/kjAfPodJxnAbNRXXF36IGi2FsWcOzjKF3xkW7aUAhrIrvDN56XAD/HPjO1yLrIYQ4Lf00HJpbdGqNa2RP4K6r9zXT/3I4WKsP1zrmwGepdvx/6zijW98QKlDb9ITDfm1uAHP8lKBNpJXIk+6LFAF/EcQh6rf3R6x5O8cqP/PZud3vbE+TuzF4KnEj0/8F3kr9AuhmVWgZkVAnmLBtGjZBMJvstWrToFWAdANd1vz9t2rTrYjbLSKc/sBWdzvOuKG2vnLSj9DjfeX4MRX3zdRS3Q453PvViQWYCTSgSXg76IAd929D8VOS8VwurkL1NTTEWQ0DKwFtleXwW1XMTug5ymMNKxscDfyq/OUaVMhj4A3AU2WuXt6Dr61pf4AA6HeS1C7TpQ3Q9fw2lPr+PFkg/QfXExe6qMIBOx3k1ZPcGKItjQ/R9Vkg50heoBeHdwG1kjjpvgv62UedwgaVIYPJ8KkuMzDAMw4ibRCLx/UDf5Xknn3xyphY/RmWxJvAttAo+Czk65e79vMA79xTPlmE52n5xN8/bjvpdr5vj+brDNRHnP78M5y02u5P9b1qsnuvndnGerxXpPOVibeBdUl/DUpTqahj5MA5dM9tJ/1x0IOX1KBqBvVCbqYURx2bblqBr9DTkrG9LeilJJdAbRckPQ+nrdyGHPZ/XugK4HTgCLSoEeT5i/3bvmD8iJ94wDMMwokkkEg8FHObz4rbHKIgeaPX8BNQG57/oBqzcDvSb3vlPJXNaYD9vv+6eazFy1IcU+DfriokR57yF8guyFYOunOWHinCOBtIdy2p3lkHCbuHFqLnYDbaRP+ugz5rvIIedveCC45ZAK+rdnOs18X/o+nsSyiQJq3JXEw6KPB+LIvPPEb3QELUtRIsLeyGdgfDCRAdacA2XWRhGUbA0bMOoMRKJxKZIqbgHsKKjo2OzCy+88PWYzTK6zxAk+OLXP++M0nHLyWK0qu/XPz+MVE/3Q9GDYuC3m/odivoVg68je/sE5l4BdqQ60/R2Bx7I8riL0iLf7MY59kXpkNlYB3ivG+eIi0ORyFzwHuh24OB4zDGqGAepMreg79zge+o8VId7CnL0umIpKomZ6W213v93VXQt2ws4iNwEzNpI7XF9O2oD9UrRrTMMwzBql0QicUkgunxP3PYYJaERRZ+PRanQz5D7Sn0xt7nohuW5Ij/veyiyXmg9dPDv9GzouT9D/Uirla4iyy5Kf+wO1+VwjmqMLPucT/rrOSJWi4xqZlPUlz34fmqj68/QXOAiYB9SF/PqjQakgXEmyqTq6u+2ErWFGxmHsUZ9YZFlw6hBTj/99GErV658jc40sENaW1tvjdMmoywMRMIyfuuq7ZEASzXzCrqBKrQ35k+R0nWQw4m/fVV3iIosv0PqjePbaEGgkL7WQ5BAUPDmPfz8UL2RZdDN+cPoc+LzAepduygWi4xqZ22UYbNpF/t9CfwTaSg8gLUvi2Ir1O/+CLJHnNtQG7gk6uNsGIZhGLnR1NT0k0B0+T0T+6pb1kRiNBfTqV5c7uhzMbbHyb8dyyCkBht8nuvzfI5KJCqyfDFqCxOc273A5/9J6HmeQ+q0tRRZBtiY9M9Dvr3DDaM/8BtU0pHp+tWByhqiBKuMzPiCaC+gOvBskebLSe+nbhjdphqFTQzDyIGBAwdehuqfANbu1auXiX3VJ3NRFNUX6RqIap9PQz1D58RnWl7sgMR0bkE1yLlwMrB6YLwEaC6uWRXDUtIXAo4r8LnGh8ZXFvg8lc4rqNdykNMovxaAUZ30RKUibwBn09mbOMxtwOZI2+HvFE+LoR5oR/XbW6PI/S/Qd1qYHsCPgbeQIna1Z1QZFYSlYRtGDdPU1DTGcZznUSuHDsdxdm1paXmsq+OMumNNOvs+7wzshFSuK5V24CqUnv1+hn36o4WA4E3TWXS/lrcSiErDngrcinos+ywGRqC0z1zZiNR0xhXoJvXPqKVYkGpOw/YZiFSHg4sqZwC/jccco0rYHfg/1Fc4inlI9XkeKof4VZnsqgf6oNZZp6IWVVHMR8Jff6Kw8h3DMAyjXkgkEmcH0rFfOfnkk3vHbZNR8fitq45Fq/Rxta7qaluClLOjIoE/DO27gNK1pSo3UWnYU7zHXgnN59tzeWro+H9487WYhu3zC1Jf1zyqu02PUToGo5KHTGKKnyInLVjvv2qZbawn9gL+Q+bviEfRAqBhFIylYRtGjbN8+fJzgZe94ZjevXtPitMeoypoQw7y1cCJyHFeBdgbRWf/hVo8xU1flFb9JjCJ1FrAH4X2vRT4okx2xcnVoXE+qdg9kKhOkKu6Z05VcAnq5eqzBnBATLYYlcuBwIuoFVT4/nkJWmhaz/u5LPDYp2Wxrj6ZiXpYH45EDcOMRa00J9P9zgqGYRhGrdLc3LxLIpHo8KLLy5qamnKt+TSMTDQgJ/oilOYbd5TZBT5G4lTrheY7gPVL82eIhWyR5bVIbVnTQe5tsg4MPedHdEZYazmyDMqgCL62WhCCM4rDAOBaoq857Sgde43YrDN8+qEskUzfR09iraaMArDIsmHUAS0tLY86jnO5N+ztOM7/YZoFRvdYFa3Wn4puJiuB1VAEOVyX/xiqS60HPgDuC4wd4Jgcjx0fGv8VqczWA1eGxvsCvWKww6gs1kc6AN+PeOx/wJ5oge7DchplRLIELRpuhPQbwmwHPIt6WhtGzpizbBh1wsqVKyfTqSK5S1NT04lx2mNUNeNQmnauTli5GR4a11uP8XDq9Hi6/r4fSrqAVzilu5Z5EkXSfQYjBXajfjkAeIp0Eak2lGq9KVLoNyqLucAhKDU7nAI/DLgTCT2aD2TkhL1RDKNOuOiii75wHOcUf+w4Tmtzc/MGcdpkVB3rol6hN1BdojWHorq2euEWpAbrM5Kue1R/H6nm+zyLhHPqhQ7g3tDcjnEYYsSOg5T2byddPPA/qGvAL4DlZbbLyI8b0YJGeLG0Ef1/r6eyuz4YFYI5y4ZRR7S0tPwDfYEA9Hdd94pkMmnXAaMrGoEJwEsoPbXa2AF4BvgLandU6ywj/57L40PjK4tlTBXxRGi8bSxWGHHioN7bSdLvka9DbfXqaRGp2vkI+A7qn94WeuwwtPg7sNxGGdWF3SQbRp3Rs2fPn6LWKABjFy9enIjTHqPi2Rx4HLgQ9S6uVhqRQ/ga0IrS8WqZK0Pjw8h8U7gpipb5rAD+XgKbKp1/h8aWeVNfOMDvkQ5DkDYUSf4+6l1uVBcuave1NxKBDLIL6lk/tNxGGdWDOcuGUWecf/75nzmOMx59geC67tmJRGLzeK0yKpA+wLkoIltLEbY+QBMS5zmd2k3DexL1XPbpjxzm7ZZsrgAAIABJREFUKMK9mG+jPtvdvBUarxuHEUYsNKJa/5+F5j9BvXynlt0io9g8BGwPvBCa/wZwD+YwGxkwZ9kw6pCWlpZ7Hcf5szfsDVyVTCZN+dXw2RV4Hvglna2DqpVFyGl8HLgL+Bvqq3sJihjtHp9pJScs9BWVit2DdKXfeuitHMXHpKZqDsIUseuFK0gXLPwQXR8eLr85Rol4G/gmWkwM8g3URaCas6eMEtEjbgMMw4gH13VPQ18a6wNbLlq06Azg17EaZcTNYOBs4OdUzmLqMiRWlet2LHB84PipwDlltLeSuAq9dv+7flfUc/nNwD4Hktoj9iMUZalHXJRmOzgw1x+lpRu1y2R03QgyD0WUXy6/OUaJ+QK1/Lqd1MXSrVG7vEPxMu8MA8xZNoy6pbW1dXFzc/N413UfRilopzc3N9/T0tLyaNy2GbHwbeAPwFpFfE4X3ZjM934Gt6i58GOF1AeG2x+FRV3qiQ9RtGR/b+z3XE4G9hkfOqaeeitHEX7t1Z5ZYWRnH1RuEuRdYA9SF5Uqnd2I7gUNug5PAJYW4TzfRi21olhGer13pbIYOAgpZe8VmP8O8Bu0aGwYgL44DcOoY5qbm1td123yhq8DW7e2tpqISf2wBjCDzPWsC+naqc302IJSGp6BJGoL4nN2aFwL7I5EaYJMRSJEYQ4nVRn7HWA0apU0DPUkDaYabw68GPE8t5G+ELEO8F7OVlcHS4C+gfEATNSpVtkQpeMOCcx9jupa/xeLRYXzM7TYmYnvIzXv7vI8sEWGx75EpQvVRH9gFqmtBV0UXf5nLBYZFYdFlg2jzlm2bNmv+vTps4/rupuhm4cZwA9jNssoD31QGu49yKEKO8DzkVNVTSwMjcN9UuuNW4DP6FT/9nsuP4iizEFH+WmiHeV6oS/6TPi0I+fZqD36ADeT6ii3Ad+j+hzlXDiO7jvLW5PZUa5WFgOHoGvfat6cA1yNXmtY9M+oQyqlJs0wjJiYMWPG8vb29uOA5d7UDxKJxNFx2mSUjWXAn4E/ATcBM4FnUfrhZ1Sfowzwfmi8bhxGVBAryNxzeXxovl6FvXxGkppx9z5Wu1ir/BbYJDSXQNfAWmRv4GvdfI7xRbCjEnkHpV8HtQkGIIfZ/CTD3gSGYcCFF174b2BSYOrSpqamMXHZYxjdYE5obO/jdCf4MCT2FYwSRTnV9Ub4vRJ+Lxm1wTaohjfIX1Ev3loiuNDTAHRnEbwXcGSW5692HiP9PbEz8OMYbDEqDHOWDcMAoLW1dYbjOLd4wwENDQ03TJgwoW/Wgwyj8vgvqaJe69OZglyvPIX+Lj79UdQkyK3UZ2/lIDuExuF+rEZtcCEStfT5ADglJltKSVisczyFaxV9C1g1MP4AtWGqJS5B7QWDnENqqr5Rh5izbBiGj9vY2PhD13XfBnBdd7PGxsYpMdtkGPmyhFTH0AF2icmWSiIcXR4ZGl9ZJjsqmV1D46discIoJd8h/f98PNJoqDWuR33mfTYkfUEoV8aHxleimv5a40RSBf1WQ63FjDrGnGXDML5iypQp8x3HOYLO9iknNzU1HRKnTYZRAA+HxmEV53rkajK30foIuLeMtlQiqyEVZB8XeCQmW4zS8cvQ+F+kRxNrhcVIxCzIcVE7dsFwYN/Q3F8LsqjyeQ91Fgjycyw7qa4xZ9kwjBRaW1ufdF036Q0dx3Gu+MUvfrFujCYZRr7cHhp/G+gdhyEVxEdI9TyKq6jvftSgOu7gPdG/SReLM6qbfYFvBMYdRLdbqyXCGSVHAP3yfI5jSe03Pgt4rTtGVTit6HrpMwD4aUy2GBWAOcuGYaQxcODAKY7j3OcNV2lra7vmhBNO6Jn1IMOoHB5Gat4+w1B7kHrnygzz15TTiArl+ND4lsi9jGrmhND4n6SWbNQiD5La/mgwWjzMh2NC41pXzV+K6tqDnEhqnbtRR5izbBhGGslkssN13aOBed7UzoMGDWqN0ybDyIOVwLWhubDSaT1yK4qsBbetqO/eyqC+01sFxh2kC6AZ1c1qwEGhuelxGFJmXNIXw/JJxd4O2CwwXoraDNY6fyS1x/raqP2WUYeYs2wYRiStra0fd3R0HEOniMcpTU1N4RVmw6hULie1tcn2wB4x2VIprER9tIPb87FaVBmcHhrfiXqvGrXDt1H7I5/XgNkx2VJu/kLqtTCfnsvjQ+ObqE0xtDALgBtDc4fGYYgRP+YsG4aRkQsvvPB+x3G+EkRxHOeypqambeK0yTBy5CXk9ARpwb73jFT2IF286II4DDFKSjiq/Ddqq09wNt4mVayuATgqh+N6AYeH5mo9BTvI30LjAym89ZZRxdhNg2EYWWlpaWkBbvCGfRzH+UdTU9Oq2Y4xjArhnNB4a+DoOAwxKpIepKfiPkx6f1qjumlAqfZBbovDkBi5MjT+AV07ft8hVQX6feCh4plU8TwEfBkYjwA2jscUI07MWTYMoyvcJUuW/AhF6gBGNjQ0/H3cuHEmdmFUOk8A/wjNXQisEYMtRuUxGdg8MHaBSTHZYpSOMcCQwPhz4IWYbImLG0l1/DYktVVaFOND4yuozd7KmVgBPBaa6+pvZtQg5iwbhtEll1xyySLHcb6LV6vkuu6e66yzTjhqZxiVyCRgWWA8DIm3WDpdfbM58JvQ3DXAUzHYYpSWLULjp6mfFGyfxaQvHGYT+lqLVEErl/oUvXsyNN4qci+jpjFn2TCMnGhpaXnDcZxjkFIsjuNMbm5uHhezWYbRFW8BZ4XmDgZ+HYMtRmUwDLiZ1N7bnwLN8ZhjlJjRofHLsVgRP+F64yOBvhn2PZbUVkmPAm+WwqgKJ/xeGRWLFUasmLNsGEbOtLS0/MtxHD+i7Liu++dEIrFprEYZRte0kB4hSFK9vZefIb0F1IwSn3NixDk/KvE5S0FPpOi7Xmj+JKrz9Rhds05o/HYcRlQAD5Pq8GbruXxsaHxlKQyqAuaExiNjscKIFXOWDcPIi/79+58F3OENBwL/OvXUU4fHaJJhdEU78D3g48BcA/BXUnuIVgtfkt4C6oMSn/N/EedcUeJzloKLgW+G5n5Pp4ihUXsMCo0/jcWK+HHRNS9IVCr2TqjO22cx9dFbOYrweyX8XjLqAHOWDcPIi2Qy2dHW1nY08Ko3NbJnz57/TCaTfeK0yzC64B1gHOo17DMALfxsEItFRrk5E/hpaO5BFDU3apd+ofGSWKyoDK7EK6Xy2If0nsvjQ+ObSBUHqycWh8b9Y7HCiBVzlg3DyJuLLrroC+AAOiN1Oy5atOhqTDTJqGweAU4MzX0N1eNZOUFtcxZKvQ/yNso4WBne2agpOkLjer73fQelY/uEey73RYuKQeqpt3KY8Hsl/F4y6oB6vmAYhtENWltb57iueyiw3Jsal0gkzozTJsPIgb8Al4bmhgP3k9pGyKgNHGAacEZofiESevuk7BYZ5caig6lcGRoHey5/l9Q2W2+T6lzXG+H3Svi9ZNQB5iwbhlEw06ZNm+U4znF0tuE4I5FIHB2nTYaRAz8H/hyaWx14ANil/OYYJaIXahMWTrP+AtgXeLHsFhlxMD80rneNjX+Quefy+NC+V1Hf0dQRofHnsVhhxIo5y4ZhdIuWlpbrXdc93xs6wOUTJ07cKU6bDKMLOoDjgd+F5ochh3ly2S0yis2aqB75+NC87yg/UXaLjLh4OzQOt5KqNxYDN4bmjgPWBnYPzEUJgtUb64bGb8dggxEz5iwbhtFtpk2b9mvHca7zhn0aGhpunTBhwvqxGmUY2XGB04ALQ/M9gCnA37B0zWplLGqvFV60+wQpYT9VboOMWAn3B7Zyi+ieyz8jtbdyuNVUPbJFaFzvf4+6xJxlwzCKgdvW1vYjx3H8aM2qjY2NdyUSidVjtcowsuMCTaie1Q09diTwGPD1chtlFEwDkEDZAeH0ybeQo/xCmW0y4ufZ0Hhr1G+7nnkUtYPzGYw+O0HqWdjLZ4fQ+JlYrDBixZxlwzCKwvTp05cC3wHe9abWdxzn1kQiYdE5o9L5LXAQ6bWNWwDPo0hzr3IbZeTF+shJbiHdEbob2BZ4udxGGRXBe8DcwLgvyj6oZ1zg6tBc8HNTz72VfQbTWcvtY+UbdYg5y4ZhFI2WlpYPXdfdH08Ew3XdHYBbksmkORpGpXMnsB3pok89UQ3zM97jRmXRA/1/XgJ2Cz3mAlPRQogJ89Q394XGB8diRWXxF6A9w2M3AIvKaEslsh+pCwivooUXo84wZ9kwjKIybdq0lxsaGvans8XCXosXL74C68FsVD7/A3YErot4bDNgNqpxHlpOo4yM7IYWMaYAvUOPfQEcAvyCzA6BUT/cFhofiaVivw88lOExS8GGY0Pj8HvIqBPMWTYMo+hccMEFTwHfA9oAXNc9qqmpaUq8VhlGTiwGvo8iT3NDjzUCE4B3kIM2sLymGR6boMjXQ6QL8ADcgRY37ObW8LkbWBAYDwe+FZMtlcQfUT1/cJsNPBKnURXASKSaHySsIG7UCeYsG4ZRElpbW+8AfoAnnOQ4zqREIhHud2oYlcrtyCm7LOKxASj19xXgBJQKbJSeddDN/QvAuIjHP0YtcA5CUTPD8FkC/D00NykOQyqMG4H1QtvOpAse1hsTSVUG/w8m7lW3mLNsGEbJaG1tvcZ13V8Hp5qamo6LzSDDyI8vgBOBA+gUrguyFnLeXgF+jpxoo/hsCVwJvIEWJxoj9vkrUi4PixYZhs+lpDqB2wN7x2SLUbmsAfw4NDcjDkOMysCcZcMwSsq0adPOc13X/6JxHMe5rLm5eb9YjTKM/LgL2AA5zp9EPL4+upn6ALgYpfAZ3aMB2AtF+J9DEeMoocDHUe3yscBnZbPOqEZeQCn6QaZjmSFGKucB/QLj97FFuLrGnGXDMErOtGnTTkWRH4BeruvenEgk9ojTJsPIkxUoJXsj4AJgacQ+g4BTkFDY9cA+REdBjcysgerCX0UKxgcRLQ74H+BAYCesvtLInbNJjS5vghbBDAOUbRDOfjsfXf+NOsWcZcMwyoG7fPny44F7vXFf4Lbm5uadY7TJMAphPqpX3hC4HFgesU8P4HDgHtRq5EJgm3IZWIUMAI5BIkzvo7/XBhn2fcPbdyvU7ssw8uFp4JrQ3FSUHWLUN72BP5PqG72CrvNGHWPOsmEYZWHGjBnL29vbD3Fd90Fvqr/rundOmjTJetca1cj7qH52XeC3RKdnA4xAkdJngJeBX6Ma3HpvpTYI+C5wLfARSnPcl8yR+AeBbwNjkLPTUQYbjdrkl8DCwLg/apUUleZv1A+tKNPAxwVOBVbGY45RKdT7l7VhGGUmmUz2W7Ro0d3ALt7UF67r7jVt2rRn47TLMLpJb9QuLYHaFnXFJ6j10b9QXe78kllWOYxG7XoOAnala+dkBXArMA14srSmGXXGCUicL8hVwPjym1IUjgbOCs01AzeX8JwPkKrPsBjYvITnKyXjgb+E5v4EHF9+U4xKw5xlwzDKzuTJkwe3t7ffB2zrTX3S2Ni4+9SpU/8bp12GUQQc4JsoVfhQFEHtinbkDD7q/XyS9B7P1UZPdOO8vbftBayZ47EvII2Da4EPS2KdUe84SOxr/9D8KZjycb0xFrif1MW7t1EG0IKoA4z6wpxlwzBi4bTTThvSo0ePB1DtIcDHruvuNm3atFfjtMswikhfFEk9CtiP/NI830dO8xNIDfo1pLZdifRGNcabIMd4O2Br9Ppz5V3gbyjF2hbNjHKwCvqMBevj21CruPtiscgoN+uia+zwwNxi1Gv6hTgMMioPc5YNw4iNU089dXjPnj0fQnWIIDGk3VpbW+fEZ5VhlIRhKNJ8ALAnhfVk/hJ4HTnOr3q/vwPMAz4mWqG7WKwGrI5qsNdHquBjkNDZSApT/X4VteW6BZiF1SEb5Wcj5CwNCcwtAQ7BHOZaZ10UUR4dmHOBI4Ab4jDIqEzMWTYMI1YmTJiwVmNj48PAet7Ue47j7NnS0vJGnHYZRgnphVL/9kNpoJsW6XkX0ek4f0yn6NgC5IguQmI1y5BjPQTdBwxECt59gT7ethpyjFf3tmL0ol2M6hzvQsrXtihmVAL7opTs4ILPcqRBcGssFhmlZkPkKK8dmj8LSJbdGqOiMWfZMIzYaWpqGuk4zsN0ioXMdV13T0vJNuqEtZHg1XYojXkrlNpc7XwAPEVnOvkTRLfaMoy4OQ2YHppbgRzmW8pvjlFCvo4c5TVC839DQmlu2hFGXWPOsmEYFcHkyZPXaW9vv5/Ofpcft7e37zV9+vQX47TLMGKgFxKX2R6J4G2M0kUHxmlUFjpQzfHrqM7vCeQkvx+nUYaRJ6cAF5F6b9yGVKUvisUio9jsi4QDh4XmrwZ+iMQWDSMFc5YNw6gYmpub13Bd93608gswv6GhYZ8LLrjgmTjtMowKYU3kNG8Y+DkCidOsjhSoS8VClOL9CaqTfhXVTr/ubaWslzaMcnE88H9AQ2j+78CPUSmBUX04wCTgXNL1Ff4EnIhpJhgZMGfZMIyKwhP9mklnHecXDQ0N+15wwQVPxWmXYVQBqyKneTXkWA9EN/2DvcfDNcqLkCLwAcjhfgr4M4qmfeJtc1H987JyvQjDiJnvo57L4Tr9V4HveD+N6mEg6qF8aMRjfwR+hjnKRhbMWTYMo+JIJBKrIyXSzb2pBcD+ra2tj8dnlWHUJENRCnV/VE88CkWQDaOeORC1MRsSmv8CmICcaattrXx2Aa6gs7zLpx34FXAB9n80uiCcZmIYhhE7ra2tH/fs2XMP4N/e1GDg7ubm5l1iNMswapHPgb96v/cGTorRFsOoFO5AegH/Cc0PQVHKB0ntz2xUFv2AKcBDpDvKn6FsmqmYo2zkgEWWDcOoWE477bQhPXv2vMt13R28qeWu635/2rRpN8dqmGHUFhug1NIG5DyPRCnahlHv9AEuBcZHPLYEOBtoxYShKon9UHr1OhGP/RulY1vbOiNnwkXuhmEYFcMTTzyxbJdddrnJdd1d0BdfD8dxDt15553fmT179gtx22cYNcLnwNbAGNRreS7wdKwWGUZl0IZ6LX8O7EaqiF5PYC+Usv0m5oDFzRjkJJ9Lp06Dj4sWPb4HfFpmu4wqxyLLhmFUPMlkst+iRYtuRKlToC++ya2trS0xmmUYtcQuwCPe73NQtNmiZYbRySjgMuQgRzET+AXwbNksMgDWAs5ArZ/ComwAbyGV8wfKaZRRO1hk2TCMiuehhx5auc8++9ywfPnytR3H2Qot9O290047DZ09e/Y9cdtnGDXAu8D+wNpIIft5TPXXMIJ8gUS/PgB2RSnaQUYjp2wM8CKqjTVKx6rAmeh/sgPpOkxtwDTgcOCN8ppm1BIWWTYMo5pwEonEBUAiMHf1gAEDfpRMJtviMsowaoTvoX6yAI8BY2O0xTAqmbWA6cBhRN9LdyCRsOlIDMwoHl8HTgWOQWUjUTyFxAqfKZdRRu1izrJhGFVHIpGYDJyPdw1zHOe2tra2I6ZPn740XssMo6ppBF5HETKAnQBr12YYmdkWfRftmWWfF4BLkOq8fUcVzlhgMqoRz+S/vAb8BrgJU7o2ioSlYRuGUXXMnj37sZ122ulDlDbaAGzkOM5O22+//a1PPPHEspjNM4xqxUX3Bft648HAjfGZYxgVz1zgauAJYBNgRMQ+awDfAn6MItKfYr3Mc2VdFCG+DJgEbEi0o/wByjg7AXipXMYZ9YFFlg3DqFoSicS3UdqoXzv2co8ePQ6cMmXK2/FZZRhVzUBUvzwECXxthJR+DcPITgMwDjlt3+hi31eBa73NVLRTGYrqjI8Cdia7r/I28Dvg/7CovVEizFk2DKOqmThx4p4NDQ03A4O8qQ9d1z142rRp1vrGMArjAqDZ+/13qD7QMIzcGQucBhxC9ixOF0Wl7wTuQn2AO0puXeWxPuqPfABKae/Vxf6zUT34PzHVfqPEmLNsGEbVk0gkNgX+BYz0ppY5jjO+paXl+hjNMoxqZS3UbqUXsBh9rkzZ1zDyZxRwMmprFO79G8XHwD3Icb6X2v3c9UN9q/dHTvIGORyzArgZOclPlc40w0jFnGXDMGqC0047bUSPHj1uozP9zQXObm1tTcZnlWFULdegNEhQ79ipMdpiGNVOb2AfpOD8bbqOnPrMA2YhdfpngaeB5aUwsMSsiVKqxwLbeFu49VYmnkXiaNehxQTDKCvmLBuGUTMkEon+juP8zXXdgwPTlw8YMOBn1lrKMPJic9Rr2UEiRqNQZMcwjO4xFLVpOwopzudzL74UeA59Nl9B6vWvI52BSlB/Xg3pHPjbZsB26DXng9V0GxWDOcuGYdQU48aNa1xnnXWmO45zcmD6nuXLlx8+Y8aMhbEZZhjVx/3AHt7vxyHVX8MwisfaKA15P2BvOrU38mUJcppfQ8rQc1EU9hPv90+8rTuLxqsBq3s/1wyM10Yq1RuSv1PsswJ4FLgbpaD/txt2GkZRMWfZMIyapKmpaYLjOK1IoRTg342NjYdMnTr13TjtMowq4gDgDu/3F4EtqIzolWHUIj1RpNl3njej+C1ev0Cf4UXAShSpXub93gb09ewYgHyEId5xg/n/9u48Pqry+uP4505C2EFcwAX3vVJrtYoE1LphabWtVVyqFVotVVsVCUGs2k5brUISwNJq3VrrXrHVn1uxbqgQsErrioqKSlUUFAWCQJa5vz/OpLlzZ7LPzDPL9/16zYs7995MzmSS4Z55nueclv9L02UZti57LvbBXF2aH18kLZQsi0jBqqio+K7nebdjxUQAPolEIidNnz79SZdxieQJD+tZ+qX4/aOBx9yFI1JU+gH70bLG9xCs73A+Wge8hK0/ng88DXzsNCKRDlKyLCIFraKi4kDP8+7Dpo0B1Pu+f15NTc31LuMSyRNnATfEt+di1WtFxI0dsCKWe9GyLngPYJDLoAKap4M3Twl/HWuH9QbF2RJLCoCSZREpeBMnTtymR48ef/d9/+DmfZ7nXb9mzZqfXX/99Q0uYxPJcT2Bd4Gt4/f3A150Fo2IpDIYS5x3xT4YHhy/bRPY3rKb32Mdtv55I1bJ+uX47WNsSnUuFRoTSRslyyJSFM4777yePXv2vBb4YWD3fOCE6upqtaMQad1lwK/j23/GesaKSH4poaWA2GZYDtAfKMWWKpVgCTHAZ/F/1wJNWL/1euBM4Mb4sVnAhRmPWsQxJcsiUlQqKiomeJ73e6yICcB/fd8/vqamZrHLuERy2ObYiFFfrMfrzlj/VxEpLtsC72P5wzJsJFukoKW7sp2ISE6rqam53vO8Y2n55Hx7z/PmVVRUnOgyLpEcthq4Nb7dE/ipw1hExJ0PsTXIALtga6dFClq6S9KLiOS82trat0eOHHmP7/tHep43GCjzPG9seXn55sOGDXt88eLFKkQikmgpliR7wDDgD1i7GREpLtsBh8W33wNqHcYiknEaWRaRolRVVfV2fX19OfBAfJcHnD9gwIDHJ06cuI3D0ERy0ZvAg/HtzYHx7kIREYceDGwf6ywKkSzRmmURKXbe5MmTpwBX0DLbZhXw/erqavWUFWlxKPBUfPsdYHes+I+IFA8PW7e8Lfb3PwT41GlEIhmkadgiUvRqa2sXlJeXP+153higH1bI6LTy8vLI6NGjn543b55aYYjYlMsxwFCsr+sLWB9VESkuXwL2x2aovgC84jYckczRNGwREaC6unqe53lfo2X9VQnwy7q6uvumTp06yGFoIrlkZmC7wlkUIuKSpmJL0dA0bBGRgGg0WlpXV3c5cFFg93u+74+tqal5zlVcIjmiBCv2tUv8/ghgkbtwRMSBvsAnQC+ss8RgoNFpRCIZomnYIiIB8+bNi9XW1j42YsSIpZ7nHQOUAZt5nnf6iBEjPl24cOHzrmMUccjHrh2Oid8fCMxxF46IONAAjAJ2A3oDj2PLNEQKjkaWRURaMWXKlD1jsdg9WKscADzPu6+xsfHMmTNnrnYYmohL/YHlwGZYgZ89gbedRiQi2fYzYHZ8ezqJs7FECobWLIuItGL69OlvfPHFFyM8z7u9eZ/v+98tKSn5T0VFxSiXsYk4tA64Ib5dApzvMBYRceP+wPZxzqIQyTCNLIuIdEBlZeUZvu//AauWDTaidvl77733mzlz5qh9jhSb7YBl2DKF9cCOqH2MSLF5CfhyfHt34C2HsYhkhNYsi4h0QG1t7YuHHnronb7vH4y1zokAXx80aNDo8vLyJ2praz93HKJINq0D9gL2xRLm1cACpxGJSLZtDxwS314GPOswFpGMULIsItJB8+fP/3zYsGG39OzZMwYcis3OGQqMLy8vX1ZbW/uq2whFsuot4Gzs7+BLwO+xGRciUhw2Aj+Kb5cAtzqMRSQjNA1bRKQLKisrv+n7/s3AVvFdPvBHoLK6unq9s8BEsutx4Ij49jjgFoexiEh2RYAVWOuoBuz/wzVOIxJJMxX4EhHpgqqqqoexKtlz47s84Bzg5crKykNa/UKRwlIT2J6MPoQXKSYxWv4P7AEc7TAWkYzQNGwRkS6qra1dP3r06DsbGho2YtOyS4BBwA/Ky8t7Dhs27JnFixfH3EYpklFvAWOxEaUhwHxs7aKIFIcy7D0A4Avg/xzGIpJ2+gRYRCQNJk+ePAxbr7Vf8z7P815uamr6wYwZM150F5lIxp1FSyupucAYh7GISHYNAFZhSfMqYGtsxFmkIChZFhFJk2g02quuri6KTUdtnrmzEYj269evKhqN6gJCClFP4F3sIhngK1hLGREpDsHaBSOARQ5jEUkrTcMWEUmTefPmNdbW1j5WXl7+KHAYsDlQChxVX19/9MiRI5+qra39zG2UImnXBPQFDo/f7wnc7y4cEcmyLYBj4tsfA084jEUkrTSyLCKSAeedd96AsrKymZ7n/Siwe63v+xU1NTU3YdWzRQrF5sByLGneBOyMVckVkcK3K1a/AOBlkNi1AAAgAElEQVRFAsuRRPKdkmURkQyqrKz8hu/7NwHbBnY/43nemVVVVW+6ikskA67F+i4DXAFc6jAWEcmu14E949s7Ae+5C0UkfTQNW0Qkg2pra9867LDDbo3FYrsCe8d37wj8aMSIEeuPOeaY5+bNm6dRZikES4GfYh/EDwP+gPVeFZHCtxNQHt9eCjzvLhSR9FGyLCKSYfPnz/+itrb27vLy8n9ja5n7A2We532jvr5+9KhRo2oXLFjwieMwRbprNXAANrrUG/gAeM5pRCKSLfXA+Pi2B9zhLhSR9NE0bBGRLJo6deqgpqamq3zfnxDY3QDM6Nev3y+i0Wi9q9hE0uBQ4Kn49jJgD6wAmIgUtlJgJTAI6wKxJbDeaUQiaaBkWUTEgYqKijGe510HbB/Y/ZLv+z+qqalZ7CoukTRYBAyPb38PuNdhLCKSPXcCp8S3v4Oq4ksB0DRsEREHFi5c+NYhhxzyJ9/3twD2xz68HOJ53g9HjBjR55hjjlk4b948rfeUfLQOODG+PRT4k8NYRCR7emEfkAHUAQ84jEUkLTSyLCLiWGVl5SG+79+ITVlt9oHneRdUVVX9zVVcIl1UghX42SV+fwQ22iwihW1zrM9yKdY6bjvUJlHynEaWRUQcq62tXV5eXv5noA9wIBABBgAnlZeXf3n48OELFi1atM5pkCId52PXF8fE7w8E5rgLR0SyZANwNNbxoT82sqx+65LXNLIsIpJDJk2a9JWSkpI/+r5/cGD3F8Cv+/XrVxONRhtdxSbSCf2B5cBmWIGvPYG3U5zXG7vAFpHCcBFwVXw7CvzKXSgi3RdxHYCIiLSYMWPGi3379h3ped44rBUP2IjzVXV1dc9PmjTp4Da+XCRXrANuiG+XAOe3ct652QlHRLIkuE75W86iEEkTjSyLiOSoiRMnblNSUjLT87yTA7tjvu9fW1paesm0adPWOAtOpH3bYe2jyrAWMjsCnwaOD8OKfx2U/dBEJIPeBHbDlmRsj/VcF8lLWrMsIpKjFi1aVLdw4cJ7Ro0a9bTv+8OxvpWe53kH+b7/4xEjRmwaOnTo80uWLFEBFclF64C9gH2xhHk1sCBw/GLgKFqmbIpIYdgNax/nAa8B/3YbjkjXaWRZRCQPRKPRXnV1dVOBqUDPwKF/x2Kx82bMmFHrKDSRtuwLvIBdb3wI7AzUAz2Aj7DquYOBVa4CFJG0Gw08Et++DzjeYSwi3aJkWUQkj1RUVOwViUR+5/v+0YHdMeAvDQ0NF1999dUfu4pNpBVPAIfHt88AbsUunv8e3zcS0Ic9IoWjDPgEK/S3HpsVtdFpRCJdpGRZRCQPTZ48+TjP82b5vr9LYPd6oHrTpk1Xzp49e5Or2KRo7QCsid+CvgU8GN9+GfgKcH98vwf8ELg5OyGKSJbcA5wQ3x4DzHUYi0iXqRq2iEgeqq6ufqBv3757+74/EaiL7+4L/LJnz54vVVZWftNheFKcVgL3ArcBR9JyjfEwsCS+/WVgLHbx7GEFgHbPbpgikgUPBbaPdRaFSDdpZFlEJM9ddNFFOzQ1NVVjSUjQvU1NTVNmzpz5lou4pCgNAZ7FKl+/D9wE/AVLnptbSb2B9V0G68F8L8m/uyKS3wYDK7APzZZj7wkieUfJsohIgaisrDzM9/3fYUWVmjV4nvdn3/cvq66uXukqNikqX8IS5n6BfS8AuwAD4vdjtIw8v4RNzRaRwrIIq4oN9v/Syw5jEekStY4SESkQtbW1740ePfrG+vr6T4GDgd7Y+/wBwFnl5eUNo0ePXjxv3rwmp4FKoVsFvAicGr/vYaNMvQLnBD+sHwD8NjuhiUgWbQ0cEd/+LzDfYSwiXaKRZRGRAnThhRduXlJSMgWYSGKrqeWe511WVVV1K7ZeVCRTLgBmdfDcocAHGYxFRLJvP+A/8e1arPK9SF5RsiwiUsAqKyt3933/CuBEAu/5vu//KxKJTK6qqnrGXXRSBK4Fzu7AeYcD8zIbiog48C62XjkGbIMVAhTJG0qWRUSKQGVl5SG+79cABwZ2+8Ccpqami2fOnLnMUWhS2EqBf2AFvtq65phASwEwESkcwQ/MxgG3OIxFpNOULIuIFA+vsrLyRN/3pwE7B/Y3eJ7354aGhuisWbNWuApOCtYArNDPHqSuleID1cCUbAYlIlkR7LM+BzjJYSwinaZkWUSkyESj0bJ169ad43ner4CBgUNfALMbGxuvmjVr1ueOwpPCtDPwPPb7Fk6Ym4AHgOOzHZSIZFxv4BOgD7AW2AqodxqRSCcoWRYRKVIXX3zxVg0NDZdiU+TKAodWe543rbGxcfbMmTM3OApPCs8o4AksWY6Ejr0O7J31iEQkG+4HjotvHwU87jAWkU5RsiwiUuQmTZq0fUlJyaW+759J4qjfKqBm06ZNs2bPnr3JUXhSWMYBN6fYX4+NPKmtmUjh+Qnwx/j2TGCSw1hEOkXJsoiIADBlypS9Y7HYr0iunP0ucOXy5ctvmjNnjpIZ6a4rgakp9u8EvJfdUEQkC7YF3sf+X3kb2M1tOCIdp2RZREQSTJo06eBIJHIl8PXQoReBX1VXV9+HejRL13nAXSQX+jkaeCz74YhIFiwG9o9v740tvRDJeamqUoqISBFbuHDh+7W1tX8pLy9fAOyDjQoAbA2cXF5efsLIkSM/qa2tfc1dlJLnHgK+ifVdbfYm8LSbcEQkw7YDDotvvwfUOoxFpMM0siwiIm3xJk+ePBaIklyA6d+xWOxXM2bMeACNNEvnbQssxdYqe8As4EKnEYlIphwEPBvfngcc7i4UkY5TsiwiIu2KRqOR9evXn+D7/q+BvUKHX/I87/Kqqqp7UNIsnXMA8AzWXuYh4NgOfE0PoD+wWfx+f6A0/hi9Auc1Ya1qANYAMWA9sA5rkyYi2eNh65a3xf42hwCfOo1IpAOULIuISIcFkubfAHuGDitplq44Hvgb8A62pnGP+G0nrCfrYOwCu3l7yzR8z/XAR8DHwMrA9gpstPsN4MM0fB8RaXEjcGZ8+/vAnQ5jEekQJcsiItJpY8eOLdlpp51O833/UmD30OHnfN//bf/+/e+PRqMxF/FJzhuITcvcF0uMjwJ2cRpRsnVY4rwUK0b0GjaNdLnLoETy2PHA3+PbtwOnO4xFpEOULIuISJcFRpovx5Ke//F9/9VIJDK9b9++d0Sj0UZHIYp7JdjU/QPit5HAV4GIy6C64SPgeay672JgAbDaaUQi+aEv8Am2XGI1NhVb/zdITlOyLCIi3RaNRkvr6up+AFwC7Bo85nneMt/3p2/atOnm2bNnb3IToWTZbsCY+O1Q7CI5XXzgc2w9chOwAdiIXXSvC5zXEyseVgIMiO8biK1xLktjPE3AC8A/gLnAovg+EUk2Fzgmvn0oVrNAJGcpWRYRkbSJRqORurq6bwG/xEYRg1YC15aUlMycNm3amuxHJxnUG6tuOwb4BpYsd0U9sAyb9rwM+ABYhf3urAhsdzcZHYS1Qtsq/u8QbD30LrSsmR7Q6le37TPgUSwp+Ac2Ei0i5mfA7Pj2dOAih7GItEvJsoiIZMTkyZOPAn4NjAgdWgtc29TUNH3mzJmavpq/SrAE+XTge9iIbUc10bIGeAlWUOsN4F1yZ1rmNlgRuz2waeRfwz4A6tOJx4gBTwK3YWs117Z9ukjB2wHrswz2t7+Pw1hE2qVkWUREMmry5MlHeJ73c9/3jwwdWud53o2RSGTWtGnTVDQpf+wDjAXGYRWrO6JQ1vl2Z/31RuAx4Bbg/7BRdJFi9DIwLL69O/CWw1hE2qRkWUREsqKysnI/3/cnAaeRmFzEfN9/2PO8y6urq591FJ60rSdwCjAR2K8D528C5mNTkecCr2QuNOcGY2swxwBH07HWViuBPwLXomnaUnyuBKbGty8AfucwFpE2KVkWEZGsmjx58jDP86b6vn8yUBo85nne40B1VVXVI6hXcy7YCjgbOBdb29uWD7ER038AT2C9jItNBDgQW7f9baxvdFs2Yb1mZwEvZjY0kZwxEvswDWx9/2iHsYi0ScmyiIg4MXHixG1KS0t/go0sbBY85nney7FY7Pf9+/e/JRqNbnQTYVHbBbgYW4/cq43z1gL3Ymtyn0RVoMP2wmZSnAbs3M65TwDTgH9mOigRx0qwGRVbYssRtiSxkn3wPL2niFNKlkVExKmJEyduFk+azwe2DR3+wPO830UiketUQTsrBgOXARNovb1SEzZ6fBtwP9a6SdrmAeVY0nwqoQ+HQh7HPqh4LgtxibhyC/CD+PaJwN9Cx4cDa7DK+CLOKFkWEZGcEI1Gy9avX3+K7/uVtBR/aVbned4dkUjkd9OmTXvVRXwFri/W0uXntN4yaR02ZXgGVrlauqYf8H1s/ffebZz3IFABLM1GUCJZdhLw1/j2zcAPQ8fnAz8B9H4vTilZFhGRnFNRUTEKuMjzvG+R+H+VDzzued7177777t/nzJmjKXrd42EXqVdio8qpvIMV4PkTan2UThHgWOBC4OutnNMAXAdcgn72UlgGYH3Ty+L/bo21WgP4DnAfVkxQa/nFKSXLIiKSsyoqKg7wPG8SNk0vPC34Dd/3f7dhw4ZbrrnmmjoH4eW7XYDrgXBLr2bLgF8Ad6F1g5n2NeC3WDXtVP4LnAM8lLWIRNLHI3XBxseBI+LbI4BF2DrlV7Ee5/sD/8lGgCKtUbIsIiI574ILLhjSo0eP8cB5wHahw2s9z7srFovNrKmp0fq29kWAs4AabEpw2KdAFXA11htYsmckVuRrZCvH5wA/xUbiRPLFSOAoYDqJNQ4mYe9DAJdj9RJ+jH2IB1ZZ/vksxSiSkpJlERHJG+edd17PsrKyUzzPO5/ktjwxz/Pu933/D9XV1Y+j1lOp7ArcARyU4th67MK1mtSVaSU7PGAscAWwW4rjq7ACbPdlMyiRbroVOAz4JVbcqwl7P3orfvxFrAje27S0qTsYeDa7YYokUrIsIiJ5KT5F+wLgFKBH6PBbwI2+799UU1PzSfajy0ljgNuBQSmOPY2NNr+Z1YikLb2AqVhl7PASBB8bpbsETZGX/LAVVtl68/i/lVgRu9exKddgv9NTAl8zEqjNYowiSZQsi4hIXgv0a/4p1q8zaBPW3uj66urqx7IeXG7wsAvQ32JTsIM+By4CbkAj8blqX+AmbF1z2DzgZGBlNgMS6aIfYKPKzeYDy7Hq8GDv12W05CeHAs9kLTqRFJQsi4hIQYhGo33q6upOxQohHZDilOc9z7u2b9++d0Wj0S+yHJ4rA7Hpj8elOHYvcC7wUVYjkq4oxUbioiSPMr8DfA94IcsxiXTFP7GighFsVkRJG+cejn0gJOKMkmURESk4FRUVB0QikQm+75+G9RAOWut53l2+78+urq5+xUV8WTIImEvy+uQmbPrudDSanG++BvwN2CG0fz32gciTWY9IpHN2BpYAPWk/DzkKq5gt4oySZRERKVgXXnjh5iUlJeOBnwB7hA77wFO+79/Yv3//v0Wj0UKq/DwYeBSbwhv0CXAqUKxT0gvBVlg7ryNC+7/A+tPqtZVcV4EVEmzPaOx9TMQZJcsiIlIUKioqRsWraH+X5IJgazzP+2tTU9MfZ8yYke99PbfFEqa9Q/ufA07AevZKfuuBVS4/L7R/A3A88EjWIxLpuBLs/Whf2p6GPQabHSPijJJlEREpKhdeeOF2paWlE3zfP5Pkns0Az/m+f2N9ff1ds2fPXpvt+LppCFY0J9xy6HFs1HF91iOSTJoKXBnatwn4NrY2VCRX7Qssxtbjt+ZbwMPZCUckNSXLIiJSlKLRaKSuru4IrGdtqtHmjcADWCXtfOjb3AObsnhYaP9crADUhqxHJNlwLvB7Eq/p1gEjgFedRCTSMdOxwnWt+Tb2HizijJJlEREpehMnTtymR48e4+KjzeFRWYAlvu//KRKJ3F5VVZWr1aOvwxL/oIeAE7HEP1+cjVXxTuV50lfw5wKsl3Eq84EFafo+2TABuJbE1mBLgeFYezCRXNQb+0BnB1JPxz4euC+rEYmEKFkWEREJCFTSPh3oEzocA57wPO/Wvn373pNDLajOBf4Q2vcoNo2xIfvhdMsyrGJuKkuAfdLwPQ4Cnm3jeBT4VRq+TzZNwtYxB/0Dq5LdlP1wRDrk68ATpM5JTsSqv4s4o2RZREQkhYsuumhgY2PjyZ7nnQ18NcUpn2NTBG9xPE17BPAUidPI38JGFVc7iah72kqWwZ7Xv7r5Pa7B+nG3Jkr+JcsAfwbGh/ZdDlyW/VBEOuxm4AyS85KTgbuzHo1IgJJlERGRdkyZMuVrsVhsHNZ2aYsUp7wG/KWpqem2mTNnfpDF0PoA/yGxLdY6oBzI1x7S7SXL1wA/7cbj9wI+xPpQtyZKfibLZVgl9EMC+xqBkXT/AwaRTNkceAN7bw3mJt8H7nQSkUickmUREZEOikajZevWrTs2EomM831/DMlFwZqwZOWWfv363ZeFadrVWM/SZj5WrOz+DH/fTGovWf4Ma4/V1XXYp9D+BXiU/EyWAbbB1nZvG9j3CjY7otFJRCLtOxW4I7TvGmzt/SDsQ65SoH/82GZYHtOLlveCz7H3wPVAffz2GbAS+Bj4CFgVv5/rBRslRyhZFhER6YKpU6cOamhoGOt53hnYyF3YBuBB4Na1a9fOvf7669O9dng3rDhOWWBfd0ddc0E4Wf4U+ALYPrCvO9Mz5wLHBO6/AOwXOidK/ibLAEdjvZaD13nnA7PdhCPyP72wmTDNtz3jt+2BrUksUpcpTVjS/AGWjL8Rvy2N3+qyEIPkCSXLIiIi3VRRUfElz/PO8DxvvO/7Q1KcstrzvHtisditNTU1C0jPqMZ9WO/kZu8Bw8j/C71wsrwKuAH4eWDfw1jxss7aDvs5NVfe9bHWNdWh86Lkd7IMtg50XOD+amBXVB1bsmcgVkxvePw2DKt8nY2EuDvexz6I/BdWCPBf2PuQFCElyyIiImkyYcKEHgMGDPiG7/uneZ73baw1StjbwB2+799RU1Pzehe/1VewtcrB/8cLpRhOqmR5JDby0/x8m7CL7g87+dgXA78N3H8C61H899B5UfI/WR6MjZIF23BdClzhJhwpArsDRwAHY0nyXuR+YtxRb2OJ87NYQcUX3YYj2aJkWUREJAOmTJnS3/f9433fPw04ktR9RBf7vn97LBa7u5OFwf4KnBS4vwgr6lUI6/BSJcuDsd7HwenuFwHTO/nYr2EX8M3GYQXRCjFZBrgEq4bd7BNgR2xau0h39QEOB8YA38BmLnRXHbY2uRe2RGIBNhtiXfz4Z4HzgktbSoAB8e2BWJLeB1vvvDUwBHsf2Rpb79xdH2BLOuZidSo0Y6NAKVkWERHJsIsvvniL+vr6E+Lrm8tJ/v83Biz0fX9OU1PT3bNmzVrRxsMNAf5LYnGxY4GH0hq0O60lyz8Grg/s72zP5XLswrtZHVYM62gKN1kegE07DyYH44Bb3IQjBWAwNovlOKzqeq9Ofn0M+50MrxV+GyvAtQHYFytSNxGrw5BuPbHnsSvJa6d3xpL1zmgEFmI1Ku4ClqctUnFOybKIiEgWVVRU7Oh53inAD7GLs7D/Jc6RSOSvVVVVH4WOV5I4oroEWwtYCKPK0HqyPABYgY0WNetMz+XrsYS72Z+AM4HjKdxkGex3pTJwfz6JraVE2tMXq49wGjCajieTMez96Vls9stzwOvApg587RVY8pztonQ9sOKJB2LvLwdjyXtnnvMzwO3APbSMhEueUrIsIiLiyJQpUw6KxWKnAicCQ1Oc0uh53pOxWOzuWCz295kzZ67GRlwOCJxTAczIQrjZ0lqyDHAbdsHerKPVv3tj65uDI6yHYhe1hZ4s74ElKM3XfD623vt9ZxFJvjgSm4lwPNCvA+dvwv6mnqQlOV7X5le0rhe2/vnhLn59OvXG3nOHA4dhcfXtwNdtwuK/GRt1jmUoPskgJcsiIiKORaPRyLp168o9zzsZOAGbHhzW0NDQMP/RRx/9+pIlS5r//45hFZ7Do8/5rK1k+Sjg0cCxz7CfVXsjVacDtwbuv4NNwfQp/GQZbIrowYH7ZwPXOYpFclsZ1ou8AhtRbc872N/kY1i7srWZCy1nlGJ/T8di70n7035O9Q72N3cdWt+cV5Qsi4iI5JBA4jwWK+K1dfD4u+++yz333NN8dxEwIsshZlpbyXIEu+jcIXD8JGBOO4/5GDZK1uwXwG/i28WQLP+cxCrY9wBjHcUiuWkIcE78Nridc1/Gphnfi603LnY7AN/GZr0c3M65a4CbsOnl72Y2LEkHJcsiIiI5Kpg4e553su/7Qx555BFefvnl5lN+gyV+haStZBnsOV8auP8QNsLTmqHYRWmwt/KuWNINxZEsH0ji2u73ge0dxSK5ZTD29zQBK3zVmveBO7GlEC9lIa58tRuWNJ+GtdJqTRNwB/b+/W7mwxIREREpYBMmTOix5557vtarVy8fS/h82k4S89UyWp6fD6wMHd8Zm37efLyB1NPWm10WerzHQsePDx33gV926xnknjKsynDwOQ5xGpG41g/7PV9L8u9/8G/rbqw9VKH0S86m4dgocvhvL3jbCMwEtnQUo4iIiEjBWEnihdaObsPJiPaSZYCnQ+dUpjgHbAbdm6FzTw+dUwzJMljP2uBzLLTp+9IxPbBR5BW0nsCtBa4GdnITYsHZCusL/z6t/8zrgKto6RUtIiIiIp3Ql8SLq3paphYXko4ky2eGznm1lcc6NHTeGhJbT0HxJMv3kvgcT3UbjjhwMPAKrSds7wDnA/1dBVjgemItA9t6DZYD33QVoIiIiEi+2pbEi6oP3YaTMR1JlvthLWmC5x2Y4rw/hc65IcU5xZIsX0viczzXbTiSRX2wUctGUidon2Ajn71cBVhkIliBvbdpPWm+GxuRFhEREZEO2J3Ei6m33IaTMR1JlgFuCZ33+9DxviSvxxyV4nGKJVmupmNT16WwHIZVrG5r6u9AZ9EVtzJsSvxHpH59VsePi4iIiEg79iDxQupNt+FkTEeT5SNC531KYjXfcaHjS0ndBaRYkuUqEp/jFLfhSIaVAjNInYTFsMJTKvKWGwYA07GCaqler9tJXj4iIiIiIgFDSbyA+sBtOBnT0WTZI3kaY7B38JOhY5e08jjFkixfQ+Jz/KnbcCSDtsSqvqdKvJYBR7sLTdqwL/AcqV+3F7GWdyIiIiKSQn8SL542UpjtXDqaLIP1Qg6e+2B8/05YH9Pm/U3ADq08RrEky3NIfI7hquBSGPbH+vaGf6ebgOuw9f6Su0qBC4D1JL+Ga4DvugtNREREJLetJvHiaTu34WREZ5Ll1nouR0OP8c82HqNYkuXnaX/9tuS3H2AfooV/n9/DKmFL/tgLeInUH3poCYWIiIhICuEpese4DScjOpMsAzwVOn8KydOzv9/G1xdDslyKFXMq9A9aitmPSZxN0Xx7Cq1Nzle9gJtJPS37KndhiYiIiOSmG0i8YLrMbTgZ0dlkeXzo/DWh+5/TdnGcYkiWv0ri8/vYbTiSZmeTOMOi+XYd0MNhXJIeE4B6kl/fKpdBiYiIiOSas0i8WHrSbTgZ0dlkuS/JPZfDCUNbiiFZriDx+d3nNhxJo0qSf383Aae4DErS7nDsg7/wa301qav8i4iIiBSdXUi8UGoANncaUfp1NlmG1qcq+kB5O19bDMnykyQ+v/PdhiNpMoHk390NwLdcBiUZcyDJdSt84DcugxIRERHJJS+TeKF0tttw0q4ryfLhpE6UW+utHFToyfJQoJHE57ez04gkHcqxEeTg67oetYUqdPth74nB1z2GZhKIiIiIAJbIBS+UnncbTtp1JVn2gLdITnov7sDXFnqyHCXxuS12Go2kww7YuvPg61oHHOYyqC56FSvIl+o2NU3fYxvs/aG173Nqmr5Ptgwj+fVfjyXSIiIiIkVte5JHCvPxIrk1XUmWu6OQk+VewAoSn9s5TiOS7uoN/IvkkcWTXAbVDV/Q+hKK/wIlafgeU9r4Hj7wkzR8j2wbQXKbsPeAwS6DEhEREckFD5J4kfSo23DSSsly+lxA4vNaAwx0GpF017Uk/77+wmlE3dNWsuwDR6Xhe6TqV5zvyTLAGSQ/lwedRiQiIiKSA0aQfJE0xmlE6aNkOT0GAatIfF5XOo1IuutIkltE3Ut+V0NuL1m+tZuPP7ydx8/nZBmsGnb4+YxzGpGIiIhIDnicxAukJUBPpxGlh5Ll9AhfRK9DUzTzWU/gTRJf06VAP5dBpUE4Wd4Qur8eGNCNx7+mncfP92S5FJhP4vP5hMLrkiAiIiLSKfuRvHb5cqcRpYeS5e4bQfLvxs+dRiTdFV532wQc4jSi9Agnyy+RPG36rC4+di+SWy3dSWEly2AtBcO95q92GpGIiIhIDriBxAukBmCU04i6T8ly9wzERhyDz2cZVhhK8tNA4DMSX9M/OI0ofVIly+EPBp7p4mOfEnqcd4BzKbxkGazyf/A51QM7uQxIRERExLWBwHISL5JWkd99dJUsd10EeIDE5xIDRrsMSrrtEhJf09XAlk4jSp9UyfIQ7IO/4P49u/DYc0n+uz6HwkyWy0iepn+N04hEREREcsA3SS76sxjo4zIocWIayYnAtU4jku4qI7mn7qVOI0qvVMkywEOh/b/p5ONuR+JShBg2XblQk2Wwwl7B57UB2MJpRCIiIiI54FckXwDeRX5XyZXOOZ3k34GF2LpNyV8nkPiafg5s5jSi9GotWR4b2t/Znss/D339E/H9hZws98Cmmgef24VOIxIRERHJAR7wV5IvAm/ApuZKYTsRW6MYfO0/xEbXJL+Fp9UXylrlZq0ly2Uktz7rTM/l10Jfe0Z8fyEnywAXkfrnKSIiIlLU+pFcRdYH7sDai0hhOpnk9Z0bgINcBiVp0Y/kVkf7O40o/VpLlgF+HzrW0Z7LI0Nft46WFjjIN3sAAAvmSURBVFuFnixvTfL7wS5OIxIRERHJETsDK0i+GLwTJcyFaDzJLaKagFMdxiTp810SX9u33YaTEW0lyweGjn1Bx6agh7sE3BQ4VujJMsCTJD6/892GIyIiIpI79gTeJ/mC8CEKa61jMfOAqVhiHHyNG7G1y1IYqkl8fWe5DScj2kqWAV4MHW+v53JvkttsBftRF0OyXEHi87vbbTgiIiIiuWVHbBQqfFH4JvBlh3FJ9/UH5pD82ipRLjwLSHyNT3AbTka0lyxPDh1vr+dyuNDdMhILHRZDsnwQic9vudtwRERERHLPDiT33WxezzreXVjSDXsAr5D8mm4CvucwLsmM1SS+zju6DScj2kuWh5BcvK6tnsuPhc69LHS8GJLlMuw9IfgcBziNSERERCQHbYu1DwpfHPpYVd1+rX+p5JjxwFqSX8dP6FyVYMkPA0n+kKsQK9u3lyxDckXw1nouDyW5t/LOoXOKIVmG5A9K93UbjoiIiEhuKgWuInXC/AHwHXehSQfsCDxC6tfvPyQnA1IY9iHxtX7DbTgZ05Fk+cTQOa31XP5F6LzHUpxTLMnyP0l8jt90G46IiIhIbjsdWE/qpOtuYEt3oUkKHjCB1KPJPnAb0MdZdJJpw0l8vf/lNpyM6Uiy3JGeyx7wVuicVGv4iyVZDtc1OMltOPmrEKdziIiISLLbgEOxC8qwsdha2AmoxVQuGAnUAtdhBb2CvsAu+E+Pb0th6h26v8FJFLmhHmt/FzQudP9QYNfA/bXA3zMZVI5bH7rf10kUBUDJsoiISPFYjK1dm4a1HQoagiVnr2LJs4dk297YKP8zwMEpjs8H9gf+mM2gxIn60P0yJ1Hkjr+E7p9AYiu88aHjd1PcHyb1Ct3f6CQKERERkTx1EDblMdUUXx9YBBzuLLrisiNwM4nFiYK3z7FRf32AUTy+QuLvwMtuw8mYjkzDbtZaz+W+JC9XGNXKYxTLNOwHSXyO33YbjoiIiEj+KQN+SfIFa/C2EFvvpunZ6fc14HaSW+MEb/cC27kKUJzZhuQPTApRZ5LlSaFzm3sujw/tX0rrHywVS7Ic/mBhuNtwRERERPLXdtgU7AZaT9o+BKLA5m5CLBgR4DjgUVr/WTeP7H/dTYiSAzySE8ktnEaUGZ1JlgeTuufyvNC+S9p4jGJIlj2SR9qHOI1IREREpADsBfwN60/aWhK3DkusD0HTgjtjF+AyYBltJ8mvoCmTYsLLJI52G05GdCZZBvi/0Pl3kPh+1QTs0MbXF0OyvCeJz+8z9F4tIiIikjYHAw/TdtLsA+8CVwBfchJl7tsCOBdYQPs/yyXYdNJU/WOlON1I4u/IL9yGkxGdTZa/R9t/R/9s5+uLIVkeT+Lze8RpNCIiIiIFai/gWlrvzxy8/Rub/ngAxd1tY3vgx8ADtL0W2ccS6EeAMWjkR5KdSeLvy0K34WREZ5PlMmAlrf9NndbO1xdDsvxXEp9f1Gk0IiIiIgVuIHABsJz2k2YfWIW1bpmAFSoqZKVY5d2rgOdpfwTZx9q43AJ82UG8kj+2I3mK8bZOI0q/zibLAFeT+u9qDdCnna8t9GS5F/ZzCD4/FfcSERERyYIewLewdYIdGW32sXZI/wauAcZho9X5PIo6BFtTfAXwGLaGuyM/hxjwNHZhriJp0lHPkfh7dLHbcNKuK8nyV0n9N3ZdB7620JPl00l8biso7pk+3aY2ECIiItJRDcBD8Vt/4Hhs2uORtL7WtgS7uP0qdqEKNvLxLJYIvIq1enkDqMtU4F1QihXl2hNL8A/A1nLv2MnHWYK1h7oDW+Mt0hm3YS3Gmp0FTMdGmYvVf4BpwKDQ/j84iCXXnB2631wATboonz/ZFRERkdywDTbiPAY4ChjQxcd5H0ual8ZvK7GRkZXY1O6V3Y60RT8s7sHAVtj01h1pSY53wUbSO6sRa/v0D6xI2gvpCFaK1hbY30WvwL7vA3e6CSftvgB6B+6/DOybwe93DjbLJehsOjYqnesOBZ4K7dsH+8BOukgjyyIiItJdK7DKvTdiCWY58A0sef5KJx5naPx2ZCvHG2lJmhuBz7GphnXYqPcGbD1wCS0J+2bY4EB/7LpnC2wqdfACvbtWAHOxBPkxrFWLSDp8is1MODOw7zJgDvY3INIsGrr/CEqURURERHLaQGy0OYpViF5Fx9b45uqtAZs6fgtWwGwftCZQMmtX7Pcu+Hv4M6cRpU9X1ix3R6GuWT6e5Od1iNOIRERERKTTPGya8zigGngQm3IdTgZy4fYB8ATwR+A8bM1yz/T/SETaFe65vJrCqIytZLn7+gHvkPic1Fs5TTQNW0RERLLJB16P34LKsHXCe2Hrhodia4qHYGuKt6Hra6FT2YiNcn9Iy5ro92kpNrYUWJvG7yfSHZcCJ2HLCcCKW10HHOcsIskV04CdAvcbgUluQik8SpZFREQkF9STOokO6kVLQa7guuTmqrjN65LBKm7HSF7PXIetMVYiLPnkI2wpQ01g37FARWifFJeTaOky0Gw2tlRE0kDVsEVEREREcl8EmEfiWtQY1vf7IRcBpcHvSaw6/wHw6wx+v8OwauJBt2M90PPNfsB8oG9g3zKsmvh6JxGJiIiIiIg4shu2Xjm4PvUzYA+XQUnWDQbeI/H3YCNWV0FERERERKQoHYOtSw0mSq9hCZQUvn7AApILlf3IZVAiIiIiIiK5YBLJydLrwHYug5KM6wc8SfJrP9NlUCIiIiIiIrkk3E7Kx9as7uwyKMmYzYCFJL/mj6KizSIiIiIiIv/TC3iY1AnzLg7jkvTbClhM8mv9LyyJFhERERERkYAy4G8kJ1GfAKMdxiXp81XsA5Dwa/wcsLnDuERERERERHJaCXAryclUI9abOeIsMumu07E2UOHX9mmsr7yIiIiIiIi0oQS4heSkysdGnpVY5Zcy4FpSv57/BPq4C01ERERERCS/eMAFQAPJCdY7WMspyX37kXp9sg9cB/RwF5qIiIiIiEj+Ogz4iNTJ1t3Alu5Ckzb0Bq4iuYe2D2wAxjuLTEREREREpEDsCDxP6oR5BTDWXWiSwuHAm6R+vd4C9nUXmoiIiIiISGHpBVSTeqSyuUjUCGfRCcA+wH2kfn184GbUGkpERERERCQj2loD6wOPopHLbBuKrT9u7YOMD4DvOItORERERESkSJQBvwQ2kTo5awRuAvZ0FWCR2B6Yga1BTvU6NAGzgX6uAhQRERERESlG+wAP0/oocxPwEHAUVl1b0mM4cBepK5U332qBclcBioiIiIiICHwdWEjriZsPvAScia19ls4rxQqp1dL2z/lV4LuOYhQREREREZEUjgJepO1k7nPgFuA4LAGUtu2DtYBqrX1X8+2/wAT0MxUREREREclJpcDJwCLaTu6aE7zpwFecRJq7dgeiwFLa/xm+ApyFRuxFRERERETyxgHYKHJba2ubb8uwis5jgQEugnWoFBiFjSA/D8Ro/+c1Hxud11pwERERERGRPLUjNoK8gvaTQB/YiLWgqgC+SuFNLY5g06vPAe4H6ujYz2U1cA2wd/ZDFhERERERkUyJYCOo1wFr6ViC6AP12Ijr1cAZWKKZTzbD1nNHgQeAT+n4c98Y/5ozgD5ZjlvSTNMARERERESkPX2wqs3fB46k82tuPwaWYOt63wBej//7HtayKts8rO/xHlh/6b3i23vH93dGI/AMcAdwD1YUTQqAkmUREREREemMPsDhwBjgG8Cu3XisTdj655VYBemPgVXYFPCV8VsjNvW5ARu53RDfrovH0jN+6wOUYGuoI8BgYCtga2BI/H7z9s50b+T3A2Bu/PYosKYbjyU5SsmyiIiIiIh0x+5Y4nwEMBxLSAvNauBZ4CksQX7RbTiSDUqWRUREREQknbbFKmsfAIwEysmv9buN2HTx+cACYDE2hdx3GZRkn5JlERERERHJpFJsqnbzuuA9A7etHMb1GZYUN6+fbt5+EytSJkVOybKIiIiIiLgyCBhK4rribWhZbzw4ft5mWO7SD+gB9MaKjNUD67Fex83rhj+npXVTcA108/aH8duqjD4zyXv/Dxtn8wlpVQK7AAAAAElFTkSuQmCC" width="65%" style="display: block; margin: auto;" /> ??? If you were interested in linkages with an exposure on the network, you could stop here. --- ## 5. Initial links formed with outcome-side <img src="data:image/png;base64,iVBORw0KGgoAAAANSUhEUgAAA+oAAAJSCAYAAABUcD9+AAAABHNCSVQICAgIfAhkiAAAAAlwSFlzAAASdAAAEnQB3mYfeAAAABl0RVh0U29mdHdhcmUAd3d3Lmlua3NjYXBlLm9yZ5vuPBoAACAASURBVHic7N132Fxlmfjx75seSEIJTZqEEqSDqKAgokhTURERFURQjLoqWFaz6q6LumrUtWDdoKsIFsQCAhZEBFkElN5774FAQoD05PfH/c5vzjnT3jIzz5Tv57rOlcy8M2fuM+fMzLnP8zz3M4AkSUM3BtgU2ARYH9gI2BDYYPDfjYA1gEnAZGAsMG3wuWsNPv85YCmwDHgWWA0sAFYB84DHgUeBxwr/vw9Y1OLtkyRJSm4gdQCSpI60DrAt8AJgZmGZmDCuh4HbgNsHl9sGl7uJRF+SJKnrmahLksYRSfnuwF7A3sB2dNdvxDPAdcBVg8vfiBZ4SZKkrtNNJ2GSpOYYTyTjBwIvB15IdFXvNfcBlwF/Af4EPJQ2HEmSpKExUZek/vB84KDBZT9g6ijW9QSRBD9KjCF/hPJ48oeJceRLgMXASuDpwectJLqnr0F0n58ArEn8Fq1NXEBYnxjr/jzyY+A3AmYMPmakricS9vOAS4gx8pIkSR3HRF2SeteWwJHAEcAOw3zuCuDWweUOymPBbweebGKMwzGeSNa3JT9mfidg+jDX9Qzwe+BnRPK+vHlhSpIkSZJUtg5wNHA+0Xq9eojLw8A5wInAq4lW726yMXAIMIdoLV/M0Lf9SeBUYru9gC1JkiRJGrUBYH/gd0R37qEm5j8iWts3bn/ILTcR2BP4dyJxX8HQ3pc7gU8y/BZ6SZIkSZKYSLSeX0/jBHQFcCXRYr47/ddyPIVocZ8L3E/j92sJ0cq+Y4pgJUmSJEndZUPgc8A86iebq4CLgfcC6yaJtDMNEFXvvw/Mp/F7+EfggCSRSpIkSZI62hRgNlFFvV5yeTPRcr5Vkii7y1hibPqpRIG5eu/r5cAr0oQpSZIkSeokE4GPENOf1UoilwO/BF6aKMZesDbwMeBe6ifsZ2OXeEmSJEnqW0dSP3F8CvgKsHmi+HrROOBw4O/UH/P/Y3qzEJ8kSZIkqYotgPOonSg+ChxPdIdX67yEaEGvd6HkOPqvOJ8kSZIk9Y0BYBa1x6EvIuYKn5YqwD61J3ARtRP2i4FtUwUnSZIkSWqN7YBLqT1d2DeA9ZJFJ4gp3m6g+j56Fvgotq5LkiRJUk84nGgtr5YAXoAV3DvJGOBD1N5f5xCF6SRJkiRJXWgs0ZV9FZUJ3wKiG7wttJ1pC+BPVE/W7wB2ShaZJEmSJGlE1gP+TO1W2U3ThaZhOJzqU+ctBo5JF5YkSZIkaThmAHdTfSz6exLGpZHZFLic6hddPp0wLkmSJEnSEMwEHqAyoXuQqC6u7jQROInqyfqchHFJkiRJkup4AfAQlYnc34CNEsal5jmKqABf3MdfTRmUJEmSJKnSzsA8KhO4nwDjEsal5nsZUQywuK9PShmUJEmSJKlsQ+B+KhO3k4npvtR7Xgg8QeU+/1jKoCRJUudzyh9Jar3xwPnAKwr3/w/wAWJqtm6wFrB1nb9fDyxvw+tcB6xowuu0w/bABeSHNawCXg/8PklEkiRJkiR+QGWr6reSRjQyB1O9UFppObRJr/OlBq+zXpNep112AJ4kvw1PEUUFJUmSJElt9gEqE80/AWNTBjVCjRL1s5rwGmOoPkSgmxN1gP2J3gbZ7bgVmJoyKEmSJEnqNy8AniOfnN0GrJ0yqFFolKgvZ/SV6w9q8BrdmqgDfJjKbZmbNCJJkiRJ6iNjgMvJJ2ULiOS9WzVK1FcDHxnla5w+hNfo1kQd4Efkt2UV8OqkEUmSJElSnziOygTz7UkjGr1qiXpxTvjrRrH+dYDFDdbf7Yn6ROBG8ttzC1FwUJIkSZLUIlOBR8gnY2cmjag5qiXq36Sye/8LR7j+fyms5yrgz1Ves5sTdYCXAivJb9PxSSOSJEkdxbl7Jan5PkB+rPZiYnxyL1oA/K5w3ztHuK5jCrdPGeF6Ot1lRBf4rE8BkxPEIkmSJEk9bw3gMfKtpZ9LGlHzVGtRP5HKAnBPEF28h2OHwjqWEi3nvdiiDrAhsIj8dn0waUSSJKlj2KIuSc31VmCDzO2ngW8kiqVd/gw8kLk9HXjtMNdxTOH2OUTC36seA75fuO9DwECCWCRJUocxUZek5np34fZc4KkUgbTRKuCnhfuG0/19HHBk4b5TRhNQl/g6sCxzeyawT6JYJEmSJKknzaBy6q2tk0bUXLW6vkMkmasy9w9nTvVDCut8lHIV9F7t+l7yS/Lb9oO04UiSpE5gi7okNc/rC7cvAe5MEUgCtxPzxpeMY+jT0R1TuH0akej3gx8Xbr8Wu79LktT3TNQlqXkOLtw+K0kU6ZxSuP2uITyn2nj2U5sSTXf4K1HHoOR5wK6JYpEkSR3CRF2SmmMAeEnhvj+mCCSh04k51Ut2AHZv8JyjyFeIvwK4oclxdbJlRLKe9dIUgUiSpM5hoi5JzbENsE7m9gLg1kSxpPI0lb0IGhWVO6Zw+5RmBdNFLivcLl7wkSRJfcZEXZKaY5vC7WuJ4mD95pTC7SOpPaf6TuS7eS8jiqv1m6sLt2cmiUKSJHUME3VJao4Zhdt3J4kivQuA+zO31wVeV+OxxanszgLmtyKoDlc8VrZIEYQkSeocJuqS1BzFKcMeTBJFequIqu1Z1bq/j6eyKvwprQioCzxUuL1+kigkSVLHMFGXpOZYs3D72SRRdIafkO/2fzBRzTzrdeQT0oeJOdP70VLy09GNo/ZwAUmS1AdM1CWpOcYXbvfLPODV3AH8PXN7HDFWPevYwu3TgJWtDKrDLS3cNlGXJKmPmahLUnMsLtyenCSKzvGTwu1jMv/fADioweP7yQCwRuG+fu6RIUlS3zNRl6TmWFS4vXaSKDrH6cAzmds7AC8a/P87yPdAuBy4pU1xdaJp5H+Pn6O/exdIktT3TNQlqTmKBcG2SBFEB3kGOLNwX6mo3NGF+09peTSdrThjQL8WIpQkSYNM1CWpOe4p3N42SRSdpdid/e3AXsDOmfuW0J9zp2cV500vHkuSJKnPmKhLUnPcSL7S+Q7AlESxdIoLgXszt9cFflx4zJnAgnYF1KH2LNy+PkkUkiSpY5ioS1JzPElUOy8ZC7w0USydYhVwauG+bQq3T2lPKB3t5YXb/0gShSRJ6hgm6pLUPP9XuH1Ikig6yynkexpkPQRc0L5QOtLGwO6Z26uASxLFIkmSOoSJuiQ1z9mF228iWtb72T1UXsAo+QlWN38zMT1byRXAY4likSRJHcJEXZKa5y/kpyTbBDg4USyd5JQa95/WziA61HGF22cliUKSJHWUcakDkKQe8hxRwfzdmfuOB85NE07HOB14oHDfMuDWBLF0krcAO2Vur6ByTL8kSZIkaZT2IMZkZ5eXJI2oeQ6mcttObPFr/rnKa67X4tdsh0nAQvLbVZx3XpIk9Sm7vktSc/0DuLhw35wUgaijnQFMK9z33ykCkSRJncdEXZKa7/OF268EDk0RiDrSOOAFVe4/Ghjf5lgkSVIHMlGXpOb7C3Be4b7vAOskiEWd55NUzicPMIuYrm6j9oYjSZIkSf1hB2A5+THIP00a0eg5Rn30dgGWkt+eK4lCcqXbDwJ7pgpQkiRJknrZF6hMMk9IGtHomKiPznTgTvLb8jiwAbAvMX966f4lwHuSRClJkiRJPWwicD35xGw58OqUQY3CdCL27LJli1/zhVVesxvHcY8HLqLyosNhmcdsBlxR+PtcYEI7A5UkSZKkXrct8BT55OtJYOuUQantvk9lkn5SlcdNAn5ceNyVwObtCVOSJEmS+sPB5McgrwZuJ1pQ1fv+k8ok/S9E9fdaZgHLMo+fB7yqtWFKkiRJUn/5OJXJ2n3AVimDUst9lsr9fjdDG2f/cuCRzPOWA7NbE6YkSZIk9adTqUzaHgBmpgxKLTEAfJ3K/b2QmBFgqDYBLius42fAGs0MVpIkSZL61XjgdCqTt4eBFyWMS801ATiZyv38FLDHCNY3scr6rgFmNCNYSZIkSep3Y6ksFuZ0XL1jY+DvVO7fJ4GXjHLdRwPPZdb5BHDAKNcpSZIkSSK6RX+HymRuNdE9fnK60DQKexO9I4r79DFg5ya9xu7AvZl1ryDGrQ80af2SJEmS1LcGgK9QPVm/jNbPT67mGQN8lHyV9tJyF7Bdk19vfeCCwuv8Dlirya8jSZIkSX3pCOAZKhO854iW0rHpQtMQbA1cSPULLn8A1mnR644D5hRe71aaf1FAkiRJkvrSC4CbqZ7sXQ3sli401TCOuJCymMp9topIose0IY63A89mXvtp4NA2vK4kSZIk9by1gauonqwvBT4PTE0WnbL2IaquV9tXTwGHtDmeXYm52VNcKJAkSZKknjQB+C7VE7/s8gTRijspTZh9b3vgDGrvn3OBTRPFNh04rxDP72ld13tJkiRJ6lmbAJeST7D+CPyC2gnhfcAsHL/eLpsDc4kK69X2x2PE1GmpjQVOJFrUS7HdAeyYMCZJkiRJ6ir7AI9QTqqWk59q601Un+6rtNwIHIct7K2yCzHn/VJq74NTidbsTvJ6YAHlGBcBhyeNSJIkSZK6wCzy03nNA/ar8rg1ieQ9m3gVl3nEmORU3a57yRjg1cA55Fumi8ulxIWWTrUt+QKFq4CTiCJ4kiRJkqSMKcAvySd9VxDdq+vZgEi06rXuLgV+QiSQFhIbng2BE4DbqF8n4DrgNYliHK6pwG/Jx38RcSxJkiRJkoCZRHf1bOI0lygmN1QzgNOIbvL1Esr7gC/h+OR61gSOIuY7b/R+3jH42G67ADJA9MhYSXlb7gdenDIoSZIkSeoExXHDi4F3jWJ9mwNfIaYDa1Qt/lrgE0TV8n43FXgj8FPgGRq/dxcCb6D7EvSi1wBP0rzjT5IkSZK61lhi/HixEvfOTVr/FGK8e3Y8cr3lEaIA2uHAWk2KodNtSXRrPx9YQuP3aCkxDdseKYJtoa2B6xldjw5JkiRJ6mrrEclhNjE6l9bMbT0GOBj4GfAsQ0valwIXAJ8FXjsYb7cbB+wKvJeo2H4/Q3svSuPPPw5s1Pao26dajYRLgOelDEqSJEmS2mF34F7yVbfn0J4u1FOAdwB/ovbc37WWO4lk/3jg5cD6bYh3pCYA2xG9A/4buJihX6TIjteeQ/+N5S/OOvAQ8NKkEUmSJElSC80iX519PnBQolg2Aj5IFEx7juElsaXlSeBy4BTgk8BhRDGyTYGJLY5/OjG2/lVEK/nXgN8TwwcaFYCrtdwKfBN4Bd0/9nw0XgE8Sr6HxQlJI5IkSUBUg5UkNcck4DvAuzP3XUsktncniShvMjF128GDy8wmrfcp4DFiPvdHiaJ5y4libauAhYOPWzh4ew0iwZ84+P9xRJG3CcTUYRsSFxjWpznjp58lisL9kehl0An7olNsCvwGeEnmvtOIiyKLk0QkSZIkSU2yOfBP8i23pxGJaKfaEjga+C5wJfmu0N28PAycRbT+70vrW/273STgf8m/h1cBWySMSZIkSZJG5WCie3spyVlCd3YhngzsBXwE+AVwDcMf893OZRUxX/z5xBj1w4kLJhqZ4pCNx4H9kkYkSZIkScM0AMwGVlJObh4A9kwZVJMNEMnvq4F/Ab5FjBG/jmi5Huk48aEui4hu9aXb5wFHEJXdO7m3Qrfai9ivpfd7OXGMS5KkNnKMuiSNzDTgJ8AbM/f9jUgiH0sSURoDxFjy9SmPLZ8CjB/8dwzl+drXGrz9HNFyu3Tw/yuIhHwlMc79cWK+93nEOOl9iTHmEFOuvau1m9T3NgZ+Tb4K/C+A9xA9LCRJkiSp4+xCTGOW7YJ9ElEUTc23IeX3+rLEsfSLccS0ddneDdcSdQ0kSZIkqaMcSX7c9tPAm5NG1B+eIN7vhY0eqKY6ivy0fvOBA5NGJEmSJEmDqrUw3krM8a3Wu4Ty+/68xLH0m92Ae8j3IJlDf89BL0mSJCmxTYBLySfpZ1Eee63W+wHl995K5O23HlFdP/sZOBs/A5IkSZIS2IcobFasgm0xzvb6KOV98MHEsfSrsVT2KrkNe5VIkiRJaqNZwDLKSck8bM1N5WDK++G7iWPpd28FniFfp+GwpBFJkiRJ6nlTgDPItxxeQcwnrjS2oLwv/po2FAE7UznzgePWJUmSJLXETOBG8kn6XGBCyqDEADHX+mpiKILSm0bUash+Vv4ArJMyKEmSJEm95fXAAspJx2Lg2KQRKesqyvtm3cSxKAwQNRtWUt43dwI7pQxKkiRJUvcrFclaRTnZuIPo3qvO8VPK++dliWNR3uuAp8hf5Hpn0ogkSZIkda1q006di913O9GnKe+jdyeORZVqDRsZnzIoSZIkSd1ld+BeLIjVLd5EeV/9d+JYVN1U4Nfkk/W/ARumDEqSJElSd5gFLKWcTMwHDkoakRrZjvL++n3iWFRbadz6Csr76wFgj5RBSZIkSepck4Afkm/xuxrYMmVQGpLxlC+u3J04FjV2EHEBrPQ5WwIclzQiSZIkSR1nc+Cf5JP0U4E1UgalYbmZ2G8rgTUTx6LGNgeuwOkOJUmSJFVxMJWteyckjUgj8RvK+3C3xLFoaCYBPyafrP8d2DhlUJIkSZLSqTbP8wPAnimD0oj9F+X9+PbEsWh4ZgHLKO+/ecArk0YkSZIkqe2mAWeSb8m7CCtQd7MjKe/LzyeORcP3cuARyvtwOXEhTZIkSVIf2BW4i/zUaycB41IGpVHbjfI+/XXiWDQymwCXk7+A9lOsFSFJkiT1tCOBZyknAU8Db04akZplDcrDGG5KHItGbiLwAypnX5iRMihJkiRJzTcOmEP+5P9WYPuUQanp7iH27TJiyjZ1r1mUp9xbDTwB7J80IkmSJElNswlwKfkk/UxinLp6yx8o7+MXJI5Fo7c7cB/lfbqCGLc+kDIoSZIkSaOzD9ULVHmi35u+RnlfH5o4FjXH+sBfyV9oOx1YM2VQkiRJkoZvgJgLvTjl034pg1LLHUd5f38qcSxqnmpDV27BXhOSJElS15gCnEH+pP4SYOOUQaktXkZ5n5+WOBY1X7EY5ELgjUkjkiRJktTQTOBG8kn6XGBCyqDUNmtT3u9XJo5FrbErcDf56RXnAGNSBiVJkiSputcDCyifwC8Gjk0akVJ4lNj/z2Ly1qumA38mf0HuXOJCjSRJkqQOMJZoUVtF+aT9DmDnlEEpmQspHwfPTxyLWqfa5/52YMeUQUmSJEmC9YDzqWxZWydlUErqu5SPhYMSx6LWez0xVr20zxcBb04akSRJktTHdgfuxbGqyvsg5WPiI4ljUXtsC9xM5XfB2JRBSZIkSf1mFrCU8on5E8CBSSNSp9iP8nFxcuJY1D7TgN+S711zIbBByqAkSZKkfjAJ+CH5k/GrgRkpg1JH2Zj8tHzqHwPAbGAl5WPgfuDFKYOSJEmSetnmwD/JJ+mnAmukDEod6Sni+JifOhAl8RrKx4AzQEiSJEkt8hrgScon3kuAE5JGpE52OeVjxa7P/Wlr4AbyF/bmAuNTBiVJkiT1gmpdWR8A9kwZlDrejygfL69IHIvSmQKcQT5Z/z9go5RBSZIkSd1sGnAm+ZPsi4ANE8ak7vBxysfM+xLHorQGiN43yykfEw/ixT5JkiRp2HYF7iI/3dJJwLiUQalrvI7ysXNS4ljUGV4BPEZ++MyspBFJkiRJXeRI4FnKJ9RPA29OGpG6zVaUj5/zE8eizrEZ1QtSTk4ZlCRJktTJxgFzyJ9E3wpsnzIodaUxwHOUuzlLJZOA/yX/PXMV8PyUQUmSJEmdaBPgUvInz2cS49SlkbiW8rG0VuJY1HlmAUspHyOPA69KGpEkSZLUQfYBHqF8wrycqPQ+kDIodb1fUD6mXpI4FnWmvYGHqfzukSRJkvpWqRrzMsonyvOA/VIGpZ7xGcrH1TFpQ1EH25jK3jw/B9ZIGZQkSZKUQrX5jS8hTpqlZjic8rH15cSxqLNNJGYHyH4fXQtsmTIoSZIkqZ1mAjeSPymeC0xIGZR6zo6Uj6+zE8ei7vAOykUIVwPzgQOSRiRJkiS1weuBBZRPhBcDxyaNSL1qAjHmeDVwR+JY1D1eCNxL+TtqBXAiMZOAJEnJWcRJUjONBb4AfILy98udwGHA9amCUs+7jejBsRLYlKjyPZWYChBgbeJ4HBj8P4N/mzr4/8XAkirrfW5wXUsGH7MMeJZI6hY1eyPUdusBp5Ovl3EO0eK+MElEkiQNMlFXL5oCbACsSbS2TRj8/xjy0zetJlp9IU66VxAn5M8QU/g816Z4e8V6RAXuV2fu+z1x0vtUkojUTSYRx9A6RDJdXBrdn8IC4vuitCwgvksWEt2p5wNPZP5fuj0PWJUgXlUaB/wX+SrwtwFvAm5OEpEkSZioq/usB2w7uMwANgI2BNYHnkck6JOb9FrPAo8CjxGJ+yOD/78TuH1wWVDz2f3lRcCvgecP3l4NfAX4FCYk/W4c8bnclPi8lv7dLHP7ecD0VAEmsIJI1h+psjxAdMm+j0j+1R5vA35IuQr8ImIWgd+mCkiS1N9M1NWp1gH2AHajnJjPBNZNGVQV84BbiaT9NuAK4Cr66wR7FvBtykXi5gNHAucli0jtNImomr01sNXgsjmRfG9CXEhr1bjf54iLZaWLAQAPAbcM/r/UVR3K3ddLz1taZ73TiGEcUO6ZM4m4CDie6LWzxuC/U2ldi/584H4iab8PuJsYh38nkcwvb9Hr9qtdiMS8VAXeC46SpGRM1NUJxhGJ+O7AXsDewAvo3qI+K4mk/arB5RLgGrrrRG8CUWzp8jqPmQR8F3hX5r5riPHo97QuNCUwjUjAs8l46f+bMvrfknlE75VHicR7ATFcIvtvtaWUbL8Y+Ofg/88AjhhlPCMxlXLivi7RQ6C0lG6vR1y4KPUwmDKK11tBJOt3Di63ERcobiR6/mhk1iXmVz8wc98fiYuPDuGRJLWNibpS2RI4aHB5FdFq1SzPEV3VnyZa1EqtaqvIFwjKFpYqFZ6aRCQlGxBz7jbLk8BfgD8NLo80cd2t8HUi3j/U+PvmRFf3F2fuOw14H47t72abATsAOw3+O5NIyNcf4foWAQ8SCfiDxHH/EPDw4PLQ4N/qtW4PxRTi8z4A3ADsPMr1tcsalHsdbEz0QtiCGEKy+eC/641gvfOBm4gx1jcSCfy1xPeQGhsgCmJ+kfIF4zuJces3pApKktRfTNTVLhOBVwIHE8n5zBGuZzHl8eG3Eyf+DxMtcqUxn83qdr42MYZ2/cF/NyJOqmcSPQC2ZmRzgq8GrqOctF9CtMJ3ioOJInA7Eyf5Ra8BfkoMT4BIsmYDJ7UlOjXDdMrJeOnfHRl+F+4lRHfsO4G7Mv/eR4y1frb2U5vufuJCw1IicV/RxtdupTUoJ+9bAdtklucTXfGH6j4iYb9mcLmWeN9U3SHEBchSEdJngHcTvTYkSWopE3W10gDRjf0o4HDKid1QrCBahC4npvUqJeYPEIluJxhLnEDPJLrq70KMq9+W4X22HiamCPopcfKc0kZEcj6d6FmQnYKqWivTg8S+rddFXulMIC647Ew5Kd+RaLkdqoXkk/C7MrcfonM+j+cBBwz+fyb9Maf6eCJZ3wbYbnDZEdie+PwOxXxiiM4/gX8M/juv6ZF2r5nAmcR7WnIy8EGsESBJkrrM9sRc2vcSJ/FDWR4mivh8AtiH5naFb7e1ifGNnwHOJaZjGur7cBPwScrV09tpDHA++WnrSqYRJ6vZWC8iuuyqM4wHdgWOA/4HuJJoXR7qsfckcDHwfeD9xOdwJN2uU/km5W15feJYOsHmxPfQvwI/Ji4CLmNox8LdxFSLHwZeRnOHAXWjqcBvyL9Hf8PvP0mS1AXGEN0ES4leo2U50eV7NlFErpd7d4whtnE28f4M5WR51eBjD6F9780nMq9/beb+XYkW1GxsJxFj+pXGWKKF/GhiX1xCdDMfymdvKXFB6AzgROIY25Lu/wy+l/I2zm7w2H41jvxxcz5Du5C4nLjwcxLD7x3VKwaI42ol5fflAeAlKYOSJEmqZRrwEaIFptHJ3jyite4NjK7acbebDryVGPv4NI3ft+uIyuqtbNV6EXEBYRUx7ODMwfuPIp8APg28uYVxqLoZxL74JpGUP8PQkvL7iZ4q/w4cStRV6NbZFBrZh/J2n5I2lK4yhugFdSzRE+NaIjGvd1ytILrLf5MosNZp02a20sFE75PSe7GEGLcuSZLUETYgKoM3SjSfJaa6eS22wFYzmZhK6mwat7Q/BvwH0Q2zmaYQ441XDL7OSuBbROtZ9vVvIT9OU60xjrhwcgLR6l0aB95oeZg4jj5DfN76rVvuepTfi38kjqXbrUlc+PhX4kLPY9Q/9lYSLe5fJuoErNH+kNtqa6J2SvY9mMvIiotKkiQ1xVSiu+wi6nfb/gvRvbLZSWUvWw/4F6KYU6OE/UM076TwNGKfZV/jnsLtX9DddQM62VQiufks8bmp99nK9k75A/A5Yjz2Jm2PujM9Trw/C+n+rvyd5gXAe4Cf0LgH1RLgr8Cnia7hvdiLYzLxXmS3+xKGV6hRkiRp1MYDs4jpz+qdnJ1KVJbW6OxOvJf1uqDeR+yTsaN4nXfWWf/qwdd3vG9zbQa8Dfg2UeCr1JOh1rIUuBT4b+Aw0hQa7BYXU37fNk0cS6/blDiOTyZfw6LWhaXTBh8/PUWwLTSLfG+oh4gCfJIkSS13MPVPxB4humSvnyrAHvZ8IkFbQO33/xpGVtBoG2JoQrY4UvHCy8+IIl0HEd3ee71LaytsRCQoPyCGGDRqLZ8PnENU/3850XKnoZlL+X3cP3Es/WYG0eJ+OpGY1zq+VxAXnv6dGOLRCz0f9gEexQuckiSpTaYTrbr1WkmOx6l72qE05KBWTYAVwNcYeiI9Abia+q25K6jsEr+a6F58BfBRYNJoN6wHTScKbH2HqK7eKDG/k+hCO4uoyN0LiUsqH6b8vh6fOJZ+NgDsQoxx/wv1pwl8kCgyeiDdTwGLqAAAIABJREFUPcZ7U6I2QnbbTsMLbZIkqckOp3YBoWeAOcBayaLrX9OJ934x1ffN3QytJfFrNZ7faFlEJKAObyibCryOeE+voXYPhdWDf7sK+AbRjX2jBPH2sgMpv9ffTxyLyqYQs338DzFkp9bnYyHRIv9WuvP3ZSLwQ/LbdDWwRcKYJElSj1gHOIvqJ1HLiXG1GySLTiUziG7p1Vq8VxEnxLVauw+s8bxaiWXpZHMW/T2tXskE4JXAF4HLqF9HYBVwI1FF/1D6axqrFDaj/N5flDYU1bED0dp+EbV79SwFziO606+XJMqRm0W+F8HjwKuTRiRJkrraztQeQ3s98OJ0oamGfYDbqb7PriYS+qwNiCEL9RL1UnK+lJgezBNM2JI4+T6D+vUCVhP1HE4dfLwFzdprgPLwkMcSx6KhWZfowXUq9Yf2XEJMWdgt0w7uRUyZmN2G2Ti0RZIkDdPbiC7txROkZURX624eO9jrJhP7qFrL1BOUu8IPAOdXeUz2RHI1cAdxQtnPrb9TgEOIbv53UD8xvw84hZiOcLMEsSrvCsr7pttaYvvdZKKL/I8oT7VXrWfXn4B30/nfUesDF5KP36kuJUnSkIwFvkn1E6LLiLlz1R32BG6legL+ceArVf5WallfRrn1vB9bfErFrz4BXED94lcLgd8S1fC3ThGs6soWwNw7cSwaubHAq4haA7WqyC8Ffge8hc4t2jaOuJCajfs6YKuUQUmSpM42lqhKW+0EaC62onejqcCvqN+lvdhN+1/pz5bHtYEjgB+T76Ja7X27EvgCMdRgXIpgNWSfpLzv3pM4FjXHWGA/ovZGraR9IdES/ypgTJow6zqKmA4zG+8bkkYkSZI60gTgN1Se7CwmuhSqew0QXddrFWlaBfyB/mw93xb4GPBX6heBe4y4iHUkFk/sNm+kvB+/njgWNd9Y4rvrZOBJqn9+HyB6EHXa7BS7AfeQ/y6eQ2deWJAkSQlMJhK1amNtX5gwLjXXAcB8KvfzT+mfE8OxRPfnOdSf03wF0Wo+Z/Dx/fL+9KKZlPfrHxPHotaaSNSSOJV8a3V2uYm4cNkpvYamU1kr5Byih48kSepj44kpb4onM7djIaxe9ElgCZX7+wcpg2qx9YnCbo0qtD9CzHl8KDFkQL1hHOVj/r7Esah91gaOI6Z8qzbMZzHwc6I1PvWFuLHERcHs7Bu3E9PWSZKkPvUdKk9gbgE2ThmUWmZdoiDgg1Tu948kjKvZdiYuSlxK9ZP0UjfTK4HPAi+i/7r995MbKe/zKYljUfttTnwf3Eb174J7gROBLZJEV3YE+dlWFgFvThqRJKljeKLaX44hCmdl3Uy0MDzS9mhG7o1EhfNq5gNfbdLrHAXsWONv9xHViLvFDKKaeXZe9ZXA64lhEN1mDDHe8xDgrcTY82oWA38HziVqMjzYluiU2q8oJzwvAq5KGIvS2h2YRUxBWuw5s4qoV3EycCYxBKbdXjD42qUZVlYT4+s/TXxHS5KkHrc3lVNO3U3njNsbju9RuzvzapozpdwkahcqWk0kf91mJvAU+e14EtgmZVDDMIm4qHQS9au0P0qMWT0cW1P71WcpHw9HJY5FnWEy8Z1wPvku56XlYaI7+uYJYptGJOvZeP5I588TL0mSRml9InnJngQsovMq4g5Vo0T9C014jbc1eI1uTNQBDqSyGvyNRBLcidYhEq1fk+8iWuzS/g/gU8Rc6NJbae73gXrLTCIpf4zK75PlxPfN/rS312Fpto7s0J37iB4hkiSpRxXn1V4FvCVpRKPTKFF/kCjWMxrVCu71QqIO8FEqt+fLSSPK2xT4ANHytYzq7/8y4M/A+4FN0oSpDrYL5WPltzUe49AvTSDGiV9I9Vb224ETiBbvdnkt+Z5Pi4lha5Ikqce8icqTj88ljWj0GiXqq4mW45HalNpzkPdCog4xR3h2e1aQdmq+bYniT1dQ/YS51AvkDODtOJWR6ptE+TN8a5W/jyO+G6WS7YBvUn3I09PAt4mW+HbYhnJBxNIyl5i1RZIk9YAJwB3kf+yvovt/7Ksl6pcWbv98FOv/FJVJea8l6tOIbpXFbWpnK+NOROXlG6h8f0vLY0Shp9fSud3z1ZnuotyVeWLhb3OAD7Y9InWDyUQL9j+oPszmj8DBtH6KtylU9oa7GNioxa8rSZLa4F+p7C7cC/O0VkvU31u4vZiRt7reUljXrCqv1+2JOsBrqNyuVrcy7g58kdrTJq0G7iQq9+9N+vmO1fleRPXk5VzKx9T2mftfTXnGA6mePYCfUVmIdTXxHfYhWluwsjRuPdvD60Fqz3oiSZK6wCQqK2N/I2lEzVMtUd8duK5w33tHsO69C+tYRBTj68VEHfLJzGrgWprbqj5AnFR+lZhloFZyfj3wn3RvgUOlM41Img4u3P8VysfXYYP3rU+5gJiFBzVUzyNmEigWZV1NdJX/4uBjWmVf8oXvlgDvaeHrSZKkFvog+ZOJhfTOVC+1EvWPFe67dATr/mFhHT8gukL2aqK+Pfkqw6upTHiGawxxwWMO0TpeKzm/iej6vt0oX0/6ItEt+RuUu7kfS/lY+w/iolH2wpR1DjRcE4gp3qoNh1pKTAu5Y4teezOihkf2NecOxiRJkrpIsXX5S2nDaapaifoGVFYJH86c6pOBBYXn701vJ+oQUxFlt+2sEaxjLNHq8z2qtzqtJhKpy4ghGTNGG7SUsSHl7snXE5/7PSkfez8Hjs/cfiZNmOohewG/ofJC5yrg98ArW/Cak4AfF17vStLM/S5JkkZgD/I/5Mtpbbe8dquVqAOcXbj/i8NY7zsKz72daIXr9UQ9m9AM53gZA+wDfAd4hMr3aDVxEvs3IknatNmBSxnZ74XniPHDpdu3EhfxSjMK3JQoRvWerYlj7zkqv/+uJFrgm11rYxb5i9LzgFc1+TUkSVILfJXRt5B2snqJenE6uuHMqf7XwnM/PXh/ryfqEGPTs9v3/hqPK3VrPwl4iMr3ZTVR+OgSYv7hjVsatVQ2g3LRrVJCvjhzu/S3FcA5iWJU71qPKPxW7XvxLuL7sJkzV7yc/AXS5YOvL0mSOlixovZb0obTdPUS9QlE60L2b0OZU/355LswrqTcnbAfEvV/I799fyj8fQdizPmDVL4XpferlJxv2J6QpQo/J18hu9aFpO+mClA9byJwNHAzlcfefcR35BpNeq1NiOFE2df4WRPXL0mSmmhT8j/aS4mqyL2kXqIO0dqb/dsvhrDO/yw857zM3/ohUd+e/PY9C+xMFHurVRAum5w7t686wS6UW9PrLbY8qtXGAIdQfT72ecR36zpNeJ2JwMmF9V+DdUAkSeo4h5H/wb4kbTgt0ShR363wt8XUPyEaoDIZfXvm7/2QqEPtruzF1sgLgX/BlnN1pj/SuFX9rcmiUz86CLiYyuPwKeDzNGdGlqPJj5N/AjigCeuVJElN8nnyJwK9Mnd6VqNEHSrHXL+vzvr2LTx2Afmug/2SqJ9F45Zzx5yr072CxhecXposOvWzvYj6CMVeH4uIoUWjTdh3B+7NrHcF0XtkYJTrlSRJTfAz8icAx6UNpyWGkqh/uPD3y+qs75TCY/+n8Pd+SdS/QH4b7ycqZ5ucq9tcRv1WdY9ppbQbcAaVU7stZPQt7OsDFxTW+ztgrVGsU5IkNcEl5H+g90sbTksMJVGfTnle5dKyXZV1TSFaM+q1tvVLon4c+W08LW040oi9kdpJ+jKaP12WNBLbAadSeVFptC3s4wafn13nrVT/DZQkSW1yA/kf553ThtMSQ0nUAc4sPOZLVR7zrsJjbqOym2C/JOrF5OZ3acORRmwAuIXqrer3pgtLqmp7ouhptRb2zwFrj3C9bycKg5bW9zRw6GiDlSRJI1MsirZ12nBaYqiJ+hsKj3mIyjnViwV+/q3KevolUX81+W28IG040qgcQ+XndhVRDFHqRLVa2J8kqsSPZAaXXYG7yX8G5mCvEkmS2q6YqG+TNpyWGGqiPg54pPC4gzJ/n0G+qM9KYLMq6+mXRP0A8tt4ftpwpFEZDzxA5Wf8xymDkoZge6on7I8TxeEmDXN904kpR7Pr+j3NmR5OktQiXlHtPc8Ubq+ZJIrOsILKOdSPyfz/XeS7uZ9HnNj3q+Kx8mySKKTmWA58jfxnfAxRJFHqZDcT063tQnkIF8B6RGv4rcCxVPYQq2U+8Brgs5l1vQb4J7Bjc0KWJEmNXET+qvmBSaNpjaG2qEOchGQfV5pTfQxwX+FvR9RYR7+0qL+P/Dba8qhutwYx3WL2uH5X0oik4duJqBJf/B26lUjoh9Po8nryn4lFwOHNDFaS1By2qPeeewu3t0wRRAe5Ebg6c3sS8FaiGv7mmfsXAme3Ma5ONKNw+94UQUhN9BxwUuG+obSojyUu6K0DbEF8j+40+O+WwAaDf5vcrEClOm4A3gLsS/4i8bbAT4ArGPoML2cDexDFFiFmPvkl8TkZ14RYJUlN4pdy77m7cHunJFF0lp8AL8zcfidwV+ExPyNa2/tZcYaA4nskdaNvA58kxqxDdCfeCdgI2JCYd7r0/3UZWfK9kij49fjg8vDgv/OIOhmPArcTF79WjGwzJP4G7E0U/vwy5d+1FwJ/GVw+DlzbYD23Ecn6T4gq8APA8cRn4y3EcStJkprsteS7xl2VNpyWGE7Xd6g+p3rx9kvqPL8fur4PEIlGdhudc1fdZoBoZXwz8CkiEbkcWELlZzjFspRoyTyTSLTeDbwUW+Y1fAPEcK3byR9jpYKJ1QqjVlvHbPLTwt0PvLgF8UqS1PfWJ1/leMXgfb1kuIk6wK+rPKe03NTguf2QqL+Y/PY9hUNj1PmmES2Ms4FziJbA1Mn4SJblxPfQqcAsYAf8/GloxhPHTHGGk6VEd/ahzMH+GvIXahdjLQdJklriavI/2MemDafpRpKoH1LlOaXl4w2e2w+J+ufIb9+v0oYjVbUm8Vn+HlFIK3tRspnLCiJxeRK4hxgGcv3gv3cBjw3+7dkWvf5qolL3WUSRxy2a8N6pt00FPk/M/JI9jp4APgxMaPD8rYljPPvcuUN4niRJGoYTyf/YXpA0muYbSaI+jhg3Wq0l63kNntvrifoAcCf57Ts6aURS2Y7AvxLjb0fThf05Irn+FfAt4N+JrueHAHsSxSVH2gV9LDHGfSdgf+Ao4KPAV4ju9xcR49RHk7jfAnwdOIDhz6Ot/rE+0ZK+nPzxcweNq7uXCstln3cJjX8jJUnSEG1P/od2FbBN0oiaaySJOsBnKLeIlZZTh/C8Xk/UD6QyoVknaUTqdzOJC453MPyEdgHwZ+CbwPuJatil8br7tGsDaliLGGZyJNH6+Uvie2i427iI+O46kKHPpa3+8gKqT+n2V2DXBs+dBSzLPOchopaCJElqgkvJ/zj/T9pwmmqkifpI9XqifgH5bTstbTjqU+sSCcIlDL1L+wrKY7tPIL4HunFs91pEJe8TibH2TzD0pH0+0UV5b6J3jJT1CqKobPaYWQn8gJjtoN7zsr1AlhKfMUmSNEpHkf9hXsLQqsB2AxP15tmLym2z5UTttC9RBb3YVbfWcjvRdf1gYsx6LxogpsqaDVxIvnWz3nIzMaa9V98XjcwA0e39PvLHyzPExaFaQyk2Bf5ReM6pOEuBJEmjMpbKbqM/TRpR85ioN8cAsJDe2y51vglEHYRi4ctqy2LgXOADwFYpgu0A04j5rucCDzK0VvYvEYmWVDIF+C9ieFP2eLmb2uPXJwH/W3j8VVjgUJKkUXkX+R/XVUT3yG5not4c76b6Sf7DxNjGbu5KrM60NvAfVE4lVVxWEmNp30V0C1fZGOCVRPK0gPrv4zLg5zQek6z+sinRMl4cYnIRsHON58wiur+XHvs4Uf9BkiSNwBjgSiqvnE9JGVQTmKiP3vNofJJfWp4AzgY+QXSVn5ggXnW3SUTl9vnUP9auJ44zW4KHZhLwZmLoQDaJqnbh4+f0b48EVfdioiZE9lhZTvTcWK/K4/ciP3vKcmJ4hiRJGoFXUHnV/IdJIxo9E/XRGUtUxc5uz1Ki4OAlNB4Pu5y4AHQS0V2y2gmdBHGx8HDiAmG9JPIcopiaRm5DYrzx49RvYZ+L022pbAB4B5XTlz5G9Loq9qjamMpitb/AugiSJI3Id6g8YftQ0ohG53NUTrO2Ywtfb1KV1/tVC1+v1b5O5fHwkczfpxBDJGYTCdRQWt5LU93NAnbA6tOKbrE3UvuYeZqYQm3LVAH2qMnE5/Amar/3i4jpKu0do5I1iQs9S8gfK1dRWWB0HDCn8Ljr8LMsSdKwrQncRmWr6KtSBqUkjqHypP1v1B+HPpZIvmcRyXixcnC15VEiyZ9NJP0mBP1jHWL8dK0p1hYAnyYKpKl1BoDXAtdQ+3N6I7BnqgDVkbYBfk/+OFlFfPdvWHjsUeQL080HDmxbpJIk9YjtqKzw/QSwdcqg1FZ7Udlach+wwQjWtTHRpfkkohv8Suon7s8S3ernAIcQyZx6zyHAA1Q/BpYS3a5Hcrxp5EpTcxVnAckmYXPp/tolaq5DqByy8hRRZHRs5nG7AfeQP57mYBFSSZKG5UBgBfkf3keI1lL1tr2ovFDzHPCiJq1/GjHG+ETgfGJKrXqJ+wqia+5cYpquGU2KQ2msC/ya2mPQTwE2TxWcgJgS74PE2ONq++luYJ9k0akTTSZ6RS0if6xcDeyRedx6xPd+9jHn4IwNkiQNy2yqd1PeKWVQaql9iPHAxVa0t7bwNccRBf5OIKZ6q1fgqrQ4LVx32pWoUWDX6u6wFnGBrNrQhBXEb4Q1JpS1BTGzQPFY+RblZHwslePWb8OGAEmShmyAGD9aPEGbh3Pt9qIDiG7nxf396QSxbEm0ns8lWtNrjWEuLYvId5e3dabzHEn142sZsd+sTdC5Xk5l7ZLScjpW8Val11HZHf4R4nu9dHHnrcAzmb8/DRzW9kglSepSY4hkqXhytoBIiNQb3kPlmPTVwCdTBpWxIXG8zSES8mqxZpfl5LvL25U6nfHAt6m+n/5Ba2dhUPOsQcwCUa3GxA1EYTEpazLVq8NfBGw/+JidgTszfyuNWx+LJElqaAD4BtULC1kIprtNBE6m+r79SJ3npbYG+WnhnmT43eXtstt6E4GzqL4/5hJjodVdDqb65+1J4MUJ41Ln2g64kPzxsoSYPnUSUbek+D3xRywkKknSkH2e6ifcf8Af1G60KXA51ZP04xPGNRLFaeGKXS6rLQuJokYnEsXtJrU76B63BvBnKt/3xcCxCePS6G0OXEHlvn0K6wyotsOJOjfZY+ZO4vt3gLjwurLwt52TRCpJUhc6nhhTWjxBuxvnWu8mR1C9aNsi4mSqFwx3Wrjlg487afB567U/5J6xJnABle/xfTRv9gClNQn4EdW/Q/ZNF5Y63NrA98h/H68Cfkhc8H8dccEne2HvnUkilSSpC+1NdCOu1hJ7KjA9XWhqYCNqT411O709XngK5WnhziHqLDRqdb+LOKZnES32dpdvbE3g71S+l9fjvOi96MtUT9b3ShmUOt4LgavIHzePEjVFZhKzQBSHyoxPEqkkSV1mY6qfjK8mKrtaubWzDBAnQPOpvs/OIVo6+sk48t3l76Nx4v4I8V7NJi5YOcY6b4CoAl58367GHgq9rNpUnk8QszdItYwjaoZkK7+vBn5PFJsrXlT+G1FYVJIkNTCB2lWAS8lfL7fQdou9iUrp1fbRYuAT2FJcUuwu32hauGfITwvX77Ua/oPK9+hKYN2UQY3AgcRxUG15YRNf53V1XmenJr5OO3yCyn1/LU7dpsa2IuqFZI+dZ4F/I2YeWZG5/wFgjzRhSpLUfV5KZTe10rKSqLS9VbLo+tf2xHtfK8n8O1GNV7VNo9xd/nziwka9xH0F+WnhZrQ94nTeQOVFu2uAqSmDGqGbqL2Pb2rSa2xD/QtBX2zS67TTf1K5Hb/CC4FqbIAoMlns9fVP4H2F+5cAx6UJU5Kk7jMR+CzVC82Vfli/gd3W2mEroit3rZ4Oi4AP4rR6IzGemNrtBOIiSLWCfMWlOC1cL77v2wJPk9/uecDzUwY1CvUS9dU0ZxqyLzR4jW5M1AeAX1K5LZ9MGZS6yobE71f2+FlGFKC7snC/UzxKkjQMOwEXU/vkcwlRKdgpV5rv5cBvqV/d/Cy6N3nqVFsSredziQSvUXf5p8lPC7dG2yNurjHA/1F5Yr1vwphGq1Gi/p1Rrn8McH+D1+jGRB1gMpVTty0nLlJJQ3UwcC/54+h64OzCfX8nhixJkqQhejXR7bXeieglxFjMsYli7AXjiffwUuq/15cBr0gUY7/ZiBivPoc4xpdSf99kp4U7GtishbEdSvPHDP8rldv0via/Rrs1StTnE72IRurABuvv5kQdYp71x8hvz9X4Xa/hWYP4Hs1efF5OTP2Y7b03D3hlohglSepKY4jE417qn5DeQYxt3LrB+rxqXrYr8FXgIeq/tzcQSaPSWZMo6DebKLD4JI2TtGJ3+WaN8X0p8CDwbpqTNG0ALCQf+++asN7Uion6M0QRq+x9h49i/b8orOtWKo+Bbk7UIb53eu0CjtLYi8rPyIPkx60vJ75jJUnSMEwkxkTfSeME5TLgQ1TOtzybSAiaWXG52zyfGOtZq3BfdrkGOIreHA/d7caSnxbubhrvz4Xku8tPGsXrf39wnTcSLbujcTKVLc29UIeimKg/TeWY8nNHuO61gOcK6/o3Kvd5tyfqUHlBYh5RoFEarslE63q2AvwqKntu/JTuH04kSVLbjQHeCFxE48RkOfBn4KPk51J9hMokvlcNALsBnyLG4TUa+7ySaM20C2D3KU4LV6/OwGqi22epu/zhwPRhvNZawKOZdV1A9NAYrs2p7Nb/wRGspxNVS9SLVdqXM7JePu8vrPtqYjxuLybqm1A5R/a/JY1I3W4PKi9WFwtZXkN/zbghSVJT7Ua0Ji6hcdKeXc4GpiSIt13WBY4AfkxclBjKe/I08C0aDx1Q95hKflq4YgtsteUu4jM1i2ixr+dtmeetIC4M/JDhJZ7fK7z+bcC4YTy/k1VL1CFqDmTv/8QI1v2PwjqOp3cTdYhjOLtdjxGto9JIjSd62WUvFK4if4HzCWD/VAFKktQL1iLGsZ9DvktbvaU0V/WpdPeUV6Uu0Ecz9JbU7Htw/uBze/nChcI48tPCFbt7VlseIT5Xs4kx8sVpjP5A/nhbScwTP4fG3ZOnUNmK9c5RbF+nqZWov6dw/3DnVN+28PylwPr0dqK+FrCA/LYdnTQi9YpdqSxamy0yt4L4/mtWjQ9JkvrW5sSP6m0Mr5V9NXEi+Gfgv4kWxX2B57U1+toGiPHl+wMfAL5NTGH3LMPbxlVEdfcPAOu1dQvUiYrd5RsNjXiGaBGeQxT62p3o0VLteU8SFwVqFZx7d+HxDxKtXL2iVqI+jcrP7UuGsd6vFJ77m8H7ezlRh/hezm7b/6UNRz1kHNVb17PH2+k0f7YLSZL6zovJtxYuo/H0VvWWhcScvj8jqqR/lCiytj8x5/tGjK41fhyRMO1KnGy/k+gO+3Xgl8TV/qF0W64X/2+JlrxWTtul7jeNfHf5xdQ/tlZQu2W+dKJ7K9Wrm/+l8PjPt2aTkqmVqAOcVvjbd4e4znFENf/sc0szMvR6oj6TfPK0Ctg0aUTqNTtTf0rYW4AXJItOkqQutz/57rT3ED+saxAnst+i+jRGzVgWEq2IDxBjfW8mWikvB/45+P9bB//28OBjF7UgjpXAtUSr5770Viul2ms8+e7yTzD847E0HOV84kQYoitz8eLZtu3YoDaql6jvV/jbAoY25vp1hec9Rvnz3euJOsSMHtntc6o2Ndt44kJlrWF0C4mCtpIkaRjeSX5s2fVExeBqppGfl3oe1X+Uu2lZSlTMXnfY75w0dFsS44PnMryLXqsGl1OAdxX+dks7N6BN6iXqA1ROqfeWIawzO3vFaqI7eEk/JOqfJL99v6n/cGnEXgbcTu3vsjl0Z10bSZLabjb5bpF/Yfhz7c4kqqV/Bvg50QJeLHaVcplPtCj9mDhhfROwBVHFvvSYe4jCUlKrHU60sDca115tKRY8PKnNsbdDvUQd4HOFv/++wfqmUznDxc6Zv/dDov5C8tv3UNpw1ONK867XKtB6LrB2sugkSepwY4Hvk//xPJXmdvfemJhX/F3Ap4mk4ufEXO43AY9T/Ud8OEnLo8ANRPfgnxLj02cTvQT2pn7ht6nk54S9GLu7q3W2BS6kesI90uWotm5BezRK1GeQv8ixgvpjro8vrO+Kwt/7IVEfT2Uhvlq9pqRm2Ze4CF7tu+t2YMdkkUmS1KHWIN+aXGqZS9UdbRqwDnGyvSWwHTG+dw+iwN3uRJKzJVFNfh2aNy3aDPLjh7/ZpPVKJZOIsZvZ4SXFZQWwnPK86kNN1O8gLkj1kkaJOsDfCo+pN6f61YXHfqDw935I1CEuUGS3cZ+04ahPTCWG+1T7/loEvDldaJIkdZZ1iWmisglC8cS13+xPvgDOcWnDUQ/Zj8opDxcS86zfQSSR5wNnEbMjzAW+TAwj+Rgx3eERRDG0/YhkvrSeVfTmtEdDSdSLY/VvrbGunQqPW0plL5t+SdTPIL+NzqeudjqU6r3oSuPWa01HKUlSX5hBvojVEoZWiKkffJzy+7IMeHnacNQDxgG7ET1BNmD0SfXaVCb8vWgoifoUKmd+qDan+jcKjzmjymP6JVE/ifw2fjhtOOpDG1DZm6+0XDj4d0mS+s7OwIOUfxSfxGS06EeU359HcK5hdZZNyJ/YPpw2nJYZSqIOUQU/+7jvFf4+nspZKV5TZT39kqh/kfw2fjptOOpjR1N9itUHiOFukiT1jf2I1rfSj+G9xDhw5U0i5m0vvU9XMbQ5mqV22Iz8Se2DacNpmaEm6vsWHlecU/3Qwt8fJXo5FPVLov558ttudHN6AAAgAElEQVT4mbThqM/NBP5B5WdvCXBswrgkSWqbo8kXsboBW4rr2ZiYuqj0fp2aNhzp/1uX/Antk2nDaZmhJuoDwJ2Fxx6R+fvvCn/7co319EuiXhwG8NG04UiMI2ZpydbeKC0n4ywskqQedgL5aYwuANZKGlF3eBn5eZc/kjYcCYAJVE5L1osnskNN1CGq6Wcf+4fB+zegssr+TjXW0S+J+qnkt9GimeoUewB3Ufk5vBzYKGFckiQ13Vjgu+R/8H5FdO3W0BxDPiE6OGk0UniY/Od6q7ThtMRwEvUtyE9nt5LoMfQxKk/4a+mXRP3/yG/j/mnDkXJqTeP2BLBnwrgkSWqaiVROw5NyjvRulr3YMR/YOm04Um5qxdXAIWnDaYnhJOoAfy08fjZwXeG+99d5fj8k6gNEwpPdxq8DBwLrJIxLKjqMfE2d0sXyf0kZlCRJo7UOcDHlH7dVxLRjGpnx5JOAm4FpSSNSv/sO+RPY/0obTksMN1E/uvD4YkK6mPrJaD8k6ttSuY3Z5WHgHOIix95YRFNpbU70gikep2fjsSlJ6kJbALdQ/kFbArw1ZUA9Yjr5sXNnYe8EpfMO8ieul6QNpyWGm6ivOfiYWknoLxo8vx8S9VnUT9SLyzLgSmLKu2OA7fF7T+01lphCMDu0ZTUxhdvzE8YlSdKw7ET8eJV+yJ4C9kkaUW/ZBXiG8vt7YtJo1M82I19QbiVROK2XDDdRB/hfaiedBzV4bj8k6ueQ375vEj0RTiIS8mIyVG1ZRFwYOmnwuTu0dQvUr14CzCN/LC4F3pAyKEmShuJVxPzBpR+wh4jEUs31JsoJ0irgLWnDUR+7ivxJ6wfThtN0I0nUX0715PJBomWunl5P1KeTn8ViNbBj4TFTiC7vJxDV4Yv7oFGX+ROJegnTW7ol6lfTgPOoPP6+lzIoSZLqOYq4slz60bqRaHFTa3yBfOtSremepFb6FPmT1WvThtN0I0nUB4A7GFnC3euJerEC/m1DfN7ziOT7RCIZf5z6SXs2eT+DSPr3xtlG1DwnEIXlssfbNcTwF0mSOsYJ5LsrXgisnTSi3jeGfBfSe4D1kkakfrQJlSerr0oaUXONJFGHOFlfp7BMGMLzejlRH098T2W37ROjWN/GwOFE9/dLgOdonLgvJ/bpqcRY+R1wvLtGbicqu8IvwincJEkdYAD4Kvkfqd9gq0W7TCV6LpTe+78A45JGpH70O/LfAX9NG05TjTRRH6leTtSPJb9di2luTYNxROJ9NDEH9k0Mbbz70+THu89oYkzqfROp7Aq/kuhtJElSEhOB08n/ODlHevvNJAr2lfbBN9KGoz60B5XJz2uSRtQ8JurNMRm4j/x2fbcNrzuV/Hj3u2mcuFcb775uG2JVd/solReGLifOlSRJapt1gL9R/jFaRcx5qzQOIN/9+N1pw1Ef+jP5E9Rb6I0TVBP15vgs+W1aSrpprTYmP959PkNL3u8ikn3Hu6uWXclfOF8NPAnsnDIoSVL/2Bi4jvwJ19uSRiSICyXZLqV7pA1HfWYXKseq90KCaaI+ejuTLzS6Gvhy0ogqbUl5irhLiO/QRon7MuL4mEt5ijh7lGky0ZKePVZWAO9NGZQkqfftCNxP/qT1gKQRKetnlPfNI0ShL6ld5lJZuOvlSSMaPRP10ZlMVMLObs+jwFopgxqC8UTiPYvyFHFDGe++kPJ498OBjdoduDrGFyhPo1paziWOLUmSmuqV5OdIf5jo5qXOMQn4J+V9dCVxoiy1w1rkL+StJqbR6ubiXCbqIzdA/uJhaTksZVCjMI3yePcziIuhIxnvvk6b41Y6+1E5G8H9eBFdktREh5HvCngTsHnSiFTLxsBDlPfVqWnDUZ85iMpWpKuANVIGNQrTyE+x1uppJ8dTOa1bt15s+ySVSevPkkbUfMXx7k/SOHFfQXmKuNJ4916o56Dq1gfupLJGw5tSBiVJ6g3FOdIvxfm6O93LgCWU99kJacNRnzmRyuTkdKKFVf3hECq7it8ATEkZVBuMpTxFXGm8e/a7uN549yspTxG3A35eeskAcDKV+/1/iWNGkqRhGSAK/mR/VH5L97bu9JtjyLfgHJQ0GvWTAeCXVJ6UnozFtvrBvsAi8vt+PrBVwphSGg/sTnmKuJuo7HVSbVlAJPpziAsfzZxzXmkcS2XRzRuxloEkaRgmAr8g/2PyLTzJ7jbfJ3+ivHXacNRHppCfHSLb9XlcwrjUWgdTWTF9GVHjRGWl8e6ziS7zjzL88e6vpnuHlPSz7YB55PfrM8QFLkmS6lobuIjyD8gq4qRA3Wc8cCHlfXkzcYIotcMMqhfcMlnvTYdSOQ3bKuB9KYPqItnx7ucTydtIxrtPaHPcGr6pxDDC4mflszjkQZJUw8bkp9JZChyZNCKN1nTgbsr79Ew8EVD7bAs8SGWCcQ6dP0WXhu59RMt5MfH4QMqguly18e7FCyHVlmcoTxHnePfONQB8icr9dzGtL1opSeoyOwD3Uf6xWITjmnvFruRbZz6TNhz1mS2Au6g8Ib0D2CldWGqCicAPqNy3JumtsSblKeKGO979fMpTxK3f5rhV2+upLDj4KH43SpIGvRR4gvw4uN2SRqRmO4zyCd0q4PC04ajPbE4k5sUEYhFwRMK4NHKbAv+genfsY9KF1XfWIj/evTj+udF499mDz3e8ezpbEfOrFz9Hs1IGJUlK71DyxX9uBp6fNCK1Srab3SJgx7ThqM9sTPXEbhXRTddEoXu8EXicyn359ODflFZxvPuzNE7cl5Mf7747Th3WTpOIoWnF/XIa1h2QpL50PPm5bi/DOdJ72RiiBaW0v+/B/a32mkgk5dUShbuB/dOFpiHYgEjkqu2/24ghVOo846gc716sKVBtWUTleHe1VvG8bDVwKzGESJLUBwaIK+3ZH4KzcI70fjCVaDUp7ffzsQK32u9o4DmqJwdnEEUQ1VkOp3or+mrgbCwO2G2mUDnefShd5h+hPEXcIXixtxVeQfROyb7vzwAHpAxKktR6E4jpkbI/AD/EZK2fzASeorz/v5Y2HPWpl5CfkSC7PAQci11vO8EuwHlU309LiTHOVhXvDWvz/9g773C5qrJv3ycnFdIhCb0EpIMgHYJYIkUBFYyvogKvKDYEO9gwNgQriA27COoH2AgCigUFVBBQKdJLpIQACSQhpJ/z/fGb/c5u08ua8ruva13J2TN77WeXWXs962mqzz4XKeOlFmby4t0vplgizov+jbMZ2cWTIeAzyDvOGGNMjzEeuIrkoD83pEAmGIegZDXRs3BiWHFMnzIOOIvksxhvdyJLrhXB9rMFcD6l780/gRcEk860iyje/SzkBl/KE6ZUvPtJyGXeymXtjCFrWBkGfgdMDCiXMcaYJrMxcAvJF+lbgkpkQvNhis/DCmDfsOKYPmY/4HZKT/yvBw4KJl1/MQP4GqVrdi8H3o+9HfqVdLz7TWRjqvPaUpLx7jPbLXgX83ayC2aP4JwBxhjTE+xItkb64UElMp3AAPATku6LmwaVyPQzo4FPkqxCkW7XolKDVhKbz07AtylvMf09KiVlTJx6492jEnFzkdXeuSlKcwDJkLVhVH/91SGFMsYY0xj7kYwzW4DKrhgDcj3+B8Xn42/I3c6YUGyGXK7XUHqC/wCKjZ4SSMZeYhZSloYofb1vRyEIxlTLxhRLxM0DnqI65f1+iiXiZqGyZUZMA/5O9pp9CYcWGGNM1/EqktaR+4Btg0pkOpEtgIUUn5MfhRXHGEBunb+i/KR+CXLT3j+QjN3KxsB7qWz5fAB4A1YCTHPYBC34RCXiynnPON49n5HAV8hep2vwwqUxxnQNJ5OMG7sBrcYak8eBJGNS3x1WHGP+jwNQ8qRyFt9oIfKTqKqByTIROB5dy1IJ4qL2EHqHjA4hqOkb4vHu5yOFvJ54963aLHcn8EZgNdlQgl1DCtUA44AXAh9Ez8KvUfnYq4FfAt8A3oO8REcFktEYYxomr0b6r4H1AspkuoN3kLRivCSsOMYk2AX4DtVZ4W5ArvG7BZG0c5iGJvT/j+qydV+PLJ7OAWBCMYFkvHupEo6V4t2ntlnuEOwBPE7yOqwCXhtSqBrZB93nZVR3n4eBRcA3UV4NY4zpGkaimujxAe17uEa6qZ5vkXwZOnGU6TSmISX8Yaqb1C1ENZ6Po/ddQ0egHCSnIYtjNdbJdUjBOSCAvMZUQ1Qibi56VhdR3W8/He/ei/lXNkALbOlz/xadPffbGric6pXzUmPXBXS3t+hvKXoOXI1L5RrTs4wHrqQ4gLlGuqmHUSjWLXqO7sD1Wk1nMhrl4biE6qzskafIX4FzgNfT/SWiJgEvAz4GXEb1CbuGgZtRnPrGbZfamMaZSbFE3HUoA3qlZ341eqedX9h3Z+SF2O0Mkh+3fh0wOaBcpXgd8CyNKenx9gTwonaeQBNJhyF9Lqw4xphWsBGadEU/9LUo6Yox9bABSXfDX9AbkxnTu0wC3gz8keqsyOlJ3jzg46j02650nuVtANgSKeXvBn4I/IfKcfvp9iDwGVSy05heYhRSvE+iWCKumt/HEorx7nOAGe0WvIm8jmSumWFU6aeTfu+nUf6+/APlGjkROAJ5UpwEfBa4rcx+a9D5dxtW1I3pcbYB7qX4I38WeEVQiUwvsDuwnOJz9fGw4hhTkSOAg1F5t3eg3By1xD2mXSofAK5CE/h3A68BDgK2RwsDzWQMsDmK1zwCLTp8Brns/4vq4svz2hBwU6Gv/fGCm+kvJlKMd7+YbDx3qZaOd+9Eq3QpdiAbFrQSODykUAVOpvQ1/zFyh6/ETsBvSvSxBjiq6VK3Fivqpun4Rd857IteJlF8ziL0UvlbMIlML/EaNLkZQC+Q1wKXBpXImHyOQs/qOuAw4NrC9jFoon4Ymqju3MRjrgSeRBb55citdiVyw1+LFgkiRgPrIxfVKJRkMgpZ2ojmKgKLUKzjlSj+cWET+zam29kE5XGI2oFUzl2xDrgbeS5G7Ub0m+9EJgI/B2bHtg0DnwY+EUQiJaf9Ldm4+aeB/0FjVi0cC3yfrPfTMrTgeVcdMoZgLcnknWcBHw4kizGmSiZTuU7oK0laPO8HntdiuUz/cTbFZ2wZyrxtTCfxJpJWiZ+U+e7GaOw8E7nJL6W8Za3T21rgVpQN/y3Ibd8Z242pnkGKJeJqjXe/iWKJuE6Ldx9ALuRpN/MrUSm0drI++dn7n0FKdb0cRv69uoHuGQdtUTemC/lqhc/fjFx8oh/2jcD0Vgtl+pIRJDOzPgBsGFQiY4q8lWRM+mXUFl8+iBafTkQT7qvQM15rnHs72iLkLfUD4EMoedL4Gs7VGFMdo5C1PSoRV228+zNI0T8LeTd2wrzsKLLK7L3Ik6ddfI7stRoiafGvl7fm9D2MFi67ASvqpul00ophL7If8EvyM/EOILelT8S2XY0SIC3L+b4xzWAiUhCimqVXAy9HLxhjQvFO4GsU30k/o2hdb5SxwHaxtgWa2E4r/DsDWYmaxVqKbvSPFf7/GJpQ311oTzXxeMaY2piEPFYOROE0+1CdIr6Aorv8dajqxHMtkrEUz0PVXDaJbVuOFOW/t/jYG6IklulFxe/TnFJkA8g76kWp7fPRea9pwjFaSTtd33cAXoj0iw1QyMdzwGLklXsrcAuNv0NHot/IC9C7cjryQFlYaNegxa9WMxN4KfJ2mYwWh55Axs2rqO53OIB+6weg52k8WpBbAPwBJT8cbrLcg4Vj7oPmHDMK2x5H1+8vKHdNs49rqmAMmpDdn/PZSODbJFfefoBWfo1pNdujWLLo2ftCWHFMn3MaybHwO1QOF2o266Nknnuiyc9slAhuDir9dlKsnVDYfnThe7OBvdAEohOsbsaY2onXd7+aZDhiuXCVO0jWdx/dBlnHI4U2Lss6tODZSt5P9hosA6Y28Rg7ke/x8KoK+22ds8+b6pThilQ/pXJFHZ5zzFrbDTXINR3N1/5bZd9LUH6DI6k9fGALtADzTBXHeQj4KLBejccAvUfT/e0Z+3xfss96nvfL6WRzJkQMAMcD91Xo527g0DrOIY+ZSK9bVOGYw2gh/2PUd/1MA8xFN+CW1Pb1yWa5PKutkhmjwSjupvXmsOKYPiWtpH+T9ivpxhiTJi/ePV0uLa89S7FEXCvj3QdQfo60UnsBrRtD/032fL/XguNck3OcX1bYp5cV9UHkfdtIvfq3VXfqDKDKQPVUJ3mY2isSlFPUTycbTlCu/Y6st8dklMuhlvP4UI3nEGcM+u1XM1ak26PAixs4tqmB7SnepD/Ets9ACUuim7IWeHvbpTNGfITis7iCxhLBGFMLA8CXSL6kzg4qkTHGlGd9iiXiaol3fxpZ6eci6+Y0msfhZOPW/02xIkWz2Jz8czugyccBKdjp4yylvLdCryrqE1BFqEaPU42uMRJZgRs5zhrkdVYtpRT1j9d5/F/E+p5EUueqpdXz7EwF/lzn8aK2ElVO+D9KuQmY+hlAK4yDaABfWtg+E8VRRNncn0M34/J2C2hMgc+hOL3XoTjeXwF7o1U9Y1rFAHAOcEps29lo9dwYYzqV5chafl1s22QU+jILKRj7klXEJ1MMk4mI4t2vA64v/H9FHTJdiXJvXA9sVti2G4ol3w/lxmgGB+dse5rWlBC+MmfbBHR9O6lk8d0k31tnkvRmuAbN+8uxoMxnY5CVeL+cz5Yh79yrUdLUxWgetwG6//sBh1CbO/XXyFeyFwI/RS7ojxaOsyWquvJqkgsoI5HL/FKSSnMtHI6qHEQ8jDwq/oFyvqwH7AG8ES3QxHk1Kj98CVpMi7vR/wMlqb0HXa/pwEGFftKW+C+jBZtFVcq8Prrfu+Z89g+02BLJvxbYtHDsN6BFsIgxwIUovKGTnvWe4m0kV5Z+iCyVC2PbF6EkJsaEZhzFBBrDKDlOLZm2jamFQfQSj68gfyyoRMYY01yiePezkCJejRvxGpLx7ntSmwv7WOBPOX2+tuGzEWkPqGHg903qO4+8EnDlYvBDWNTTNDvr+9fJf1a+SXVeGesjRfBfVLaoH1XiWD9ElulS7EIxGVq8PUUy4WEp8izqkUfyOjQ/KDUnHQv8KGf/21A4Z/T3AuAVZWTYHP320v18vAr5I/LkeAglwCvHOJR3IL3vg5S/7qZONkLJG6JyQGvRKlC8vu8DyDXemE5hC5ILST8MKo3pVQbRJDR6zobQhNQYY3qZkSTj3W+iurKRy8jGu1fi02Td8ZuRMDbP/bqV+ZUuyTneuWW+32uK+svIns864B119DWAvDpKMRolNEsf73yqy68wlXxF96Iq9s1T1KP5wRur2H+QfPf2aHHsMWDbKvrZhvzSh9Wc/zE5x78NeTdUy0dz+vhoDfubKrmU7OAb//sfKE7dmE5jFsnkFyeHFcf0GKORG1z0fK3FCQyNMf3LeLLx7pUU90jxmEcx3j1PGXgVKqEV3+8GGqss9M8cWd7VQH+VyLPg/7zM93tNUf8T2fNpVV32Y3OOdRu1PS87kX3mVlPZql5KUf9GDcd+VYk+hqkti/t3cvafWcV+N6f2WYoMYLUwgDxU0r91e7g2kSMpP7jeiOJ/NyUbC2FMJ/Auis/rGpx90jSH9VCcXlxJPy6oRMYY03lsTLFE3DwUz1qt8n4xxRJxY5Hi+kTO96pxR84jryRYNRbPejkj53jXlPl+Lynqe5M9l7toXem/a3OO97I6+jkvp58zKuyTp6ivpLZSp2PJDy/5cw19ALw8p49jKuzzkpx96g3nm5XT18vr7MukmIASLNRSRmAdqv33APAtpMAbE5rzKT6ji6huNdGYUqxPcpV4FXoxG2OMqcwmwByKJeJqiXe/CLg/9dlK6lPC0kr/MFpUaBWn5ByvXIb0XlLUP0n2XFrl5TierNwPUF9JwV3Iyn1thX3yFPVL6zj2LTn9nFBjH5vk9PGRCvt8I/X9tTTmOf1Iqr/PNtCXifFVqivRkbfaMzunP2NCMYpkDdN/IWXLmFqZjJITxieIRwWVyBhjupvRKEHxu1ACqzupLt493oaoXYlclNPPYY2dSlnekXO8m8p8v5cU9WtS/awDptTZVyVeTPa6fbGB/u5M9bWC8p4AeYp6PblrfpnTzzZ19JN2369UNva21Pf/Wscx4/y/VH9/qiWjpMlnbzRgVrP6tK7w719RJsCDaW3WTGNqZQ0qG/hw4e/no/i5elZXTf8yBfgtsH/h7+XAEag0ijHGmPpYjUIpvw4cD+yIFkUPAt4D/BhljC7HAPBuanuvr8zZ1sqs1HnJz55r4fE6hUFU4i/OXagUXivYM2fbLQ30d3Pq77HI0l4L99Rx3GWpv9chz4BG+5lY5rtTyCZ4vL2OY8ZJl0fe1HXUG2MkSj5QiXWozMZVyKXlH60UypgGWYjqY16H4ouPRu4/dsEx1TAD1XaN6okuQXFWja40G2OMyRJlh38g1g5ESlgpS+x9yGJXLUvJxre3UlHP63tpC4/XKUxGym2cf7fweHmx4P9poL87qjxGOZ6p47irU39H1bYa7adcQr3NyS52vRTNf2plEC0KpH9jU62oN8YHkcWxFEPoJv4CJQdp5OE3pp38EzgJuLDw96eAW1GCG2NKsTnwB+B5hb+fRu6RNwaTyBhjeosxyEq5B7B7oT2f2hIV74QUwjxLeR6PAjuktlVTy7te8pS7x1p4vE5has62xS08Xp7nQj2KckSe5b9cabg80spyPaxqQh+VyLtXM2lubqfJVtS1arcRKnGxHrJ8T4p9NqKwfTWKP1mJYi6mIeUljyim5KfAZ6jPjcOY0FyEXv4fRL+Di5Arc96KqTFboVCeKC5sIXAIWuAxxhhTOxOB3ZB1fCfkarsXjZVteha926tV0iHfnX6PBmSoRF7f9bgydxt5HhCt9CTI81xIu3/XQp6stSrq3UKeot5sBntdUV8P2C7WtkardNNRGYxpZF1MmsESpJyPQ5ka7wPuLmybjyztxnQDp6OJwctRdYN5KC/DopBCmY5je6Skb1b4ewHKLOxFHWOMqY5NSCrke6IY9GpiyZ9BscyrkRJeyi39GeQWX6vS+6+cbXvV2Ee1jCMb+1tKhl4jLw6/FXpKxIqcbY0sAuXJmneMXmBdzrbbgcebeZBeUdQH0WC2L/ACiop5XvxAO5iMsnLuk/PZKqSw34MG1X+gkhNNvbHGNIkh4PXA39FvbGvkKXI4+YOU6T92QjFZUWzVfBSndX8wiYwxpnMZBLakqIzviRbAqy3rtAAl7boZLYY+iGown16hj2VoAbWeMMy8zOdbojlBpeR1tXIA2djgdZQvz9ZMBtt0nDzy3NxbaZHOc3OfRP1hBnkLRK1KhBeap3K2fReVUmwa3aqozwD2Q4r5fmhVb0JQiapnDEqytGtq+3w0CP0dxXPeTG1uSca0iqXAq9HzOQm96D8HfCikUKYjeAHK7r5h4e97kJL+SDCJjDGmcxiNcnbsGWt7II/PSqxFY+odSLm+Gc0Rnyx8vj7wFuA8Kiv5q4HXUL7EWTn+icKZ0sc5AfhEnX2W4n9ztt1IeYUvz1O13spWIV21F6Pw2biRcesWHi/vmm6ByqzVw1ZVHqMXyPMsbfq96hZFfRRaLTys0HZrYt/LkTV7IXI5WUcxxmIJ+vE/hwbbkaju+Uw06M1HE9IpyJ1+GuUzBJZjy0J7beHvFaiW4lWF5jh3E5K7gdcBl6PV5g+igfwHIYUyQdkbjU1RnNadaHzsh4Q/xhiTZjJK8hZXyrenOgvtMjTPixTyqOW5DUcK+ukox1Il1gHHAr+r4rulGEY1nk9JbT8B5WtqlofdFFRpJs1PK+z3bM62WpLrxUln3m4nK5G37Y6xbXsj/WNtC46Xp5Dvjhbg62H31N/D9G4i7QeRfhhfdNsvkCxB2Bh4GypiH6XZr7WtRfHhVwBfQfXOX4ncarZBg10tHAr8F9WfHFfiO9OQO9OL0cB4GvA9VDrjyTrPYxi5kX4deAVaNDAmBB+j+EyuQC8Q038cTHJcvoWiVd0YY3qdTYAjUUWfi5HVe4jq5nOL0ZzwXOA4NGesxvq7PnAqcn0v1/+y2P+HgDc3dqr/x67kn+OpTeof5B2Q7v9ZKifuGpmz36frOP7mOf3kuf3nsSq13xfrOD7At3JkeGmdfVVii5xj1aukT0KKa7yvuyvsc3TO8fNqu1fiu6k+FtTRB2R/W9+t8P3fpb6/jupDWLqSCWjQ+i1SsmtRZJ9Aia4+htx0d6L5Cu1LmtDnVLTicgIapP+GVtBqOddFwDeRl0GIGHzTvwwAP6P4LD5K2NVn034OJ/kyvpH2ZD81xph2MxIp0scBZ6F5Zi1Gl8cK+8xFiv3GdcgwHhl90scdIqsc3pL6+311HK8cvyR7jsuQwtco+yBFJ93/56vcf2Fqv1/XIcN7co5fraK+KLXfD+s4PsCcHBkurbOvavhv6lhrqe85fQtZub9XYZ9uV9Q/RFb+z9R57I5lJLIS/wS5oVcz8K1FcTrnIqt1M2vWhWA0irc/BZXAeojqXwIPoodix3SnxrSIcSjOLXoGr6exLKGmezgSeVJE9/7PdE9+EGOMKccEZAA5Cc0vryNrISzVViOr+gXIwjybxhcwx1Pagv5Y6u+ngW+QVHTnNnj8PHYg+Q6I2k00Ftu9BZrPpvt9nOqv45WpfZdRm+fseuTPv6tV1B9I7Xd5DceOMwp4ONXXOuBFdfZXiU+RPeev19jHWLLnPwwcVGG/blfUNyTpwTKMvA23rPP4HcVENAA9RHWD4BPIveg4+sN6MxO9LOaRPyjmtevQSlzIjJWmP9gS/SajZ+8HYcUxbeB1aDIa3fMrKR0CZIwxncwUpJSfipTrO8i35ua1JRRd108q9NPMxeoJyIKettAOI4v5E6ltV6EEbGti22pVtGrhtBy5hpF31QZ19LclClPN6/OYGvr5RM7+769h/94/mEgAACAASURBVLSyV6uifkVqv/k1HDvN+3LkmI9c85vNpiSfnWFkEH1hDX18iay8/6xiv25X1AHOJnsO/6JxI0Y9Xg1NYTs0gDxL5cHwZpQsY3f628V7PeR18A2qc7m6G3gntcfgG1MLs0i63b0zrDimASrlGngjyRf5PFpb29UYY5pFPJ58HllrdLn2GCo/GY8nb9V8dCqybj6TkmEI+A0KO4vHiC9HCw0vIWnQuZD6M55XwwDwC/Kv10LgDVX2MwLlfCqVh6rWGO9tycbQL0N5qcoxmvy48FoV9U/n7Pv2ms6gyDik7KX7+y+1KbLj0DV+ZYXvnZNzrGeofO0GgI/n7DsEvLwK+XpBUZ+C8oilz+M/KLlkreyBFg/zssq3lBegAbLSiuVDwGdRjLnJMgo4AmXArOSStRi5xYcsNWF6m5MpPm+rURLFUhzVFolMrWwLfKfM528nOW7/jPqrWxhjTKsYRTGePHJdT7ullmpr0WQ7Hk/erqRQGxaO+XRKpqGCPP+LDDDxz/6Kyr7tTVLRvYz2VHQahxYwSl3P/6BFh/3RQskguj+bIkvt58l3lY7aD6hvsSFt1R5Gc+VPkQ2T3QwlrY4rWU8gS3A9ivrOJc7lBpRX6ozC8U6KtVeV6W878hcx1iFF7qXke3JMBg5Biw9PUd2CwTh0z/J+F18hG147iMI7/ljinL9V4XgRvaCog6qR5Rmg1yJ97RBKVyGYjpKVn0nyd760xPebzjZIyHLZMJcD30axDP1sOa+VCcDxqIxbuRfQIuAD2AJmWsO3KT5rT1E6Z8TtOCyjE7mC0glfPkhyLLmQ7inraYzpXSZRdF0/Hynl1SbmXUUynnwW1dU1bzbTkIK+JCXfOqSg71v4PJ5ceXVh2yBS1B+PffYn2jvPGwf8mOqu+Tqqy4o/BHyO+j0CZlLeY3cJioXP+84qpPymlf1qFXWAS6o4x7QSX46XkF3AibcVKGzgRmSB/y/517kay/5OqOR0qWMtQJ7Ot1O+Gtfvqd6jt1cUdYCXIQNpqeuyBhmjb0H3/W7Ke0m3XFHfEGXILDdwLkADTj0xLSbJ7uhlVS6e/WG0gueJtmkmo1BSseg5+xfZQXpi4bNK7lemvbwC3Ze81e90HOK3aK07pTHG5BG5rp9GMZ682lJoT5MthRZ6wXg6mh+nEyhHCvoeSM6bU5/fXvgMFKv8EMn3bijvyRMoWm4bafPRO6lRjqL63E5Re46idbsRRX0KyflQo4o6SIHOc62upVXrgr8l+Zb1attPqa06Vi8p6iDjdF7IQj1tSZ3nUJFBlAShnLvRjcDrsftkK9gYxcmUW6W5DZWIM6ZZzCCZpfTnJL1jdihs/0P7RTMlGE0xgc95qc/SWWC/hr2djDGtJSqFNodiPHm65Fa5Fi+FNofWxpPXwwykoKfDFtehZMk7IHlPJWnkWocWGiI35+kkXWTvRNb5kExF1z1u4a+2PQS8l+Z6NRyEFnSqOf7NyNgV0YiiDtKD3oTmO5UWDKpR1EGeEu+nttKAw+g5ugi50VfLWLQols6VUK7dRX2GmF5T1EHj2JvQNalHQb8N+AjNKXmYYRdUNq3UwW9Hg6dpPeuhH1opl5khZIEvFTNhTK3sQdJC8OHYZ7Nj23dtv2gmh7hb+5cL2wbIJpU5K4h0xpheZjTZePJqS/Suoei6fhqytm/YXvFrYgt0jmmlbTU6h0iJ2opsKOODJLNvr4cUx+jzh+msUlCDyAX4TKSozieZiHQ1sg7/Fin2B9K6xZSRqFrJhSgePnq+1gL3oDrnL885/kFIV4naSxqUYQeUv+fIVL9zkKt9LayH8lSdh1yoH6VYjWU1Wij5C0o+fQyNeVlMAv4H+BH6vUXVBtagRbGbUMK/l1K/p+6mZK/JlDr62SvVx5F1ypO+R3vV2Q/IA3F/4KMon8M9JBc/lqHf7+/R/ToeXY+WMAoNlqXc3Ocjl+vQ7kb9yFTyV3DjL4FDgklneo03Uny21qEXCmgyFm3/ZhjRTIwZ6CUxVGhno/E5vSp9RigBjTE9w2SypdDicdfl2lKkEMTjybulLORWSEFPz40jBX3b2HePIxv3ewFJY8ooVIot+vwJYPtWnkATGUln6AC96snbrvNy6GzjtP0Z3BFZyvMG2EUoK3Qt8QqmNWyBVsXy4rqGkPLUzDqgpn/5IslJ1s7Iuh5tW4EWkEw4fkgyk/tn0fgQHxPeE0o4Y0zXki6FVkts7WKy8eTdmBdjBzSepmtTr0BhRPFa2NOBX6W+9zjZKikDKBN69J3lyFpnjDElOYrSMQzzaKEJ39TNQWTLfETtZrQCbEwjjEA1X6Pn6m5U/ituQXlfMOnMnmQX7OKxfEPAu4JJZ4zpBqJ48uOQ1948aksoFo8nPxLl1+l2dkVW8LS3wEoUapieEx9DNub4EvLd+M+OfWc1cHjzxTfG9AqDaHDNs84uAF4TTDJTDePQizXP9exJFE9sTCNMQfE40XO1kOLztg6VEOkEN7h+YwAl8yzldroWxUkZY0zEBORyfhLFePJqs2qvJlsKrddy4+yPFh3Sc+JnkYdZuh77RKS4x7/7DLq+ebwj9r0hVFfdGGNymYKC4vMG5B8RrjyEqZ19ybeur0EZOI2pxKHAtShpzAVoAegUlMnz9ZSvtVlvsg9TP/FcAem2DvgrGsfPL7RvoHt6FqomsVn7RTbGtJFN0GJ9PJ48HiZTrj1D0XX9JKSU93JI3SykoKevw1J0DTbK2eelaKE6/v2rSbrDx3ktyet/evPEN8Z0OrVmWpyCMjXundq+CiWTO7cZQpm2MgHFPR2T89nZ+KVgKrMJ8H2ktFfLOuBPKDtsSEagsjbT0KRqBooZnIHGu7HIA2UQWUFAi5HR2LkUncsK5N64FiVpew55ETyOvFQej/1/dYvPqRQTgHvRuebFfUaJ5YYLnw/Gtv8UJZZ7oPViGmPaxCYoFCZqe5GvXOaxAIXL3YHqLt9Msf5yrzMblbBMx4g/BXwdzYWfTn02FnmifpDi+LsC+CTwBTTOpjkYzbmjxY5vIeu6McZkmAHcSnbl8BGc0KLbGUALLXnusF+ks+qQms5kACWOXEl+SExeG0JJd1rNCJR74RDg3SiZz+9Q7dZqMw83sz0BXA98D/3uXo1iPVtteTqrBhmj6/J7VHbPGNO9jCJbCu1Zqh8L7kB1vuciT6jpbZW+MxiBzv1GstfocTSWr19i371RrfP4PjdQPmP7Lii5XvT9X+NwMWNMCTYlv3D7X8jG3pju5TDykwOeh5V1Ux07oNqe1UwA1yGluZkMoJq0byr0/Q+qj6UM3daiUJQfo0WPvWleGY9tKNZarUZB/zvwoiYd2xjTPiaRLIV2E6VL56bbKrLx5Ou1V/yOYwSqpxx5C8Tbg+g6lSoXNxIp8Kti+6xBi6blxvbNSLrH/xlZ5I0xJsOGyF0yPUBdRffUsjTVsxcqq5e+3+eEFMp0FSOR5WUdlS3Wy9HEspFjHQR8AriC/Ge3m9sKZH3/MvAq5L5eD/MoH2cafXYnmpQaYzqfqBTaaRTjyav1aHqabCk0W2yLjEbXJZ4UNWr3oRj8cjWld0ALxfH9/oPmWOXYgOSiwO0oDMsY04dUspKORC6iL05tvwLFNK9shVAmODshl9d0uZS3Ad9uvzimS9kP+AlyOy831ryH2vJbbIq8Pw5DsYKNJrBcjOLJn0Rxl9H/n0Bj3AqKsedQ9DoBKc4j0aLl2ML/JxTaJigefAaK+5xe+LuRyfAaNLm+CsUu/ruKfWajZEV5DKN78whKFvc9pLQbYzqHkchNeiekUO+JksFOq3L/KJ48anfgfBOlWB94C/ABsskzbwW+BFxE6XFyAHgrWlyNXOGHUYnS96L8JaUYh8bqAwt/PwocgKzrxpg+pJKi/nXgnalt85DFZVVLJGoNJ6P6lnnchxJ5NIOPUzor8j+A7zbpOO1gB6Ssx+t+rkHJv/4cRCLTjUxAeQ5OQpaedBKzYRQrvi35yXQidgTegKxHu9Uhx1PIrfxuZCGJ2n20dywbAWyB3PO3L7TtCm0Lag8xeQwtnP4E/S7T13AkClvaJrU9UtAXA59HHjPdNKYb06uMR+NCpJDvifJEVOOCvhaNa/Ekb39D458pzxTgXahqSXoB5K/AmWisHaY0W6DkvC+JbfsvcAJKnlqOQZQH4OjC30uAF6LFAWOMyfBWsu4+f6S8q0+ncjml3b/W0Jw4+y0p71r60yYco93shqyI8fNYiF5GxtTC0UgpLPUbeXnOPhsjC8RNJfYp1Z5B1uZPAUdQvdUpNOsjN/4PAJeQLeFTqT2MKjXEFzI+l/O9IZRI6izqd6U3xjTOFJLx5LWUQluKvGvOpxhP7jjm2pmOQrWeJnuNr6P6MqJzyIZeXUz1butfi+23At1PY4zJZQ+SyS+GkZvUBiGFaoByivowcr1tlE9UOEY3KuogBSsd83YD3blgY8Iyg+JvMf1MRV4ao4DXIUW72ozsC1ECthOQ5b2XEh9ugrLCf4X8hJ6l2q1o8pleaFuJlHfHPBrTXqJ48rnIM/F+qv89P4ZcouPx5HklFk31bIMWOdKJ9oaAy1DoVjVMRq7w8T6eQON2tcyN7buW/HK5xhgDqERQugzbMkq7jncDlRT1Rl2LBpALbS8q6pC/CPHhoBKZbmUAucGvIKusnwnMp/zvKJrI3IQswrPorwnrVuj6XUx+hYZSbQgtZmzSdomN6S/SpdCuRq7ntSjl8yiWQqu2rrmpjl2R98Iaktd9Hbrue9bQ12EojjzezxXUNs6mvVffXcO+xpg+5AyyL45uX93LU9TT7mWN1Ap+UYW+u11RHwB+RdYyt21IoUxXswPZBcFKyvnVwPHYGhwxGk0ULyBrOc8b7y5HieWMMc1hIlosPIliffJqy0GuJlsKrVQdbtM4s5Ainl4gXoXuwXY19LUeut/xvpag56AWjiC5YPCpGvc3xvQZm6LYxfgg9sOQAjWJPEX9j6m/Gyk/9sNUX3/IOV43K+qgMn0LSZ7Tr4NKZLqZbYCfUbmU0C3A+7AluBLrA8cCvyFrKUq3G0kmOzLGVGYTtNBVTzz5MyRLoe2JvBdN65mNrn36nixF92PT0rvmsj/ZksXXU7vhYn+UAT7q43v0VtiWMaYFfJ/k4LOQxksfdQJ5ivpxqb+for4X53iy1qx0372gqIOybqfP6+CgEpluYwZKmrOa0pPaZ1HFiZ0Dydjt7A98g8rJ6H4LvCCQjMZ0KoPATJLx5I9TvfdP5Lp+FsV4citg7WUEun/pOubDKH58LrV7Zo0q7BfPnbIC1bCvteTmTJKGj8tx3h9jTAW2IJtA7q1BJWoeeYr6DLKxRbUk/4h4c6qPO5Bbby8q6gPAtSTP63dBJTLdwnjgk5R30X4WTaKmBpKx1xiJEvPdQOlrPoRKu80MJKMxIUnHk19H1quwVFuD3vcXU4wnn95W6U2a0ehe3k32fj2EvCGqKXOXZhfgn6n+bgV2r6OvaSQt8n+vUyZjTJ8RLw0xDNxJ7auEnUqeor4hKmUU3/arOvr+S6qPD9K7ijoozit9bnsFlch0OodTPkncP5FCORItltWSzMdUx/7ALygdavAc8CFs1TG9yySSpdBuImucKNWWFb4fjye3ctU5TAROJ9/z4XbgTdQ3to1A9zueGX4t8pYYXUd/45CbfNTXAzSnPLAxpscZR7aG5HFBJWoupRT17VPb1lBbhtWtSU5816Daz72sqANcQ/Lcvh1UGtOpTEHlb0pNfuej5Du9siDYDewN/J7S9+RfeKHEdD/xUmgXI8t3pXwYUVtMMp7cpdA6lxnoHi8mex9vRvev3vfL1qhsaLzPB4CD6uxvBHBprK+n0BzUGGMq8kaSg9HjyCWsVyilqIPcjuLb31tDv59O7TuvsL3XFfWjyFobnK3WxHkd2eSDUVuIStDUY5EwzeFQlKgv7/6sBj6Dk12ZzmckUqTnUIwnf4LqFPJhsqXQHALSHWyLFlLyMuxfh+5lIxxHMkxrCC06j2+gz6/E+luBvDKMMaYq0qW3zg4rTtMpp6i/I7X9tir7HEHWnfc1hc96XVEfSXPi+03vMZZsUsq4x8nnaWyyY5rHCDQhfZL8+3UTsGUw6YxJMh55e8TjyeNZs8u1dCm02cAG7RXfNIFZwGVkvSPWoioijSbHnFHoP973AlRGrRHeHutvCCXmNcaYqhhHNnlKr2UCLqeoTyL7sq+mpvrLUvssomiB6nVFHVTOLn5+3w8rjukANqN04rJbkdu16TymUjpE4SngkHCimT5lCsl48lpKoS1BSvz5FOPJx7ZXfNNEogzueSXWVqLnoxku5HPQeBfv/2IaX9A5gmSm+A802J8xps94IcmB6b/0XimRcoo6SImOf1ZNTfWLUvt8NfZZPyjqLyF5fg+EFccE5mDyE/mspv7EO6a9vJz8km5rkWuw43RNK4jHk89D7ujVKOTDhe9ejePJe5Eog/t/yN73J9F7ZeMmHGcS2YXKp1ECukbZm6Qh7Pwm9GmM6TM+RHKA+lFYcVpCJUX90NRnlWqqTwKWp/aJJ2DqB0V9DMlMqMPUlojP9A7vJGkxiNqdwE4B5TK1Mwllh89Tin5Cb+UuMe0lXQrtauSJVo1Cvha4n2Q8ud83vclE5AnxCNnn4MHCZ83KifMy4OHUMX6LvMMaZWuSi9eulW6MqYsLSQ5SJ4cVpyVUUtRHkLUklYu5flvqu+m49n5Q1AFuJHmOh4YVxwTg/eRnU74MKX2m+xhAk+E1ZO/r5diN2FRmIkXX9fOR23Je4q+8lo4nn4WTlfYDG6FFmHQFomFUjeI4mqfojkMW+Xg4xXPoeWuGR+kGwF2xvm/Cz7Axpk7+SnJAfGlYcVpCJUUd4MzU5+Vqqv8t9d33pT7vF0X9ApLn+Law4pg2cxr5lq+52P20F3gR+Zn7r0QTXWNAruuzScaTV1sK7RmSpdD2xGEy/cauaDGnXAb3ZoZj7kNSiR5G1X+2a1L/Y0nG0z+IvT+MMQ2QdvvZJqw4LaEaRX07sjXR8wbXar7XL4r6XJLneGZQaUw7SZcmHEbhIIeHFMo0na2Au8ne699jC1G/MYjKl8XjyfPyUpRq8VJoc5AbfK/lwzHVMxu4guyizhqUA2j3Jh9vJFpcXp061lzqr7WeZoCkl+ozwC5N6tsY06csITlI9qK7ajWKOsD1qe/k1VQ/K/WdX+d8p18U9VNInuN5YcUxbSJ934dRwpyXhBTKtIwZKGt/+p7/CntO9CqjScaTX0e2OkyptgZZ1S+mGE8+ra3Sm05lFFqkyasO8iyyrD+vBcfdCbmfx493O82vcPTFWP+r6E0PVWNMm0nHIfai21m1ivpbU99Jx54PkvVAyItl7xdF/c0kz/EHYcUxbeClZMeMZ4ADQgplWs40FCeaHtc+HVIo0xQmky2FlpccMq8tQwpQPJ7cYREmzWRkzc5LEPc48DFUJrLZDAAnkUz+O4QWn8olDK6Hk1LHaEbWeGOMYRXJQbMXEwVVq6hPJJvNPb7i+vLUZ6Wyw/eLoh5/MQ0D3wkrjmkxM8nWmX0G2CukUHWSV4Ysahc26RgbkK2MEG8fadJx2sVU4BaS5zAEvDakUKYm4qXQLqa2ePLFJOPJXQrNVGIr5IWYlyDuHrS406qFnS2BP6aO+RDKvdFsXkFyAfv0FhzDGNOnpMujbBBWnJZQraIO2Sz458Y+uzj1Wal66/2iqL+X6q6H6X7GIw+T+P1eixavupFyivpzyALUKO8pc4xuVNRBZYsWkDyP5TQ/ntQ0xkiKrutnodjwJ6lOIR8mGU9+JM2pV236hxcgL4u8yhGtSBCXZg5aWIof9wJgQguOtRfJsBAbLIwxTeU+koNZLya+qEVRn536XmQ1n0rWOlZqctovivrnSZ7jGWHFMS3k22Sf6Q8Flagxyinqw8hbpFHyXMW7XVEH2J/sWHgndnkOxQSULT0eT/4c1Snk6VJos2mNC7LpfUYgBfxq8p+zi4F9WyzDNOCXqWMvBF7ZouPNJJlQ8Te4VroxpsmkB9WjworTEmpR1EcA81PfPRrVl49vu7XM8fpFUb+E5Dk6Jqs3OZSse+xFdHfG5kqK+vUN9r9Hhf67WVEHOIHs+ZwdUqA+YQrZePJ4LehybQlF1/WTCv30YqibaS9j0CLRneQ/c+cCm7dBjsORJ0j8+L+hdd4gE0l6md2MPM+MMaapfIvkwPbxsOK0hFoUdciWnvo12Yyhp5bZv18U9XtJnqMTivUeo8ne516wnqYV9WfJumlu30D/56b6SlfX6HZFHeTiGT+fNSi7smkO8XjyeWSVkHLtMbQIH48n7+aFNdN5bAR8lmzekmHgATRHaoWreZqJKFt8erxthldUKUYBf4gd7yEcHmKMaRFvITnAXR5WnJZQq6K+LUkLYjoD7mrKl5vpB0V9Gtl68usFlci0gg+SfI7X0nr3xXaQVtQfRdaX+LbP1tn3aOCJVF95oQPdrqhPJOt99PugEnUno8iWQltGdQr5WuB+kvHkM9oqvek3dkeKcV54xc3oOW6X+/cBZMM3r0Mu6a0kbuBaCuzW4uMZY/qYXUkOcsvoPXe4WhV1gL/k7BO1X1TYtx8U9WPJvqBNbzGebAKq84JK1DzyFPU5qW2PoJKMtXJMqp+HgdeRHRO6XVEHhQWlz+vgoBJ1NpMouq6fj5SKcpUB4m0VyXjyWcD67RXf9Cnl4s/XoYWi2W2UZyxKkhgP+1iBSsC1uhLBR2LHXAsc0eLjGWP6nAGytS27NZNzKepR1NM1wuOtUhx/PyjqPyN5fmeGFce0gNNI3uPF9E5ViDxFfTTZhYlD6uh7XqqPT6MSZr2oqAP8ieR5/S6sOB1D5Lp+GsV48mpLoT1NthRaPYtGxjTCRFTd5X6yz+gy4GvIA7Gd7Eo2Uee/aY9Vew7JxYF3tuGYxhjDN+ltpbIeRX08+e6HC5GrYjl6XVGfTLbe/H5BJTLNZiRSXuP3+BNBJWoueYo6aOIZ335Rjf3OQKEx8T62p7cV9Vlkz23XoBK1l0HkahuPJ19IdQr5MMlSaHNwPLkJz0xksU6XOIue17m0vzrASLTotSomy5qCnKPbcPy9Sc57Pt+GYxpjDAAvIjkQr6R8DHa3UY+iDnoB3JRqH65iv15X1E8leW4P0np3M9NejiRrPZkSVKLmUkpR3yu1fQW11VRPx/T/pbC9lxV1gD+TPLevhhWnZYwmG0+eXrQs1dZQdF0/Df3GqnkPGdMuZqEyann1z2+ivfHncWYC16bkuR84sE3H34pkGbZ52MPFGNNGBoB7SA6CnwkqUXOpV1Gvl15W1EchxTx+bh8NKpFpBenSe98JK07TKaWog9wo45/Vkj341tS+Jxa297qi/j8kz+0pur+e8GSypdDSiUVLtaVIsYnHk3d7pQTTm4xGnhx/J/scr0VKaaiKLgNo/H02JtMQyu/QrvwM6TJsN7Xx2MYY83+8l+QA/Qy9E49qRb15nEjW4rhRUIlMsxmDFI34fd4/qETNp5yi/v7UZ9XWVN8ntd+zaJIHva+ojyFbpungoBLVRroUWl5Mbqm2mGw8uT2MTKczHXl2pHMURfO/c4EtgkmneUV67vYY7c2hNIpkAr1HgE3beHxjjPk/1iMbV9cr7otW1JvDeLK1fL8WVCLTCg4hq8T2WsxsOUV9Otk48x2q6PMbqX1+FPus1xV10PnGz68TYzhHUnRdPwsp5ekEguVaPJ78SFw72XQfLwB+SH61gTuAtxG+1OocYBFJ2S6m/XHxLsNmjOko0vGVa4DnB5WoOVhRbw5fIHlOzxF2xd20hrkk7/O3g0rTGsop6gCXpT6vVNVgLNnESy+Jfd4Pinr6HK8LKw4TkMv5SRTjyVdQnUK+mmwptPHtFd+YpjFI6fJq0W/1SMIvyE4GfkxStqeBNwSQxWXYjDEdx2iyser/oj0ZNVuJFfXG2ZdsfOang0pkWsUVJO/z/4YVpyVUUtTT9cEr1VRP10l/iKT7cz8o6puRPL8VVK6Q0SymkI0nj5dRKteWUHRdP6nQz5g2yW1MK5kBfIxsBY9hlATxW8COwaRLcghZN/yrUFhKu3kNLsNmjOlQjiI7oH8lqESNY0W9MaYC95I8n/nYwtSrPEDyXvdiqa1Kivpo4InUdw4t099Vqe/OTX3eD4o6ZENjnteCY6TjyReQvbalWuS6fhbFePLQVkRjms0LUMK158j+Bh5Fv51OyUE0Di2SDZFcRDiVML9Nl2EzxnQ8F5Id3N8SVKLGOIdsmbVaSi7VylY5x6vkOtupDJK1sA4Bh4UUyrSMkSRL8wwRPl6xFVRS1EGTx/h3flKir01JepsMAdukvtMvinq6hNLLGuhrFNlSaPHsz+XaWpQQLh5PPr0BWYzpdKLs7eXc2+fQWdUY9gPuJinnX2nNAl81bEWyDNvPcWJIY0wHMhVZTOOD50p6L/Ozqcw5ZF/43wgqkWklG5O810+EFadlVKOo75H6Tqma6h9Jfe+anO/0i6J+EclzPL7K/SaRdF2/ifxkV3ltFdl48l5cXDImjxkoe3t6TIvGrAvoPK+oUWgBLb7AubqwLVR9cpdhM8Z0FXuQdP+J3Aa3DCmUaSsnkX3x/4Xuz1lgSrMtyft9f1hxWkY1ijrAP1Pfe1vOd+5MfeeEnO/0i6L+TZLnmBfbGbmun0YxnnyI7PXJa0+TLYUWamJvTEj2RO7teQkS70e/r3ZnSa+GnYGbScp7G5pzhiJdhu0hXHbWGNMFHE12AjUfTeZNb/MWssmYHsLuo73OziTv+X/CitMyqlXU35P63l9Tnx+Y+vxZlG08Tb8o6l8meY5fQO62c5EberoEaLmWLoU2s21nYUxnMhYtUN1I9vcyBPwO/VY60V17AHm8xD1l1qFFt9DJG12GzRjTtXya7AthAZrQm97kbWSV9OfQCr7pbbbBFvU4wnBnogAAIABJREFUGyDX6vh34zXVv5P67Acl+ukHRX1j5HFTrSIed3n9J/B94BTghcgN1RgjtkEJENMJLoeBZciyvksw6SqzFQoJisv9IPqth+Z0ijKtQdnnjTGmaxhBNu7Qynrv8j6yXhSr0Cq96X02InnvnwwrTsuoVlEH+GXqu1FiyHHIFTv+2cEl+ugHRX1fKivlS5Hr+vkU48nHhRDWmA5nBDAbuJhsadRh4D461709znHodx+X/QI6o2rM0SSNEu8IK44xxtTHILJ2pF8US4BXB5TLNI+RaMU+fY9XAq8MKJdpL4PIwhl/BjphQtVsalHUX5n6blRT/U2p7Q9SupxQPyjq48gu8t1OMp68E11yjekkpiMr74Nkx4y1wGWo6kqn/5amA78iKf/jqARwJ5Auw/aFsOIYY0xjDADnkR8XdRad/9IwpZkO/JHsvV1OY+WVTHdyP8nn4PlhxWkJtSjqI8nW6j4U+ENq2xll+ugHRR20eBs/x3SZOmNMPlFyuLza5wvRPGurUMLVyDHIGyt+DpcAG4YUKsYmwMMUZbscJ6U0xvQAA8BXyHdp/A0wJZxopk4OREmb0vfzWeClAeUy4bic5LNwYlhxWkItijrAl1Lfv5aky+Q6yk+i+0FR34LsQl8n1Ww2ptOIksOlq0tE7SZUfaVbwkMmocWG+Dk8g86hUxgP/IuifLfgMmzGmB7jPSjpRvql8gCKqTKdz2jgk2TdnIeRRXX3cKKZwJxB8nn4flhxWkKtivou5E+ko/aHCvv3g6J+LMnz+0tYcYzpWLZDFvJFZMeFqPZ5t72DX0p2XL0a2CykUClGAL+mKN9jwOZBJTLGmBZxEFl30KhdjLIlm85kD7SKnHfvrqDzk9OY1vJSks/EAnovtKVWRR2ytX/j7U0V9u0HRf1Ckuf3ubDiGNNRjAFeD/yJbC6HYeAO4GS6r+LBWLToEPcweg4luuu098Y5JGXcJ6w4xhjTWjZFdYXzJq4LgNeEE83kMA69UPMyyDrXgIkYjdwV48/HQUElaj71KOrvJn+sW0blhHu9rqiPBRaTPL9ZQSUypjPYFr1bF5IdA9YC85AnYqlElJ3M3sCdJM/pBmD7kEKV4ESSoUqvCiuOMca0hzEofjNP+RsGrkIWXBOOQeB/ySoncSXlFcGkM53IT0k+Iz8IK07TqUdRn4qqIKR/P9+tYt9eV9TfQPLcnsDJmUz/Mhr95v9AvvX8YRRitHEoARtkJLKYr6J4TmvQgsSogHKV4hCS4ZofCCuOMca0n3Lu1EPIHf55waTrX2YD/6a0y67DFEweh5N8TpYD04JK1FzqUdQB3ogmqPG2QxX79bqi/jeS5/alsOIYE4TNgLnkW8/XoZjtOXR3ksUdgX+QPLf/oKz1nciOwNMUZf1eWHGMMSYco9FLKr7KGm+rga/RWclFepXZZCfP8XY/zupuSjNIVpn9TFCJmku9inq99LKi/mKyC7M7BpXImPYxiN63F5PvWbgAWZq3DiVgkxhA2dufJflbPx9YL6Bc5dgQuJeivH9G81RjjOlrdiFbYzitsP8UJ/JoNmOAE0iWHkm3FcDncTkSU5n3knx2lgAzgkrUPKyoN4cB4DqS53V5UImMaQ+bIo+avJCyuPW8E13Ba2ULsnO6+WiRrlMZC1xPUd67cAlhY4xJMJvymZKHUZ3Q4+huV7DQTEMThkcofZ3XoRX/bl/VN+1jPRRrHH+OvhNUouZhRb05vJ7seR0YVCJjWsco4GjgSpJZzuPW88/SW+/ZOWTLyF1MZyu9A8CPKcr7FA69NMaYXEagydx9lFfYH0aW3t3CiNl1jEFZSy9FVvJy1/aXwE5hxDRdzqlkF3wODipRc7Ci3jiT0bgdP6dfB5XImNawA/AFSsee/w44ht6wnkdMA35O8lyfAF4dUqgqmUvSg7OTLf/GGNMRjALeimqFllMqh4HbkIV4iyCSdi4jgBeimLB0KaS8di9wVBBJTa8wimz5nfuoXI6s07Gi3jgXkDyfVdhqZXqHsciafDX5mdsXA+cC24QSsIUchsbE+PleAWwSUqgqeS3J+3VcWHGMMaa7GAAORWXb8l5+6ZXqvwEfR/U6+7HG9wRkOf8WpcurlWurUPkoT6BNvbyI7G/153Rn3d8IK+qN8Q6y5/OpoBIZ0xz2QYvhS8ifk1wJvIbeTEo2AZ17fLxfgpLIdQMHkPQw7KUEqMYY03Z2Qi+FZVSndC5EcUfHAtMDyNsOBoBdgQ+i5C2lMuin2z3ITXk3tMqfrve8DpiHFjyMqZWvkn3mPhFUosZ4J8kSa+9q8fF2IFvWbf8WH7NVzCI7Lv2b3lRcTH8wGSmjpUrMPkJvZG4vx/4kM6QPo2Rs24YUqga2IhmacAn9adwxxpimMxY4EiUoWU311uLHkPJ5Gpo8jmu34E1gEkq6Nxedy1NUf/6L0ELHLLLWzS2Rwr48Z7/r0PU2plrWIzuJHUKJlUz/sBXwJMnnYCmwc0CZjKmHEejdeT7578lV6J08B5Vg61VGoflHvLTcCjSv6pbzngjcSlH+m+jcknHGGNPVTAPeTfka4OXcvG8EvodeMq9G9Xw7wdIzBdgPOB44EyWBe5Daz/FZ4CLg5VSXKX964XjP5PR1baGfbnZhNu1jC7LJlJahya7pfWYAt5P11HEeDNNNbAmcATxA/jv2dlSacsNQAraRXYB/kjz/W4HnhxSqRgZRSci498OmQSUyxpg+YRPgzciF6WlqV2qjtga5dP0G+D6KWzoFeB3KYL0jeinXY5GfXJBzD6T0Hg98CPgycCFShtMlrmptdwHnoNj+sXXICIo9O5VsgpjoxewSeaYaDiAbVrEceYWY3mUjskr6MPDhkEIZUyVRYrh5JC3HUXsOefTNpj8Wrkeg+UB8LF+D3Ps7wbBRC18n6d3jykHGGBOAkcBBqEbpjdTmIl+PYr8YuZbfH2sPF7bnWaeb2RYBl6E42mbHxI1BSvk9Ocd9AL28610MMP3BCWSfnZU4nKJX2Zz88eJS+kOpMd3LnigELF0HPGo3odj0CaEEDMDWwJ/JvvsPCilUnbyX4jmsxe8gY4zpGEahl/CpqEzQHVTOIt+JbU1B9gvQhGFn2pMAZRRS2P+TI9MjwPvo/hJcpnWcTr6y3g01dk31bEd+iM7leEHPdCabAx8lf3FpGOW2+TxKZttvHEcyee8QitHvxnf94SS9I94dVhxjjDGVmA4cggbsr6P6p/PpDAV+JaoPfymKGT8elYEJPdkdQKvQeTkBliBXuA2CSWc6mXeQ/W0NoWemW5IQmdK8gvyQo3nIM8eYTmEMRdf2NeS/f6PEcKMCyRiSGchLL35NFgBHhBSqAXYm6dn47bDiGGOMaYT1UAz5q4C3o7JSX0O1oK9FMeCLyc/8Wqk9jV54twK/Q+Xjvozi1I9HceUz6Q7FZRaazKTP8VnkPrhZONFMh3ISSiiWfmauBKYGlMvUzwBKxpl3X/8f/anomM4kcm0vVS3lDvQsTwslYAcwh+z1uZjuXYDfGPgvxXO5CufXMcaYvmIkytI+FSnZUdsMWemjF8SbQgnYYvYFfkXWWroSrVx3S11V0x7+l/ycEfejBTLTPWwIXEG+0vM9umPB0fQ2WwMfRwvsec/pAuCLKKN5PzMJubWnDQvdPG8ZB/yd5ELM5KASGWOM6SheQfElcSu9nUxpZxQ7n3YlXIcs73uHE810GAei2M/0pDnKJGxX6c7nSPKrQqxBVkljQjEZxVdfTX4oW7zmuT0+4GUo+W38Gv2W7i5bNgD8lOL5PAlsE1QiY4wxHccAWsWNXhb9UJZqa+AbwAqy8ci/wXW0jZgG/JF8K9e9wIuCSWbKsRHKnZF3354AXhxONNPHjEHJKX9BtiRk1G5COWm61Y272YxDC6PxsJXnUNLdbjcqnEXxnFYA+4cVxxhjTKfyVpKxuP3CNGAuiudPT5iuQxa5bp8MmMYYjfI/5E2q16F4UrsqdgaDKMdAXsK46De9cTDpTL8SxZ0/Qf5z+XDh891DCdih7EM2HODvqHJDt3MCSQPBG4JKY4wxpqMZg+LgohfHbmHFaTsT0Ap9npvsrchF0cld+ptXkEz4E2+LkCv1uGDSmdnAv8i/P6uR9Wp0MOlMv7ElGhPuJf+ZfAaFYR2J8ySkGYmuXTxPyGq0qN4L1+ogkh4VHw8rjjHGmG7gDJJJlvqRccDJwENkJ1b3IWudY5P7l0nI8pWXPXwYeAQ9I17UaR/7A38m/34MA38FdgwmneknNgJOIZkcLL1gdBmKOw9dxrRT2Qm5/8ev2+3AC0IK1URmolj06Nx+hr32jDHGVMFUVLZsGK329rOL6Ag0mYrH7kdtIVrZt7tz//JiSlvKhoH/AG/GizqtZDYqY1TqHixDsb4jQglo+oIoKVypeufDFEuqTQ8kYzcwgBY54yVlh9DCaK+Mo1OBu0mG4vTKuRljjGkD36T4EvlMYFk6gQHkmvg3spOvJWgSsVEw6UxIxgGnUzoeehh4HC3qzAgjYs8xBi2A3Erpa74W+D6weSAZTe8zFr0XLiCpWMbbf1G4RS/EU7eaLckm7XyI3krWOQr4A8XzewAv3BhjjKmR51F0610EjA8rTkcxC1lN8mqxn48Vg35lKvB5lIm4lPK4EimPewWSsdvZDPgEWvgodY2HgV8h11ljms0g8uK4AFhK/vP3FHoXzMLuzNUyh2wy1wtQ3pheIl7/fQmwS1hxjDHGdCu/ovhCeVdgWTqR3dFEYi3Z+MMLcDxsv7I5yu1Qyv01ancCH0MlAk1pJgMnAn+idE6AqP0ZlzYyzWcEUrrPRSFPec9ePCmc651Xz3Tgl2TDyl4ZUqgW8WGK57gG1YQ3xhhj6uIgku5ZvZBltRXsAPyAZGbaYaRUXIpK8pj+Y0vgi2gCX065HAKuB96Jwyci1geOBn6O6gqXu35rgYuBA4JIanqVAfRMfRklh8x79pajJGCvxDHG9XA48BjJa3opsGFIoVrE0SQXGt8RVhxjjDG9QDxr7asDy9LpbIRiEfNiFV2LvX8Zj5Ij3Ul5hTNqd6DnaDb9VUZsJrpO86isnA8jt+Nzga0CyGp6l51RPon7KL0wdDVKHNdrbtntYiJJF/DIDfykkEK1kL1Jzgu+FFYcY4wxvcL/kFQ2TWU2RBO9RWQnefegOu2us91/jECWt18Bq6hOaX8auARZX/agt8q9bQMcixJXPkR112MYLR6+CytJpnlUUs7XofffqTjxV6McQPY6/57eze2yKUmPjN9g70RjjDFNYhC4n+JLxvGf1TMB+CD5bpOPoUzhU4JJZ0IyFXg7cC3ZpITl2vLCPl9EyZe2arPc9bIBcChwBnA5yfrB1bR7kSL1vDbLbXqTyK39Kygreynl/C+otN8mYcTsKcYiT6G4+/cKVK6uV0snTiRZmeIWnJjXGGNMk3kPxRfNJYFl6UZGIzfJf5OdDC4DzqF7FC7TfLYCPoIsxZWSpeW1Z9EE8GfAJ5GVei+UhK2djEUJFF+FJt/fQ1bIWpXyqN2Hfhv7tfMkTM8ygqJy/jBWztvJrsC/SF7rfwO7hRSqxQwCl5FcnO9VrwFjjKkZxwI3jwnI6jAZTWS2R1Z2UzuzkBLzCpLP6BBwBXAmqtVu+pMNgUOAw5AFulE325XAE2iS+CTKpryg8P9VKAHicvS7XlrY5xlk9RmJQjTGosWm9dHkc1qhbYzqwk9DSk2jrugrgGuAK4GrkBXdmEYYRF5gc4BjkBtymiE05l5SaI+1TbreZyTwfuBTFPNtrEUx2meg8adXOQ84ufD/54AXAzeGE8cYY0wvczbFleGvBpalF3g+KuWTzhQ/jJSVo3EcW78zAlnGTwN+ATxKfZbpTm1LgT+gxalDcd4G0xzGAkehShxPYct5KGaiMJ34db8fODCkUG0i7oW4jt4sNWeMMaaD2JRiAqzlKObUNM5mwOfJL+H1EIpxdxy7idgcWQa/gBSNpwmvcFfTVgL/RJme34ySd/VqXKppPxOB16EyfcvIfwbXokVQK+etZQBlb3+W4rUfQr/99QPK1S4OR89adO7vDSuOMcaYfuHHFF8+Hw4sS68xEXgfMJ/sBHM58C2k3BiTZjrwQuAtaNHn18BdyN2yncr4ahQiczXwdeAUZCnfGivlpvlsCJyIkhOuJP+ZXIVCit6Ks7W3g43Q/Yjfg8eAl4cUqo3sQnLR/TthxTHGmM7FMerNZzeUEGYAxbpuhSZIpnmMRMm4TgEOSn02jFyFv4pKvAy1VzTThYxHseTTKcaST0MeMWNIxp5PLOwzGVnD1qK48ZUkY9kXoxj3g1HJuIkoidwT7Tgh09dsi9zaj0TjY1540HKU5+AXSElf0jbp+ps5aEF5amzbJai6xeIgErWXjYEbKCaM+y1wBBpHjTHGmLbwe4qrxccHlqXX2R25Cy4naym6H8UuTy25tzGtJT4WuGyjaQUjgD1Rab6bKO3NsRi5vR+Hy1+1m8kkve2GUUjOG0IK1WbGISU9Ov87aH/VDWOMMYbDKb6MbsOeC+1gOvAx8pOJLUVuxrsGk870K/GESZ8NLIvpHSagPAw/pHxpv0eBbwCzkSeSaT+HAI+QvC9X0V85AEYAv6R4/k8C2wSVyBhjTN8yQLIe+CFhxekrBpHL59XkT1xvQkl8+iFhjwnPTIrP3r8Dy2K6mxnIGj6P0vHmkaXyLFTm0nkPwjEOOBeFX8VzqZxK/y3ef4niNVgB7BdWHGOMMf3Omym+mH4bWJZ+ZW/gQoqZ+NNuoOcAOwWTzvQLd1J87rYKK4rpIkYiZftMVA2glGK+Cr1jTga2DCKpSbMfcA/J+/RX4HkhhQrEiSQz278+rDjGGGOMklA9RvEF9fyw4vQ1M1Cs+n2Ut7KvF0pA09N8nuKz9q7AspjOZhpKOHYBWkwspZwvohhvPimIpCaPUShXQLz02OrCtrykfr3OIcAaitfi9LDiGGOMMUU+RvEF9YPAshi5gR4C/Jzk5CFqTwFfBLYLJaDpSQ6m+IxdEVgW01kMoiSDn0YLhnE36XS7Cy36vJD+VPo6nZ2BW0jes9tQ1Yd+ZEeUMC+6Ft8PK44xxhiTZCoq4RStqm8WVhwTYxO0kPIQ2QnxEPAnlLHf2ZFNowyiRaBhFFs8Iaw4JjDTgDcCP6H4XOS159DCzsk48VYnM4DizuN5A9ah+PQxAeUKyYYkPdj+jMpcGmOMMR3F1ym+rM4MLIvJMgJlRL6YfCv7isJnR+KsyaZ+LqL4TL0qsCymvYwkWT5tHaWV8wdQuck5eEGnG9gKuIbkPXwQeT30K2NRPH50Pe4EpgSVyBhjjCnBTIrxaouxhbaT2QxNph8mfxL9X7TYskMg+Uz3cizF5+g7gWUxrWUA2A14PyrDtZzSivkKlAjuVBxy020ch0p/xu/nBfT3O34ALWxH18Nl2IwxxnQ8v6D44np3YFlMZQZRLPuFlJ5k/x14JwpvMKYSUyh6bDxG/5Vn6nU2Bt6EFLUFlFbMh4F7kafVETiBZTcyHfg1yXv6OPK66nfOIlmN4EVBpTHGGGOq4ECSro1OBNQ9TESl9v5MfqKnlcClwFE4Bs+U5xqKz83eYUUxDTIZ/ebPAW6lvGK+GI0RJwFbhxDWNI1jkJU4fn8vQTHZ/c5JJPO8HBtWHGOMMaZ6/kbxJfaawLKY+tgclXlL18eN2tPIonYk/ZtEyJTmgxSflblhRTE1sj7KZXEWcB1KDlpKMV+LYtHPKuwzKoC8prlMQrkD4vf5GaScGjiUZI6Xj4UVxxhjjKmNORRfYjcGlsU0zp4oq2+prM2R0j4Hu7casQPF5+PmwLKY8owDZqGFuauRG285q/n9FJPAua55bzEb5SeJ3++rcRWXiJ3RokV0bVyGzRhjTNcxSLJcyQFhxTFNYhzwehSzGC/Pk7a8XIBcZW1p728ib4whPNHvJMYDLwM+icJcSv2W45m9v4/i0n0fe5OxyCsinqX/ObR4MyKgXJ3ExsB8itfnGhwCZowxpks5heIL7eeBZTHNZz3k9n4B8Cz5E/zlwDyUMXj9MGKagHyF4rNgt9lwTEe/1ciVvZLFfAHKZn0SsiCa3mZvVFYs/gzcAGwfUqgOYzxwC8Xrcwcuw2aMMaaLWZ+iq/Q6YNuw4pgWMgEl0/klKsGUN/lfihZsTgQ2CSOmaTMvpXj/LwssSz+xDXA88F3gLsor5VEW758Bb8Nl0/qJkchiHl+4WYMWdJxroMggycz3C4Atg0pkjDHGNIHPUXy5fS2wLKY9TEDu8T9HrpN5isEQilv+DLA/rgzQq4xC+QsiN1rnL2g+UXz5+1G29ceorJg/BPwYKeY7tV1i0wnsCPyD5HNxB8pHYpKcRzIcYL+w4hhjjDHNYROKq/XLcVmXfmM88D+opM8ySisOT6I67scCGwSR1LSKiyne51cElqUX2AZ4A/BVlKizXEb2yJvpVlTL/FgcY97vDKCQhni40hBKDuiFtCwfIPlbelVYcYwxxpjm8kOKL7qPhhXFBGQMSmD1FeBuSisWa4HrgY8Ae6CJpelejqN4b78ZWJZuYxLwEvRbuAxYSGVr+UoUh34WcASOozVFtgD+QPJ5mQ+8OKRQHcwxJJPrnRJWHGOMMab57IJW7IfRRHNsWHFMhzATWXbmUTqufRh4AlllT0VumVbcu4sN0OLLMPAovn+lGI9c2E9FCRrvIKkklGqPod/QXFRaa1yb5TbdwRxgEcln52K8kFOKvZAXYHStzgkrjjHGGNM6fkfxhfe/gWUxncf6qJzbt8jW8E23h4EfoWRZM0MIa2rmeor3b/fAsnQC44GDgPeikI87qU4pfw64FvgCcDSwabsFN13HNJQvJL34+eqQQnU4W6MEi9H1uhznUTHGGNPDHErxpXcbtqqZ8uyKshH/ico1nh9FGavfjZRAT6g6jw9TvF8fCyxLu9kClUb7KLJg3k11Svk6ZFW/AD3be+NM3KY2DkPjY/y5ugJX3SjHVJKVEm7CpUWNMcb0Af+i+PI7LLAspnsYh8p8fRb4KyofVE7BWYImo58ADsfJ6TqBXSnen78HlqVVjEGhGW9GbrJ/AhZTWSGPlPI7kXX9PcjaPr694pseYgJKDpceF08KKVQXMAr4PcVr9iCwUVCJjDHGmDZxAsUX4NVhRTFdzASUPfwLSHGvlPV6GLgXuAjF/+6PsxuH4EGKSmk3T37HAS9AGdQ/C/wCWckrLSBFbS2y2P0EeB9wMHqmjWkG+6PxLv7MXQ9sG1KoLmAAea/EFzZ2DSqRMcYY00ZGofji6EW4R1hxTI+wHvAi4OPAVcBSqlOW7kDK+wdQEq6pbZa73/ga3ZWnYiqwL5L1bBSnej/Vua1H7RkUU/414K3APniRyLSGMSjTf5S4cRgl6TwNGBFQrm7hUxSv22pUocQYY4zpKz5C8WV4QWBZTG8yCDwfuXl+D+VEiE9ey7X5SCE7GyWr2wu7IDeLwyhe558HliViErrHr0MLPT9GrvlPUb0yHi383IvO6xOo1vLWbTwP09/sAvyT5DN5KxoHTWVeT7EyzTDy/jPGGNNmnMAsPFNQVu/xyFV0G2RlN6aVjEcK2T4oKdceKGN8NWPCMPAQiiH+D1LI7iu0yEPEVGYM8CRy834W2BBY1eJjjkbJ3LYCtoz9uw1yBZ5eY39r0H3/D3oe7kBu7HehpIfGtJMRKNHg59GzDlo0+hJwBrIMm/K8DPgNxUSNn0bXzhhjTJuxot4ZnAecXPj/2cDpAWUx/ctEZHHaA2WL3wPYmdoya6+kqLTfhxT6+YV//4vc8E2RXyJrM6gSxO8a6GsAmIEyWG9aaJshZTxqG1Of2+/TFBdk7iK5SLOmAZmNaRZbAz8EXhjb9gCyBl8bQJ5uZFfgL8Dkwt//D1nXvfhqjDEBsKLeGWyNJryDSJHZHCs0pjMYBWwH7FRoOxf+3Y76SmM9gxT2+YW2EHgE1TF+tPD3QvpnYngi8N3C/88DTsn5znhk6Z6OrO4bouRzkUK+MRozZtBYubKlaByKFPJ7gXsK/3+qgX6NaTXHAV+nGJYzDHwHeD/yVjGV2Qz4W+FfgGtQeE6rvXyMMcaUwIp653ApcEzh/+8Bzg0oizGVGIVcpXco/Pu8wr/booleI2PLWqS4P45cwxcX2qKc/y8Flhf+XYLiKjuRMUiJmFxoUwr/bo7ccgeAZcCvkSIeV8zHNkmG55BnQ9Tmx/59EF1zY7qJGUghPzK27XGUqPDyIBJ1J5OQ10GU1f0OYBZaWDXGGBMIK+qdw97A/2/vvsMlq6qEjb+3E53obkLT0CRBkkjOgooiICqooKCDiiMOplFhMGAaZUQdBJVBx4BpFDNmUcAhiAqCBAkDAgqCIE2Tmm5C5+77/bGqvjqh6saqu2+den/Pc55bt+I651Q46+y9176mdvkeIvFZXbjPPGAS0fIojVfTaCTtWxA9RrbILHM7+NrLiBa0J4iDzDVEAg+R0K8kkvnidUMxnUi46+bQ+A6dSYyJnVP7O6NwXactI3omPEDUCVhY+/9+Gsm4ibiq5Gjgi8AGmet+CLyVOJGnoZlKDLl5Tu3/+4kp7ayVI0mJmaiPL1cC+9cuvwo4r3D7u4AbgUvHMiipzaYR46U3I7pt15d5RFfujWp/e726/FKiy/nC2t+Ha38fpNHj4B/AAmIMudQLZhPF4t6UuW4xUUTu20ki6l4TiHHor6z9/ziRsN+cLCJJ0v9noj6+HEVjmqbriFb2rJuIFoQvjWVQUiLTibm7N8j83TBzeQ5RMX0O0YI9o/D/OuWnHDNLibGdS4gW+ydq1y3OLEtqf2cTU6FBnIh7GZGQLx3bkKVx7xDg6zTGUUO0Bh+PPc1G4ixiqB3E99SLsSFAksYNE/XxZQJwB9FlGOLM9hW1y/Wu8WcBJ499aFLXmUQk7hDJ8AQieZ9eu269AR47lWj5hxiC8kTmtid4zR9EAAAgAElEQVRoDEtZRaNYVTYpH44+osDeZkRX/Y2xeJuUNQ34CPAeGrMWLAPeD3yW3ik+2U7vBs6sXe4nCvLZI0GSpAG8nfjR7Cembqr7Qu2681MEJamjvkTjc/+6xLFI48m+xJSA/ZnlKmLmCY3Mq4iTgvXt6cl/SZKGYDrRmtZPFL3agWjdW1K77q/pQpPUIUfQOGj+fuJYpPFgEnAK0Uul/tlYCZxKTGWqkTkQWE5jm34hbTiSJI1PbwNeQ6OLbd3Hyf+Ivjrz/yriAEZSdUwjKtD3E+PWp6QNR0pqR6JOS7YV/RZgj5RBVcAziWkt69v0PBpDCSRJUsYM4HqitfxLwH616+cR4+/6iYP3y4kxsfUf122KTySp651P4zN+UOJYpBT6iGru9ZNW9Z5lZ5O2OGQVzCemaaxv198RvfUkSVILmxJzH9d/PO8A3kt0f80eqGRbFl6UJFJJnfQWGp/xTyeORRprWwKXkf+tuxt4XsKYqmIWMaNEfbveSsyeIUmSBrEHMRVTNinPFnopLu9ME6akDppP4/N/Z+JYpLF0NPku2f3AuTRmbdDITSamsKtv1/uJkyKSJGmIjiQO0rOt58WW9HoC/7lEMUrqrD/R+KxvnzgWqdM2ImY3yf7GPQi8LGVQFdJHnPCob9vHgd2SRiRJUpd6L61b0bOJ+q9TBSipoz5K47P+rsSxSJ30YmAB+d+3HwEbpgyqYs4kXzH/0LThSJLU3c5h8GT9nlTBSeqofWh8zn+TOBapE2ZR/p1bQhSRU/tka16sBV6fNhxJkrrfZKKgTrNu79lWdSvgStUzgUZxyVXAemnDkdrqAKL+Qvb37BJg85RBVdDR5OvcvC9tOJIkVcf6xMFMdkq24rJDsugkddLXaHzOX504FqkdpgKnk08elwGn4Dze7fZ8YDmN7fyltOFIklQ9Tyeq4LZK1l+aLjRJHXQkjc/5txPHIo3WLuSnBusHrsGTzZ2wF1Ewrr6dfwZMTBqRJEkV9VyiAEyzqdosNCVV0wyitbEfeBSYlDYcaUQmES3mK2j8bq0iWtanJIyrqrYBFpKvcTE1aUSSJFXcGygn6WuxO5tUZRfR+Lw/J3Es0nBtDfye/O/WbcDeKYOqsPnA3TS29U3AnKQRSZLUI7JTrNSX3yWNSFInvZ3GZ/2TiWORhqqPqN7+JPkTy+cQPUXUfrPJDy24E9g4aUSSJPWQCcDPKU9nI6matqDxWb81cSzSUGwM/JL879QCYr50dcY08j0X7ge2ShqRJEk9aDrRdTB7EGQLhVRd/0fjs75N4likgRxN1FPI/j6dR8xgos6YCPyExvZeDOyWNCJJknrYZsSPcf2Hede04UjqoP+k8Vl/Z+JYpGbmAN8in6A/BrwmZVA9oA/4Oo1tvhRrWUhS5fSlDkDDtgdwBdHl7ZXAjzO3zQDmEV0Q59b+zqtdngbMqt1vNtGdfgb56ruP1f4+SVTnXUEcADwCPAQ8DDyQufxQW9dMUtYBxGcd4GLg0ISxSEUvBL4GbJq57tfA8USXd3XOp2jM/LIGOIZoXZckSQmtC7yHKNBzCfA94Hryc6eO1bKKKFxzAXAW8BbgIKLlX9LoTCROiPUTJ83WTRuOBMRJ37OJ36D6b8FTwIl48n8svI98ob43pg1HkqTeNBN4HjEX7U+B+xj7ZHykyxNEa+CnifGLW7R300g94Vwan6lXJI5F2g/4C/nv+j9gDYWx8jryJ0jekzYcSZJ6xzzgOODLxDyoq0mfcLdzWUCccDiF6MJv64s0sGNofH7+J3Es6l2TgVPJ/yatrF03MVlUveUIohdbfft/Nm04kqROM1FKaxLwLOCw2rI7o9snK2iMI3+wdnkBMZb8SaKVGxoF6Z4iDrbq5tRefyZxYLZO7fJcYCPyY943Ica6j8aDxJjGC4kxuI+O8vmkqplFfI6n1P5uTLSoSWPlmUTBuN0z191CtO7emCSi3vMsYqjb9Nr/3yZO6vcni0iSpAqaSnRhPY8o3jbcVumVwO3EvOpnACcABxIJ9FibTkwHcwzwIeJg7hpGtl6rgauB9wKbj+VKSOPcpTQ+J/sljkW9o48Yd76cxvtvDTE+fZ2EcfWanYFFNPbB+cRJfkmS1AYTiGT6KwwviV0L/Bn4BvBWokWjW36gNyNOSJwJ/I5o0R/qeq8BfkMUyRltq73U7f6NxmfjY4ljUW94GnA5+e/lvwHPTRdST9qa6BVX3wdXEbO1SJKkUXoaMRfyvQw9Qf0j8B/EVExVSlInEa3vbyWmlFvC0LbJMuCHxFRADtVQL3o6jc+DXY3VacdRnkXkXGIYlMbOxsBfaeyDm4H1kkYkSVIF7Ekc2GQLv7RaHia6wb+JGPfdKyYS2+lU4DrylWxbLX8humJOLz+dVGm30/gcPC1tKKqoecSQqux37kKiiJnG1hzgTzT2w704JEySpBGbDBxLjNEeLOFcCPwXsC/RLV5xkuKdxDj1oZzc+DgwP0mk0tg7k8b7/22JY1H1vIL4Xs1+z/4Q2CBlUD1qFvnjiAeBbZNGJElSl5pAzBN+JwMnl0uJlvMjiKRerW1JTOF2BwNv05XAOUQXQanKnkfjff+rFvfZfsyiUVXMJr5Ds9+ri4keXhp708jXBlhMTGUqSZKG6UXEmNGBksmbgOOxAMxIPQv4LpGUt9rGjwMfxjGUqq6JwCPE+3055ff6HOBrYx2UutrBlOunXEwUAtXYm0KchKvviyeBZyeNSJKkLrQPUZV8oKJw5wMvSBVgBW0GnE7MtT7QkIK3Y48FVdN3abzXX1a47QzgF2MekbrRNOK7dA35Hl+n4FCsVCYSPe6y++N5KQOSJKnbrAt8jvwBTnZZAXwB2C5VgD1gBlE5/m5aJ+w3YHdBVc9raLzHv5y5fnOilf2CFEGpq+wN3Eb++/KPOGwipQnAd8gfR7wkaUSSJHWZw4B7aJ4YriXOhm+TKrgeNJkYR/kAzffJKuBsHHKg7jOB5l1e16cxk8QCGtMV/k/tuovGJDp1o0lEi3l2CNEqomXdHkjp9AFforFPVgPHJI1IkqQusgHwLVq33l5IzA+uNNYl5p1/gub75y/Agcmik0bmzcAfgOcUrv8tjff2XsDONHr4XDyWAaprPAO4lvz34q3E9JhK6wzyJ/zfmDYcSZK6x77AfTRPAP+KY9DHk3nE3PWtagZ8mEYLpDTe9RFJ+VrgxzSG07yXxvv6VKL41Ora/S4b8yg1nvURvY6eJJ8MngNMTxiXwmnk98tb0oYjSVL3OI4o6GKX6u4y0BCFXwHrJYtMGp7taXRVXg18kWhhr7+fi9MX/jZNmBqHtgAuJf/+uAd4fsKY1HAS+X3z3rThSJLUHdYhCjU1S/RuJorxaHybTrmqcbYnxM7pQpOG5UPkW92Wkp/5YHXm8hWJYtT4cjSwiPz33nl4knK8eBv5ffORtOFIktQd1geupnmxuDOw6E63eT7wIOX9+STwooRxSUM1hRhPnE3IWy1XJ4pR48NcYphE9j3xEHBkyqCUcxz5E8j/lTYcSZK6w1zgRsoHv08QLRTqTpsSRbmK+3UFcFTCuKSh2o84WThYon5dqgCV3GHA/eTfDxcAm6QMSjlH0Zi1oR/4GtZNkSRpUPOIbu3NKobvlDAutcc6RF2B4v5dDbw2YVzSUH2WwZP1PyWLTqmsSxSHy74PlhBF5DR+HAosp7GPziWmYZQkSQPYjHJRpn6iEM+shHGp/d5Kedy6ybq6wQzg7wzcBf6mZNEphWcRNTey74ErgaenDEolBwHLaOyjnxLz2kuSpAHMpHlL+oXAtIRxqXOOJd/9sJ9I3l+cMihpCF7EwC3qt6YLTWNoKlEsM3vSZhlwCrbSjjfPIobP1ffTr4i6E5IkaQB9RCXc4sHuL4kDIVXXMTSmvcp2F90xZVDSEHyf5rMZ9AO3J4xLY2Mn4Aby+/1mYNeUQampPchX378Cp3WVJGlIPkr5QPfHdF9l91cycCtbu6aT+49BXqfbeiAcRbkb8W3A7JRBSYPYkDj4bzX1oKppItFivoLG/l5FtKzbQjv+7AU8RmNfXUX04JMkSYN4BeXCTNfSfckmDJ6o/3cbXqMPuGuQ1+nGbXcy5fX4FXYf1fj2Bpp/Bu9KGZQ6Zivgd5T39bNTBqWWdgMeIV/kcf2kEUmS1CU2Jd8drR9YCGyeMqhRGCxRf5Soej4aBw3yGt2aqAN8hfK6nJQ0ImlgfcAllFvV/54yKLVdH1G9PTvGeS1R5d3W2fFpV/JJ+g2YpEuSNGQXkj+4XQ7snzSi0RksUe9n9PPAf2sIr9Gtifo6RLfE7Lo8BWybMihpEFsBS8m/bxfUbpsGrEeclNwa2B7Ys7YcXFv2qf2/Y+0+W9QeYwI4PswDfkF+/z4AvCRlUBrQLsDDNPbXjcAGSSOSJKmLNEtq35k0otFrtk6LC///ahTPP4tIXAd6/m5O1KF5L4sLk0YklfURCfXBwJuBi8m/Z9eQb30d6bIUuJs4gfVzotfJR4F3ELMjbIPTS3XS0eRbZfuJwqcmfePXDsSJlPr+uomoJyFJkoZgKvA38gc/v6H7xyM3S9S/TH4M/mpg/gif/18Kz30L8Icmr9nNiTrA6ymvk1O2KYU+ohX8WOA04AdEF9riCbOUywrgz8Sc0J8EjifG5prAj9xsolt7djs/Brw2ZVAa1Pbkk/TbiB4RkiRpiN5B/gBoJfED2+2aJervJaaCKV43EsXnOZlqJupQbqG8kUiapE6aRRQGOwU4n3z32W5bniS+M84GjiO61WtwhwD3kd+WvyZ6+2j8egZR4yZ7InujpBFJktRlpgD3kj8I+q+kEbVPq0S92BJ+6wiee1vyLfOrgI2pbqK+G+UCXY4JVbtNA14InEW0SneitXsR8A+iOvjtwHW15eLa8sfa/7fW7vP32mOK497bsdwLfJX4rprThu1XJdOIKday3ztLgRPxJOF4twNRF6K+324HNkkakSRJXeg15A8cl1KdrmmtEvVmY8v3GeZzf6Lw+J/Xrq9qog7wQ/LrdWnacFQR2xL1MC5gdMnwIuBq4JtEt/h3Ei3XC4nx6eu1IdaZwHZEK/+RwL8CpxJj1S8n34I43GUVMdXYB4A96O1kdF/gDvLb5ypi22t82xl4kMZ+u404iS1JkobpMvIHQ59LG05btUrUoVyt/fPDeN4JlHshHFm7rcqJ+u7k12stUTxLGq75wLuI8eXDTWgfAy4iWluPJ5LmuQO81kFEkcexMhvYmzgJehrwM/LjdIe63EUUqqvCMKShmkQMc1hJYzusJE6GTEwXloZoV/LDU25n5DVgJEnqaVuQ777dT0xJVBUDJeovKFy/mKEn1C8sPPYRGvOxVzlRh/L6fTRtOOoi04iq3ecTrcdDSVZXE13QzyXmzX4mIyty+cVRxt4O84EjiKTzYoZXAO9WIoGtcsvkjsD15Nf7FqJ3gca/3cgn6bdhd3dJkkbs7ZS7FlbJQIl6H+VK98cM8Xm/X3hcdkx/1RP1N5JftxvShqMu8ExitoWhdmu/B/gS8HJg3TbFMB7Hf69DnDA8k0hIh3ri4kdEL4Kq6CNOwmRPXKwliu6tM8DjNH7sTn7avD9T7ZNKkiR13IXkDwLfkzacthsoUYdoDc7eNpQ51WdTTjh2y9xe9UR9QyJZyB5QW31ZRX3AYUR17mKvnWbJ58XAvxGVonvV5sAJwE+AZQyetF9LdK+fnCLYNtmSmAo0u153AwemDErDsgfwKI39dyPOky5J0qhMIMZ6Zg+QqnaQPFiivhXlOdU3G+Q531p4vpsLt1c9UQe4kvz6HZ02HI0jk4hZFYZSsf064CRseWtmNvAGomBjcbaF4vIP4H1EobtucjRRADC7LufSvl4U6rw9ySfpN2CSLknSqO1A/gBpEdWrMjxYog7w20FuL/pj4f4nFW7vhUT9TPLr9+m04Wgc6CM+b8VK3c26tZ9GfP9oaDYF3k2cFBxo2y4kqtCP9xb2jYCfko/9QeBlKYPSsB0ALCF/4q0dMytIktTzXk7+QOnitOF0xFAS9TcUbr99gOfbsXDflZSrTfdCov4qhj9kQNW1P/B7Bm89P45ocdfIPRs4j/zwk2YnQ97E+KyS/mLy82v3E2PubYXtLgcCj9PYh9cD6yeNSJKkCjmZ/MHSeKiK3G5DSdRnEHMsZ++zb4vnK7Yk/6TJfXohUd+T/PrdmjYcJbI1Mfd5q4RxJfBdYqoytdc2xFSaxe+u7HI9rb/Lxtos4Bzy8S0hTiiouxxOvobCH4ihGpIkqU0+Sf6g6QNpw+mIoSTqAN8o3OcLTe4ziXJLULOumr2QqM8lv36L0oajMTaRONHXanqxVUSV981TBdhD5hBDCZ6k+b5YDZxFnJBM5QDgzkJcl+D7oxu9DFhOYz/+FmsKSJLUdv9N/sDp7WnD6YihJurPK9yn2Zzqhxfu8yDNx4L2QqK+DuWWU/WGnYCrad2KezGwc7LoeteGwOnkk6js8jfgkDGOaWotpmwxvGXEfPATxjgWjd6xxEm4+r68gOr9tkmSNC58lfyB3Alpw+mIoSbqfZRbfF5VuM+PCre3KqDWC4k6lKfcGo/jYdU+E4B/J07KNEsEL2f8dLPuZdsA36P5lHhrga8wNt9HuwA3FV7/Giwi2K3eTP6Eyy9wjntJkjqm2KL+jrThdMRQE3WAUwv3uyBz2waUW6p2bfE8vZCoTyW/fivShqMOm0W5Snd9eYwYZ1y1GSO63XNpXYH/RqK+QCdMIlrMV2RebxXRsj6lQ6+pznoP+RM/38WikJIkddTp5A/ePpg2nI4YTqL+NPItBmtozKn+zsJzXD/Aa/ZCor4R+fV7NG046qBdKPc2qS/nE1OHaXyaRnzPN6sQv4T2T4e2NeXq/7cBe7X5dTR2TiG/P8/BYQuSJHXcSZR/gKtmOIk6wGWF+55Su/5PhesH6n3QC4n63uTX7//ShqMOOZbmBeMWEtM7qjvsB/yZ8n5cA3yI0feG6CN6VWQL2q0lflNSFrHTyPVRnuXkv7HnjCRJY+Kl5H+EL0sbTkcMN1F/feG+dxCFsYrdvAea87cXEvV/otyyqmp5G83HOV8LbJEwLo3MTOAHNO8Z8WVG3kq6MfDLwvP9HTholPEqnYmUa9icnjQiSZJ6zLbkf4gXU70ubcNN1GcAjxfuX+zK+aNBXrMXEvXPkF+/T6YNR232XpondOfgOONu9yaaFwT8DsMfd3w0Mewl+zznAeu3K1iNuanAz8j3jHhX0ogkSepBfZQPsnZJGlH7DTdRB/hak8dkl8MHeXwvJOrXkF+/o9KGozY6lfL7dxnR20TV8HzgIUaerM8Bvl147GPEUAl1r5nA/9LYp6uBf0kakSRJPex88gdb708bTtuNJFF/bpPHZMfmNps7PavqifrG5Ivura1dp+73Ucrv3SexG3MVbQ/8g/L+/gED96x6YZPHXQjM72Sw6rj1gavID/F6ZdKIJEnqcW8mf8B1Xdpw2m4kiXof8Ncmj+sniusMpuqJ+lvIr9u1acNRm7yG8vv2CaL1VdX0NOAuyvv9tCb3nQ6cTb5uwVPAiVhgrNvNJwqCZk/OHZo0IkmSBtEL84SeD3yBRgvKnsT84Dcliyi9fuBconWx6NwxjmU8KnaF/HmSKNROuxMFxbIWAy8Crh77cEZsL+AFLW5bQ9RWWNuG13k2cECL25YBn23Da4yFe4gTMZcC22Su/yBwK/D92v/7Ed9922bucxVwHDF1n7rXDsCvaRSIXAS8hO763EuSVFm/Jt+aUjxg72YjaVEHmEcUXcourx3ia1a5RX0/8uu1BtgyaUQarY2B+yiPSd8nZVAjVJxysrgc3KbX+eMAr/Fwm15jLG0OLCC/Hk8R0zCeSn4e9pW16yYmiFPttQ/5WgX3As9IGpEkSco5mvwB2nJgs6QRtc9IE/XRqHKi/gvKY1PVvSZSntWgn+gG340GS9S/1YbX2H6Q1+jGRB3gWcR3f3ZdVhT+vwHYKVWAaqtDyM9wchewddKIJElSySTK4xTPSRpR+5iot8++lOfVblcLpdI4hfJ7tZun2hssUV9KVCwfjTMGeY1uTdQBXkfzdVpDjE9fJ11oaqPjgVU09u+1wNykEUmSpJZOIH9gtppqTNVmot4efeQrAvfjGMZutx3Rxb3YQ6KbuzQPlqj3M7rppiZR7iJepUQd4CzK6/RvSSNSO51C/oTrxcCspBFJkqQBTQZuJ39wdj2DT0U23pmot8fbKK/TIUkj0mgVp2Z8hO6fZq9Zon5D4f/fj+L5X1J4roXAA4Xruj1RnwLcTH6dbqP7fwt63UTgi+T36zdxv0qS1BUOp3yQ2+3zqpuoj95W5Mcy9gM/SRqRRusgyu/RY5NG1B7NEvV3kS+G1k+MMx+JHxWe50yiAFeVEnWIInJryK/X25NGpNGYCvyQ/P48G6fVkySpq/yE/I/5KlpPd9QNTNRHZwblFsnHsdJ7t/st+X16JdU4aG+WqP8TcEHhuo+N4LnXp1xsbReqmagDfJX8et2HY9S70frkC0auJnpISZKkLjMXeJD8Adqj5OfY7SbbE2Pyssu+HX7N1zV5zUkdfs1O6CPmUS4mPiekDEqj9mzK+7Qbp2JrplWifkzhun8w/LH47yg8xzW166uaqG9MTNGWXbfRjO/X2Nua/JC2pcCRSSOSJEmj8mLK3R5vAdZNGZTG3IcpJz3nUY2W1172HfL79KK04bRVq0R9CjEGP3v9cGssXF94/L/Wrq9qog7RPTq7bjemDUfDsC9RQ6G+7xYBz0kakSRJaotm0zZdRIx1U/W9nvJUbDcQXeHVveYQrWqjSVjHs1aJOsDnC9d/exjPu1PhsSuADWu3VTlR35Ly+P49kkakoXgl+c/5PcAzUgYkSZLa6+uUD3ovB2YmjEmd9y+Ue1Q8QnSjVHd7Pfn9+jdgQtKI2mugRH3vwvXDmVP9M4XH/iBzW5UTdYgp+7Lrd3racDSIE8l/f98AbJY0IkmS1HZTgT9SPvD9Pc67WlVvoZykrwQOTBmU2qZY+XkkRdXGs4ESdShPOzaUeguTKE/B9qLM7VVP1P+J/PrdmjYctTCJcq+RnwDTUwYlSZI6ZyPgJsoHv1fR6Pqpangf5e7uK4CXpwxKbdNHjFPN7t+9k0bUfoMl6u8u3HbFEJ7z5YXHPEC+OGTVE/U5xMm67DrOTxqRimYCvyK/j86mWr1lJElSE3OAqykfAN9H5yuoq/OmAv9Def8uB16aMC611/bk9+8iqlcYcLBEfR7lpHOHQZ7zZ4X7F7t+Vz1Rh/KUk1YOHz82JT+F5mqc816SpJ4ym+bzgy/HKXu62TaUuwP3E9MyVanImOBY8vv4f9OG0xGDJeoA5xduH6j7/0aUE/tiUa5eSNTPotpDJrrVPsACGvvlcfLDMiRJUo9YF7iU8oFwP/AlYFq60DQCRwCPUd6Xj+GY9Cr6EPn9/Om04XTEUBL1VxRuH2hO9ZML972qyX16IVE/nvw6fjdtOAJeRX6e+/uxIr8kST1tEtH1sziWuR+4EzgoXWgaojnAOTTfh7cxeFdgdaevkN/XVeweO5REfQqRTGfv06r3yI2F+72lyX16IVF/Pvl1/EPacHraBOI3OLs/rse6AZIkqebVwJOUD4rXEkmgVeHHpyOIFsRmvSK+i/OkV1mx4vvRacPpiKEk6gCfLdznO03us2fhPsuA9ZrcrxcS9WeSX8c/pw2nZ80Efkp+X5yHld0lSVLBzsBfaZ703Qu8LF1oKtiCmKqn2b5aRcy9q2q7gPx+f0nacDpiqIn6HpST8OKc6p8r3KdVd+9eSNS3pPz9rrG1NXAL+ZPip2Nld0mS1MJ04mChOPd2dkyn453TWZ/YP8tovn9upnpTdKk5E/W8Yrf2N2Vua9Y9/tAWz9MLifoW5NfxvrTh9JznAA/S2P5PAkcljUiSJHWN/YnukM2SwX7gYmDXZNH1nhnAKTQvFtdPVLI+nUhI1Bt+hF3fs4qF4q7M3HZ04baBCs71QqK+I/l1vC1tOD3lTeRnHriPGJYhSZI0ZFOBTxBdqZslh6uBb2Nl2k5an0jQF9L6pMmVlKeYUvV9jfz74G1pw+mI4STqzaZeqxdS/FXh+o8P8Jq9kKgfSH4dr04bTk9Yh6j3kt3uvwPmpgxKkiR1t52AX9A6UewHLgdejuPr2mU74PM0L/BXX/4O/DNu8171YfLvhzPThtMRw0nUAX5GOSGfR/lk40AzIfRCov7P5Nfx+0mjqb5NiWFj2W3+FewBJUmS2uTZwBUMnLDfSRQy2yBRjN1sIvBC4Hxa1wioJw7/RrTQqHe9lvz74sK04XTEcBP1Iwv3/Qfw/sJ1Vwzymr2QqH+K/Dp+Im04lfY88uPRVwLvSBmQJEmqrpeSr1bbbFkB/Bw4BpiWJsyusQfwGWABA2/TJ4HTgNlpwtQ4Uxxn/CjQlzSi9htuoj4ZeKhw/ycK/58wyGv2QqJ+Jfl1fEXacCqrOB79IeCgpBFJkqTKmwAcDlzCwMllP7AE+B/gEOzqV7cd8CEGLthXXB4jDvzs6i6I98ES8u+R3ZNG1H7DTdQBzmrymPqylPK0bUVVT9RnESdSs+u4edKIqmcG8D3y2/hKYH7KoCRJUu/ZHjibOAgeLNl8iqgYfyIxl2+vmAYcTFRmH2pyfj3wdeBxygd8O41t+BqnimOyP5I2nLYbSaK+c5PH1JdvDeE1q56oFyvg35E2nMrZhpgmM7uNz8GT1JIkKaF5wAcYXivxrcCniTlkNx37kDtmOjFX7ruBX9N63vNmvQ++TkyPV7c5MYwge7+VxLhShxX0tuMpJ11V6v4+kkQd4IYmj+sHXjCEx1Y9US9+l1SxCGEqR5CfQnMpcFzSiCRJkgr2ZGjjrovLfcCPgfcQUwjNGuvAR2ASMV74DcAXiSSh1ZR2zZahjuc/gqj0nn3sXcBh7V4hdeSE4u8AABAfSURBVI0NKXdjPjBpRO010kT9xCaPu4/Wc6dnVTlR35Tyd9N+SSOqhknAqeSLgN4L7J0wJkmSpAFNBA4Fvsrwk/b6cj9wGfAl4GTgJUT3wrGuer4x0Ur+RuCTwE+B2ynP3TyUZRnwv8BbGV6F/HWJYQarC8/3HaJHg3rPeeTfC79IG05bjTRRH40qJ+pnkl+3m9OGUwmbUy7O92uc+USSpGGpUpfQbtQH7Eq0AB8GHEC0RIzGYmAhUU13ITENzkPAIiKZXUq0OK4kxsevqV2eVnvtdWtx1QtMzQI2IpLeuUTxn41ql0cb613ARcQ0Wr+pxTZSuxHjHvfJXLeYaNX5HLB2FM+t7nIQcGnm/35iJoEb04TTVicRxeGyjiUKdXXKveSLqz1CfP673QbA3cR3Xt2/Al9IE04lHA58g0ZS3g+cAXyQ+K2RJEnqSrOIsemfIeY2Hkoxum5a7iCKV/0rsG2btlnWBKIKfLHY3BVYbK7XFFv0LksbTtvYot4+nye/XguxxsVINevq/iDRe0ySJKlyJhNj298GfJMoNDeSbuUplgXABUTV7cOA9du8bQbSqtjcGeRbz1RdL6b8njwqaUTtYaLeHrtQHpt+ctKIulezru6X4NAjSZLUYyYRrdGHA+8iuntfTiTGxXHanV4WE4XifgB8lOiCuxfjp+DdkUTBrOIJhNfhsI9ecBH5ff8QMWyjm5moj94k4Fry63QXY1/jowoOJ4ZC1LfjauLk7FCKFEqSpAGMdoyxxt5q4K+15ZeF2/qIsaMb1ZZNav/PJcacTyK6dk4l5rCdQRxQTSGKua0GnqCRhENMj7aESHAfJpKdB2qXl3dg/drpp0TLzmlEd/tJxDY5F3gL8A7gT8miU6e9mxivPrn2/1yiAOMriPe4etOpxAnFrJOJ2h0amqlEAdF30DjpuRB4DdUZZiJJkqQxsD3lFtY1RNLe7a2sau1Uyi3QH0kZ0CjZoj46RxGFJbPr8/2kEXWfHYmeVNlt+BviJKgkSZI0IkdTnnt9EfB27GFSRZOB68jv77XEsAj1lt2JmS6y74UHqEYF+7HQR7SgL6Ox/VYBH8Ku7pIkSWqDaURLa/aAsx+4DasUV9GWRAXq7L5+gigopt4wj/IJuuXA/imD6iJzgfPJb7+7iSlFJUmSpLbanOj6XuxKfD6R3Kk69icSs+x+fhDYNWVQGhPzgJspf85PSBlUFzmEqFGS3Xbn4gwakiRJ6rAXAreTPxB9EvgAVoKukn+mnKwtAvZJGJM6axNiWsvifv9MyqC6xFTgc+TH9C8CjkkZlCRJknrLFKJK+OPkD+jvJMa1O51bNXyGctL2GLBfyqDUEU8jpl1r1mPGMdUD2xf4M/ntdjnRC0mSJEkac5sQc9OvIX+Qeg3w3IRxqT36gE9TTt4eJyqCqxr2Ae6jvJ9/hr1kBjKZqN+xmnzBuFPx5IYkSZLGgf2Baykf6P8Y2C5hXGqPUynv27XA2Vj9v9sdByylvH/PIxJRNbcXcAv5bXYr5TnnJUmSpKT6iG7vxe6zK4lW93npQlMbnEI5mat38XXfdp91gC/TfJ9+D0/AtDKZ+CyspLG91hAnraYmjEuSJEka0BTgTcBDlKf4OpWY7k3d6c3ACsqJ3T3AC9KFpWHaCbiO5r0kzsBu263sAvyJcl0Oh/lIkiSpa6xPFCMrJnb3Et1tJ6QLTaOwJ5GYN2uJPQ/YIFlkGky9Nbg49V79RNrR6UIb16YA/0G5Ff1TeOJRkiRJXWoLYh7h7LRF9fGcL0kYl0ZuQ+ASmifrDwCvSBeaWtiNcmtwffkL0cqusgMoj0W/CzgwZVCSJElSu+wL/J5yknAhkUSou0wCzqRc8T9bMfwZyaJT3SbAF8lXJs8u3wdmJYtu/JpD1NbInmBcA/wXMD1hXJIkSVJHvBy4nfLY2B8COyaMSyOzH+UWx2xicx6wVbLoetdMopv747Tu+fDKZNGNb0cDCyj3ADogZVCSJElSp00iCs49QPPEzindukt9DG+zQnP9wDKi9d3x6503HXgP8CjN98Va4GvAeqkCHMfmE1NKFmetOB3nk5ckSVIPWRf4d+Ax8gfHq4CvAlumC00jsDNwBc0TxH7gKaIb9g6pAqyw+cDHgUdovf3vAA5OFeA4NhE4iSiol91ev8GThpIkSeph6wEfo3ygvAL4PLBputA0AgcDN9E6YVwLXAwcAfQlirEqdiPGUi+j9fZ+iOgGPyVRjOPZ/sD15LfXo8Ab8L0pSZIkAdE1+nSi5bWYsJ9DFMZSd5gIHE9Mx9cqgewnxre/G9gsTZhdaT3gBJoXZ8wujwHvx+JnzWxMfKcUiyGeB2yUMC5JkiRp3NoE+Czl+Z6fBP6TmKNd3WEqcDLwdwZOKtcAlxHJ/ewkkY5vU4lp735C83nQiy3Cn8DPSTOTgXcBS8hvs78CL0wYlyRJktQ1NidavVaSP6heApxGzOet7jAJeBVwFQMnmfXicz8G3khvD3tYn9hm36Bcx6HZcjvwVmBGgli7waHAbZRP/r0fi8VJkiRJw7YlkbCvIn+Qvbx2/ebpQtMI7Af8gPL+bLXcRAyJeB7RIlpVE4C9gA8BV9J67vPicjHwktrjVbYZcC7l7XY+FqyUJEmSRm0H4HuUx5WuIKrEW6G5u8wDTgSuYWgJaT9RcPAyomv3y+nuugXrEd2tPwz8EniYoW+HO2qPe/qYR909ZhGV8JdSPvFzYMK4JEmSpEp6OnA25bG6a4hWsr3ThaYR2p6Yi/1Ohp6s1pe/E0XAPgC8EtiFGNM9XkwmTiK9lJjj/Fyim/pahreeC4n3/T5jG37XmQy8idhexeJ6JxLDMCRJUoU4VYs0vmxBVAx/I/nK1v3ARUThud8niEsj1wfsCRwGvAjYl6ggP1xriYrzf6kt9wIPEq3WC4gpyx4mupePNt6NgLlED4GNa/9vSiTn2wNbMfIu+zcT7+ULiTnqRxtv1R1FfO6zvWvWAF8HPkjsc0mSVDEm6tL4tCHwduCdRHfirCuBTxJdi/vHOC6N3vrAIUTifhiRCLfTQ0RBsZXEtIBrgMdrty0m3jOzifHfM4i5yKcC02rLRozsREIri4FLiOT8IuD+Nj53le0DnAk8t3D9JcTJvJvGPCJJkiRJQCRU7ydaTovdhm8AjqXaxch6wXzgCKK43BWUxx9307IauJXoCn8i0ZPAgnDDsy3wI8rDCK4HXpAwLkmSJEkF04gW9nsoJ0f3E2OZndqtGqYQralvIcZvXwT8jXLBwdTLP4BLgS8CJxEtv06jNnJb0HzqxnuA1+IJD0mSeopd36XuMhn4J+B9wDMKt60gCpCdCfzfGMelzluHGKe8HVF8cBNiHPnGxFjy+rjydnyvL6Ix/v0Bojv9Q8BdNMbIP9GG11FMtfYeolhctmDgk8CniWEuyxLEJUmSEjJRl7rTBOBwonvxQYXb+omWzrOBC4gutOoNE4mEfRrRMj+jdt2s2u1ziO/9JcT74imiBXc5kQyuIJLzlWMadW+aTwxrOYE4CVO3nOil8HHg0QRxSZIkSWqD7Ymk/CnK3ZPvBE4hEjRJ6c0l6hEUaxGsILq+b5ouNEmSJEntNhf4EDFVVzFhXwx8CnhaquCkHrcJ0ZW9WYL+BaILvCRJkqSKmgIcDfyBcsK+Bri4drvV4qXOezrR46WYoK8kquM/PV1okiRJklLYk0gGVlFO2hcQXXC3ThadVF17EJ+91TRP0LdJF5okSZKk8WBL4AyiUFizVvYLgaOwlV0ajT7gxcBvKX/OlhJd3D0xJkmSJClnHeDVwGVE1e9iMvEA8J/YHVcajsnA64CbKX+mHgVOI6rxS5IkSdKAtiG6vi+knFysBa4g5naelipAaZzbmJhV4V6an/Q6FZidKjhJkiRJ3WsicDBwHuXxtP3AImJM7cFE116p19VrP6yk/Hn5C3AiMDVZdJIkSZIqZSvgY8D9lBOQfuDu2u07pgpQSmQm8Gaad29fS8ymcDiezJIkSZLUIZOAlwG/oHmrYT/wJ+BkYH6iGKWx8AxierXFlD8DS4DPAjski06SJElST1oPOI5oMWxWgG4NMZ79RGCDRDFK7TSbgd/ztxNj0+ekClCSJEmS6rYG/h24jeat7MuAHxKV5WclilEaiQnAocB3ifdx8b29inhvPy9RfJIkSZI0qL2As4jq1s2S9uXABcAJwLxEMUqD2Zaou9Cscns/cBfwYWDTVAFKkiRJ0nBNJFoivwk8TvNkZw3we2JM+9ZpwpT+v02BdxJDNpp1bX8C+AZwIBaHkyRJktTlpgEvJ5L2R2metPcDNwIfAXZNE6Z60GZEHYUriBNHzSq3/xb4Z6LCuyRJkiRVziTgBcDngPtonbTfBXyeqDK/bpJIVVWbAycBV9K85bwfuAf4KPD0NCFKkiRJUhp9wN7AJ2hdiK6fmArut8AHa/efkCJYdbWdiYrsf6B1cv534NPAs7BruyRJkiQBMe/0+4FraN4Nub48DHwfeAMW81JzM4nhFufQuiBcveX8U8C+mJxLkiRJ0oA2Ao4linctoHWi1Q/cQlSaPxIryfeyHYjChBcTswu0er/cDZwJ7IPJuSRJ6iIeuEgaT/qIrsuH1pbnAFMHuP9fiC7Ov6/9vb3TASqJrYn3woHEPOZbtbjfGuBqYlrAC4EbxiI4SZKkdjNRlzSeTQMOAA6uLXsw8PfW40R3+iuJ6t5XEC2u6i5bA88m9v0htE7MAR4BfgNcAvwCWNjx6CRJkjrMRF1SN5lPtKjuT7Sw7sTABeeWAX8iWlbryy3Aqo5GqeGYAewG7EUk589h4GENa4DrabSaX0cUjZMkSaoME3VJ3WwmsB+N1tcDiFb4gawmusxfn1luAJ7qXJiqmQJsC+yZWfauXd/KGuBGGr0kLgUWdTZMSZKktEzUJVXJFCL5O4BI3vcH5g7hcWuAvxIJ4W1EIv+X2nVPdCTSapsAPA3YDtgR2JXYLzsAEwd57HLgWuByGrUHPIkiSZJ6iom6pKrbkuhavXtm2XwYj19AI3GvL3cQFcV7vQv9TGD72vKMzOXtGLgIYF0/cCeNng3X1BbrCkiSpJ5moi6pF21IPnHfneiSPdB496LVRBJ/H3B/7fK9wAPAP2rLAmBl26IeW5OBjYEtgM1qy+a1ZdPa3/nDeL5+4C7yQw6uB5a0L2RJkqRqMFGXpDATeCbRGlxvFd629nf6KJ53IZGwPwwsBh6r/S1ezl63muhyv3oUrwvRzXxW7fJMYL1BlvVry2ZEkj6cExdZ9xG9Du4gpsz7MyblkiRJQ2aiLkkD6yNaj+tJezaJ34yhdfEerWU0uoOvpPmY7dk0Euv1xiCmJ4kW8npCXh/bf0ftNkmSJI2Qibokjc40YBNi7u/5TS7PJ1qnu+n7djnRC+CB2t+/tbgsSZKkDuimA0dJ6lbTiXHxcwrLek2umwOsW3tcX+3/ujk0vrdnEV3b1wCP165bASytXV5a+5/a7auIbvWLan9bLYuw0r0kSVJS/w+x+o8fQhlyGAAAAABJRU5ErkJggg==" width="65%" style="display: block; margin: auto;" /> --- ## 6. Direct effects estimated between network and outcome <img src="data:image/png;base64,iVBORw0KGgoAAAANSUhEUgAAA+oAAAJSCAYAAABUcD9+AAAABHNCSVQICAgIfAhkiAAAAAlwSFlzAAASdAAAEnQB3mYfeAAAABl0RVh0U29mdHdhcmUAd3d3Lmlua3NjYXBlLm9yZ5vuPBoAACAASURBVHic7N132Bxlufjx75uekIQSmqFIKEE6iAoKIoqUqKiIiAoiCEY9KtixnIJ61FiOisdygh7FYEEsIGBBRJCDgFKl9w6BQCAhQHry++N+97czs+Vtu/ts+X6ua65k992dvWdndnfuecrdhyRJgzcK2BzYDNgI2BTYBNi4/99NgUnABGAiMBqY2v/cdfuf/xywHFgBPAusBRYBa4AFwOPAo8Bjhf/fDyxp8vZJkiQl15c6AElSW1of2B54ATCzsIxPGNcjwO3AHf3L7f3LPUSiL0mS1PFM1CVJY4ikfE9gH2BfYAc66zfiGeCfwDX9y1+JFnhJkqSO00knYZKkxhhLJOMHAy8HXkh0Ve829wNXAH8G/gg8nDYcSZKkwTFRl6Te8HzgkP7lAGDKCNb1BJEEP0qMIZ9PeTz5I8Q48mXAUmA18HT/8xYT3dMnEd3nxwHrEL9F6xEXEDYixro/j/wY+E2BGf2PGa4biIT9AuAyYoy8JElS2zFRl6TutTVwFHAksNMQn7sKuK1/uZPyWPA7gCcbGONQjCWS9e3Jj5nfBZg2xHU9A/wO+CmRvK9sXJiSJEmSJJWtDxwDXEi0Xq8d5PIIcB5wCvBqotW7k0wHDgXmEK3lSxn8tj8JzCO22wvYkiRJkqQR6wMOBH5LdOcebGL+Q6K1fXrrQ2668cDewL8SifsqBve+3AV8iqG30EuSJEmSxHii9fwGBk5AVwFXEy3me9J7LceTiRb3ucADDPx+LSNa2XdOEawkSZIkqbNsAnwOWED9ZHMNcCnwHmCDJJG2pz5i1vvvAQsZ+D38A3BQkkglSZIkSW1tMnAyMYt6veTyFqLlfJskUXaW0cTY9HnEBHP13tcrgVekCVOSJEmS1E7GAx8myp/VSiJXAr8AXpooxm6wHvBR4D7qJ+znYpd4SZIkSepZR1E/cXwK+AqwZaL4utEY4Ajgb9Qf8/8junMiPkmSJElSFVsBF1A7UXwUOJHoDq/meQnRgl7vQskJ9N7kfJIkSZLUM/qA2dQeh76EqBU+NVWAPWpv4BJqJ+yXAtunCk6SJEmS1Bw7AJdTu1zYN4ANk0UniBJvN1J9Hz0LfARb1yVJkiSpKxxBtJZXSwAvwhnc28ko4IPU3l/nERPTSZIkSZI60GiiK/saKhO+RUQ3eFto29NWwB+pnqzfCeySLDJJkiRJ0rBsCPyJ2q2ym6cLTUNwBNVL5y0Fjk0XliRJkiRpKGYA91B9LPq7E8al4dkcuJLqF10+kzAuSZIkSdIgzAQepDKhe4iYXVydaTxwKtWT9TkJ45IkSZIk1fEC4GEqE7m/ApsmjEuNczQxA3xxH381ZVCSJEmSpEq7AguoTOB+DIxJGJca72XEZIDFfX1qyqAkSZIkSWWbAA9QmbidRpT7Uvd5IfAElfv8oymDkiRJ7c+SP5LUfGOBC4FXFO7/H+D9RGm2TrAusG2dv98ArGzB6/wTWNWA12mFHYGLyA9rWAO8HvhdkogkSZIkSXyfylbVbyWNaHhmUX2itNJyWINe50sDvM6GDXqdVtkJeJL8NjxFTCooSZIkSWqx91OZaP4RGJ0yqGEaKFE/pwGvMYrqQwQ6OVEHOJDobZDdjtuAKSmDkiRJkqRe8wLgOfLJ2e3AeimDGoGBEvWVjHzm+kMGeI1OTdQBPkTltsxNGpEkSZIk9ZBRwJXkk7JFRPLeqQZK1NcCHx7ha5w5iNfo1EQd4Ifkt2UN8OqkEUmSJElSjziBygTz7UkjGrlqiXqxJvw/R7D+9YGlA6y/0xP18cBN5LfnVmLCQUmSJElSk0wB5pNPxs5OGlFjVEvUv0ll9/4XDnP9/1JYzzXAn6q8Zicn6gAvBVaT36YTk0YkSZLairV7Janx3k9+rPZSYnxyN1oE/LZw3zuHua5jC7dPH+Z62t0VRBf4rE8DExPEIkmSJEldbxLwGPnW0s8ljahxqrWon0LlBHBPEF28h2KnwjqWEy3n3diiDrAJsIT8dn0gaUSSJKlt2KIuSY31VmDjzO2ngW8kiqVV/gQ8mLk9DXjtENdxbOH2eUTC360eA75XuO+DQF+CWCRJUpsxUZekxjq+cHsu8FSKQFpoDfCTwn1D6f4+BjiqcN/pIwmoQ3wdWJG5PRPYL1EskiRJktSVZlBZemvbpBE1Vq2u7xBJ5prM/UOpqX5oYZ2PUp4FvVu7vpf8gvy2fT9tOJIkqR3Yoi5JjfP6wu3LgLtSBJLAHUTd+JIxDL4c3bGF22cQiX4v+FHh9mux+7skST3PRF2SGmdW4fY5SaJI5/TC7XcN4jnVxrPPa0g0neEvxDwGJc8Ddk8UiyRJahMm6pLUGH3ASwr3/SFFIAmdSdRUL9kJ2HOA5xxNfob4q4AbGxxXO1tBJOtZL00RiCRJah8m6pLUGNsB62duLwJuSxRLKk9T2YtgoEnlji3cPr1RwXSQKwq3ixd8JElSjzFRl6TG2K5w+3picrBec3rh9lHUrqm+C/lu3iuIydV6zbWF2zOTRCFJktqGibokNcaMwu17kkSR3kXAA5nbGwCvq/HYYim7c4CFzQiqzRWPla1SBCFJktqHibokNUaxZNhDSaJIbw0xa3tWte7vY6mcFf70ZgTUAR4u3N4oSRSSJKltmKhLUmOsU7j9bJIo2sOPyXf7n0XMZp71OvIJ6SNEzfRetJx8Obox1B4uIEmSeoCJuiQ1xtjC7V6pA17NncDfMrfHEGPVs44r3D4DWN3MoNrc8sJtE3VJknqYibokNcbSwu2JSaJoHz8u3D428/+NgUMGeHwv6QMmFe7r5R4ZkiT1PBN1SWqMJYXb6yWJon2cCTyTub0T8KL+/7+DfA+EK4FbWxRXO5pK/vf4OXq7d4EkST3PRF2SGqM4IdhWKYJoI88AZxfuK00qd0zh/tObHk17K1YM6NWJCCVJUj8TdUlqjHsLt7dPEkV7KXZnfzuwD7Br5r5l9Gbt9Kxi3fTisSRJknqMibokNcZN5Gc63wmYnCiWdnExcF/m9gbAjwqPORtY1KqA2tTehds3JIlCkiS1DRN1SWqMJ4nZzktGAy9NFEu7WAPMK9y3XeH26a0Jpa29vHD770mikCRJbcNEXZIa5/8Ktw9NEkV7OZ18T4Osh4GLWhdKW5oO7Jm5vQa4LFEskiSpTZioS1LjnFu4/SaiZb2X3UvlBYySH+Ps5m8myrOVXAU8ligWSZLUJkzUJalx/ky+JNlmwKxEsbST02vcf0Yrg2hTJxRun5MkCkmS1FbGpA5AkrrIc8QM5sdn7jsROD9NOG3jTODBwn0rgNsSxNJO3gLskrm9isox/ZIkSZKkEdqLGJOdXV6SNKLGmUXltp3S5Nf8U5XX3LDJr9kKE4DF5LerWHdekiT1KLu+S1Jj/R24tHDfnBSBqK2dBUwt3Pe1FIFIkqT2Y6IuSY33+cLtVwKHpQhEbWkM8IIq9x8DjG1xLJIkqQ2ZqEtS4/0ZuKBw37eB9RPEovbzKSrryQPMJsrVbdracCRJkiSpN+wErCQ/BvknSSMaOceoj9xuwHLy23M1MZFc6fZDwN6pApQkSZKkbvYFKpPMk5JGNDIm6iMzDbiL/LY8DmwM7E/UTy/dvwx4d5IoJUmSJKmLjQduIJ+YrQRenTKoEZhGxJ5dtm7ya76wymt24jjuscAlVF50ODzzmC2Aqwp/nwuMa2WgkiRJktTttgeeIp98PQlsmzIotdz3qEzST63yuAnAjwqPuxrYsjVhSpIkSVJvmEV+DPJa4A6iBVXd7z+oTNL/TMz+XstsYEXm8QuAVzU3TEmSJEnqLR+nMlm7H9gmZVBqus9Sud/vYXDj7F8OzM88byVwcnPClCRJkqTeNI/KpO1BYGbKoNQUfcDXqdzfi4mKAIO1GXBFYR0/BSY1MlhJkiRJ6lVjgTOpTN4eAV6UMC411jjgNCr381PAXsNY3/gq67sOmNGIYCVJkiSp142mcrIwy3F1j+nA36jcv08CLxnhuo8Bnsus8wngoBGuU5IkSZJEdIv+NpXJ3Fqie/zEdKFpBPYlekcU9+ljwK4Neo09gfsy615FjFvva9D6JUmSJKln9QFfoXqyfgXNr0+uxhkFfIT8LO2l5W5ghwa/3kbARYXX+S2wboNfR5IkSZJ60pHAM1QmeM8RLaWj04WmQdgWuJjqF1x+D6zfpNcdA8wpvN5tNP6igCRJkiT1pBcAt1A92bsW2CNdaKphDHEhZSmV+2wNkUSPakEcbweezbz208BhLXhdSZIkSep66wHXUD1ZXw58HpiSLDpl7UfMul5tXz0FHNrieHYnarOnuFAgSZIkSV1pHPAdqid+2eUJohV3Qpowe96OwFnU3j/nA5snim0acEEhnt/RvK73kiRJktS1NgMuJ59g/QH4ObUTwvuB2Th+vVW2BOYSM6xX2x+PEaXTUhsNnEK0qJdiuxPYOWFMkiRJktRR9gPmU06qVpIvtfUmqpf7Ki03ASdgC3uz7EbUvF9O7X0wj2jNbievBxZRjnEJcETSiCRJkiSpA8wmX85rAXBAlcetQyTv2cSruCwgxiSn6nbdTUYBrwbOI98yXVwuJy60tKvtyU9QuAY4lZgET5IkSZKUMRn4Bfmk7yqie3U9GxOJVr3W3eXAj4kE0onEhmYT4CTgdurPE/BP4DWJYhyqKcBvyMd/CXEsSZIkSZKAmUR39WziNJeYTG6wZgBnEN3k6yWU9wNfwvHJ9awDHE3UOx/o/byz/7GddgGkj+iRsZrytjwAvDhlUJIkSZLUDorjhpcC7xrB+rYEvkKUAxtotvjrgU8Qs5b3uinAG4GfAM8w8Ht3MfAGOi9BL3oN8CSNO/4kSZIkqWONJsaPF2fi3rVB659MjHfPjkeut8wnJkA7Ali3QTG0u62Jbu0XAssY+D1aTpRh2ytFsE20LXADI+vRIUmSJEkdbUMiOcwmRufTnNrWo4BZwE+BZxlc0r4cuAj4LPDa/ng73Rhgd+A9xIztDzC496I0/vzjwKYtj7p1qs2RcBnwvJRBSZIkSVIr7AncR37W7Tm0pgv1ZOAdwB+pXfu71nIXkeyfCLwc2KgF8Q7XOGAHonfA14BLGfxFiux47Tn03lj+YtWBh4GXJo1IkiRJkppoNvnZ2RcChySKZVPgA8SEac8xtCS2tDwJXAmcDnwKOJyYjGxzYHyT459GjK1/FdFK/l/A74jhAwNNAFdruQ34JvAKOn/s+Ui8AniUfA+Lk5JGJEmSgJgNVpLUGBOAbwPHZ+67nkhs70kSUd5EonTbrP5lZoPW+xTwGFHP/VFi0ryVxGRta4DF/Y9b3H97EpHgj+///xhikrdxROmwTYgLDBvRmPHTzxKTwv2B6GXQDvuiXWwO/Bp4Sea+M4iLIkuTRCRJkiRJDbIl8A/yLbdnEIlou9oaOAb4DnA1+a7Qnbw8ApxDtP7vT/Nb/TvdBOB/yb+H1wBbJYxJkiRJkkZkFtG9vZTkLKMzuxBPBPYBPgz8HLiOoY/5buWyhqgXfyExRv0I4oKJhqc4ZONx4ICkEUmSJEnSEPUBJwOrKSc3DwJ7pwyqwfqI5PfVwL8A3yLGiP+TaLke7jjxwS5LiG71pdsXAEcSM7u3c2+FTrUPsV9L7/dK4hiXJEkt5Bh1SRqeqcCPgTdm7vsrkUQ+liSiNPqIseQbUR5bPhkY2//vKMr12tftv/0c0XK7vP//q4iEfDUxzv1xot77AmKc9P7EGHOIkmvvau4m9bzpwK/IzwL/c+DdRA8LSZIkSWo7uxFlzLJdsE8lJkVT421C+b2+InEsvWIMUbYu27vhemJeA0mSJElqK0eRH7f9NPDmpBH1hieI93vxQA9UQx1NvqzfQuDgpBFJkiRJUr9qLYy3ETW+1XyXUX7fn5c4ll6zB3Av+R4kc+jtGvSSJEmSEtsMuJx8kn4O5bHXar7vU37vnYm89TYkZtfPfgbOxc+AJEmSpAT2IyY2K86C7WScrfURyvvgA4lj6VWjqexVcjv2KpEkSZLUQrOBFZSTkgXYmpvKLMr74TuJY+l1bwWeIT9Pw+FJI5IkSZLU9SYDZ5FvObyKqCeuNLaivC/+kjYUAbtSWfnAceuSJEmSmmImcBP5JH0uMC5lUKKPqLW+lhiKoPSmEnM1ZD8rvwfWTxmUJEmSpO7yemAR5aRjKXBc0oiUdQ3lfbNB4lgU+og5G1ZT3jd3AbukDEqSJElS5ytNkrWGcrJxJ9G9V+3jJ5T3z8sSx6K81wFPkb/I9c6kEUmSJEnqWNXKTp2P3Xfb0Wco76PjE8eiSrWGjYxNGZQkSZKkzrIncB9OiNUp3kR5X30tcSyqbgrwK/LJ+l+BTVIGJUmSJKkzzAaWU04mFgKHJI1IA9mB8v76XeJYVFtp3PoqyvvrQWCvlEFJkiRJal8TgB+Qb/G7Ftg6ZVAalLGUL67ckzgWDewQ4gJY6XO2DDghaUSSJEmS2s6WwD/IJ+nzgEkpg9KQ3ELst9XAOolj0cC2BK7CcoeSJEmSqphFZeveSUkj0nD8mvI+3CNxLBqcCcCPyCfrfwOmpwxKkiRJUjrV6jw/COydMigN239S3o9vTxyLhmY2sILy/lsAvDJpRJIkSZJabipwNvmWvEtwBupOdhTlffn5xLFo6F4OzKe8D1cSF9IkSZIk9YDdgbvJl147FRiTMiiN2B6U9+mvEsei4dkMuJL8BbSf4FwRkiRJUlc7CniWchLwNPDmpBGpUSZRHsZwc+JYNHzjge9TWX1hRsqgJEmSJDXeGGAO+ZP/24AdUwalhruX2LcriJJt6lyzKZfcWws8ARyYNCJJkiRJDbMZcDn5JP1sYpy6usvvKe/jFySORSO3J3A/5X26ihi33pcyKEmSJEkjsx/VJ6jyRL87/RflfX1Y4ljUGBsBfyF/oe1MYJ2UQUmSJEkauj6iFnqx5NMBKYNS051AeX9/OnEsapxqQ1duxV4TkiRJUseYDJxF/qT+MmB6yqDUEi+jvM/PSByLGq84GeRi4I1JI5IkSZI0oJnATeST9LnAuJRBqWXWo7zfr04ci5pjd+Ae8uUV5wCjUgYlSZIkqbrXA4son8AvBY5LGpFSeJTY/89i8tatpgF/In9B7nziQo0kSZKkNjCaaFFbQ/mk/U5g15RBKZmLKR8Hz08ci5qn2uf+DmDnlEFJkiRJgg2BC6lsWVs/ZVBK6juUj4VDEsei5ns9MVa9tM+XAG9OGpEkSZLUw/YE7sOxqsr7AOVj4sOJY1FrbA/cQuV3weiUQUmSJEm9ZjawnPKJ+RPAwUkjUrs4gPJxcVriWNQ6U4HfkO9dczGwccqgJEmSpF4wAfgB+ZPxa4EZKYNSW5lOviyfekcfcDKwmvIx8ADw4pRBSZIkSd1sS+Af5JP0ecCklEGpLT1FHB8LUweiJF5D+RiwAoQkSZLUJK8BnqR84r0MOClpRGpnV1I+Vuz63Ju2BW4kf2FvLjA2ZVCSJElSN6jWlfVBYO+UQant/ZDy8fKKxLEoncnAWeST9f8DNk0ZlCRJktTJpgJnkz/JvgTYJGFM6gwfp3zMvDdxLEqrj+h9s5LyMfEQXuyTJEmShmx34G7y5ZZOBcakDEod43WUj51TE8ei9vAK4DHyw2dmJ41IkiRJ6iBHAc9SPqF+Gnhz0ojUabahfPxcmDgWtY8tqD4h5cSUQUmSJEntbAwwh/xJ9G3AjimDUkcaBTxHuZuzVDIB+F/y3zPXAM9PGZQkSZLUjjYDLid/8nw2MU5dGo7rKR9L6yaORe1nNrCc8jHyOPCqpBFJkiRJbWQ/YD7lE+aVxEzvfSmDUsf7OeVj6iWJY1F72hd4hMrvHkmSJKlnlWZjXkH5RHkBcEDKoNQ1/p3ycXVs2lDUxqZT2ZvnZ8CklEFJkiRJKVSrb3wZcdIsNcIRlI+tLyeORe1tPFEdIPt9dD2wdcqgJEmSpFaaCdxE/qR4LjAuZVDqOjtTPr7OTRyLOsM7KE9CuBZYCByUNCJJkiSpBV4PLKJ8IrwUOC5pROpW44gxx2uBOxPHos7xQuA+yt9Rq4BTiEoCkiQl5yROkhppNPAF4BOUv1/uAg4HbkgVlLre7UQPjtXA5sQs31OIUoAA6xHHY1///+n/25T+/y8FllVZ73P961rW/5gVwLNEUrek0RuhltsQOJP8fBnnES3ui5NEJElSPxN1daPJwMbAOkRr27j+/2dP0iFaURb1/38JcfK9jDgRX0CcpGvwNiRm4H515r7fESe9TyWJSJ1kAnEMrU98TovLQPensAh4JrMsIr5LFhPdqRcCT2T+X7q9AFiTIF5VGgP8J/lZ4G8H3gTckiQiSZIwUVfn2RDYvn+ZAWwKbAJsBDyPSNAnNui1ngUeBR4jau/OJ06w7yJO5O6gnOj3uhcBvwKe3397LfAV4NOYkPS6McTncnPi81r6d4vM7ecB01IFmMAq4rtkfpXlQaJL9v1E8q/WeBvwA8qzwC8hqgj8JlVAkqTeZqKudrU+sBewB+XEfCawQcqgqlgA3EYk7bcDVwHX0Fsn2LOB/6Y8SdxC4CjggmQRqZUmELNmbwts079sSSTfmxEX0po17vc54mJZ6WIAwMPArf3/L3VVh3L39dLzltdZ71RiGAeUe+ZMIC4CjiV67Uzq/3cKzWvRXwg8QCTt9wP3EOPw7yKS+ZVNet1etRuRmJdmgfeCoyQpGRN1tYMxRCK+J7APsC/wAjp3Up/VRNJ+Tf9yGXAdnXWiN56YbOmKOo+ZAHwHeFfmvuuI8ej3Ni80JTCVSMCzyXjp/5sz8t+SBUTvlUeJxHsRMVwi+2+1pZRsvxj4R///zwKOHGE8wzGFcuK+AdFDoLSUbm9IXLgo9TCYPILXW0Uk63dR7uVzK1Fp4bERrLfXbUDUVz84c98fiIuPDuGRJLWMibpS2Ro4pH95FdFq1SjPEV3VnyZa1EqtamuonCBo/f5/SxNPTSCSko2JZLVRngIuBP7Yv8xv4Lqb4evAn4Hf1/j7lkRX9xdn7jsDeC+O7e9kWwA7Abv0/zuTSMg3Gub6lgAPEQn4Q8Rx/zDwSP/ycP/f6rVuD8Zk4vPeB9wI7DrC9bXKJMq9DqYTvRC2IoaQbNn/74bDWO9C4GZijPVNRAJ/PfDkiCPuDX3EhJhfpHzB+C5i3PqNqYKSJPUWE3W1ynjglcAsIjmfOcz1LCW6mZeW+cQJ/wLKYz4b1e18XeLEeSNiLO2mxEn1TKIHwLYMryb4WuCflJP2y4hW+HYxi5gEblfiJL/oNcBPKF/kWE5MxHRqS6JTI0yjnIyX/t2ZoXfhXkZ0x74LuDvz7/3EWOtnaz+14R4gLjQsJxL3VS187WaaRDl53wbYLrM8n+iKP1j3Ewn7df3L9cT7puoOJS5Artt/+xngeKLXhiRJTWWirmbqI7qxHw0cQTmxG4xVRIvQlURZr1Ji/iCR6LaD0cQJ9Eyiq/5uxLj67RnaZ+sRokTQT4iT55Q2JZLzaUTPgmwJqmqtTA8R+/bKFsaowRtHXHDZlXJSvjNxAWqwFpNPwu/O3H6Y9vk8XgAc1P//mfRGTfWxRLK+HbBD/7IzsCPx+R2MhcQQnX8Af+//d0HDI+1cM4Gzife05DTgAzhHgCRJ6jA7ErW07yNO4gezPEJM4vMJYD8a2xW+1dYjxjf+G3A+UY5psO/DzcCnKM+e3kqjiO752bJ1JVOJk9VsrJcQXXbVHsYCuwMnAP8DXE20Lg/22HsSuBT4HvA+4nM4nG7XqXyT8ra8PnEs7WBL4nvoY8CPiIuAKxjcsXAPUWrxQ8DLaOwwoE40Bfg1+ffor/j9J0mSOsAooptgKdEbaFlJdPk+mZhErpt7d4witvFk4v0ZzMnymv7HHkrr3ptPZF7/+sz9uxMtqNnYTiXG9CuN0UQL+THEvriM6GY+mM/ecuKC0FnAKcQxtjWd/xl8D+VtPHmAx/aqMeSPmwsZ3IXElcSFn1MZeu+obtFHHFerKb8vDwIvSRmUJElSLVOBDxMtMAOd7C0gWuvewMhmO+5004C3EmMfn2bg9+2fxMzqzWzVehFxAWENMeygVDv4aPIJ4NPAm5sYh6qbQeyLbxJJ+TMMLil/gNiX/wocRsyr0KnVFAayH+XtPj1tKB1lFNEL6jiiJ8b1RGJe77haRXSX/yYxwVq7lc1spllE75PSe7GMGLcuSZLUFjYmZgYfKNF8lih181psga1mIlFK6lwGbml/jOhOP6XBMUwmxhuv6n+d1cC3iNaz7OvfSn6cpppjDHHh5CSi1bs0Dnyg5RHiOPp34vPWa91yN6T8Xvw9cSydbh3iwsfHiAs9j1H/2FtNtLh/mZgnYFLrQ26pbYm5U7LvwVyGN7moJElSQ0whussuoX637T8T3SsbnVR2sw2BfyEmcxooYf8gjTspPIPYZ9nXuLdw++d09rwB7WwKkdx8lvjc1PtsZXun/B74HDEee7OWR92eHifen8V0flf+dvMC4N3Ajxm4B9Uy4C/AZ4iu4d3Yi2Mi8V5kt/syhjZRoyRJ0oiNBWYT5c/qnZzNI2aW1sjsSbyX9bqg3k/sk9EjeJ131ln/2v7Xd7xvY20BvA34b2KCr1JPhlrLcuBy4GvA4aSZaLBTXEr5fds8cSzdbnPiOD6N/BwWtS4sndH/+Gkpgm2i2eR7Qz1MTMAnSZLUdLOofyI2n+iSvVGqALvY84kEbRG13//rGN6ERtsRQxOykyMVL7z8FHgvcQzsSPd3aW2GTYkE5fvEEIOBWssXAucRs/+/nGi50+DMpfw+Hpg4ll4zg2hxP5NIzGsd36uIC0//Sgzx6IaeD/sBj+IFTkmS1CLTiFbdeq0kJ2LpnlYoDTmoNSfAKuC/GHwiPQ64lvqtuauo7BK/luhefBUxieCEkW5YF5pGTLD1bWJ29YES87uILrSziRm5uyFxSeVDlN/XExPH0sv6gN2IMe5/pn6ZwIeISUYPBgU8zgAAIABJREFUprPHeG9OzI2Q3bYz8EKbJElqsCOoPYHQM8AcYN1k0fWuacR7v5Tq++YeBteS+LUazx9oWUIkoDs3aoO6wBTgdcSFkuuo3UNhbf/frgG+QXRj3zRBvN3sYMrv9fcSx6KyyUS1j/8hhuzU+nwsJlrk30pn/r6MB35AfpuuBbZKGJMkSeoS6wPnUP0kaiUxrnbjZNGpZAbRLb1ai/ca4oS4Vmv3wTWeVyuxLJ1szqa3y+qVjANeCXwRuIL68wisAW4iZtE/jN4qY5XCFpTf+0vShqI6diJa2y+hdq+e5cAFRHf6DZNEOXyzyfcieBx4ddKIJElSR9uV2mNobwBenC401bAfcAfV99m1REKftTExZKFeol5KzpcT5cE8wYStiZPvs6g/X8BaYj6Hef2Pd0Kz1uqjPDzkscSxaHA2IHpwzaP+0J7LiJKFnVJ2cB+iZGJ2G07GoS2SJGmI3kZ0aS+eIK0gulp38tjBbjeR2EfVWqaeoNwVvg+4sMpjsieSa4E7iRPKXm79nQwcSnTzv5P6ifn9wOlEOcItEsSqvKso75tOa4ntdROJLvI/pFxqr1rPrj8Cx9P+31EbAReTj99Sl5IkaVBGA9+k+gnRFUTtXHWGvYHbqJ6Afxz4SpW/lVrWV1JuPe/FFp/S5FefAC6i/uRXi4HfAO8Btk0RrOrKToC5b+JYNHyjgVcRcw3UmkV+OfBb4C2076RtY4gLqdm4/wlskzIoSZLU3kYTs9JWOwGai63onWgK8Evqd2kvdtP+GL3Z8rgecCTwI/JdVKu9b1cDXyCGGoxJEawG7VOU9927E8eixhgNHEDMvVEraV9MtMS/ChiVJsy6jibKYWbjfUPSiCRJUlsaB/yaypOdpUSXQnWuPqLreq1JmtYAv6c3W8+3Bz4K/IX6k8A9RlzEOgonT+w0b6S8H7+eOBY13mjiu+s04Emqf34fJHoQ7ZIoxlr2AO4l/108h/a8sCBJkhKYSCRq1cbavjBhXGqsg4CFVO7nn9A7J4ajie7Pc6hf03wV0Wo+p//xvfL+dKOZlPfrHxLHouYaT8wlMY98a3V2uZm4cNkuvYamUTlXyHlEDx9JktTDxhIlb4onM3fgRFjd6JNUH2/9/ZRBNdlGxMRuA83QPp+oeXwYMWRA3WEMsIzyxUf1hvWAE4iSb9WG+SwFfka0xqe+EDeauCiYrb5xB1G2TpIk9ahvU3kCcyswPWVQapoNiAkBH6Jyv384YVyNtisxNvlyqp+kl7qZXg18FngRvdftv5fcRHmfT04ci1pvS+L74HaqfxfcB5wCbJUkurIjyVdbWQK8OWlEkqS24YlqbzmWmDgr6xaihWF+y6MZvjcSM5xXsxD4aoNe52hg5xp/u5+YjbhTzCBmM8/WVV8NvJ4YBtFpRhHjPQ8F3kqMPa9mKfA34HxiToaHWhKdUvsl5YTnRcA1CWNRWnsCs4kSpMWeM2uI+SpOA84mhsC02gv6X7tUYWUtMb7+M8R3tCRJ6nL7UtkF+h7aZ9zeUHyX2t2Z19KYknITqD1R0Voi+es0M4GnyG/Hk8B2KYMaggnERaVTqT9L+6PEmNUjsDW1V32W8vFwdOJY1B4mEt8JF5Lvcl5aHiG6o2+ZILapRLKejecPtH+deEmSNEIbEclL9iRgCe03I+5gDZSof6EBr/G2AV6jExN1gIOpnA3+JiIJbkfrE4nWr8h3ES12af878GmiFrr0Vhr7faDuMpNIyh+j8vtkJfF9cyCt7XVYqtaRHbpzP9EjRJIkdaliXe01wFuSRjQyAyXqDxGT9YxEtQn3uiFRB/gIldvz5aQR5W0OvJ9o+VpB9fd/BfAn4H3AZmnCVBvbjfKx8psaj3Hol8YR48Qvpnor+x3ASUSLd6u8lnzPp6XEsDVJktRl3kTlycfnkkY0cgMl6muJluPh2pzaNci7IVGHqBGe3Z5VpC3Ntz0x+dNVVD9hLvUCOQt4O5YyUn0TKH+Gb6vy9zHEd6NUsgPwTaoPeXoa+G+iJb4VtqM8IWJpmUtUbZEkSV1gHHAn+R/7a+j8H/tqifrlhds/G8H6P01lUt5tifpUoltlcZta2cq4CzHz8o1Uvr+l5TFioqfX0r7d89We7qbclXl84W9zgA+0PCJ1golEC/bfqT7M5g/ALJpf4m0ylb3hLgU2bfLrSpKkFvgYld2Fu6FOa7VE/T2F20sZfqvrrYV1za7yep2eqAO8hsrtanYr457AF6ldNmktcBcxc/++pK93rPb3IqonL+dTPqZ2zNz/asoVD6R69gJ+SuVErGuJ77AP0twJK0vj1rM9vB6idtUTSZLUASZQOTP2N5JG1DjVEvU9gX8W7nvPMNa9b2EdS4jJ+LoxUYd8MrMWuJ7Gtqr3ESeVXyWqDNRKzm8A/oPOneBQ6UwlkqZZhfu/Qvn4Orz/vo0oTyDmxIMarOcRlQSKk7KuJbrKf7H/Mc2yP/mJ75YB727i60mSpCb6APmTicV0T6mXWon6Rwv3XT6Mdf+gsI7vE10huzVR35H8LMNrqUx4hmoUccFjDtE6Xis5v5no+r7DCF9P+iLRLfkblLu5H0f5WPs34qJR9sKU8xxoqMYRJd6qDYdaTpSF3LlJr70FMYdH9jXn9sckSZI6SLF1+Utpw2moWon6xlTOEj6UmuoTgUWF5+9LdyfqEKWIstt2zjDWMZpo9fku1Vud1hKJ1BXEkIwZIw1aytiEcvfkG4jP/d6Uj72fASdmbj+TJkx1kX2AX1N5oXMN8DvglU14zQnAjwqvdzVpar9LkqRh2Iv8D/lKmtstr9VqJeoA5xbu/+IQ1vuOwnPvIFrhuj1RzyY0QzleRgH7Ad8G5lP5Hq0lTmL/SiRJmzc6cCkj+73wHDF+uHT7NuIiXqmiwM2JYlT32ZY49p6j8vvvaqIFvtFzbcwmf1F6AfCqBr+GJElqgq8y8hbSdlYvUS+WoxtKTfW/FJ77mf77uz1Rhxibnt2+99V4XKlb+6nAw1S+L2uJiY8uI+oPT29q1FLZDMqTbpUS8qWZ26W/rQLOSxSjuteGxMRv1b4X7ya+DxtZueLl5C+Qrux/fUmS1MaKM2q/JW04DVcvUR9HtC5k/zaYmurPJ9+FcTXl7oS9kKh/kvz2/b7w952IMecPUflelN6vUnK+SWtClir8jPwM2bUuJH0nVYDqeuOBY4BbqDz27ie+Iyc16LU2I4YTZV/jpw1cvyRJaqDNyf9oLydmRe4m9RJ1iNbe7N9+Poh1/kfhORdk/tYLifqO5LfvWWBXYrK3WhPCZZNza/uqHexGuTW93mLLo5ptFHAo1euxLyC+W9dvwOuMB04rrP86nAdEkqS2czj5H+zL0obTFAMl6nsU/raU+idEfVQmo2/P/L0XEnWo3ZW92Bp5MfAv2HKu9vQHBm5Vf2uy6NSLDgEupfI4fAr4PI2pyHIM+XHyTwAHNWC9kiSpQT5P/kSgW2qnZw2UqEPlmOv31lnf/oXHLiLfdbBXEvVzGLjl3DHnanevYOALTi9NFp162T7E/AjFXh9LiKFFI03Y9wTuy6x3FdF7pG+E65UkSQ3wU/InACekDacpBpOof6jw9yvqrO/0wmP/p/D3XknUv0B+Gx8gZs42OVenuYL6reoe00ppD+AsKku7LWbkLewbARcV1vtbYN0RrFOSJDXAZeR/oA9IG05TDCZRn0a5rnJp2aHKuiYTrRn1Wtt6JVE/gfw2npE2HGnY3kjtJH0FjS+XJQ3HDsA8Ki8qjbSFfUz/87PrvI3qv4GSJKlFbiT/47xr2nCaYjCJOsDZhcd8qcpj3lV4zO1UdhPslUS9mNz8Nm040rD1AbdSvVX9vnRhSVXtSEx6Wq2F/XPAesNc79uJiUFL63saOGykwUqSpOEpToq2bdpwmmKwifobCo95mMqa6sUJfj5ZZT29kqi/mvw2XpQ2HGlEjqXyc7uGmAxRake1WtifJGaJH04Fl92Be8h/BuZgrxJJklqumKhvlzacphhsoj4GmF943CGZv88gP6nPamCLKuvplUT9IPLbeGHacKQRGQs8SOVn/Ecpg5IGYUeqJ+yPE5PDTRji+qYRJUez6/odjSkPJ0lqEq+odp9nCrfXSRJFe1hFZQ31YzP/fxf5bu4XECf2vap4rDybJAqpMVYC/0X+Mz6KmCRRame3EOXWdqM8hAtgQ6I1/DbgOCp7iNWyEHgN8NnMul4D/APYuTEhS5KkgVxC/qr5wUmjaY7BtqhDnIRkH1eqqT4KuL/wtyNrrKNXWtTfS34bbXlUp5tElFvMHtfvShqRNHS7ELPEF3+HbiMS+qE0urye/GdiCXBEI4OVJDWGLerd577C7a1TBNFGbgKuzdyeALyVmA1/y8z9i4FzWxhXO5pRuH1fiiCkBnoOOLVw32Ba1EcTF/TWB7Yivkd36f93a2Dj/r9NbFSgUh03Am8B9id/kXh74MfAVQy+wsu5wF7EZIsQlU9+QXxOxjQgVklSg/il3H3uKdzeJUkU7eXHwAszt98J3F14zE+J1vZeVqwQUHyPpE7038CniDHrEN2JdwE2BTYh6k6X/r8Bw0u+VxMTfj3evzzS/+8CYp6MR4E7iItfq4a3GRJ/BfYlJv78MuXftRcCf+5fPg5cP8B6bieS9R8Ts8D3AScSn423EMetJElqsNeS7xp3TdpwmmIoXd+hek314u2X1Hl+L3R97yMSjew2WnNXnaaPaGV8M/BpIhG5ElhG5Wc4xbKcaMk8m0i0jgdeii3zGro+YrjWHeSPsdKEidUmRq22jpPJl4V7AHhxE+KVJKnnbUR+luNV/fd1k6Em6gC/qvKc0nLzAM/thUT9xeS37ykcGqP2N5VoYTwZOI9oCUydjA9nWUl8D80DZgM74edPgzOWOGaKFU6WE93ZB1OD/TXkL9QuxbkcJElqimvJ/2AflzachhtOon5oleeUlo8P8NxeSNQ/R377fpk2HKmqdYjP8neJibSyFyUbuawiEpcngXuJYSA39P97N/BY/9+ebdLrryVm6j6HmORxqwa8d+puU4DPE5VfssfRE8CHgHEDPH9b4hjPPnfuIJ4nSZKG4BTyP7YXJY2m8YaTqI8hxo1Wa8l63gDP7fZEvQ+4i/z2HZM0IqlsZ+BjxPjbkXRhf45Irn8JfAv4V6Lr+aHA3sTkksPtgj6aGOO+C3AgcDTwEeArRPf7S4hx6iNJ3G8Fvg4cxNDraKt3bES0pK8kf/zcycCzu5cmlss+7zIG/o2UJEmDtCP5H9o1wHZJI2qs4STqAP9OuUWstMwbxPO6PVE/mMqEZv2kEanXzSQuON7J0BPaRcCfgG8C7yNmwy6N192vVRtQw7rEMJOjiNbPXxDfQ0PdxiXEd9fBDL6WtnrLC6he0u0vwO4DPHc2sCLznIeJuRQkSVIDXE7+x/l/0obTUMNN1Ier2xP1i8hv2xlpw1GP2oBIEC5j8F3aV1Ee230S8T3QiWO71yVm8j6FGGv/BINP2hcSXZT3JXrHSFmvICaVzR4zq4HvE9UO6j0v2wtkOfEZkyRJI3Q0+R/mZQxuFthOYKLeOPtQuW22nKiV9idmQS921a213EF0XZ9FjFnvRn1EqayTgYvJt27WW24hxrR36/ui4ekjur3fT/54eYa4OFRrKMXmwN8Lz5mHVQokSRqR0VR2G/1J0ogax0S9MfqAxXTfdqn9jSPmQShOfFltWQqcD7wf2CZFsG1gKlHvei7wEINrZf8SkWhJJZOB/ySGN2WPl3uoPX59AvC/hcdfgxMcSpI0Iu8i/+O6huge2elM1BvjeKqf5D9CjG3s5K7Eak/rAf9GZSmp4rKaGEv7LqJbuMpGAa8kkqdF1H8fVwA/Y+AxyeotmxMt48UhJpcAu9Z4zmyi+3vpsY8T8z9IkqRhGAVcTeWV88kpg2oAE/WRex4Dn+SXlieAc4FPEF3lxyeIV51tAjFz+0LqH2s3EMeZLcGDMwF4MzF0IJtEVbvw8TN6t0eCqnsxMSdE9lhZSfTc2LDK4/chXz1lJTE8Q5IkDcMrqLxq/oOkEY2cifrIjCZmxc5uz3JiwsHLGHg87EriAtCpRHfJaid0EsTFwiOIC4T1ksjziMnUNHybEOONH6d+C/tcLLelsj7gHVSWL32M6HVV7FE1ncrJan+O8yJIkjQs36byhO2DSSMamc9RWWZt5ya+3oQqr/fLJr5es32dyuPhw5m/TyaGSJxMJFCDaXkvlbqbDeyEs08rusXeRO1j5mmihNrWqQLsUhOJz+HN1H7vlxDlKu0do5J1iAs9y8gfK9dQOcHoGGBO4XH/xM+yJElDtg5wO5Wtoq9KGZSSOJbKk/a/Un8c+mgi+Z5NJOPFmYOrLY8SSf7JRNJvQtA71ifGT9cqsbYI+AwxQZqapw94LXAdtT+nNwF7pwpQbWk74Hfkj5M1xHf/JoXHHk1+YrqFwMEti1SSpC6xA5UzfD8BbJsyKLXUPlS2ltwPbDyMdU0nujSfSnSDX039xP1Zolv9HOBQIplT9zkUeJDqx8Byotv1cI43DV+pNFexCkg2CZtL589dosY6lMohK08Rk4yOzjxuD+Be8sfTHJyEVJKkITkYWEX+h3c+0Vqq7rYPlRdqngNe1KD1TyXGGJ8CXEiU1KqXuK8iuubOJcp0zWhQHEpjA+BX1B6DfjqwZargBERJvA8QY4+r7ad7gP2SRad2NJHoFbWE/LFyLbBX5nEbEt/72cechxUbJEkakpOp3k15l5RBqan2I8YDF1vR3trE1xxDTPB3ElHqrd4EV6XFsnCdaXdijgK7VneGdYkLZNWGJqwifiOcY0JZWxGVBYrHyrcoJ+OjqRy3fjs2BEiSNGh9xPjR4gnaAqy1240OIrqdF/f3ZxLEsjXRej6XaE2vNYa5tCwh313e1pn2cxTVj68VxH5zboL29XIq5y4pLWfiLN6q9Doqu8PPJ77XSxd33go8k/n708DhLY9UkqQONYpIloonZ4uIhEjd4d1UjklfC3wqZVAZmxDH2xwiIa8Wa3ZZSb67vF2p0xkL/DfV99PfaW4VBjXOJKIKRLU5Jm4kJhaTsiZSfXb4S4Ad+x+zK3BX5m+lceujkSRJA+oDvkH1iYWcCKazjQdOo/q+/XCd56U2iXxZuCcZend5u+w233jgHKrvj7nEWGh1lllU/7w9Cbw4YVxqXzsAF5M/XpYR5VMnEPOWFL8n/oATiUqSNGifp/oJ9+/xB7UTbQ5cSfUk/cSEcQ1HsSxcsctltWUxManRKcTkdhNaHXSXmwT8icr3fSlwXMK4NHJbAldRuW+fwnkGVNsRxDw32WPmLuL7t4+48Lq68Lddk0QqSVIHOpEYU1o8QbsHa613kiOpPmnbEuJkqhsMtSzcyv7Hndr/vA1bH3LXWAe4iMr3+H4aVz1AaU0Afkj175D904WlNrce8F3y38drgB8QF/xfR1zwyV7Ye2eSSCVJ6kD7Et2Iq7XEzgOmpQtNA9iU2qWx7qC7xwtPplwW7jxinoWBWt3vJo7p2USLvd3lB7YO8Dcq38sbsC56N/oy1ZP1fVIGpbb3QuAa8sfNo8ScIjOJKhDFoTJjk0QqSVKHmU71k/G1xMyuztzaXvqIE6CFVN9n5xEtHb1kDPnu8vczcOI+n3ivTiYuWDnGOq+PmAW8+L5diz0Uulm1Up5PENUbpFrGEHOGZGd+Xwv8jphsrnhR+a/ExKKSJGkA46g9C3Ap+evmFtpOsS8xU3q1fbQU+AS2FJcUu8sPVBbuGfJl4Xp9roZ/o/I9uhrYIGVQw3AwcRxUW17YwNd5XZ3X2aWBr9MKn6By31+Ppds0sG2I+UKyx86zwCeJyiOrMvc/COyVJkxJkjrPS6nsplZaVhMzbW+TLLretSPx3tdKMv9GzMar2qZS7i5/IXFho17ivop8WbgZLY84nTdQedHuOmBKyqCG6WZq7+ObG/Qa21H/QtAXG/Q6rfQfVG7HL/FCoAbWR0wyWez19Q/gvYX7lwEnpAlTkqTOMx74LNUnmiv9sH4Du621wjZEV+5aPR2WAB/AsnrDMZYo7XYScRGk2oR8xaVYFq4b3/ftgafJb/cC4PkpgxqBeon6WhpThuwLA7xGJybqfcAvqNyWT6UMSh1lE+L3K3v8rCAmoLu6cL8lHiVJGoJdgEupffK5jJgp2JIrjfdy4DfUn938HDo3eWpXWxOt53OJBG+g7vJPky8LN6nlETfWKOD/qDyx3j9hTCM1UKL+7RGufxTwwACv0YmJOsBEKku3rSQuUkmDNQu4j/xxdANwbuG+vxFDliRJ0iC9muj2Wu9E9DJiLOboRDF2g7HEe3g59d/rK4BXJIqx12xKjFefQxzjy6m/b7Jl4Y4BtmhibIfR+DHDH6Nym97b4NdotYES9YVEL6LhOniA9Xdyog5RZ/0x8ttzLX7Xa2gmEd+j2YvPK4nSj9neewuAVyaKUZKkjjSKSDzuo/4J6Z3E2MZtB1ifV83Ldge+CjxM/ff2RiJpVDrrEBP6nUxMsPgkAydpxe7yjRrj+1LgIeB4GpM0bQwsJh/7bxuw3tSKifozxCRW2fuOGMH6f15Y121UHgOdnKhDfO902wUcpbEPlZ+Rh8iPW19JfMdKkqQhGE+Mib6LgROUK4APUllv+WQiIWjkjMud5vnEWM9aE/dll+uAo+nO8dCdbjT5snD3MPD+XEy+u/yEEbz+9/rXeRPRsjsSp1HZ0twN81AUE/WnqRxTfv4w170u8FxhXZ+kcp93eqIOlRckFhATNEpDNZFoXc/OAL+Gyp4bP6HzhxNJktRyo4A3ApcwcGKyEvgT8BHytVTnU5nEd6s+YA/g08Q4vIHGPq8mWjPtAth5imXh6s0zsJbo9lnqLn8EMG0Ir7Uu8GhmXRcRPTSGaksqu/V/YBjraUfVEvXiLO0rGV4vn/cV1n0tMR63GxP1zaiskf3JpBGp0+1F5cXq4kSW19FbFTckSWqoPYjWxGUMnLRnl3OByQnibZX1gbcAPyIuSgzmPXka+BYDDx1Q55hCvixcsQW22nI38ZmaTbTY1/O2zPNWERcGfsDQEs/vFl7/dmDMEJ7fzqol6hBzDmTv/8Qw1v33wjpOpHsTdYhjOLtdjxGto9JwjSV62WUvFK4hf4HzCeDAVAFKktQN1iXGsZ9HvktbvaVUq3oenV3yqtQF+hgG35KafQ8u7H9uN1+4UBhDvixcsbtntWU+8bk6mRgjXyxj9Hvyx9tqok78HAbunjyZylasd45g+9pNrUT93YX7h1pTffvC85cDG9Hdifq6wCLy23ZM0ojULXanctLa7CRzq4jvv0bN8SFJUs/akvhRvZ2htbKvJU4E/wR8jWhR3B94Xkujr62P2LYDgfcTrd+XAs8ytG1cQ8zu/n5gw5ZugdpRsbv8QEMjniFahOcQE33tSfRoqfa8J4mLArUmnDu+8PiHiFaublErUZ9K5ef2JUNY71cKz/11//3dnKhDfC9nt+3/0oajLjKG6q3r2ePtTBpf7UKSpJ7zYvKthSsYuLxVvWUxUdP3p8Qs6R8hJlk7kKj5vikja40fQyRMuxMn2+8EPg58HfgFcbV/qAl5Mf7fEBcfmlm2S51vKvnu8kupf2ytonbLfOlE9zaqz27+58LjP9+cTUqmVqIOcEbhb98Z5DrHELP5Z59bqsjQ7Yn6TPLJ0xpg86QRqdvsSv2SsLcCL0gWnSRJHe5A8t1p7yV+WCcRJ7LfonoZo0Ysi4lWxAeJsb63EK2UVwL/6P//bf1/e7j/sUuaEMdq4Hrgy0SvgG5qpVRrjSXfXf4Jhn48loajXEicCEN0ZS5ePNu+FRvUQvUS9QMKf1vE4MZcv67wvMcof767PVGHqOiR3T5LtanRxhIXKmsNo1tMTGgrSZKG4J3kx5bdQMwYXM1U8nWpF1D9R7mTluXEjNkbDPmdkwZva2J88FyGdtFrTf9yOvCuwt9ubeUGtEi9RL2PypJ6bxnEOrPVK9YS3cFLeiFR/xT57ft1/YdLw/Yy4A5qf5fNoTPntZEkqeVOJt8t8s8MvdbuTOBI4N+BnxEt4MXJrlIuC4kWpR8RJ6xvArYiZrEvPeZeYmIpqdmOIFrYBxrXXm0pTnh4aotjb4V6iTrA5wp//90A65tGZYWLXTN/74VE/YXkt+/htOGoy5XqrteaoPV8YL1k0UmS1OZGA98j/+M5j8Z2955O1BV/F/AZIqn4GVHL/Wbgcar/iA8laXkUuJG4wPATYnz6yUQvgX2pn3xPIV8T9lLs7q7m2R64mOoJ93CXo1u6Ba0xUKI+g/xFjlXUH3N9YmF9VxX+3guJ+lgq5+uo1WtKapT9iYvg1b677gB2ThaZJEltahL51uRSy1yq7mhTiRrmmxPdg3cgxvfuRUxwtyeR5GxNzCa/Po0rizaD/PjhbzZovVLJBGLsZnZ4SXFZBaykXFd9sIn6ncQFqW4yUKIO8NfCY+rVVL+28Nj3F/7eC4k6xAWK7DbulzYc9YgpxHCfat9fS4A3pwtNkqT2sgFRJiqbIBRPXHvNgeQnwDkhbTjqIgdQWfJwMVFn/U4iibwQOIeojjCXmMzw34GPEhUHjiQmQzuASOZL61lDd5Y9GkyiXhyrf1uNde1SeNxyKssr9kqifhb5bbSeulrpMKr3oiuNW69VjlKSpJ4wg/wkVssY3ERMveDjlN+XFcDL04ajLjAG2IPoCbIxI0+q16My4e9Gg0nUJ1NZ+aFaTfVvFB5zVpXH9Eqifir5bfxQ2nDUgzamsjdfabm4/++SJPWcXYGHKP8oPonJaNEPKb8/87HWsNrLZuRPbB9JG07TDCZRh5gFP/u47xb+PpbKqhSvqbKeXknUv0h+Gz+TNhz1sGOoXmL1QWK4myRJPeMAovWt9GN4HzEOXHkTiLrtpffpGgZXo1lqhS3In9Q+lDacphlsor5/4XHFmuqHFf7+KNHLoajPGkTZAAAgAElEQVRXEvXPk9/G3xPzgIxLGZR61kzg71R+9pYBxyWMS5KkljmG/CRWN2JLcT3TidJFpfdrXtpwpP9vA/IntE+mDadpBpuo9wF3FR57ZObvvy387cs11tMriXpxGEBpWUm853OJ34udUgWonjOGqNKSnXujtJyGVVgkSV3sJPJljC4C1k0aUWd4Gfm6yx9OG44ERMtnsSxZN57IDjZRh5hNv9hKDDHWtTjL/i411tErifo8qifq1Zb5wHnE+3so1rxWc+0F3E3lcXglsGnCuCRJarjRwHfI/+D9kujarcE5lnxCNCtpNFJ4hPznepu04TTFUBL1rciXs1tN9Bj6KJUn/LX0SqL+fww+US8uq4j9Mo+oRLAT6cp5qjvVKuP2BLB3wrgkSWqY8VSW4UlZI72TZS92LAS2TRuOlCutuJZo7ew2Q0nUAf5SePzJwD8L972vzvN7IVHvIxKe4Sbq1ZbFxPE4hzgOi2XvpOE4nPycOqULRf+SMihJkkZqfeBSyj9ua4iyYxqeseSTgFuAqUkjUq/7NvkT2P9MG05TDDVRP6bw+GJCupT4bqylFxL17Wlskl5reYS4UHwSsC9OVKfh2ZLoBVM8vs7FCV4lSR1oK+BWyj9oy4C3pgyoS0wjP3buHOydoHTeQf7E9bK04TTFUBP1dfofUyt5/PkAz++FRH02rUnUi8szxDF6KnFB5fnN3lB1jdFECcHs0Ja1RAk3jyNJUsfYhfjxKv2QPQXslzSi7rIbccJZen9PSRqNetkW5CeUW01MnNZNhpqoA/wvtZPFQwZ4bi8k6ueR375/EMMDVlE/0W7Gch9x8eQkYuzx+OZttrrAS4AF5I+h5cAbUgYlSdJgvIqoH1z6AXuYSCzVWG+inCCtAd6SNhz1sGvIn7R+IG04DTecRP3lVE8KHyJa5urp9kR9GvkqFmuBnfv/tg7RRf0kosv6owycaDd6qVYerq/xb4M62FTgAiqPne+mDEqSpHqOJq4sl360biJa3NQcX6D8Xi+hdrknqZk+Tf5k9fq04TTccBL1PuBOKk/kB5Nwd3uiXpwB//YBHj+dmBxuDtFtfSn1E+1mLMXycPXmGFDvOInKXiDXERecJElqGyeRH7t1Mda6bbZR5LuQ3oszHav1NqPyZPVVSSNqrOEk6hAn6+sXlsFMZtbNifpY4nsqu22fGOI6xgB7Er8586jcP61YsuXhTuqPx7lCetMuVHaFX4Il3CRJbaAP+Cr5H6lfY430VplC9Fwovfd/Jk5kpVb6LfnvgL+kDaehhpuoD1c3J+rHkd+upTRmToNNiZbuU4iLl08xtMS7EcvTWB6uV42nsiv8aqK3kSRJSYwHziT/42SN9NabSf7E9Btpw1EP2ovKxOU1SSNqHBP1xpgI3E9+u77TpNcaTYwtP4YYa34zlbN1t2KxPFxv+QiVx9mVOEGhJKnF1gf+SvnHaA1wctKIettB5LsfH582HPWgP5E/Qb2V7jhBNVFvjM+S36bltLas1VQiWT6ZaHV/nKEn3iNdiuXhtmrmBiuJ3ans0fEksGvKoCRJvWM6UUone8L1tqQRCeIEtLRPlhKtnFKr7EblWPVuSDBN1EduV/ITja4Fvpw0orA1kTCfSiTQxRhb1ep+HvH9vS/R80CdbSLRkp7dz6uA96QMSpLU/XYGHiB/0npQ0oiU9VPK+2Y+MdGX1CpzyZ+criRKlXUyE/WRmUjMhJ3dnkeBdVMGVUO2PNw8osZ6qxN3y8N1jy9QLqNaWs4nJlWUJKmhXkm+RvojRDcvtY8JwD8o76OrsYVGrbMu+Qt5a4kuxjNSBjVCJurD10f+4mFpOTxlUEPUDuXhFgEXYnm4TnQA8Bz5/fkAXkSXJDXQ4eRPUG4GtkwakWqZDjxMeV/NSxuOeswhVLYiXQNMShnUCEwlX2Kt2WUnx1JZ1q1TL7Z9isqk86dJIxq5MUQr92zK5eGKx3uzF8vDdZaNgLvI78PlwJtSBiVJ6g7FGumXY+mZdvcyYBnlfXZS2nDUY06hMrk4E7vw9pJDqZwB+0ZgcsqgmqQdy8Nt1MwN1pD1AadRud/+l6hSIEnSkPQRE/5kf1R+Q+e27vSaY8m3wBySNBr1kj7gF1SelJ6GLX+9YH9gCfl9vxDYJmFMrdSu5eG6oQpDpzuOykk3byIu9kiSNCjjgZ+T/zH5Fp5kd5rvkT9R3jZtOOohk8lXh8h2fR6TMC411ywqx3GvIOY46WVTSF8ebgUxb0mpPFwnzx3RyXYAFpDfN88QF7gkSaprPeASyj8ga4gufeo8Y4GLKe/LW4gxt1IrzCCqD5is94bDqCxxtgZ4b8qg2th04AjaozzcKcCrscdcq0whhhEWPyufxSFCkqQappMvpbMcOCppRBqpacA9lPfp2XgioNbZHniIygThPNqzRJeG571Ei20x8Xh/yqA6TLE83L20PnG3PFzr9AFfonIfXErzJ62UJHWYnYD7Kf9YLMFxzd1id6JrXWnf/nvacNRjtgLupvKE9E5gl3RhqQHGA9+nct+apDdGsTxcsdRXKxbLwzXX68lP/roWeBS/GyVJ/V4KPEG+O9weSSNSox1OuYzQGqLLpdQqWxKJeTEJWAIcmTAuDd/mwN+p3KeriMks1XiWh+tO2xD11Yvv8+yUQUmS0juM/OQ/twDPTxqRmiXbzW4JsHPacNRjplM9sVtDjNPt1FrrveiNVJ8Q7en+v6l1NiFfHu5JWt/qXioPdypxEdjycEM3gRiaVnxvzwDGJYxLkpTIieTLxlyBNdK72SjiRK60v+/F/a3WGk+czFc72b8HODBdaBqEjYmW1Gr773aitVdpZcvDnUrM9G55uM5RPC9bC9xGDCGSJPWAPuLqe/aH4Byc8bUXTCG6LZb2+4U4A7da7xhqj7c9i5gEUe3lCGqXFTsXJwdsZ8XycMXyYK1YLA9XaRawQZX7X0H0Usi+f88AB7UuNElSCuOI8kjZH4AfYLLWS2YCT1He//+VNhz1qJeQr0iQXR4GjiNaB5XWbsAFVN9Py4nkz5nBO4/l4dLbiihhOYe4mJK1OfmL6muJYUL/iXMCSFJXmgz8kfyX/ikpA1IyBxGT1ZSOhePThqMeNZE4Sc0ei9nlViKZMBFsvS3/H3vnHXZHVb3t+00P6YEk9BKQDtI7ohhpAioYCypgw0axYwOxgIANBBVULAjqD1CkiEhRkCYd6R0ikBAggSSkJ+/7/fGc851pp5d9ynNf176Sd86ZmTXlzOy191rPQmW6il2be4HtgllnGo3Lw4XhixQGJ48iPmkynPTEygBwDTC2tWYaY4xpJmsA9xB/IX4sqEUmNF+lcD8sBnYOa47pYXYBHqR4B/4WYM9g1vUWU4CzKT7DuhD4Ao526AXy5eFOQmlS7VAeLitUvJMZDNxJ4XgfJT04+UnSA2bPY00IY4zpCjYjXSN9/6AWmXagD/gD8TDEtYJaZHqZYcC3iFehSLabUKlBO4mNZ3PgF5R2xq5DpaRMb9IO5eEGgKforvJw26DJk+gx3oZy1fPsRjxlbQDVX39XSy01xhjTUHYhLgA0C73YjAGFHkdH82/D6rwmLGuj0NdkxzXanka50RMC2dhN7IHyhEs5XA+iWT5jkoxD+eUnEa483ALi5eEmN/OAm0S0fGq0/Q3YKvedScB/Mr7zQzp/sMIYY3qOdxKfHXkS2CioRaYdWReYTeE++V1Yc4wBNHP3V0p30OehMO1dA9nYqawBfI60WFXWgMgHsBNgKsfl4WpjOPAw2ceyEh3LVBTV8OOM79yABy6NMaZjOJr4y/F2NBprTBa7E89JPSasOcb8f3ZD4knlQmyfRKHzG4cxs+0ZCxyBzmUxgbh8exa9Q4aFMNR0He1YHq4d87v3ovRzbimKNpoCfBAdU3JwYqvUVjuDkcCbgC+hY7wMaRNcC1wK/Az4LIoSHRrIRmOMqZusGumXAasEtMl0Bp+icM8sB/YOa44xMbYEfknpHPbowOTxwNZBLG0fJqEO/f9RmRDYLSh02BoAptkky8MtIcyse7Q8XDv0k35BebsXoGoZuwMvJj5bCryn5VbXzk5Ib2ABlV+3OcDPka6GMcZ0DENQTfToA+08XCPdVM45xF+GFo4y7cYk5IQ/R2WdutkobPRwuj80dBDSIDkeOT+VhByvRM7KbgHsNSbPUHTv5svDPU3rHfd8ebjzkWBeiPJw45CieyX2voSiiG7N+Owc2rvvtwFwJfVdr5XoWnVytOg/KEQOXItL5RrTtYwG/k7hAeYa6aYWhqJct/x99BCu12rak2FIh+NiKptlz3fEbwXOAN6Pcj47mXHA24BvAJcDr1B5J/dulKe+RsutNqYy2qE83DxaXx7ukCptfIxsp/dmYHwL7K2W9wGv07hr9BLw5lYeQANJpiF9L6w5xphmsDrqdOV/6CvQaLAxtbAq8dmMv9D6WQVjqmEc8BHgn1QvXPUSmlE+AZV+24r2E57qA9ZDTvkxwG+R8FS1pbGeAb6LSnYa02m0a3m4ZqSK/LkGu54kXS1jFu31ez+e0tfsThQl8FHgQDQ4chRwMvBAifWWowGATsOOujFdzobAExR+5K8Dbw9qkekGtgEWUrivTghrjjFlORCJMa2N9BYuo7q8x2hbiQarrkY5tMcA7wb2BDZBAwONZDiwDsrXPBANOnwXhezfR+0zif1IQOu7SBnfA26m20iWh5tD6x33ZpSHW53aS90l8/2XAPs3wKZ6OZriNv8ehcOXY3NUsq6Ys35ww61uLnbUTcPxi7592Bm9mPL5OXPQ6ONtwSwy3cS7kaPQh14g7wEuCWqRMdkcjO7VlcB+wE255cOR2vR+qKPaSKXnJcDLaEZ+IVJgXoLC8FegznueYcAoNPOWTyUZj1KWVqex4alzUKju31H+4+wGbtuYTmAq+t1vjwTXtqX1JQZnoUjHm5FI413o+VANRyEF9EYwAHwH+GaDtlcte6PnUTJv/lXgveiZVQ2HAb8mHf20AA14PlqDjSFYQTwi41Tgq4FsMcZUyHjKv1TeQXzG8yngDU22y/QepxGfNdgyrDnGpPgQ8VmJP5T47hro2XkKCpOfT+mZqXZvK4D7kRr+x1DYvhXbjYkzGjnux6EBvU4pD9cHXNdgO/6OSqG1klFkiwO+hpzqWtmP7GoBt9M5z0HPqBvTgfykzOcfIZ6HdAeNCbUyJskg4iI1TwOrBbXImAIfJ56TfjnV5ZcPRoNPH0Ud6KvRPV5tnnsr2hwULfUb4MtIPGl0FcdqjCnQKeXh3kDjRfSeQJE8reJ7GTb0o2Oul49nbHsADVx2AnbUTcNx6Htz2QW4lGwl3j4UtvTNyLJrkQDSgozvG9MIxiIHIV+z9FrgAPSCMSYUnwbOpvBO+hOF2fV6GQFsHGnroo7tpNy/U9AsUaNYQSGMfmbu/zNRh/qxXHulgfszxsQZCmxNPGS+1VUhVqLfejRk/mHgKygKqJEsAt4K/KfB202yGhKxTA4q/prGlCLrQ9FRb04sn4EGOZY3YB/NpJWh75sCb0L+xaqoXOkipIXwFIrOuof636FD0O9oO/SunIwiSmbn2g1ICLLZTEX3+BYoUrkfvWPvQIPyiyrYRh+K+tgN3U+jUSTILOB6JH440GC7B+f2uRPqc0zJLXsRnb9/I+2aRu/XVMBw9JB+KuOzIcAviI+8/Qa9XIxpNpugXLL8vff9sOaYHud44s/CX9L6HNRRSMxze9T5mYaE4Kaj0m9HRdqRueWH5L43DdgBdSAcDWVMe5IsDxdNN2xVm4fC32c2Ydv9wGcadbKK8IWM/S6gsWXuNidbSf6dZdbbIGOdD9Vow1WJ7RTTito/Y5/VttursGsy6q/9r8Jtz0MVBw6i+vSBddEAzGsV7OdZ4OtkR5GUI6t84faRz3dGgzel9v8aGgBLaibk6QOOQJUUSm3nMWDfGo4hi6nIr6tEEHMmKstay/kzdXASugD3JJaPIq1yeWpLLTNGD6NomNZHwppjepSkk/5zWu+kG2N6j3x5uMORyFuo8nCNbtfTvGfofzP2d14T9nNDxn4uLbNONzvqg1H0bT316j9R2aHThyoD1ZKi8RzVVyQo5ah/hXQ6Qal2Deloj/FIy6Ga4/hylccQZThKwVla5T4HgBeAt9Sxb1MFm1C4SNdHlk9BAiT5i7IC+GTLrTNGfI3CvbiY+oRgjKmGPuCHxF9SpwW1yBjT67RDebhGtAUoMqiRrFNkX7s1eD8gBzu5n/mo4kYxutVRH4PuxXr3U4mvMQTNAtezn+Uo6qxSijnqJ9S4/79Etj2OuM9VTavl3pkI3Fjj/vJtCaqc8P8pFiZgaqcPjTAORqOz83PLp6I8irya+yJ0Ma5stYHG5PgeUpZ+H8rj/SuwIxrVM6ZZ9AFnAMdGlp2GRs+NMSYU+dD06yLL2qE8XLWMRg7DlcBnyU7BrJa9Mpa9SnNKCP89Y9kYdA3aqWTxY8TfW6cQvzduQP3+Uswq8dlwNEu8S8ZnC1B07rVINHUu6setivQZdgH2obpw6rPJdrJnA39EIegv5PazHqq68i7iAyhDUMj8fOJOczXsD3wr8vdzKKLiTqT5sgr6HX4QDdBEeRcqP3wxcD7xMPo7kUjt4+h8TQb2zG0nORP/IzRgM6dCm0eh671Vxmd3osGWvP0rgLVy+/4AGgTLMxy4AKU3tNO93lV8gvjI0m/RTOXsyPI56IFvTGhGUhDQGABupTqlbWOqYTB6iUdHkL8R1CJjjKmcZHm4aN+uHdty4Czq189IRkANEB/QaDRZJeA+XeL7IWbUkzRa9f2nZF/TnyNhsnKMQo7gfZSfUT+4yL5+g2ami7ElBTG0aHsF6UKUI2tGPR+RvBL1D4r1SUcAv8tY/wGUzpn/exbw9hI2rINSX5LbOaEC+/Nk2fEsEsArxUikO5Bc9xlKn3dTI6ujEdl8OaAVaBQoWt/3aRQab0y7sC7xzsZvg1pjupXBaIQ7f5/1o86uMcZ0Mu1QHq5cW4BC+sfUeIxZ4dfN1Fe6OGN/Z5b4frc56m8jfTwrgU/VsK0+lKtdjGFkCxyeS2XVwSaS7eheWMG6WY56vn/wwQrWH0x2eHs+x34msFEF29mQ9O/2CSo7/kMz9v8Aim6olK9nbOPrVaxvKuQS0jV7o3/fifLUjWk39iAufnF0WHNMlzEMhcHl768VWMDQGNOdDEUht8ehwcksJyZUmw0cQ+l87yzuzdhWM1Xms2bw/1zi+93mqP+L9PE0qy77YRn7eoDqKlFtjsq2RbexjPKz6sUc9Z9Vse93FtnGANWpuP8yY/1KyjrenVhnPpoAq4Y+FKES3c5MHOHaUA6i9MPxDpT/uxbpXAhj2oHPULhfl2P1SdMYVkF5elEn/fCgFhljTGvZCEVYtouy/GxUbrLSsl1ZJcEqmfGslRMz9ndDie93k6O+I+ljeZTqB1cq5aaM/b2thu2clbGdE8usk+WoL6G6VI0RZKvU31jFNgAOyNjGoWXW2TtjnVrT+fbI2NYBNW7LJBiDBBaqKSOwEtX+exo4BznwxoTmXAr36BwqG000phijiI8SL0UvZmOM6RUOAGYQ3jnPag+isP1yvJSx7kG1nIwKOTZjf6UU0rvJUf8W6WNpVpTjaNJ2P01lId9JtiRt901l1sly1C+pYd/3ZGznyCq3sWbGNr5WZp2fJb6/gvoip59PbO/kdleu7BROBtag8pFJkDLkf9GI5iex0rZpD46mMAo5EYUrjwpnjulgxiNF2ryYylLUIaxVCdYYYzqJKSj0/W9UHwrbKrZAong3U1rkOKt/u7wpFhXfdjV97E5mr8Tf/VSW710LO5I+r/k0tWp5EM38R9mB6iMByjn3WcxowHZmkr7vygm67Zn4+w4UrVIrtyT+3s2Oev3siEKGKxl9Wpn791bUed2L5qpmGlMty1HZwOdyf78RdTRqGV01vcsE4B/Arrm/FwIHotIoxhjTzfSh2byHqX1Wt9Xsjhyb/6NQRjjKkoxlzVSlzhI/W9TE/bULg4GdE8seRaXwmsH2GcvuqWN7dyf+HoFm2qvh8Rr2uyDx90oUGVDvdsaW+O4ENNAV5cEa9hklOWm7luuo18cQJD5QjpVoBv1qFNJyZzONMqZOZqP6mDej/OJDUPjPySGNMh3DFDSTnq8nOg+Fft4azCJjjGkNG6IUsnJlmdqRPlSD+l3Ar4BvAy/mPptPWhismY561rbnN3F/7cJ45NxG+W8T95eVC/5wHdt7qMJ9lOK1Gva7LPF3vtpWvdspJai3DulJrLei/k+1DEaDAsnf2EQ76vXxJTTjWIx+dBH/gspi1HPzG9NK7kVpGRfk/v42cD8q0WJMMdYBrqcwI/MqsB8KBzPGmG5lCPA51NdbJawpdTMUlQE7HPgR8AM007dp4nuV1PKulSznbmYT99cuTMxYNreJ+8uKXKjFUc6TNfNfqjRcFklnuRaWNmAb5ci6VlNprLbTeDvqGrVbHdW7WwXNfI+LfDYot3wZEglYAixGD6hvF9lmXizuj8B3qS2Mw5jQXIgGor6EfgcXolDmrBFTY9ZHqTwb5v6eDeyDBniMMaZb2RbNQG8X2pAGMwo4AekoPZXx+bZN3HfWtmsJZe40JmQsa2YkQVbkQjL8uxqybK3WUe8Ushz1RjO42x31VYCNI20DNEo3GYm/TSIdYtII5iHnfCQS53oSeCy3bAaaaTemE/gKysE5AFU3uALpMswJaZRpOzZBTvraub9nofIuHtQxxnQrI9EM+ufRjHonshTNoBZrr+b+3QnYJbHuDk2yaSTp3F+A+5q0v3YiKw+/GX5KnsUZy+qp3Z1la9Y+uoGVGcsepJAu0hA69cGSZDCwGRJg2I6CY56VP9AKxqOH2k4Zny1FDvvjSCDiTlRyoqEX1pgG0Q+8H/gP+o1tgCJF9if7IWV6j81RTlY+t2oGytPKmoExxphuYE+kUbRJaENQv3Ipmpwq1a9fjMqe3YYc8MVULlJ2J/CRxLL1UJ/gmWqMrYDdSOcGr6R0ebZGElJdPivMvZkz0llh7uOoPc0ga4a+WUJ4oXklY9mvgDMbuZNOddSnoJG9nXP/7oBm+zqB4UhkaavE8hnoIfQflM95N9kqm8a0mvlIXOZ29BB+G6oP+uWQRpm2YDuk7r5a7u/HkZP+fDCLjDGmeYwHTgM+TvMngvpR9NoLwLNotu52FLG0BDnnB6LIt3K1m5cB7wSuqdGWe1E6U3I/RwLfrHGbxfhwxrI7KO3wZUWq1lrZKmSo9lyUPhu9tzZo4v6yzum6wCM1bm/9CvfRDWRFljb8WnWKoz4U2AOJEu0HbN3AbS9Es9mzUcjJSgo5FvPQj38RqgM4BJiGhAKWIef6eZRTMhmF0pdSCCzFern2ntzfi4EbkFL81TjP3YTlMeB9wJVotPlL6EH+m5BGmaDsiJ5N+TytR9DzsRcEf4wxvcd7gJ8Qd1ZXUggTX4C0jIai/O4JuVaJw7gA9fMeRhM1+ZYVNjwK+Bhy0FevYNsrgcOo3UkHOY//h2bkoxyJ9JoaFWE3AVWaSfLHMuu9nrFsdI02JJW3W8kSFG27WWTZjsj/WNGE/WU55NugAfha2Cbx9wDdK6T9DPIPo+KRyfSQrmYN4BPApRRk9qttK1B++FXAj1G983egsJoN0cOuGvYF/gccg3JospiEcmvegh6MxwPnoVJXL9d4HAMojPSnwNvRoIExIfgGhXtyMXqBmN5jL+LP5XsozKobY0y3sQZwEAp53wpFcr4H5ahfhPQ4+qmsPzcX9QnPRMrqW1CZMz8KOA7NqJfa/oLI//tJh6zXylZkH+NxDdo+wFkZ23+d8sJdQzLW+04N+18nYzu3Vbju0sR6P6hh/wDnZNjQrHJ/62bsq1YnfRxyXKPbeqzMOodk7D+rtns5fpXYxqwatgHp39avynz/msT3V1I+uqWjGYMeWv9ATnY1juxLSOjqGyhMd3Ma79Du3YBtTkQjLkeih/RtaAStmmOdA/wcRRmEyME3vUsf8CcK9+ILhB19Nq1nf+Iv4ztojfqpMca0miHIkT4cOBX1M6uZdJmZW+ck5OivUYMNo9GkT3K//aSdw3sSf3++hv2V4lLSx7gAOXz1shNydJLbP73C9Wcn1rusBhs+m7H/Sh31OYn1flvD/gGmZ9hwSY3bqoT/Jfa1gtru04+Rtvu8Mut0uqP+ZdL2f7fGfbctQ9As8R9QGHolD74VKJf7TDRr3ciadSEYhvLtj0UlsJ6l8pfAM+im2Cy5UWOaxEjgLgr34C3UpxJqOoeDUCRF/trfSOfogxhjTCnGoAmQo1D/8mbSM4TF2jI0q34+mmGeRv0DmKMpPoM+M/H3q8DPiDu6J9W5/yw2Jf4OyLe7qC+3e13Un01u90UqP49/T6y7gOoiZ1chu/9dqaP+dGK9K6vYd5ShwHOJba0E3lzj9srxbdLH/NMqtzGC9PEPoAiUUnS6o74a8QiWARRtuF6N+28rxqIH0LNU9hB8CYUXHU5vzN5MRS+LK8h+KGa1m9FIXEjFStMbrId+k/l77zdhzTEt4H2oM5q/5n+neAqQMca0MxOQU34ccq4fIns2N6vNoxC6flRuO40crB6DZtCTM7QDaMb8pcSyq5EA2/LIsmodrWo4PsOuARRdtWoN21sPpalmbfPQKrbzzYz1v1DF+klnr1pH/arEejOq2HeSz2fYMQOF5jeatYjfOwNoQvRNVWzjh6TtvbeC9TrdUQeJSyaP4T7qn8SoJaqhIWyMHiCvU/5heDcSy9iG3g7xXgVFHfyMykKuHgM+TfU5+MZUwx7Ew+4+HdYcUwfltAY+SPxFfgXNre1qjDGNYk0UDXQSenYlZ6NLtZmo/GQ0n7xZ/dGJaHbztYQN/cDfUNpZNEd8IRpo2Jv4hM4F1K54Xgl9wF/IPl+zgQ9UuJ1BSPOpmA5VtTneG5HOobq0ac8AACAASURBVF+AdKlKMYzsvPBqHfXvZKz7yaqOoMBI5Owlt/c/qnNkR6Jz/I4y3zsjY1+vUf7c9QEnZKzbDxxQgX3d4KhPQDpiyeN4GNiyBhu2RYOHWaryTWU79IAsN2L5LHAyyjE3aYaichx/pHxI1lwUFh+y1ITpbo6mcL8tQyKKxTi4JRaZatkI1QMuxieJP7f/RO3VLYwxplkMpZBPng9dT4alFmsrUGc7mk/eKlGo1XL7fDVhU3/Ong+jCZjoZ7cCb0CDrFFH93JaU9FpJBrAKHY+H0aDDruigZLB6PqshWZqTyc7VDrffkNtgw3JWe0B1Ff+Nuk02bWRaHXUyXoJzQTX4qhvUeRYbke6Uifm9ndUpL2zxPY2JnsQYyVy5N5KdiTHeCR+fS6FqIxyAwYj0TXL+l38mHR67WCU3vHPIsd8Tpn95ekGRx1UjSxrAnoF8tf2oXgVgsnoep1C/Hc+v8j3G86GyMhSapgLgV+gXIZenjmvljHAEaiMW6kX0Bzgi3gGzDSHX1C4116huGbEgzgtox25iuKCL18i/iy5gM4p62mM6V7GUQhdPxc55ZUK8y4lnk++B/ESS61iEnLQ5yXsW4kc9J1zn0fFlZfllg1GjvqLkc/+RWv7eSOB31PZOV9JZar4/cD3qD0iYCqlI3bnoVz4rO8sRc5v0tmv1FEHuLiCY0w68aXYm/QATrQtRmkDd6AZ+P+RfZ4rmdnfHJWcLravWSjS+UFKV+O6jsojervFUQd4G5ogLXZelqPJ6HvQdX+M0lHSTXfUV0MKmaUenLPQA6eWnBYTZxv0siqVz/4cGsFzR9s0kqFIVCx/n91H+iE9NvdZufAr01rejq5L1uh3Mg/xHJobTmmMMVnkQ9ePp5BPXmkptFdJl0ILPWA8GfWPkwLKeQd9W2Tn3YnPH8x9BspVfpb4ezdU9OSRaJC+Ggc1q81A76R6OZjKtZ3ybRGF2e16HPUJxPtD9TrqIAc6K7S6mlZpCP56ZM+sV9r+SHXVsbrJUQdNTmelLNTS5tV4DGUZjEQQSoUb3QG8H4dPNoM1UJ5MqVGaB1CJOGMaxRTiKqV/Jh4ds2lu+fWtN80UYRgFAZ+zEp8lVWDPxtFOxpjmki+FNp1CPnmy5FapFi2FNp3m5pPXwhTkoCfTFlciseRNkb3HEZ/kWokGGvJhzpOJh8g+gmbnQzIRnffoDH+l7VngczQ2qmFPNKBTyf7vRpNdeepx1EF+0IdQf6fcgEEljjooUuILVFcacADdRxeiMPpKGYEGxZJaCaXao9Q2EdNtjjroOfYhdE5qcdAfAL5GY0oeptgSlU0rtvMH0cPTNJ9V0A+tWMhMP5qBL5YzYUy1bEt8huCrkc+mRZZv1XrTTAbRsPYf5Zb1kRaVOTWIdcaYbmYY6XzySkv0LqcQun48mm1frbXmV8W66BiTTtsydAx5J2p90qmMzxBX314FOY75z5+jvUpBDUYhwKcgR3UGcSHSZWh2+B/Isd+d5g2mDEHVSi5A+fD5+2sF8Diqc35Axv73RL5Kvu1dpw2bIv2egxLbnY5C7athFaRTdRYKoX6BQjWWZWig5N9IfPpQ6ouyGAe8F/gd+r3lqw0sR4NidyHBv7dSe6TuWqTPyYQatrNDYhsH1WhP8hrtUON2QBGIuwJfR3oOjxMf/FiAfr/Xoet1BDofTWEoelgWC3OfgUKuQ4cb9SITyR7Bjb4E9glmnek2Pkjh3lqJXiigzlh++c/DmGYiTEEvif5cOw09n5Oj0ieGMtAY0zWMJ10KLZp3XarNRw5BNJ+8U8pCro8c9GTfOO+gbxT57uGk837PJz6ZMhSVYst//hKwSTMPoIEMoT18gG6N5G3VcTl1tn5afg9uhmbKsx6wc5AqdDX5CqY5rItGxbLyuvqR89TIOqCmd/kB8U7WFmh2Pb9sMRpAMuH4LXEl95PR8yH6TPhsKOOMMR1LshRaNbm1c0nnk3eiLsam6HmarE29GKURRWthTwb+mvjei6SrpPQhJfT8dxai2TpjjCnKwRTPYbiCJk7hm5rZk3SZj3y7G40AG1MPg1DN1/x99Rgq/xWdQfl8MOvM9qQH7KK5fP3AZ4JZZ4zpBPL55IejqL0rqE5QLJpPfhDS1+l0tkKz4MlogSUo1TDZJz6UdM7xxWSH8Z8W+c4yYP/Gm2+M6RYGo4dr1uzsLODdwSwzlTASvVizQs9eRvnExtTDBJSPk7+vZlO431aiEiLtEAbXa/QhMc9iYacrUJ6UMcbkGYNCzo+ikE9eqar2MtKl0LpNG2dXNOiQ7BO/jiLMkvXYxyLHPfrd19D5zeJTke/1o7rqxhiTyQSUFJ/1QP4d4cpDmOrZmezZ9eVIgdOYcuwL3IREY85HA0DHIiXP91O61matYh+mdqJaAcm2ErgVPcfPzbWfoWt6KqomsXbrTTbGtJA10WB9NJ88miZTqr1GIXT9KOSUd3NK3R7IQU+eh/noHKyesc5b0UB19PvXEg+Hj/Ie4uf/K40z3xjT7lSrtDgBKTXumFi+FInJndkIo0xLGYPyng7N+Ow0/FIw5VkT+DVy2itlJfAvpA4bkkGorM0k1KmagnIGp6Dn3QgUgTIYzYKABiPzz8756FgWo/DGFUikbRGKIngRRam8GPn/siYfUzHGAE+gY83K+8wLyw3kPh8cWf5HJCz3dPPNNMa0iDVRKky+7UC2c5nFLJQu9xCqu3w3hfrL3c40VMIymSP+CvBT1Bd+NfHZCBSJ+iUKz9/FwLeA76PnbJK9UJ87P9hxDppdN8aYFFOA+0mPHD6PBS06nT400JIVDvsD2qsOqWlP+pBw5BKyU2KyWj8S3Wk2g5D2wj7AMUjM5xpUu7VS5eFGtpeAW4Dz0O/uXSjXs9kzT6dWYWP+vFyHyu4ZYzqXoaRLob1O5c+Ch1Cd75NQJNTkllrfHgxCx34H6XP0InqWjyqy7o6o1nl0ndsprdi+JRLXy3//MpwuZowpwlpkF27/N+ncG9O57Ee2OOBZ2Fk3lbEpqu1ZSQdwJXKaG0kfqkn7ody276TyXMrQbQVKRfk9GvTYkcaV8diQQq3VShz0/wBvbtC+jTGtYxzxUmh3Ubx0brItJZ1PvkprzW87BqF6yvlogWh7Bp2nYuXihiAHfmlkneVo0LTUs31t4uHxN6IZeWOMSbEaCpdMPqCupnNqWZrK2QGV1Ute7zNCGmU6iiFo5mUl5WesF6KOZT372hP4JnAV2fduJ7fFaPb9R8A7Ufh6LVxB6TzT/GePoE6pMab9yZdCO55CPnmlEU2vki6F5hnbAsPQeYmKoubbkygHv1RN6U3RQHF0vYdRH6sUqxIfFHgQpWEZY3qQcrOkQ1CI6FsSy69COc1LmmGUCc7mKOQ1WS7lE8AvWm+O6VB2Af6Aws5LPWs+S3X6Fmuh6I/9UK5gvQKWc1E++cso7zL//5fQM24xhdxzKESdgBznIWjQckTu/2NybU2UDz4F5X1Ozv1dT2d4OepcX41yF/9bwTrTkFhRFgPo2jyPxOLOQ067MaZ9GILCpDdHDvX2SAx2UoXr5/PJ8+0hrDdRjFHAx4AvkhbPvB/4IXAhxZ+TfcDH0eBqPhR+AJUo/RzSLynGSPSs3j339wvAbmh23RjTg5Rz1H8KfDqx7Ao047K0KRY1h6NRfcssnkRCHo3gBIqrIt8J/KpB+2kFmyJnPVr3czkS/7oxiEWmExmDdA6OQjM9SRGzAZQrvhHZYjp5NgM+gGaPtq7BjldQWPljaIYk356ktc+yQcC6KDx/k1zbONfWpfoUk5lo4PQP6HeZPIdDUNrShonleQd9LnA6ipjppGe6Md3KaPRcyDvk2yOdiEpC0Feg51pU5O029PwzpZkAfAZVLUkOgNwKnIKetQMUZ10kzrt3ZNn/gCOReGopBiMdgENyf88D3oQGB4wxJsXHSYf7/JPSoT7typUUD/9aTmPy7NejdGjpHxuwj1azNZpFjB7HbPQyMqYaDkFOYbHfyAEZ66yBZiDuKrJOsfYamm3+NnAglc86hWYUCuP/InAx6RI+5dpzqFJDdCDjexnf60dCUqdSeyi9MaZ+JhDPJ6+mFNp8FF1zLoV8cucxV89klKr1KulzfDOVlxGdTjr16iIqD1s/O7LeYnQ9jTEmk22Ji18MoDCpVUMaVQelHPUBFHpbL98ss49OdNRBDlYy5+12OnPAxoRlCoXfYvKeykdpDAXehxztShXZZyMBtiPRzHs3CR+uiVThf0y2oGexdj/qfCYH2pYg5905j8a0lnw++UkoMvEpKv89z0Qh0dF88qwSi6ZyNkSDHEmhvX7gcpS6VQnjUSh8dBsvoed2pZwUWXcF2eVyjTEGUImgZBm2BRQPHe8Eyjnq9YYW9aEQ2m501CF7EOKrQS0ynUofCoNfTNpZPwWYQenfUb4jcxeaEd6D3uqwro/O30VkV2go1vrRYMaaLbfYmN4iWQrtWhR6Xo1TfgWFUmiV1jU3lbEVil5YTvy8r0TnffsqtrUfyiOPbucqqnvOJqNXj6liXWNMD3Ii6RdHp4/uZTnqyfCyemoFv7nMtjvdUe8D/kp6Zm6jkEaZjmZT0gOC5Zzza4Ej8GxwnmGoo3g+6ZnzrOfdlUhYzhjTGMaiwcKjKNQnr7Qc5DLSpdCK1eE29bMHcsSTA8RL0TXYuIptrYKud3Rb89B9UA0HEh8w+HaV6xtjeoy1UO5i9CH225AGNYgsR/2fib/rKT/228S2rs/YXyc76qAyfbOJH9NlQS0yncyGwJ8oX0roHuDzeCa4HKOAw4C/kZ4pSrY7iIsdGWPKsyYa6Koln/w14qXQtkfRi6b5TEPnPnlN5qPrsVbxVTPZlXTJ4luofuJiV6QAn9/GeXRX2pYxpgn8mvjDZzb1lz5qB7Ic9cMTf79CbS/O0aRns5Lb7gZHHaS6nTyuvYJaZDqNKUg0ZxnFO7Wvo4oTWwSysdPZFfgZ5cXo/gFsF8hGY9qVwcBU4vnkL1J59E8+dP1UCvnkdsBayyB0/ZJ1zAdQ/vhJVB+ZNTS3XlQ7ZTGqYV9tyc2pxCc+rsS6P8aYMqxLWkDu40EtahxZjvoU0rlF1Yh/5PlIYhsPobDebnTU+4CbiB/XNUEtMp3CaOBblA7Rfh11oiYGsrHbGIKE+W6n+DnvR6Xdpgay0ZiQJPPJbyYdVVisLUfv+4so5JNPbqn1JskwdC0fI329nkXREJWUuUuyJXBvYnv3A9vUsK1JxGfk/1OjTcaYHiNaGmIAeITqRwnblSxHfTVUyii67K81bPvfiW18ie511EF5Xslj2yGoRabd2Z/SInH3IodyCBosq0bMx1TGrsBfKJ5qsAj4Mp7VMd3LOOKl0O4iPTlRrC3IfT+aT27nqn0YC3yF7MiHB4EPUduzbRC63lFl+BUoWmJYDdsbicLk89t6msaUBzbGdDkjSdeQPDyoRY2lmKO+SWLZcqpTWN2AeMd3Oar93M2OOsANxI/tF0GtMe3KBFT+pljndwYS3+mWAcFOYEfgOopfk/vwQInpfKKl0C5CM9/l9DDybS7xfHKXQmtfpqBrPJf0dbwbXb9a3y8boLKh0W0+DexZ4/YGAZdEtvUK6oMaY0xZPkj8YfQiCgnrFoo56qCwo+jyz1Wx3e8k1r0it7zbHfWDSc82WK3WRHkfafHBfJuNStDUMiNhGsO+SKgv6/osA76Lxa5M+zMEOdLTKeSTv0RlDvkA6VJoTgHpDDZCAylZCvs3o2tZD4cTT9PqR4POo+vY5o8j21uMojKMMaYikqW3TgtrTsMp5ah/KrH8gQq3OYh0OO+7c591u6M+hMbk95vuYwRpUcpoxMnp1NfZMY1jEOqQvkz29boLWC+YdcbEGY2iPaL55FHV7FItWQptGrBqa803DWAP4HLS0RErUBWResUxp+S2H932LFRGrR4+GdlePxLmNcaYihhJWjyl25SASznq40i/7Cupqf62xDpzKMxAdbujDipnFz2+X4c1x7QBa1NcuOx+FHZt2o+JFE9ReAXYJ5xppkeZQDyfvJpSaPOQE38uhXzyEa013zSQvIJ7Vom1Jej+aEQI+XT0vItu/yLqH9A5kLhS/Bfr3J4xpsd4E/EH0//ovlIipRx1kBMd/aySmuoXJtb5SeSzXnDU9yZ+fE+HNccEZi+yhXyWUbvwjmktB5Bd0m0FCg12nq5pBtF88itQOHolDvlA7rvX4nzybiSv4P4w6ev+MnqvrNGA/YwjPVD5KhKgq5cdiU+EnduAbRpjeowvE39A/S6sOU2hnKO+b+KzcjXVxwELE+tEBZh6wVEfTlwJdYDqhPhM9/Bp4jMG+fYIsHlAu0z1jEPq8FlO0R/oLu0S01qSpdCuRZFolTjkK4CniOeT+33TnYxFkRDPk74Pnsl91ihNnLcBzyX28Q8UHVYvGxAfvHatdGNMTVxA/CF1dFhzmkI5R30Q6ZmkUjnXn0h8N5nX3guOOsAdxI9x37DmmAB8gWw15cuR02c6jz7UGV5O+rpeicOITXnGUghdPxeFLWcJf2W1ZD75HlistBdYHQ3CJCsQDaBqFIfTOEd3JJqRj6ZTLEL3WyMiSlcFHo1s+y58DxtjauRW4g/Et4Y1pymUc9QBTkl8Xqqm+m2J734+8XmvOOrnEz/GT4Q1x7SY48me+ToJh592A28mW7n/76ijawwodH0a8XzySkuhvUa8FNr2OE2m19gKDeaUUnBvZDrmTsSd6AFU/WfjBm1/BPF8+mdw9Icxpg6SYT8bhjWnKVTiqG9MuiZ61sO1ku/1iqN+EvFjPCWoNaaVJEsTDqB0kP1DGmUazvrAY6Sv9XV4hqjXGIzKl0XzybN0KYq1aCm06SgMvtv0cEzlTAOuIj2osxxpAG3T4P0NQYPLyxL7Oonaa60n6SMepfoasGWDtm2M6VHmEX9IdmO4aiWOOsAtie9k1VQ/NfGdyzK+0yuO+rHEj/GssOaYFpG87gNIMGfvkEaZpjEFqfYnr/lfceREtzKMeD75zaSrwxRry9Gs+kUU8skntdR6064MRYM0WdVBXkcz629own43R+Hn0f09SOMrHP0gsv2ldGeEqjGmxSTzELsx7KxSR/3jie8kc88Hk45AyMpl7xVH/SPEj/E3Yc0xLeCtpJ8ZrwG7hTTKNJ1JKE80+Vz7TkijTEMYT7oUWpY4ZFZbgBygaD650yJMkvFoNjtLIO5F4BuoTGSj6QOOIi7+248Gn0oJBtfCUYl9NEI13hhjWEr8odmNQkGVOupjSau5R0dcD0h8Vkwdvlcc9eiLaQD4ZVhzTJOZSrrO7GvADiGNqpGsMmT5dkGD9rEq6coI0fa1Bu2nVUwE7iF+DP3Ae0IaZaoiWgrtIqrLJ59LPJ/cpdBMOdZHUYhZAnGPo8GdZg3srAf8M7HPZ5H2RqN5O/EB7K80YR/GmB4lWR5l1bDmNIVKHXVIq+CfGfnsosRnxeqt94qj/jkqOx+m8xmNIkyi13sFGrzqREo56ovQDFC9fLbEPjrRUQeVLZpF/DgW0vh8UlMfQyiErp+KcsNfpjKHfIB4PvlBNKZetekdtkNRFlmVI5ohEJdkOhpYiu73fGBME/a1A/G0EE9YGGMaypPEH2bdKHxRjaM+LfG9/Kz5RNKzY8U6p73iqJ9O/BhPDGuOaSK/IH1PfzmoRfVRylEfQNEi9ZIVKt7pjjrArqSfhY/gkOdQjEFq6dF88kVU5pAnS6FNozkhyKb7GYQc8GvJvs8uAnZusg2TgEsT+54NvKNJ+5tKXFDxb7hWujGmwSQfqgeHNacpVOOoDwJmJL57CKovH112f4n99YqjfjHxY3ROVneyL+nw2AvpbMXmco76LXVuf9sy2+9kRx3gSNLHc1pIg3qECaTzyaO1oEu1eRRC14/KbacbU91MaxmOBokeIfueOxNYpwV27I8iQaL7/xvNiwYZSzzK7G4UeWaMMQ3lHOIPthPCmtMUqnHUIV166jLSiqHHlVi/Vxz1J4gfowXFuo9hpK9zN8yeJh3110mHaW5Sx/bPTGwrWV2j0x11UIhn9HiWI3Vl0xii+eRXkHZCSrWZaBA+mk/eyQNrpv1YHTiZtG7JAPA06iM1I9Q8yVikFp983jYiKqoYQ4HrI/t7FqeHGGOaxMeIP+CuDGtOU6jWUd+I+AxiUgF3GaXLzfSCoz6JdD35VYJaZJrBl4jfxytofvhiK0g66i+g2ZfospNr3PYw4KXEtrJSBzrdUR9LOvrouqAWdSZDSZdCW0BlDvkK4Cni+eRTWmq96TW2QY5xVnrF3eg+blX4926k0zdvRiHpzSQ6wTUf2LrJ+zPG9DBbEX/ILaD7wuGqddQB/p2xTr79pcy6veCoH0b6BW26i9GkBajOCmpR48hy1Kcnlj2PSjJWy6GJ7TwHvI/0M6HTHXVQWlDyuPYKalF7M45C6Pq5yKkoVRkg2pYSzyffAxjVWvNNj1Iq/3wlGiia1kJ7RiCRxGjax2JUAq7ZlQi+FtnnCuDAJu/PGNPj9JGubdmpSs7FqMVRT9YIj7Zyefy94Kj/ifjxnRLWHNMEjid+jefSPVUhshz1YaQHJvapYdtXJLbxHVTCrBsddYB/ET+ua8Ka0zbkQ9ePp5BPXmkptFdJl0KrZdDImHoYi6q7PEX6Hl0AnI0iEFvJVqSFOv9La2a1pxMfHPh0C/ZpjDH8nO52Kmtx1EeTHX44G4UqlqLbHfXxpOvN7xLUItNohiDnNXqNvxnUosaS5aiDOp7R5RdWud0pKDUmuo1N6G5HfQ/Sx7ZVUItay2AUahvNJ59NZQ75APFSaNNxPrkJz1Q0Y50scZa/X0+i9dUBhqBBr6URW5bn7BzWgv3vSLzfc3oL9mmMMQC8mfiDeAmlc7A7jVocddAL4K5E+2oF63W7o34c8WN7huaHm5nWchDp2ZMJQS1qLMUc9R0SyxdTXU31ZE7/v3PLu9lRB7iR+LH9JKw5TWMY6Xzy5KBlsbacQuj68eg3Vsl7yJhWsQcqo5ZV//wuWpt/HmUqcFPCnqeA3Vu0//WJl2G7Ake4GGNaSB/wOPGH4HeDWtRYanXUa6WbHfWhyDGPHtvXg1pkmkGy9N4vw5rTcIo56qAwyuhn1agH359Y96O55d3uqL+X+LG9QufXEx5PuhRaUli0WJuPHJtoPnmnV0ow3ckwFMnxH9L38QrklIaq6NKHnr+vR2zqR/oOrdJnSJZhu6uF+zbGmP/P54g/oF+je/JR7ag3jo+SnnFcPahFptEMR45G9DrvGtSixlPKUf9C4rNKa6rvlFjvddTJg+531IeTLtO0V1CLqiNZCi0rJ7dYm0s6n9wRRqbdmYwiO5IaRfn+35nAusGsU78i2XebSWs1lIYSF9B7Hlirhfs3xpj/zyqk8+q6JXzRjnpjGE26lu/ZQS0yzWAf0k5st+XMlnLUJ5POM9+0gm3+LLHO7yKfdbujDjre6PG1Yw7nEAqh66cipzwpIFiqRfPJD8K1k03nsR3wW7KrDTwEfILwpVanA3OI23YRrc+Ldxk2Y0xbkcyvXA68MahFjcGOemP4PvFjWkTYEXfTHE4ifp1/EdSa5lDKUQe4PPF5uaoGI0gLL+0d+bwXHPXkMd4c1hzGoJDzoyjkky+mMod8GelSaKNba74xDWMwxcur5X+rBxF+QHY88Hvitr0KfCCALS7DZoxpO4aRzlW/j9YoajYTO+r1szPp/MzvBLXINIuriF/nD4c1pymUc9ST9cHL1VRP1kl/lnj4cy846msTP77FlK+Q0SgmkM4nj5ZRKtXmUQhdPyq3neEtstuYZjIF+AbpCh4DSATxHGCzYNbF2Yd0GP7VKC2l1bwbl2EzxrQpB5N+oP84qEX1Y0e9PiYCTxA/nhl4hqlbeZr4te7GUlvlHPVhwEuJ7+xbYntXJ757UuLzXnDUIZ0a84Ym7COZTz6L9Lkt1vKh66dSyCcPPYtoTKPZDgmuLSL9G3gB/XbaRYNoJBok6yc+iHAcYX6bLsNmjGl7LiD9cP9YUIvq4wzSZdaqKblULetn7K9c6Gy7Mpj0DGs/sF9Io0zTGEK8NE8/4fMVm0E5Rx3UeYx+5w9FtrUW8WiTfmDDxHd6xVFPllB6Wx3bGkq6FFpU/blUW4EE4aL55JPrsMWYdiev3l4qvH067VWNYRfgMeJ23kpzBvgqYX3iZdj+jIUhjTFtyEQ0Yxp9eC6h+5SfTXnOIP3C/1lQi0wzWYP4tX4prDlNoxJHfdvEd4rVVP9a4ns3ZHynVxz1C4kf4xEVrjeOeOj6XWSLXWW1paTzybtxcMmYLKYg9fbkMy3/zDqf9ouKGooG0KIDnMtyy0LVJ3cZNmNMR7Et8fCffNjgeiGNMi3lKNIv/n/T+ZoFpjgbEb/eT4U1p2lU4qgD3Jv43icyvvNI4jtHZnynVxz1nxM/xqzcznzo+vEU8sn7SZ+frPYq6VJooTr2xoRkexTeniWQ+BT6fbVaJb0StgDuJm7vA6jPGYpkGbZncdlZY0wHcAjpDtQM1Jk33c3HSIsxPYvDR7udLYhf84fDmtM0KnXUP5v43q2Jz3dPfP46UhtP0iuO+o+IH+P3UbjtSSgMPVkCtFRLlkKb2rKjMKY9GYEGqO4g/XvpB65Bv5V2DNfuQxEv0UiZlWjQLbR4o8uwGWM6lu+QfiHMQh160518grSTvgiN4JvuZkM8ox5lVRRaHf1utKb6LxOf/abIdnrBUV8DRdxU6ohHQ17vBX4NHAu8CYWhGmPEhkgAMSlwOQAsQDPrWwazrjzro5SgqN3PoN96aL5CwablSH3eGGM6hkGk8w7trHcvnycdRbEUjdKb7md14tf+5bDmNI1KHXWASxPfzQtDjkSh2NHP9iqyjV5w1HemvFM+H4Wun0shn3xkCGONaXMGAdOAi0iXRh0AnqR9w9ujHI5+91Hbz6c9qsYcQnxS4lNhzTHG6pforAAAIABJREFUmNoYjGY7ki+KecC7AtplGscQNGKfvMZLgHcEtMu0lsFohjN6D7RDh6rRVOOovyPx3XxN9Q8llj9D8XJCveCojyQ9yPcg8XzydgzJNaadmIxmeZ8h/cxYAVyOqq60+29pMvBX4va/iEoAtwPJMmzfD2uOMcbURx9wFtl5UafS/i8NU5zJwD9JX9uF1FdeyXQmTxG/D94Y1pymUI2jPoR0re59gesTy04ssY1ecNRBg7fRY0yWqTPGZJMXh8uqfT4b9bPWD2VclRyKorGix3AxsFpIoyKsCTxHwbYrsSilMaYL6AN+THZI49+ACeFMMzWyOxJtSl7P14G3BrTLhONK4vfCR8Oa0xSqcdQBfpj4/k3EQyZXUroT3QuO+rqkB/raqWazMe1GXhwuWV0i3+5C1Vc6JT1kHBpsiB7Da+gY2oXRwH0U7LsHl2EzxnQZn0WiG8mXytMop8q0P8OAb5EOcx5AM6rbhDPNBOZE4vfDr8Oa0xSqddS3JLsjnW/Xl1m/Fxz1w4gf37/DmmNM27IxmiGfQ/q5kK993mnv4LeSfq5eC6wd0qgEg4DLKNg3E1gnqEXGGNMk9iQdDppvFyG1ZNOebItGkbOu3VW0vziNaS5vJX5PzKL7UluqddQhXfs32j5UZt1ecNQvIH583wtrjjFtxXDg/cC/SGs5DAAPAUfTeRUPRqBBh2iE0SIkdNdu740ziNu4U1hzjDGmuayF6gpndVxnAe8OZ5rJYCR6oWYpyFprwOQZhsIVo/fHnkEtajy1OOrHkP2sW0B5wb1ud9RHAHOJH98eQS0ypj3YCL1bZ5N+BqwArkCRiMWEKNuZHYFHiB/T7cAmIY0qwkeJpyq9M6w5xhjTGoaj/M0s528AuBrN4JpwDAY+TNo5iTopbw9mnWlH/kj8HvlNWHMaTi2O+kRUBSH5+/lVBet2u6P+AeLH9hIWZzK9yzD0m7+e7Nnz51CK0RqhDKyTIWjGfCmFY1qOBiSGBrSrGPsQT9f8YlhzjDGm9ZQKp+5H4fBvCGZd7zIN+C/FQ3adpmCy2J/4fbIQmBTUosZSi6MO8EHUQY22TStYr9sd9duIH9sPw5pjTBDWBk4ie/Z8JcrZnk5niyxuBtxJ/NgeRqr17chmwKsUbD0vrDnGGBOOYeglFR1ljbZlwNm0l7hItzKNdOc52p7Cqu6mOINJO7PfDWpRY6nVUa+VbnbU30J6YHazoBYZ0zoGo/ftRWRHFs5CM80bhDKwQfQh9fbXif/WzwVWCWhXKVYDnqBg742on2qMMT3NlqRrDCcd9j9iIY9GMxw4knjpkWRbDJyOy5GY8nyO+L0zD5gS1KLGYUe9MfQBNxM/riuDWmRMa1gLRdRkpZRFZ8/bMRS8WtYl3aebgQbp2pURwC0U7H0UlxA2xpgY0yitlDyA6oQeTmeHgoVmEuowPE/x87wSjfh3+qi+aR2roFzj6H30y6AWNQ476o3h/aSPa/egFhnTPIYChwB/J65yHp09P5nues9OJ11G7iLa2+ntA35Pwd5XcOqlMcZkMgh15p6ktMP+HJrp3TqMmR3HcKRaegmaJS91bi8FNg9jpulwjiM94LNXUIsagx31+hmPntvRY7osqEXGNIdNge9TPPf8GuBQumP2PM8k4M/Ej/Ul4F0hjaqQk4hHcLbzzL8xxrQFQ4GPo1qhpZzKAeABNEO8bhBL25dBwJtQTliyFFJWewI4OIilplsYSrr8zpOUL0fW7thRr5/ziR/PUjxrZbqHEWg2+VqyldvnAmcCG4YysInsh56J0eO9ClgzpFEV8h7i1+vwsOYYY0xn0Qfsi8q2Zb38kiPVtwEnoHqdvVjjewyaOT+H4uXVSrWlqHyUO9CmVt5M+rf6Zzqz7m8eO+r18SnSx/PtoBYZ0xh2QoPh88juk/wdeDfdKUo2Bh179Hk/D4nIdQK7EY8w7CYBVGOMaTmbo5fCAipzOmejvKPDgMkB7G0FfcBWwJeQeEsxBf1kexyFKW+NRvmT9Z5XAlegAQ9jquUnpO+5bwa1qD4+TbzE2meavL9NSZd127XJ+2wWe5B+Lv2X7nRcTG8wHjmjxUrMPk93KLeXYlfiCukDSIxto5BGVcH6xFMTLqY3J3eMMabhjAAOQgIly6h8tngmcj6PR53Hka02vAGMQ6J7J6FjeYXKj38OGujYg/Ts5nrIYV+Ysd7N6HwbUymrkO7E9iNhJdM7rA+8TPw+mA9sEdAmY2phEHp3nkv2e3IpeidPRyXYupWhqP8RLS23GPWrOuW4xwL3U7D/Ltq3ZJwxxnQ0k4BjKF0DvFSY9x3Aeegl8y5Uz7cdZnomALsARwCnIBG4Z6j+GF8HLgQOoDKl/Mm5/b2Wsa2bctvp5BBm0zrWJS2mtAB1dk33MwV4kHSkjnUwTCexHnAi8DTZ79gHUWnK1UIZ2EK2BO4lfvz3A28MaVSVDEYlIaPRD2sFtcgYY3qENYGPoBCmV6neqc235Sik62/Ar1He0rHA+5CC9WbopVzLjPz4nJ3bIqf3CODLwI+AC5AznCxxVW17FDgD5faPqMFGUO7ZcaQFYvIvZpfIM5WwG+m0ioUoKsR0L6uTdtIHgK+GNMqYCskLw11BfOY43xahiL5p9MbA9SDUH4g+y5ej8P52mNiohp8Sj+5x5SBjjAnAEGBPVKP0DqoLka/FsZ+LQsufirTncsuzZqcb2eYAl6M82kbnxA1HTvnjGft9Gr28ax0MML3BkaTvnSU4naJbWYfs58Ul9IZTYzqX7VEKWLIOeL7dhXLTx4QyMAAbADeSfvfvGdKoGvkchWNYgd9BxhjTNgxFL+HjUJmghyivIt+ObXnO9vNRh2ELWiOAMhQ57A9n2PQ88Hk6vwSXaR5fIdtZ74Qau6ZyNiY7RedKPKBn2pN1gK+TPbg0gLRtTkditr3G4cTFe/tRjn4nvuv3Jx4dcUxYc4wxxpRjMrAPemD/FNU/nUF7OPBLUH34S1DO+BGoDEzozm4fGoXO0gSYh0LhVg1mnWlnPkX6t9WP7plOESEyxXk72SlHV6DIHGPaheEUQtuXk/3+zQvDDQ1kY0imoCi96DmZBRwY0qg62IJ4ZOMvwppjjDGmHlZBOeTvBD6JykqdjWpB34RywOeSrfxarr2KXnj3A9eg8nE/QnnqR6C88ql0huOyB+rMJI/xdRQ+uHY400ybchQSFEveM38HJga0y9ROHxLjzLqu/0dvOjqmPcmHtherlvIQupcnhTKwDZhO+vxcROcOwK8B/I/CsVyN9XWMMaanGIJU2iciJzvf1kaz9PkXxIdCGdhkdgb+Snq2dAkaue6UuqqmNXyYbM2Ip9AAmekcVgOuItvpOY/OGHA03c0GwAlogD3rPp0F/AApmvcy41BYe3JioZP7LSOB/xAfiBkf1CJjjDFtxdspvCTup7vFlLZAufPJUMKVaOZ9x3CmmTZjd5T7mew055WEHSrd/hxEdlWI5WhW0phQjEf51deSncoWrXnuiA94GxK/jZ6jf9DZZcv6gD9SOJ6XgQ2DWmSMMabt6EOjuPmXRS+UpdoA+BmwmHQ+8t9wHW0jJgH/JHuW6wngzcEsM6VYHWlnZF23l4C3hDPN9DDDkTjlX0iXhMy3u5AmTaeGcTeakWhgNJq2sgiJ7nb6pMKpFI5pMbBrWHOMMca0Kx8nnovbK0wCTkL5/MkO081oRq7TOwOmPoYh/YesTvVKlE/qUMX2YDDSGMgSjMv/ptcIZp3pVfJ55y+RfV8+l/t8m1AGtik7kU4H+A+q3NDpHEl8guADQa0xxhjT1gxHeXD5F8fWYc1pOWPQCH1WmOz9KETR4i69zduJC/5E2xwUSj0ymHVmGnAf2ddnGZq9GhbMOtNrrIeeCU+QfU++htKwDsI6CUmGoHMX1QlZhgbVu+Fc7Uk8ouKEsOYYY4zpBE4kLrLUi4wEjgaeJd2xehLN1jk3uXcZh2a+stTDB4Dn0T3iQZ3WsStwI9nXYwC4FdgsmHWml1gdOJa4OFhywOhylHceuoxpu7I5Cv+PnrcHge1CGtVApqJc9Pyx/QlH7RljjKmAiahs2QAa7e3lENFBqDMVzd3Pt9loZN/hzr3LWyg+UzYAPAx8BA/qNJNpqIxRsWuwAOX6DgploOkJ8qJwxeqdD1AoqTY5kI2dQB8a5IyWlO1HA6Pd8hydCDxGPBWnW47NGGNMC/g5hZfIdwPb0g70odDE20h3vuahTsTqwawzIRkJfIXi+dADwItoUGdKGBO7juFoAOR+ip/zFcCvgXUC2Wi6nxHovXA+cccy2v6H0i26IZ+62axHWrTzWbpLrHMocD2F43saD9wYY4ypkjdQCOudA4wOa05bsQeaNcmqxX4udgx6lYnA6UiJuJjzuAQ5jzsEsrHTWRv4Jhr4KHaOB4C/otBZYxrNYBTFcT4wn+z77xX0LtgDhzNXynTSYq7nI92YbiJa/30esGVYc4wxxnQqf6XwQvlMYFvakW1QR2IF6fzD83E+bK+yDtJ2KBb+mm+PAN9AJQJNccYDHwX+RXFNgHy7EZc2Mo1nEHK6z0QpT1n3XlQUzvXOK2cycCnptLJ3hDSqSXyVwjEuRzXhjTHGmJrYk3h4VjeorDaDTYHfEFemHUBOxSWoJI/pPdYDfoA68KWcy37gFuDTOH0izyjgEODPqK5wqfO3ArgI2C2IpaZb6UP31I+QOGTWvbcQiYC9A+cY18L+wEzi5/QSYLWQRjWJQ4gPNH4qrDnGGGO6gahq7bsC29LurI5yEbNyFV2LvXcZjcSRHqG0w5lvD6H7aBq9VUZsKjpPV1DeOR9AYcdnAusHsNV0L1sgPYknKT4wdC0Sjuu2sOxWMZZ4CHg+DPyokEY1kR2J9wt+GNYcY4wx3cJ7iTubpjyroY7eHNKdvMdRnXbX2e49BqGZt78CS6nMaX8VuBjNvmxLd5V72xA4DAlXPktl52MADR5+BjtJpnGUc85XovffcVj4q152I32er6N7tV3WIh6R8TccnWiMMaZBDAaeovCScf5n5YwBvkR22ORMpBQ+IZh1JiQTgU8CN5EWJSzVFubW+QESX1q/xXbXyqrAvsCJwJXE6wdX0p5AjtQbWmy36U7yYe0/RqrsxZzzf6PSfmuGMbOrGIEihaLh34tRubpuLZ04lnhlinuwMK8xxpgG81kKL5qLA9vSiQxDYZL/Jd0ZXACcQec4XKbxrA98Dc0UlxNLy2qvow7gn4BvoVnqHZAIWysZgQQU34k63+ehWchqnfJ8exL9NnZp5UGYrmUQBef8Oeyct5KtgPuIn+v/AluHNKrJDAYuJz44361RA8YYUzXOBW4cY9Csw3jUkdkEzbKb6tkDOTFvJ36P9gNXAaegWu2mN1kN2AfYD81A1xtmuwR4CXUSX0ZqyrNy/1+KBBAXot/1/Nw6r6FZnyEoRWMEGmwahTqfk3JtDVQXfhJyauoNRV8M3AD8HbgazaIbUw+DURTYdOBQFIacpB89cy/OtZkts677GQJ8Afg2Bb2NFShH+0T0/OlWzgKOzv1/EfAW4I5w5hhjjOlmTqMwMvyTwLZ0A29EpXySSvEDyFk5BOex9TqD0Mz48cBfgBeobWa6Xdt84Ho0OLUv1m0wjWEEcDCqxPEKnjkPxVSUphM9708Bu4c0qkVEoxBX0p2l5owxxrQRa1EQwFqIck5N/awNnE52Ca9nUY6789hNnnXQzOD3kaPxKuEd7kraEuBepPT8ESTe1a15qab1jAXeh8r0LSD7HlyBBkHtnDeXPqTe/jqFc9+PfvujAtrVKvZH91r+2D8X1hxjjDG9wu8pvHy+GtiWbmMs8HlgBukO5kLgHOTcGJNkMvAm4GNo0Ocy4FEUbtlKZ3wZSpG5FvgpcCyaKd8AO+Wm8awGfBSJEy4h+55cilKKPo7V2lvB6uh6RK/BTOCAkEa1kC2JD7r/Mqw5xhjTvjhHvfFsjQRh+lCu6/qog2QaxxAkxnUssGfiswEUKvwTVOKlv7WmmQ5kNMoln0whl3wSiogZTjz3fGxunfFoNmwFyhtfQjyXfS7Kcd8LlYwbi0TkXmrFAZmeZiMU1n4Qej5mpQctRDoHf0FO+ryWWdfbTEcDyhMjyy5G1S3mBrGotawB3E5BMO4fwIHoOWqMMca0hOsojBYfEdiWbmcbFC64kPRM0VMod3li0bWNaS7RZ4HLNppmMAjYHpXmu4vi0RxzUdj74bj8VasZTzzabgCl5HwgpFEtZiRy0vPH/xCtr7phjDHGsD+Fl9EDOHKhFUwGvkG2mNh8FGa8VTDrTK8SFUw6ObAtpnsYg3QYfkvp0n4vAD8DpqFIJNN69gGeJ35drqa3NAAGAZdSOP6XgQ2DWmSMMaZn6SNeD3yfsOb0FINRyOe1ZHdc70IiPr0g2GPCM5XCvfffwLaYzmYKmg2/guL55vmZylNRmUvrHoRjJHAmSr+KaqkcR+8N3v+QwjlYDOwS1hxjjDG9zkcovJj+EdiWXmVH4AIKSvzJMNAzgM2DWWd6hUco3HfrhzXFdBBDkLN9CqoGUMwxX4reMUcD6wWx1CTZBXic+HW6FXhDSKMC8VHiyvbvD2uOMcYYIxGqmRReUG8Ma05PMwXlqj9J6Vn2VUIZaLqa0ynca58JbItpbyYhwbHz0WBiMed8DoV883FBLDVZDEVaAdHSY8tyy7JE/bqdfYDlFM7FV8KaY4wxxhT4BoUX1G8C22IUBroP8GfinYd8ewX4AbBxKANNV7IXhXvsqsC2mPZiMBIZ/A4aMIyGSSfbo2jQ5030ptPX7mwB3EP8mj2Aqj70Ipshwbz8ufh1WHOMMcaYOBNRCaf8qPraYc0xEdZEAynPku4Q9wP/Qor9Vkc29TIYDQINoNziMWHNMYGZBHwQ+AOF+yKrLUIDO0dj4a12pg/lnUd1A1ai/PThAe0KyWrEI9huRGUujTHGmLbipxReVqcEtsWkGYQUkS8ie5Z9ce6zg7BqsqmdCyncU+8MbItpLUOIl09bSXHn/GlUbnI6HtDpBNYHbiB+DZ9BUQ+9ygiUj58/H48AE4JaZIwxxhRhKoV8tbl4hradWRt1pp8juxP9PzTYsmkg+0znchiF++iXgW0xzaUP2Br4AirDtZDijvliJAR3HE656TQOR6U/o9fzfHr7Hd+HBrbz58Nl2IwxxrQ9f6Hw4jomsC2mPINRLvsFFO9k/wf4NEpvMKYcEyhEbMyk98ozdTtrAB9CjtosijvmA8ATKNLqQCxg2YlMBi4jfk1fRFFXvc6pxKsRvDmoNcYYY0wF7E48tNFCQJ3DWFRq70ayhZ6WAJcAB+McPFOaGyjcNzuGNcXUyXj0mz8DuJ/Sjvlc9Iw4CtgghLGmYRyKZomj1/dilJPd6xxFXOflsLDmGGOMMZVzG4WX2LsD22JqYx1U5i1ZHzffXkUzagfRuyJCpjhfonCvnBTWFFMlo5CWxanAzUgctJhjvgLlop+aW2doAHtNYxmHtAOi1/k15Jwa2Je4xss3wppjjDHGVMd0Ci+xOwLbYupne6TqW0y1Oe+0T8fhrUZsSuH+uDuwLaY0I4E90MDctSiMt9Ss+VMUROBc17y7mIb0SaLX+1pcxSXPFmjQIn9uXIbNGGNMxzGYeLmS3cKaYxrESOD9KGcxWp4nOfNyPgqV9Ux7b5OPxujHHf12YjTwNuBbKM2l2G85quz9a5SX7uvYnYxAURFRlf5FaPBmUEC72ok1gBkUzs8NOAXMGGNMh3IshRfanwPbYhrPKijs/XzgdbI7+AuBK5Bi8KgwZpqA/JjCveCw2XBMRr/VfCh7uRnzWUjN+ig0g2i6mx1RWbHoPXA7sElIo9qM0cA9FM7PQ7gMmzHGmA5mFIVQ6ZXARmHNMU1kDBLTuRSVYMrq/M9HAzYfBdYMY6ZpMW+lcP0vD2xLL7EhcATwK+BRSjvleRXvPwGfwGXTeokhaMY8OnCzHA3oWGugwGDiyvezgPWCWmSMMcY0gO9ReLmdHdgW0xrGoPD4P6PQySzHoB/lLX8X2BVXBuhWhiL9gnwYrfULGk8+v/wLSG19JuUd82eB3yPHfPOWW2zagc2AO4nfFw8hPRIT5yzi6QC7hDXHGGOMaQxrUhitX4jLuvQao4H3opI+CyjuOLyM6rgfBqwaxFLTLC6icJ3fHtiWbmBD4APAT5BQZylF9nw00/2olvlhOMe81+lDKQ3RdKV+JA7ogbQ0XyT+W3pnWHOMMcaYxvJbCi+6r4c1xQRkOBKw+jHwGMUdixXALcDXgG1Rx9J0LodTuLY/D2xLpzEO2Bv9Fi4HZlN+tnwJykM/FTgQ59GaAusC1xO/X2YAbwlpVBtzKHFxvWPDmmOMMcY0ni3RiP0A6miOCGuOaROmopmdKyie1z4AvIRmZY9DYZl23DuLVdHgywDwAr5+xRiNQtiPQwKNDxF3Eoq1meg3dBIqrTWyxXabzmA6MIf4vXMRHsgpxg4oCjB/rs4Ia44xxhjTPK6h8ML7cGBbTPsxCpVzO4d0Dd9kew74HRLLmhrCWFM1t1C4ftsEtqUdGA3sCXwOpXw8QmVO+SLgJuD7wCHAWq023HQck5BeSHLw810hjWpzNkACi/nzdSXWUTHGGNPF7EvhpfcAnlUzpdkKqRH/i/I1nl9AitXHICfQHar246sUrtc3AtvSatZFpdG+jmYwH6Myp3wlmlU/H93bO2IlblMd+6HnY/S+ugpX3SjFROKVEu7CpUWNMcb0APdRePntF9gW0zmMRGW+TgZuReWDSjk481Bn9JvA/licrh3YisL1+U9gW5rFcJSa8REUJvsvYC7lHfK8U/4Iml3/LJptH91a800XMQaJwyWfi0eFNKoDGApcR+GcPQOsHtQiY4wxpkUcSeEFeG1YU0wHMwaph38fOe7lVK8HgCeAC1H+765Y3TgEz1BwSju58zsS2A4pqJ8M/AXNkpcbQMq3FWjG7g/A54G90D1tTCPYFT3vovfcLcBGIY3qAPpQ9Ep0YGOroBYZY4wxLWQoyi/Ovwi3DWuO6RJWAd4MnABcDcynMmfpIeS8fxGJcE1ssd29xtl0lk7FRGBnZOtpKE/1KSoLW8+311BO+dnAx4Gd8CCRaQ7DkdJ/XrhxAIl0Hg8MCmhXp/BtCudtGapQYowxxvQUX6PwMjw/sC2mOxkMvBGFeZ6HNBGinddSbQZyyE5DYnU74BDkRrEfhfP858C25BmHrvH70EDP71Fo/itU7oznB36eQMf1TVRreYMWHofpbbYE7iV+T96PnoOmPO+nUJlmAEX/GWOMaTEWMAvPBKTqPRqFim6IZtmNaSajkUO2ExLl2hYpxlfyTBgAnkU5xA8jh+zJXMtHiJjyDAdeRmHerwOrAUubvM9hSMxtfWC9yL8bolDgyVVubzm67g+j++EhFMb+KBI9NKaVDEJCg6ejex00aPRD4EQ0M2xK8zbgbxSEGr+Dzp0xxpgWY0e9PTgLODr3/9OArwS0xfQuY9GM07ZILX5bYAuqU9ZeQsFpfxI59DNy//4PheGbApei2WZQJYhr6thWHzAFKVivlWtrI2c839agtrDfVykMyDxKfJBmeR02G9MoNgB+C7wpsuxpNBt8UwB7OpGtgH8D43N//x+aXffgqzHGBMCOenuwAerwDkaOzDrYoTHtwVBgY2DzXNsi9+/G1FYa6zXksM/ItdnA86iO8Qu5v2fTOx3DjwK/yv3/LODYjO+MRjPdk9Gs+2pIfC7vkK+BnhlTqK9c2Xz0HMo75E8Aj+f+/0od2zWm2RwO/JRCWs4A8EvgCyhaxZRnbeC23L8AN6D0nGZH+RhjjCmCHfX24RLg0Nz/PwucGdAWY8oxFIVKb5r79w25fzdCHb16ni0rkOP+IgoNn5trczL+Px9YmPt3HsqrbEeGIydifK5NyP27DgrL7QMWAJchRzzqmI9okA2LUGRDvs2I/PsMOufGdBJTkEN+UGTZi0io8MogFnUm41DUQV7V/SFgDzSwaowxJhB21NuHHYE7cv9/Fjk+KxLfmQIMQTOPxrQrIyk47euiiJF1I21SE/e9GM2gLUCdzJXIgQc59MuQM59cVgmrIIc7z3gKz9DRKCd2fO7fUYllzWYxikyYhXQCXsz9/QIFZ9yOuOkmpgM/B1aNLLsY+BQayDOVMQKl3OyZ+/sFVNLOWjnGGBMYO+rtxS3Abrn/vxe4KPH5F4D7gOtbaZQxDWYkypdeG4Vt59sUFMo9Ofdvr6vLL0Ih5y/m/n059+9sChEHzwMzUQ65Mb3AOCQWd1Rk2WtIRO6CIBZ1LoNQHvq7/1979x0uSVUmfvx7JzEBhiEMA0OSjEjOgooiKCKooKCiouJiWgUWRUyrrKiLoCK6BsxixiwKuCCigCAZFgQEJAkMaWAIk2fu74+3+9ddVd03dt9zq/r7eZ56bt+Ob1V1qLfOOe+p/f8kkbDfmCwiSdL/Z6I+vhxKY5qmq4lW9mY3EC0IXxvLoKREphNzd6/V9HftpsuziIrps4gW7Bm5/1cpPuWYWUiM7VxAtNg/VbvuiaZlQe3v6sRUaBAn4l5JJOQLxzZkadzbH/g2jXHUEK3BR2FPs5E4nRhqB/E9dSA2BEjSuGGiPr5MAG4jugxDnNm+tHa53jX+dOD4sQ9NKp1JROIOkQxPIJL36bXr1hjgsVOJln+IIShPNd32FI1hKctoFKtqTsqHo48osLcB0VV/XSzeJjWbBnwcOIHGrAWLgA8BX6R3ik920vuB02qX+4mCfPZIkCRpAO8hfjT7iamb6r5Su+6cFEFJ6qqv0fjcvylxLNJ4sgcxJWB/03I5MfOERua1xEnB+vb05L8kSUMwnWhN6yeKXm1NtO4tqF13e7rQJHXJwTQOmn+SOBZpPJgEnEj0Uql/NpYCJxFTmWpk9gEW09imX0kbjiRJ49O7gTfQ6GJb9ymyP6Kva/p/GXEAI6k6phE/qGx8AAAgAElEQVQV6PuJcetT0oYjJbUNUaeluRX9JmDnlEFVwHOIaS3r2/RsGkMJJElSkxnANURr+deAPWvXzyHG3/UTB+8XE2Ni6z+um+efSFLpnUPjM75v4likFPqIau71k1b1nmVnkLY4ZBXMJaZprG/XvxC99SRJUhvrE3Mf1388bwM+QHR/bT5QaW5ZeFmSSCV10ztpfMY/lzgWaaxtDFxE9rfuLuCFCWOqipnEjBL17XozMXuGJEkaxM7EVEzNSXlzoZf8ckyaMCV10Vwan/87EscijaXDyHbJ7gfOojFrg0ZuMjGFXX273k+cFJEkSUN0CHGQ3tx6nm9JryfwX0oUo6TuupbGZ32rxLFI3bYOMbtJ82/cQ8ArUwZVIX3ECY/6tn0S2DFpRJIkldQHaN+K3pyo/yFVgJK66hM0PuvvSxyL1E0HAg+Q/X37ObB2yqAq5jSyFfNfkjYcSZLK7UwGT9bvThWcpK7ancbn/E+JY5G6YSbF37kFRBE5dU5zzYuVwJvThiNJUvlNJgrqtOr23tyqbgVcqXom0CguuQxYI204UkftTdRfaP49uxDYMGVQFXQY2To3H0wbjiRJ1bEmcTDTPCVbftk6WXSSuulbND7nr0sci9QJU4FTyCaPi4ATcR7vTnsRsJjGdv5a2nAkSaqezYgquO2S9VekC01SFx1C43P+g8SxSKO1PdmpwfqBK/FkczfsShSMq2/nXwMTk0YkSVJFvYAoANNqqjYLTUnVNINobewHHgMmpQ1HGpFJRIv5Ehq/W8uIlvUpCeOqqs2BeWRrXExNGpEkSRX3VopJ+krsziZV2fk0Pu/PTxyLNFybApeQ/d26BdgtZVAVNhe4i8a2vgGYlTQiSZJ6RPMUK/XlL0kjktRN76HxWf9M4likoeojqrc/TfbE8plETxF13upkhxbcAaybNCJJknrIBOA3FKezkVRNG9H4rN+cOBZpKNYFfkf2d+oBYr50dcc0sj0X7gc2SRqRJEk9aDrRdbD5IMgWCqm6/o/GZ33zxLFIAzmMqKfQ/Pt0NjGDibpjIvBLGtv7CWDHpBFJktTDNiB+jOs/zDukDUdSF/03jc/6MYljkVqZBXyfbIL+OPCGlEH1gD7g2zS2+UKsZSFJldOXOgAN287ApUSXt9cAv2i6bQYwh+iCOLv2d07t8jRgZu1+qxPd6WeQrb77eO3v00R13iXEAcCjwMPAI8CDTZcf7uiaSWq2N/FZB7gAeEnCWKS8lwLfAtZvuu4PwFFEl3d1z2dpzPyyAjicaF2XJEkJrQacQBTouRD4MXAN2blTx2pZRhSuORc4HXgnsC/R8i9pdCYSJ8T6iZNmq6UNRwLipO8ZxG9Q/bfgGeBYPPk/Fj5ItlDf29KGI0lSb1oVeCExF+2vgPsY+2R8pMtTRGvg54jxixt1dtNIPeEsGp+pVyeORdoT+AfZ7/q/Yg2FsfImsidITkgbjiRJvWMOcCTwdWIe1OWkT7g7uTxAnHA4kejCb+uLNLDDaXx+vpM4FvWuycBJZH+Tltaum5gsqt5yMNGLrb79v5g2HElSt5kopTUJeC5wQG3ZidHtkyU0xpE/VLv8ADGW/GmilRsaBemeIQ626mbVXn9V4sBsldrl2cA6ZMe8r0eMdR+Nh4gxjecRY3AfG+XzSVUzk/gcT6n9XZdoUZPGynOIgnE7NV13E9G6e32SiHrPc4mhbtNr//+AOKnfnywiSZIqaCrRhfVsonjbcFullwK3EvOqnwocDexDJNBjbToxHczhwEeJg7krGdl6LQeuAD4AbDiWKyGNc3+k8TnZM3Es6h19xLjzxTTefyuI8emrJIyr12wHzKexD84hTvJLkqQOmEAk099geEnsSuDvwHeBdxEtGmX5gd6AOCFxGvAXokV/qOu9AvgTUSRntK32Utn9B43PxicTx6Le8CzgYrLfy/8EXpAupJ60KdErrr4PLidma5EkSaP0LGIu5HsZeoL6N+C/iKmYqpSkTiJa399FTCm3gKFtk0XAz4ipgByqoV60GY3Pg12N1W1HUpxF5CxiGJTGzrrA7TT2wY3AGkkjkiSpAnYhDmyaC7+0Wx4husG/nRj33SsmEtvpJOBqspVs2y3/ILpiTi8+nVRpt9L4HDwrbSiqqDnEkKrm79x5RBEzja1ZwLU09sO9OCRMkqQRmwwcQYzRHizhnAd8AdiD6BavOElxDDFOfSgnNz4FzE0SqTT2TqPx/n934lhUPa8mvlebv2d/BqyVMqgeNZPsccRDwBZJI5IkqaQmEPOE38HAyeVCouX8YCKpV3sbE1O43cbA23QpcCbRRVCqshfSeN//vs19thqzaFQVqxPfoc3fq08QPbw09qaRrQ3wBDGVqSRJGqaXEWNGB0ombwCOwgIwI/Vc4EdEUt5uGz8JfAzHUKq6JgKPEu/3xRTf67OAb411UCq1/SjWT7mAKASqsTeFOAlX3xdPA89LGpEkSSW0O1GVfKCicOcAL04VYAVtAJxCzLU+0JCC92CPBVXTj2i811+Zu+1U4LdjHpHKaBrxXbqCbI+vE3EoVioTiR53zfvjhSkDkiSpbFYDvkT2AKd5WQJ8BdgyVYA9YAZROf4u2ifs12F3QVXPG2i8x7/edP2GRCv7uSmCUqnsBtxC9vvybzhsIqUJwA/JHke8PGlEkiSVzAHA3bRODFcSZ8M3TxVcD5pMjKN8kNb7ZBlwBg45UPlMoHWX1zVpzCTxAI3pCr9Tu+78MYlOZTSJaDFvHkK0jGhZtwdSOn3A12jsk+XA4UkjkiSpRNYCvk/71tvziPnBlcZqxLzzT9F6//wD2CdZdNLIvAP4K/D83PV/pvHe3hXYjkYPnwvGMkCVxrOBq8h+L95MTI+ptE4le8L/bWnDkSSpPPYA7qN1Ang7jkEfT+YQc9e3qxnwMRotkNJ410ck5SuBX9AYTvMBGu/rk4jiU8tr97tozKPUeNZH9Dp6mmwyeCYwPWFcCieT3S/vTBuOJEnlcSRR0MUu1eUy0BCF3wNrJItMGp6taHRVXg58lWhhr7+f89MX/jlNmBqHNgL+SPb9cTfwooQxqeE4svvmA2nDkSSpHFYhCjW1SvRuJIrxaHybTrGqcXNPiO3ShSYNy0fJtrotJDvzwfKmy5cmilHjy2HAfLLfe2fjScrx4t1k983H04YjSVI5rAlcQeticadi0Z2yeRHwEMX9+TTwsoRxSUM1hRhP3JyQt1uuSBSjxofZxDCJ5vfEw8AhKYNSxpFkTyB/IW04kiSVw2zgeooHv08RLRQqp/WJolz5/boEODRhXNJQ7UmcLBwsUb86VYBK7gDgfrLvh3OB9VIGpYxDacza0A98C+umSJI0qDlEt/ZWFcO3TRiXOmMVoq5Afv8uB96YMC5pqL7I4Mn6tcmiUyqrEcXhmt8HC4gicho/XgIsprGPziKmYZQkSQPYgGJRpn6iEM/MhHGp895Fcdy6ybrKYAZwDwN3gb8hWXRK4blEzY3m98BlwGYpg1LBvsAiGvvoV8S89pIkaQCr0rol/TxgWsK41D1HkO1+2E8k7wemDEoagpcxcIv6zelC0xiaShTLbD5pswg4EVtpx5vnEsPn6vvp90TdCUmSNIA+ohJu/mD3d8SBkKrrcBrTXjV3F90mZVDSEPyE1rMZ9AO3JoxLY2Nb4Dqy+/1GYIeUQamlnclW378Up3WVJGlIPkHxQPcXlK+y+2sYuJWtU9PJ/dcgr1O2HgiHUuxGfAuwesqgpEGsTRz8t5t6UNU0kWgxX0Jjfy8jWtZtoR1/dgUep7GvLid68EmSpEG8mmJhpqsoX7IJgyfq/9OB1+gD7hzkdcq47Y6nuB6/x+6jGt/eSuvP4J0pg1LXbAL8heK+fl7KoNTWjsCjZIs8rpk0IkmSSmJ9st3R+oF5wIYpgxqFwRL1x4iq56Ox7yCvUdZEHeAbFNfluKQRSQPrAy6k2Kp+T8qg1HF9RPX25jHOK4kq77bOjk87kE3Sr8MkXZKkITuP7MHtYmCvpBGNzmCJej+jnwf++0N4jbIm6qsQ3RKb1+UZYIuUQUmD2ARYSPZ9+0DttmnAGsRJyU2BrYBdast+tWX32v/b1O6zUe0xJoDjwxzgt2T374PAy1MGpQFtDzxCY39dD6yVNCJJkkqkVVJ7TNKIRq/VOj2R+//3o3j+mUTiOtDzlzlRh9a9LM5LGpFU1Eck1PsB7wAuIPueXUG29XWky0LgLuIE1m+IXiefAN5LzI6wOU4v1U2HkW2V7ScKn5r0jV9bEydS6vvrBqKehCRJGoKpwD/JHvz8ifKPR26VqH+d7Bj85cDcET7/v+We+ybgry1es8yJOsCbKa6TU7YphT6iFfwI4GTgp0QX2vwJs5TLEuDvxJzQnwGOIsbmmsCP3OpEt/bm7fw48MaUQWlQW5FN0m8hekRIkqQhei/ZA6ClxA9s2bVK1D9ATAWTv24k8s9zPNVM1KHYQnk9kTRJ3TSTKAx2InAO2e6zZVueJr4zzgCOJLrVa3D7A/eR3ZZ/IHr7aPx6NlHjpvlE9jpJI5IkqWSmAPeSPQj6QtKIOqddop5vCb95BM+9BdmW+WXAulQ3Ud+RYoEux4Sq06YBLwVOJ1qlu9HaPR/4F1Ed/Fbg6tpyQW35W+3/m2v3uaf2mPy4904s9wLfJL6rZnVg+1XJNGKKtebvnYXAsXiScLzbmqgLUd9vtwLrJY1IkqQSegPZA8eFVKdrWrtEvdXY8t2H+dyfzj3+N7Xrq5qoA/yM7Hr9MW04qogtiHoY5zK6ZHg+cAXwPaJb/DFEy/U8Ynz6Gh2IdVVgS6KV/xDg34GTiLHqF5NtQRzusoyYauzDwM70djK6B3Ab2e1zObHtNb5tBzxEY7/dQpzEliRJw3QR2YOhL6UNp6PaJepQrNb+5WE87wSKvRAOqd1W5UR9J7LrtZIoniUN11zgfcT48uEmtI8D5xOtrUcRSfPsAV5rX6LI41hZHdiNOAl6MvBrsuN0h7rcSRSqq8IwpKGaRAxzWEpjOywlToZMTBeWhmgHssNTbmXkNWAkSeppG5Htvt1PTElUFQMl6i/OXf8EQ0+oX5p77KM05mOvcqIOxfX7RNpwVCLTiKrd5xCtx0NJVpcTXdDPIubNfg4jK3L51VHG3glzgYOJpPMChlcA72Yiga1yy+Q2wDVk1/smoneBxr8dySbpt2B3d0mSRuw9FLsWVslAiXofxUr3hw/xeX+Se1zzmP6qJ+pvI7tu16UNRyXwHGK2haF2a78b+BrwKmC1DsUwHsd/r0KcMDyNSEiHeuLi50QvgqroI07CNJ+4WEkU3VtlgMdp/NiJ7LR5f6faJ5UkSeq688geBJ6QNpyOGyhRh2gNbr5tKHOqr04x4dix6faqJ+prE8lC8wG11ZeV1wccQFTnzvfaaZV8XgD8B1EpuldtCBwN/BJYxOBJ+1VE9/rJKYLtkI2JqUCb1+suYJ+UQWlYdgYeo7H/rsd50iVJGpUJxFjP5gOkqh0kD5aob0JxTvUNBnnOd+We78bc7VVP1AEuI7t+h6UNR+PIJGJWhaFUbL8aOA5b3lpZHXgrUbAxP9tCfvkX8EGi0F2ZHEYUAGxel7PoXC8Kdd8uZJP06zBJlyRp1LYme4A0n+pVGR4sUQf48yC35/0td//jcrf3QqJ+Gtn1+1zacDQO9BGft3yl7lbd2k8mvn80NOsD7ydOCg60becRVejHewv7OsCvyMb+EPDKlEFp2PYGFpA98daJmRUkSep5ryJ7oHRB2nC6YiiJ+ltzt986wPNtk7vvUorVpnshUX8twx8yoOraC7iEwVvPjyRa3DVyzwPOJjv8pNXJkLczPqukH0h2fu1+Ysy9rbDlsg/wJI19eA2wZtKIJEmqkOPJHiyNh6rInTaURH0GMcdy8332aPN8+ZbkX7a4Ty8k6ruQXb+b04ajRDYl5j5vlzAuBX5ETFWmztqcmEoz/93VvFxD+++ysTYTOJNsfAuIEwoql4PI1lD4KzFUQ5IkdchnyB40fThtOF0xlEQd4Lu5+3ylxX0mUWwJatVVsxcS9dlk129+2nA0xiYSJ/raTS+2jKjyvmGqAHvILGIowdO03hfLgdOJE5Kp7A3ckYvrQnx/lNErgcU09uOfsaaAJEkd9z9kD5zekzacrhhqov7C3H1azal+UO4+D9F6LGgvJOqrUGw5VW/YFriC9q24FwDbJYuud60NnEI2iWpe/gnsP8YxTa3F1FwMbxExH/yEMY5Fo3cEcRKuvi/PpXq/bZIkjQvfJHsgd3TacLpiqIl6H8UWn9fm7vPz3O3tCqj1QqIOxSm3xuN4WHXOBOA/iZMyrRLBixk/3ax72ebAj2k9Jd5K4BuMzffR9sANude/EosIltU7yJ5w+S3OcS9JUtfkW9Tfmzacrhhqog5wUu5+5zbdthbFlqod2jxPLyTqU8mu35K04ajLZlKs0l1fHifGGVdtxoiyewHtK/BfT9QX6IZJRIv5kqbXW0a0rE/p0muqu04ge+LnR1gUUpKkrjqF7MHbR9KG0xXDSdSfRbbFYAWNOdWPyT3HNQO8Zi8k6uuQXb/H0oajLtqeYm+T+nIOMXWYxqdpxPd8qwrxC+j8dGibUqz+fwuwa4dfR2PnRLL780wctiBJUtcdR/EHuGqGk6gDXJS774m166/NXT9Q74NeSNR3I7t+/5c2HHXJEbQuGDePmN5R5bAn8HeK+3EF8FFG3xuij+hV0VzQbiXxm5KyiJ1Gro/iLCf/gz1nJEkaE68g+yN8UdpwumK4ifqbc/e9jSiMle/mPdCcv72QqL+eYsuqquXdtB7nfBWwUcK4NDKrAj+ldc+IrzPyVtJ1gd/lnu8eYN9Rxqt0JlKsYXNK0ogkSeoxW5D9IX6C6nVpG26iPgN4Mnf/fFfOnw/ymr2QqH+e7Pp9Jm046rAP0DqhOxPHGZfd22ldEPCHDH/c8WHEsJfm5zkbWLNTwWrMTQV+TbZnxPuSRiRJUg/qo3iQtX3SiDpvuIk6wLdaPKZ5OWiQx/dCon4l2fU7NG046qCTKL5/FxG9TVQNLwIeZuTJ+izgB7nHPk4MlVB5rQr8L419uhz4t6QRSZLUw84he7D1obThdNxIEvUXtHhM89jcVnOnN6t6or4u2aJ7K2vXqfw+QfG9+zR2Y66irYB/UdzfP2XgnlUvbfG484C53QxWXbcmcDnZIV6vSRqRJEk97h1kD7iuThtOx40kUe8Dbm/xuH6iuM5gqp6ov5Psul2VNhx1yBsovm+fIlpfVU3PAu6kuN9PbnHf6cAZZOsWPAMciwXGym4uURC0+eTcS5JGJEnSIHphntBzgK/QaEHZhZgf/IZkEaXXD5xFtC7mnTXGsYxH+a6Qv0kShTppJ6KgWLMngJcBV4x9OCO2K/DiNretIGorrOzA6zwP2LvNbYuAL3bgNcbC3cSJmD8Cmzdd/xHgZuAntf/3JL77tmi6z+XAkcTUfSqvrYE/0CgQOR94OeX63EuSVFl/INuakj9gL7ORtKgDzCGKLjUvbxzia1a5RX1Psuu1Atg4aUQarXWB+yiOSd89ZVAjlJ9yMr/s16HX+dsAr/FIh15jLG0IPEB2PZ4hpmE8iew87Etr101MEKc6a3eytQruBZ6dNCJJkpRxGNkDtMXABkkj6pyRJuqjUeVE/bcUx6aqvCZSnNWgn+gGX0aDJerf78BrbDXIa5QxUQd4LvHd37wuS3L/XwdsmypAddT+ZGc4uRPYNGlEkiSpYBLFcYpnJo2oc0zUO2cPivNqd6qFUmmcSPG9Wuap9gZL1BcSFctH49RBXqOsiTrAm2i9TiuI8emrpAtNHXQUsIzG/r0KmJ00IkmS1NbRZA/MllONqdpM1Dujj2xF4H4cw1h2WxJd3PM9JMrcpXmwRL2f0U03NYliF/EqJeoAp1Ncp/9IGpE66USyJ1wvAGYmjUiSJA1oMnAr2YOzaxh8KrLxzkS9M95NcZ32TxqRRis/NeOjlH+avVaJ+nW5/y8ZxfO/PPdc84AHc9eVPVGfAtxIdp1uofy/Bb1uIvBVsvv1e7hfJUkqhYMoHuSWfV51E/XR24TsWMZ+4JdJI9Jo7UvxPXpE0og6o1Wi/j6yxdD6iXHmI/Hz3POcRhTgqlKiDlFEbgXZ9XpP0og0GlOBn5Hdn2fgtHqSJJXKL8n+mC+j/XRHZWCiPjozKLZIPomV3svuz2T36WVU46C9VaL+euDc3HWfHMFzr0mx2Nr2VDNRB/gm2fW6D8eol9GaZAtGLid6SEmSpJKZDTxE9gDtMbJz7JbJVsSYvOZljy6/5ptavOakLr9mN/QR8yjnE5+jUwalUXsexX1axqnYWmmXqB+eu+5fDH8s/ntzz3Fl7fqqJurrElO0Na/baMb3a+xtSnZI20LgkKQRSZKkUTmQYrfHm4DVUgalMfcxiknP2VSj5bWX/ZDsPj0/bTgd1S5Rn0KMwW++frg1Fq7JPf7fa9dXNVGH6B7dvG7Xpw1Hw7AHUUOhvu/mA89PGpEkSeqIVtM2nU+MdVP1vZniVGzXEV3hVV6ziFa10SSs41m7RB3gy7nrfzCM590299glwNq126qcqG9McXz/zkkj0lC8huzn/G7g2SkDkiRJnfVtige9FwOrJoxJ3fdvFHtUPEp0o1S5vZnsfv0nMCFpRJ01UKK+W+764cyp/vncY3/adFuVE3WIKfua1++UtOFoEMeS/f6+DtggaUSSJKnjpgJ/o3jgewnOu1pV76SYpC8F9kkZlDomX/l5JEXVxrOBEnUoTjs2lHoLkyhOwXZg0+1VT9RfT3b9bk4bjtqYRLHXyC+B6SmDkiRJ3bMOcAPFg9/LaXT9VDV8kGJ39yXAq1IGpY7pI8apNu/f3ZJG1HmDJervz9126RCe81W5xzxItjhk1RP1WcTJuuZ1nJs0IuWtCvye7D46g2r1lpEkSS3MAq6geAB8H92voK7umwp8h+L+XQy8ImFc6qytyO7f+VSvMOBgifociknn1oM8569z9893/a56og7FKSetHD5+rE92Cs3lOOe9JEk9ZXVazw++GKfsKbPNKXYH7iemZapSkTHBEWT38f+mDacrBkvUAc7J3T5Q9/91KCb2+aJcvZCon061h0yU1e7AAzT2y5PAy5JGJEmSklgN+CPFA+F+4GvAtHShaQQOBh6nuC8fxzHpVfRRsvv5c2nD6YqhJOqvzt0+0Jzqx+fue0WL+/RCon4U2XX8UdpwBLyW7Dz392NFfkmSetokoutnfixzP3AHsG+60DREs4Azab0Pb2HwrsAqp2+Q3ddV7B47lER9CpFMN9+nXe+R63P3e2eL+/RCov4isuv417Th9LQJxG9w8/64BusGSJKkmtcBT1M8KF5JJIFWhR+fDiZaEFv1ivgRzpNeZfmK74elDacrhpKoA3wxd58ftrjPLrn7LALWaHG/XkjUn0N2Hf+eNpyetSrwK7L74mys7C5JknK2A26nddJ3L/DKdKEpZyNiqp5W+2oZMfeuqu1csvv95WnD6YqhJuo7U0zC83Oqfyl3nx+3ec1eSNQ3pvj9rrG1KXAT2ZPip2Bld0mS1MZ04mAhP/d2fbkcxzuntCaxfxbRev/cSPWm6FJrJupZ+W7tb2+6rVX3+Je2eZ5eSNQ3IruO96UNp+c8H3iIxvZ/Gjg0aUSSJKk09iK6Q7ZKBvuBC4AdkkXXe2YAJ9K6WFw/Ucn6FCIhUW/4OXZ9b5YvFHdZ022H5W4bqOBcLyTq25Bdx1vShtNT3k525oH7iGEZkiRJQzYV+DTRlbpVcrgc+AFWpu2mNYkEfR7tT5pcRnGKKVXft8i+D96dNpyuGE6i3mrqtXohxd/nrv/0AK/ZC4n6PmTXsVX1e3XWKkS9l+bt/hdgdsqgJElSuW0L/Jb2iWI/cDHwKhxf1ylbAl+mdYG/+nIP8Bbc5r3qY2TfD6elDacrhpOoA/w6d99PAXMonmwcaCaEXkjU30J2HX+SNJrqW58YNta8zb+BPaAkSVKHPA+4lIET9juIQmZrJYqxzCYS42bPoX2NgHri8B9EC4161xvJvi/OSxtOVww3UT8kd99/AR/KXXdZ20eHXkjUP8vQexhodF5Idjz6UuC9KQOSJEnV9Qqy1WpbLUuA3wCHA9PShFkaOwOfBx5g4G36NHAysHqaMDXO5McZPwb0JY2o84abqE8GHs7d/6nc/29v++jQC4n6ZWTX8dVpw6ms/Hj0h4F9k0YkSZIqbwJwEHAhAyeX/cAC4DvA/tjVr25L4KMMXLAvvzxOHPjZ1V0Q74MFZN8jOyWNqPOGm6gDnN7iMfVlIcVp2/KqnqjPJE6kNq/jhkkjqp4ZxPR/+Z4cc1MGJUmSes9WwBnEQfBgyeYzRMX4Y4m5fHvFNGA/ojL7UJPza4BvA09SPODbdmzD1ziVH5P98bThdNxIEvXtWjymvvxgCK9Z9UQ9XwH/trThVM7mxDSZzdv4TDxJLUmSEpoDfJjhtRLfDHyOmEN2/bEPuWumE3Plvh/4A+3nPW/V++DbxPR4dRsSwwia77eUGFfqsILedhTFpKtK3d9HkqgDXNficf3EybLBVD1Rz3+XVLEIYSoHk51CcyFwZNKIJEmScnZhaOOu88t9wC+AE4gphGaOdeAjMIkYL/xW4KtEktBuSrtWy1DH8x9MVHpvfuydwAGdXiGVxtoUuzHvkzSizhppon5si8fdR/u505tVOVFfn+J3055JI6qGScBJZIuA3gvsljAmSZKkAU0EXgJ8k+En7fXlfuAi4GvA8cDLie6FY131fF2ilfxtwGeAXwG3Upy7eSjLIuB/gXcxvAr5qxHDDJbnnu+HRI8G9Z6zyb4Xfps2nI4aaaI+GlVO1E8ju243pg2nEjakWJzvDzjziSRJw1KlLqFl1AfsQLQAHwDsTbREjMYTwDyimu48Yhqch4H5RDK7kGhxXEqMj19Ru2VGm2AAAA9USURBVDyt9tqr1eKqF5iaCaxDJL2zieI/69QujzbWO4HziWm0/lSLbaR2JMY97t503RNEq86XgJWjeG6Vy77AH5v+7ydmErg+TTgddRxRHK7ZEUShrm65l2xxtUeJz3/ZrQXcRXzn1f078JU04VTCQcB3aSTl/cCpwEeI3xpJkqRSmkmMTf88MT/7UIrRlWm5Dfg+cTC8RYe2WbMJRBX4fLG5S7HYXK/Jt+hdlDacjrFFvXO+THa95mGNi5Fq1dX9IaL3mCRJUuVMJsa2vxv4HlFobiTdylMsDwDnElW3DwDW7PC2GUi7YnOnkm09U3UdSPE9eWjSiDrDRL0ztqc4Nv34pBGVV6uu7hfi0CNJktRjJhGt0QcB7yO6e19MJMb5cdrdXp4gCsX9FPgE0QV3V8ZPwbtDiIJZ+RMIb8JhH73gfLL7/mFi2EaZmaiP3iTgKrLrdCdjX+OjCg4ihkLUt+Ny4uTsUIoUSpKkAYx2jLHG3nLg9tryu9xtfcTY0XVqy3q1/2cTY84nEV07pxJz2M4gDqimEMXclgNP0UjCIaZHW0AkuI8Qyc6DtcuLu7B+nfQromXnZKK7/SRim5wFvBN4L3BtsujUbe8nxqtPrv0/myjA+GriPa7edBJxQrHZ8UTtDg3NVKKA6HtpnPScB7yB6gwzkSRJ0hjYimIL6woiaS97K6vaO4liC/THUwY0Sraoj86hRGHJ5vX5SdKIymcboidV8zb8E3ESVJIkSRqRwyjOvT4feA/2MKmiycDVZPf3SmJYhHrLTsRMF83vhQepRgX7sdBHtKAvorH9lgEfxa7ukiRJ6oBpREtr8wFnP3ALVimuoo2JCtTN+/opoqCYesMciifoFgN7pQyqRGYD55DdfncRU4pKkiRJHbUh0fU935X4HCK5U3XsRSRmzfv5IWCHlEFpTMwBbqT4OT86ZVAlsj9Ro6R5252FM2hIkiSpy14K3Er2QPRp4MNYCbpK3kIxWZsP7J4wJnXXesS0lvn9/vmUQZXEVOBLZMf0zwcOTxmUJEmSessUokr4k2QP6O8gxrU7nVs1fJ5i0vY4sGfKoNQVzyKmXWvVY8Yx1QPbA/g72e12MdELSZIkSRpz6xFz068ge5B6JfCChHGpM/qAz1FM3p4kKoKrGnYH7qO4n3+NvWQGMpmo37GcbMG4k/DkhiRJksaBvYCrKB7o/wLYMmFc6oyTKO7blcAZWP2/7I4EFlLcv2cTiaha2xW4iew2u5ninPOSJElSUn1Et/d899mlRKv7nHShqQNOpJjM1bv4um/LZxXg67Tepz/GEzDtTCY+C0tpbK8VxEmrqQnjkiRJkgY0BXg78DDFKb5OIqZ7Uzm9A1hCMbG7G3hxurA0TNsCV9O6l8Sp2G27ne2BaynW5XCYjyRJkkpjTaIYWT6xu5fobjshXWgahV2IxLxVS+zZwFrJItNg6q3B+an36ifSDksX2rg2Bfgviq3on8UTj5IkSSqpjYh5hJunLaqP53x5wrg0cmsDF9I6WX8QeHW60NTGjhRbg+vLP4hWdhXtTXEs+p3APimDkiRJkjplD+ASiknCeUQSoXKZBJxGseJ/c8XwZyeLTnXrAV8lW5m8efkJMDNZdOPXLKK2RvMJxhXAF4DpCeOSJEmSuuJVwK0Ux8b+DNgmYVwamT0ptjg2JzZnA5ski653rUp0c3+S9j0fXpMsuvHtMOABij2A9k4ZlCRJktRtk4iCcw/SOrFzSrdyqY/hbVVorh9YRLS+O369+6YDJwCP0XpfrAS+BayRKsBxbC4xpWR+1opTcD55SZIk9ZDVgP8EHid7cLwM+CawcbrQNALbAZfSOkHsB54humFvnSrACpsLfAp4lPbb/zZgv1QBjmMTgeOIgnrN2+tPeNJQkiRJPWwN4JMUD5SXAF8G1k8XmkZgP+AG2ieMK4ELgIOBvkQxVsWOxFjqRbTf3g8T3eCnJIpxPNsLuIbs9noMeCu+NyVJkiQgukafQrS85hP2M4nCWCqHicBRxHR87RLIfmJ8+/uBDdKEWUprAEfTujhj8/I48CEsftbKusR3Sr4Y4tnAOgnjkiRJksat9YAvUpzv+Wngv4k52lUOU4HjgXsYOKlcAVxEJPerJ4l0fJtKTHv3S1rPg55vEf40fk5amQy8D1hAdpvdDrw0YVySJElSaWxItHotJXtQvQA4mZjPW+UwCXgtcDkDJ5n14nO/AN5Gbw97WJPYZt+lWMeh1XIr8C5gRoJYy+AlwC0UT/59CIvFSZIkScO2MZGwLyN7kL24dv2G6ULTCOwJ/JTi/my33EAMiXgh0SJaVROAXYGPApfRfu7z/HIB8PLa41W0AXAWxe12DhaslCRJkkZta+DHFMeVLiGqxFuhuVzmAMcCVzK0hLSfKDh4EdG1+1WUu27BGkR3648BvwMeYejb4bba4zYb86jLYyZRCX8hxRM/+ySMS5IkSaqkzYAzKI7VXUG0ku2WLjSN0FbEXOx3MPRktb7cQxQB+zDwGmB7Ykz3eDGZOIn0CmKO87OIbuorGd56ziPe97uPbfilMxl4O7G98sX1jiWGYUiSpApxqhZpfNmIqBj+NrKVrfuB84nCc5ckiEsj1wfsAhwAvAzYg6ggP1wriYrz/6gt9wIPEa3WDxBTlj1CdC8fbbzrALOJHgLr1v5fn0jOtwI2YeRd9m8k3svnEXPUjzbeqjuU+Nw3965ZAXwb+AixzyVJUsWYqEvj09rAe4BjiO7EzS4DPkN0Le4f47g0emsC+xOJ+wFEItxJDxMFxZYS0wKuAJ6s3fYE8Z5ZnRj/PYOYi3wqMK22rMPITiS08wRwIZGcnw/c38HnrrLdgdOAF+Suv5A4mXfDmEckSZIkCYiE6kNEy2m+2/B1wBFUuxhZL5gLHEwUl7uU4vjjMi3LgZuJrvDHEj0JLAg3PFsAP6c4jOAa4MUJ45IkSZKUM41oYb+bYnJ0PzGW2andqmEK0Zr6TmL89vnAPykWHEy9/Av4I/BV4Dii5ddp1EZuI1pP3Xg38EY84SFJUk+x67tULpOB1wMfBJ6du20JUYDsNOD/xjgudd8qxDjlLYnig+sR48jXJcaS18eVd+J7fT6N8e8PEt3pHwbupDFG/qkOvI5iqrUTiGJxzQUDnwY+RwxzWZQgLkmSlJCJulROE4CDiO7F++Zu6ydaOs8AziW60Ko3TCQS9mlEy/yM2nUza7fPIr73FxDvi2eIFtzFRDK4hEjOl45p1L1pLjGs5WjiJEzdYqKXwqeAxxLEJUmSJKkDtiKS8mcodk++AziRSNAkpTebqEeQr0WwhOj6vn660CRJkiR12mzgo8RUXfmE/Qngs8CzUgUn9bj1iK7srRL0rxBd4CVJkiRV1BTgMOCvFBP2FcAFtdutFi9132ZEj5d8gr6UqI6/WbrQJEmSJKWwC5EMLKOYtD9AdMHdNFl0UnXtTHz2ltM6Qd88XWiSJEmSxoONgVOJQmGtWtnPAw7FVnZpNPqAA4E/U/ycLSS6uHtiTJIkSVLGKsDrgIuIqt/5ZOJB4L+xO640HJOBNwE3UvxMPQacTFTjlyRJkqQBbU50fZ9HMblYCVxKzO08LVWA0ji3LjGrwr20Pul1ErB6quAkSZIklddEYD/gbIrjafuB+cSY2v2Irr1Sr6vXflhK8fPyD+BYYGqy6CRJkiRVyibAJ4H7KSYg/cBdtdu3SRWglMiqwDto3b19JTGbwkF4MkuSJElSl0wCXgn8ltathv3AtcDxwNxEMUpj4dnE9GpPUPwMLAC+CGydLDpJkiRJPWkN4EiixbBVAboVxHj2Y4G1EsUoddLqDPyev5UYmz4rVYCSJEmSVLcp8J/ALbRuZV8E/IyoLD8zUYzSSEwAXgL8iHgf59/by4j39gsTxSdJkiRJg9oVOJ2obt0qaV8MnAscDcxJFKM0mC2IugutKrf3A3cCHwPWTxWgJEmSJA3XRKIl8nvAk7ROdlYAlxBj2jdNE6b0/60PHEMM2WjVtf0p4LvAPlgcTpIkSVLJTQNeRSTtj9E6ae8Hrgc+DuyQJkz1oA2IOgqXEieOWlVu/zPwFqLCuyRJkiRVziTgxcCXgPton7TfCXyZqDK/WpJIVVUbAscBl9G65bwfuBv4BLBZmhAlSZIkKY0+YDfg07QvRNdPTAX3Z+AjtftPSBGsSm07oiL7X2mfnN8DfA54LnZtlyRJkiQg5p3+EHAlrbsh15dHgJ8Ab8ViXmptVWK4xZm0LwhXbzn/LLAHJueSJEmSNKB1gCOI4l0P0D7R6gduIirNH4KV5HvZ1kRhwguI2QXavV/uAk4DdsfkXJIklYgHLpLGkz6i6/JLasvzgakD3P8fRBfnS2p/b+12gEpiU+K9sA8xj/kmbe63AriCmBbwPOC6sQhOkiSp00zUJY1n04C9gf1qy84M/L31JNGd/jKiuvelRIurymVT4HnEvt+f9ok5wKPAn4ALgd8C87oenSRJUpeZqEsqk7lEi+peRAvrtgxccG4RcC3RslpfbgKWdTVKDccMYEdgVyI5fz4DD2tYAVxDo9X8aqJonCRJUmWYqEsqs1WBPWm0vu5NtMIPZDnRZf6apuU64JnuhamaKcAWwC5Ny26169tZAVxPo5fEH4H53Q1TkiQpLRN1SVUyhUj+9iaS972A2UN43ArgdiIhvIVI5P9Ru+6prkRabROAZwFbAtsAOxD7ZWtg4iCPXQxcBVxMo/aAJ1EkSVJPMVGXVHUbE12rd2paNhzG4x+gkbjXl9uIiuK93oV+VWCr2vLspstbMnARwLp+4A4aPRuurC3WFZAkST3NRF1SL1qbbOK+E9Ele6Dx7nnLiST+PuD+2uV7gQeBf9WWB4ClHYt6bE0G1gU2AjaoLRvWlvVrf+cO4/n6gTvJDjm4BljQuZAlSZKqwURdksKqwHOI1uB6q/AWtb/TR/G884iE/RHgCeDx2t/85ebrlhNd7peP4nUhupnPrF1eFVhjkGXN2rIBkaQP58RFs/uIXge3EVPm/R2TckmSpCEzUZekgfURrcf1pL05id+AoXXxHq1FNLqDL6X1mO3VaSTWa4xBTE8TLeT1hLw+tv+22m2SJEkaIRN1SRqdacB6xNzfc1tcnku0Tpfp+3Yx0Qvgwdrff7a5LEmSpC4o04GjJJXVdGJc/KzcskaL62YBq9Ue11f7v24Wje/tmUTX9hXAk7XrlgALa5cX1v6ndvsyolv9/Nrfdst8rHQvSZKU1P8D/qaEZCg8cSIAAAAASUVORK5CYII=" width="65%" style="display: block; margin: auto;" /> --- ## 7. Repeat identifying links, delete indirect links, find ambiguous <img src="data:image/png;base64,iVBORw0KGgoAAAANSUhEUgAAA+oAAAJUCAYAAACCKdxjAAAABHNCSVQICAgIfAhkiAAAAAlwSFlzAAASdAAAEnQB3mYfeAAAABl0RVh0U29mdHdhcmUAd3d3Lmlua3NjYXBlLm9yZ5vuPBoAACAASURBVHic7N13vBxl9cfxz8zedEKAUEKv0kF6b1IEpAhoKKIIglF/gBrlZyQQfEggCYiiYgMUIoKKEem9SZPey4/epCRASCA99+48vz/Orjs7d/fW3Z0t3/frta/cmZ3dPfduyZ55nuecABERkZ4LgdWAVYEVgFHASsCKuX9HAUOBwcAQIAMsnbvtiNztFwCLgSXAfMADc4AI+AD4EJgBzEz8/BYwt8q/n4iIiEjqgrQDEBGRurQssAGwIbB+4jIoxbjeA14CXs5dXspdXscSfREREZGGp0RdRETasKR8a2BnYBdgIxrr/4h5wNPA47nLvcCbaQYkIiIi0leN9CVMREQqYyCWjO8L7ApsiU1VbzZvAw8CdwC3AO+kG46IiIhIzyhRFxFpDWsB++UuewFL9eO+PsLWi8/A1pC/T2E9+XvYOvJFwEIgC3yau90n2PT0odj0+YHAMOz/omWAAdi695WAlSleAz8KWDt3TF89iyXstwD3Y2vkRUREROqOEnURkea1LnA0cASwcS9v2wG8mLu8QmEt+MvAxxWMsTcGYMn6BhSvmd8UWL6X9zUPuAm4HLgVJe0iIiIiIiJSJcsCxwC3Y6PXvoeX94DrAQfsjY16N5JVgIOAqdho+QJ6/rt/DFyG/d46gS0iIiIiIiL9FgCfB67DRoZ7kpy+C/wROBxLcpvNQGB74DTgPqCdnv1dXgPG0/sRehEREREREREGYaPnz9B9AtoBPIaNmG9N640cD8NG3C/Eisx19/dahI2yb5pGsCIiIiIiItJYVgImAR/QdbIZAfcAY7Ap8WICrA3db7HCeN39DW/BKuSLiIiIiIiIFFkKGIdVUe8quXwBGzlfN5UoG0sGW5t+GVZgrqu/60PAHqlEKSIiIiIiInVlEDAWa39WLolsB/4G7JBSjM1gBPAD4A26TtivR1PiRUREREREWlIAfBV4k64rlp8DrJ5OiE0pA3wZqx5f7u+eBabRnIX4REREREREpIS1gdvoup3aSViRNKmebYFrKf88zAa+SesV5xMREREREWkZIVb8bS6lE8O5WK/wpdMKsEVtD9xN+YT9XmCD1KITERERERGRqtgIeJDy7cJ+jvp7p+1AyrfDmw/8EDvZIiIiIiIiIg3ucMqPot+BKrjXkxBbdlDu+boeWCa16ERERERERKRfMthU9ojOCd8cbBq81j/Xp1WAayidrL8CbJ5eaCIiIiIiItIXywO3U35UdrX0QpNeGE3p1nkLgWPTC0tERERERER6Yx1K9+teBJyQYlzSN6tSvr7A6SnGJSIiIiIiIj2wPvAfOid0/wF2SDEu6Z82bBlDqWR9aopxiYiIiIiISBc2BN6lcyJ3D7BSinFJ5XwVqwCffI7PQ/UGRERERERE6spngQ/onMBNw0ZjpXnsCMym83P9qzSDEhERERERkYKVgLfpnLhdhPpuN6stKV1k7n/TDEpERETqn6bgiYhU3wCsF/puif2/A07EkrdGMAJYr4vrnwHaa/A4TwMdFXicWtgYe+5Xju2LgC8CN6QSkYiIiIiIiPBHOo+q/iLViPpmf0oXSstfDqvQ40zp5nGWr9Dj1MrGwCyKf4fZwAZpBiUiIiIiItKqTqJzonkTkEkzqD7qLlG/tgKPEVJ6iUAjJ+oAe2OzDeK/x0vA0mkGJSIiIiIi0mo2BBbQOTlbJs2g+qG7RL0dGFXlx2jURB3ge5SuUSAiIiIiIiI1kAEepjgpm0NjT3fuSRL9g34+xpU9eIxGTdQB/kDx7xIB+6QakYiIiIiISIv4Jp0TzCNTjaj/SiXqyZ7wz/Tj/pcDFnZz/42eqA8CnqP493kRKzgoIiIiIiIiVTIcmEFxMnZVqhFVRqlE/Xw6T+/fuo/3n6+An788BtxW4jEbOVEH2AGrWh//nb6XakQiIiJSV9S7V0Sk8k7C+qbnLQTGphRLtX1C5yJyX+/jfR2b2P5TH++n3j0EXJLYdyowJIVYREREREREmt5Q4AOKR0vPTDWiyik1ou6AfRP7ZmFTvHtjk8R9LMZGzptxRB1gReBTin+vk1ONSEREROqGRtRFRCrrKGCF2PanNGbP9N64HfhPbHs54MBe3sdxie3rgI/6E1Sd+wD4XWLf94AghVhERESkzihRFxGprG8ktn8PzE4jkBqKgMsT+3oz/b0NODqxb1p/AmoQP8dmDuStC+yeUiwiIiIiIiJNaW06t95aN9WIKqvc1HeA9bHfty891Q9O3OcMLHmH5p36nvc3in+3i9MNR0REROqBRtRFRCrn4MT2/cBraQSSgpeBB2PbpUbJyzk2sX0ZVhW9FUxLbB+I/m8WERFpefoyICJSOfsntq9OJYr0JKu0J9edlzIS+EJi358rE05DuAurY5A3CtgipVhERESkTihRFxGpjADYPrHv5jQCSdHfsJ7qeZsA23Rzm69RXCH+EeDZCsdVz5YAdyb27ZhGICIiIlI/lKiLiFTGZ4BlYtuzgZdSiiUtn9J5FkF3ReWS10+rWDSN46HE9napRCEiIiJ1Q4m6iEhlfCax/TRWHKzVTEtsf4XyPdW3pHia9xLg71WIqd49kdhOvpZERESkxShRFxGpjLUT26+nEkX67gLejm0vBxxU5thjE9tXA7OqEFO9S75W1kojCBEREakfStRFRCoj2TLsnVSiSF+EVW2PKzX9fSBwVGLftGoE1ACSr5UVUolCRERE6oYSdRGRyhiW2J6fShT14RKKp/3vB6ycOOZAihPS94DbqxxXvVqC9Z3Pa6P8cgERERFpAUrURUQqY0Biu73kUa3hDayHfF4b8NXEMcnWbZcB2WoGVecWJ7aVqIuIiLQwJeoiIpWxMLE9JJUo6keyp/qxsZ9XAvbt5vhWEgBDE/taeUaGiIhIy1OiLiJSGXMT28uUPKp1XAnMi21vDGyb+/lrFM9AeBB4sUZx1aMRFP9/vIDWnl0gIiLS8pSoi4hURrIg2FppBFFH5gH/TOzLF5U7JrF/WtWjqW9rJbb/k0YQIiIiUj+UqIuIVMYbie0NUomiviSnsx8F7ApsFtu3iNbsnR6XfK0kX0siIiLSYpSoi4hUxrMUVzrfBFgqpVjqxd3Am7Ht5YBLE8f8E5hTq4Dq1PaJ7WdSiUJERETqhhJ1EZHKmAO8EtvOADulFEu98HQeVV83sT2tNqHUtd0S24+kEoWIiIjUDSXqIiKVc29i+6BUoqgv0yieaRD3LnBX7UKpS6sAW8W2I4pb24mIiEgLUqIuIlI51ye2D8N6iLeyN+l8AiNvGqpuPhprz5b3CDAzpVhERESkTihRFxGpnDsobtO2CrB/SrHUk2ll9v+5lkHUqW8mtq9JJQoRERGpK60+0iMiUkkLsP7hJ8T2fZfOI+2t5ko6t69bDLyUQiz1ZC+s6GBeB3BZSrGIiIiIiIg0re2wNdnxS7Kqd6Pan86/m6vyY95W4jGXr/Jj1sqdFP9eyb7zIiIi0qI09V1EpLIeAf6V2Dc1hTikvu0F7JnYd14agYiIiEj9UaIuIlJ5ZyW298AKy4kADAB+ldh3B/DvFGIRERGROqREXUSk8u4Ebkns+zWwXAqxSP0ZD2wc246AU1OKRUREROqQEnURker4IVYcLG9lLFmX1rYllqjHTQMeq30oIiIiUq+UqIuIVMcLdF6bfhTwgxRikfowEvgHMDC270M0mi4iIiIJQdoBiIg0sYHAo8DmsX1Z4AtYNfNGMxIbEY57PXeplq3ovGTgHqC9io9ZDQOxdei7JvYfBlxd+3BERERERERa1/rAxxS34foY+EyaQUnN/Z7ObeZ+nmpEIiIiIiIiLWxfbL16PEl7DVgzzaCkZibROUm/DWhLMygREREREZFW90M6J2tvAeulGZRUXakk/XVg+TSDEhEREREREXMpnZO2d4AN0wxKqiIAfknn53s2sFGKcYmIiIiIiEhMG3AFnZO3GcD2KcYllTUIuITOz/MsYJsU4xIREREREZESQuCPdE7iFgHfSzEuqYxVgQdRki4iIiIiItJQAuBXdE7mPHAZMDS90KQfdgPep/SMic1SjEtERERERER6IAAmAxGdE7tHUfu2RpIBxmG93ZPP5cvABumFJiIiIiIiIr11MDCHzgneQsABA1KLTHpiU+AhSs+OuAFYNr3QREREREREpK/WB56ndLL3FLB1eqFJGQOwUfRFdH7OImAqVo9AREREREREGtTSwGOUTtaXYInfiNSik7g9gWcp/Vx9BOyfXmgiIiIiIiJSCYOACymd+CUrh48DhqQTZsvbFPg75Z+f67Gq7yIiIiIiItLAVqNzO69/ULqFW/7yH2AM1pddqm9N7ERKltLPx/vAl1OLTkRERERERCpmd6x1Vz7ha8dGzPMOwpLycgn7S8B3UDu3atkauBxbelDq7x9hJ1SWSStAERERERERqYwA+B7F7bxmAp8rcexQLHmfTfmEfQ7wS2CNagfeAkLsBMntdL0M4X5gl5RiFBERERERkQoaDkynOOm7D1i5m9uNBM7D2raVSx6XAH8B9kIVx3trFeAU4DW6TtAfA/ZJKUYRERERERGpsA3o3IbtQnrXK30N4BJgMV0nlO9iif0WFYq9GS0NHAvcAXTQ9d/zBeAIbDaEiIiIiIiINIEjgbkUEr+5WOLXV6sAZ2PtwLqrFv8cMB74bD8er1ksA4wGrgQW0PXfLQJuw9qtKUEXERERERFpEm1YD/SIQgL4MrBZhe5/EHAM5ft6Jy8zsRZjxwDLViiGercJts7/drqfieCBRcBlVO45EhERERERkTqxAnAnnXttV6NKeADsDUwDPqVnSXs7cC8wGfgi3a+TbwQDge2Ak4A/Y63TevK38MCjwPex501ERERERESazM7YOvF8EtgBOGpT5G0INtX+esq3Fit3eQsbcf8hVoV+lRrE21dDsOn8XwF+gfWjX0Tvft/XgUnAhjWOXURERKQsrbkTEam8McAF2OguwCwsmbwthViWB74EfAHYE1iqD/cxF5uu/zLWu/0l4G2sB/xMYH5FIu0sBFbERrhXBdbDEur1c5c16Nv/Y88BtwDXAP/GEnYRERGRuqFEXUSkcgYDvwWOi+17AkuU30wjoISBwK7AfliBtE0qdL/zKSTtH2AF7qLc/iXYKPdCbFbBQqxFXUBhCcDSQAYYhiXmoygk6JWYgfAptgThZixB/08F7lNERERERETq3HrA0xRPq74Mm55dr1YHjqLv08br9RKfvr8LvWt/JyIiIpI6jaiLiPTfAVjhsnwV9cXAycDFqUXUNwOxvuvbA9sAG2G935dOM6guZLGk/GXgSeAR4GGsgJyIiIiIiIi0oABr+5WlMJr7NlZ1vJmMAvbA1t6fB1yHJcVvU/1R+A+B92LbNwGHYdP2B1XxdxYRERFJjUbURUT6ZiRwBbBvbN/dWLX1D1KJKD0jsNZuK+Quy2Fry4dho/SDsSUAbbl/52JJ95zc7T+hsKZ9NjYi/kHu0g7sDvwrd+ylwDeq/PuIiIiIiIhIg9kSa+uVH+WNgKlYQTSpvBUp/K0fSjkWERERERERqTPHYCO/+cTxE2wqtlTXRxT+3poNJiIiIiIiIgwCfknx+un/wwquSfXdR+HvvkrKsYiIiIiIiEjKVsOmXMeT9CuwNdhSGxdR+NvvlXIsIiIiIiIikqI9gBkUksR2rNK71NZYCs/BySnHIiIiIiIiIinIt17roJAgvgvslGZQLWw/Cs/Db1OORURERERERGpsOPAPiqe634u1IJN0rEnhubg75VhERERERESkhjYEXqA4Sb8QGJBmUEJAof/6jJRjERERERERkRo5EphHIUGfCxyRakQS9xiF52ZkyrGIiIiIiIhIFbUBUykeRX8Z2DTNoKSTP1N4fnZOORYRERERERGpkhWAOylO0q8DlkkzKClpPIXn6ISUYxEREREREZEq2AV4j0Ly1wE4IEwxJinvUArP1c9SjkVEREREREQqbAywmELi9yGwT6oRSXc2pPB83ZRyLCIiIiIiIlIhQ4BLKZ7q/jiwVooxSc+0AYuw5+yNlGMRERERERGRClgPeIbiJP0yLHmXxvA89rxlgWEpxyIiIiIiIiL9cCAwm0KCvhAVJGtE/6DwHG6VciwiIiIiIiLSBwEwDhuBzSd4bwPbpRmU9NkkCs/j0SnHIiIiIiIiIr00EriV4qnudwErphmU9MtXKDyXZ6Uci4iIiIiIiPTClsDrFJK6CJgKZNIMSvptSwrP6VUpxyIiIiIiIiI9dAywgEJC9wnWg1sa3xAKyxheSDkWERERERER6cYg4EKKp7o/BaybZlBScfmZEkuAgSnHIiIiIiIiImWsDjxEcZJ+BWrh1YxupPAcb5RyLCIiIiIiIlLCHsBMCslbO1bpXZrTeRSe68NSjkVERERERERi8q3XOigkbu8AO6UZlFTd8RSe79NSjkVERERERERyhgP/oHiq+73AqDSDkprYkcJzfnnKsYiIiIiIiAiwIVbxO56kXwgMSDMoqZllKDzvj6cci4iIiIiISMs7CphHIVGbCxyeakSShvex538+EKYci4iIiIiISEtqA6ZSPIr+ErBpmkFJau6i8DpYK91QREREREREWs+KFCdmHrgWmwItrenXFF4L+6cci4iIiIiISEvZBXiPQlLWATis4ru0rhMpvCZ+kHIsIiIiIiIiLWMMsIRCQvYhsE+qEUm92JPC6+LilGMRERERERFpekOASyme6v4YWossBStTeG08kHIsIiIiIiIiTe0zwDN0br02MM2gpC59jL0+ZqcdiIiIiIiISLM6EEu68gn6QuCEVCOSevYghdfKSinHIiIiIiIi0lQyWIG4LIXE6y1g2xRjkvr3Rwqvlz3SDUVERERERKR5jARupXiq+03AcmkGJQ3hFAqvmf9JORYREREREZGmsBXwOoVkKwKmAmGaQUnDOIDCa+eClGMRERERERFpeMcACygkWp8Ah6YakTSadSi8fu5IORYREREREZGGNQi4iOKp7k8B66YZlDSkEJiPvYbeTTkWERERERGRhrQ68DDFSfrlwNA0g5KG9iSF19IyKcciIiIiIiLSUD4HzKSQVLUD41KNSJrBXyi8prZPORYREREREZGGEGAJeQeFhOodYMc0g5KmMYHC6+q4lGMRERERERGpe0sDV1E81f0eYFSaQUlT+TKF19a5KcciIiIiIiJS1zYEXqA4Sb8QGJBmUNJ0NqHw+ro+5VhERERERETq1lHAPAoJ1FxgdKoRSbMaiNU78MCrKcciIiIiUhFB2gGISFNpA86iuEjcy8BhwPOpRCSt4MVBgwZtEARBdMghh6y+4YYbDs7tb8tms8MBMpnMIArdBYZGUTSo1B2FYeiBObFd87LZbHsmk1kILGpvb1+czWYXzJ8/f8EFF1ywuFq/kIiIiLQ2JerSbAJgRWAFrF93/st5BlsvDTACiLBRXoDZuX/nYSNzC4EPgI9qE3LTWAX4O7BzbN91wDHAJ6lEJA1j3LhxI4YMGbIC1mJtmSiKlgWW8d4vGwTBMvn9scuysZ8Hl7nbaluCfW7MwT5P5uUuHwdBMMt7P8t7/3EQBLPCMJyVzWZnZTKZmbNmzZqpJF9ERES6okRdGs3qwPq5y7pYUr4SVpwsn6BnKvRY7VjC/gHwPvBh7t/XsVHiF7F2YwK7AlcCK+e2s8BpWHEvn1ZQkr6xY8cOGTZs2CpBEKwCrAqMCsNwde/9Stj7eRSwGoXR7lYxC5jhvX8/CIL3gyB4P4qi9zOZzNvZbPatjo6Ot6dMmfJh2kGKiIhIOpSoS70ahfVE3grYgEJyPizNoEr4BEvaXwL+D3gCeJjCKH0rGAP8mkKRuI+wNep3pBaR1ExuJHzdKIrWBdYD1vXerx4EwarYLItlq/TQ+Snqs+fPnx9++umna7W3tzN06NDXR44c+XoQBPOxEW+AT4MgyOZ+nuO990EQeO99fop7yRiDIBgGDPTeDwEGe+8HAUODIBgALJW73VLAcGBIFX7HBcCbwFvA29771zOZzKvAq5988skr559//sIqPKaIiIjUASXqUg8GAJsDuwBb5y4bpxpR/70P3A88ADwOPAYsSjWi3hmMPQ8PdHHMUsAfgCNi+x4HvoQlFtIknHMrZrPZdYF1gyBYj1xCnrus0M+7b8dmprzjvf8gDMM53vvZWEI9B5iTyWTmAHOy2ezsKIrmdHR0zDnnnHPiyym2AR7N/TwdOLyfMfXa6NGjM+uuu+7SgwcPHpHNZkdkMpmRURSt4L0fCSwXhuHI3M/LYzMI8rOB+soD7wKvYEX0XgnD8HngBefcW2gmi4iISENToi5p2QTYL3fZFVtLXikfY1/8F2DJ8UJsKvanues/AUJsFAxsjWuAJZ75kbKVKaxpr4T5wJ3ALbnLGxW872q4ALgZuKnM9Z8B/glsGtt3EXAyhVFMaSDOuRBYJ4qizbz3m4RhuJn3/jNYUj68m5uXMxOYAbwTBMH73vt3vffvZTKZ94D3gPecczPpf1I5DFsjHgDPAZv18/5qYsyYMQNWWGGFFQcMGLBqFEUrYQn86kEQrOm9XxNYC/ssCnt51/OwGT7Pee9fyGQyzwHPOOfeq2T8IiIiUj1K1KVWhmJJ+b65f9fo4/3Mxqaa5y/vYGvHZ+QuH1K5RHEwNuq1CjZquCIW9/qxy1J9vO+XsIT9ZuAubFSxXhwMXIMlO6UqtR8EXIad4AA7GXIS8MeaRCf9dtppp606YMCATbLZ7OZhGG7ivd8M2IjerxP/BHgtCIJXvfevAa9571/NZDJvADOcc7U8afMW9v5cgiXuHTV87Kpxzg0EVouiaE1gvSAI1oudQFmP3k25/wB4MgiCJ6MoetJ7/2RbW9trzrmoCqGLiIhIPyhRl2rKAHsBXwUOpXdJ7WLgSWy997MUirfVW3Gl1Sgk7Vtg6+o3xdqU9dQsrBDbFcCDpDtldVXs770sNoo6L3ZdBpiQu+RH+N4Gvkxh2rHUEefcUsAWURRthp142TR36c268ZnAf5PwIAjyyfhrzrl66oxwC3YiEKyuxcspxlIrwWmnnbZaJpNZz3u/QRAEm2LLhjal50sS5gJPBUHwcBAED7e3tz989tln/6dqEYuIiEiPKFGXatgaS86PoFAFvDuvY0nqw8AjWJLeqFOoh2F/g+2AHYAdsVH5nngdS9ivwEbdaykEbgf2xIp0xZO55YG/APvE9t2MPc8f1ypAKW/s2LFDRowY8dlsNrtNEATbYOu2N6TnXRDeB54LguDZKIqeB55dvHjxi+eee+7c7m5YJ84Hvp/7+RDg2hRjSZ1zbvncMoaNcssYtsBqgfRk1sT73vuHgyB4BHhowYIFj5x33nnzqxuxiIiIxClRl0oZgH05Hoslpt1ZiBUquwO4HniheqHVhXWwKeMH0vM1+Q8Av8TWgme7ObYSTgPOyv38FLBl7uetgKuw9bJgI/7nAuOxfvRSY865tmw2uwGwdRiGW3vvt8YS8568rj7Bio+9gC1teCEMw0edczOqF3FNjAEuzP18KjA1xVjq0ujRozObbLLJmlEUbQJs7b3fOgiCbem+qF0WeCkIgvu993e0t7f/S63jREREqkuJuvTXSOwL8kl0P2r8DvAPrEDZfTRWFfRKGo4tCdgfq5A+spvjX8YS9j9hRemqYTvsxEAGS76vAw4DjgF+T2Ed7KfA17E17FIbwemnn75hGIY7AdtiszU2BwZ2czuPJeSPAU+GYfhse3v78008rXkX7HMFrIbC11OMpaE459bJZrM7BEGwHbZ8Z0u6PunjsWJ19wH3dXR03DV58uT3axCqiIhIy1CiLn21KjYC+3W6nkr5CTYaezlwDxqBTRqIFdc7GiviNriLY2djSfNPqWyf9mWAZ7ATLflE/TdYcn5C7LinsBMLr1fwsSXBOTcY2CaKop2BnYGd6P5kDlgngce8949lMpnHFi5c+HiihVmzGwnk18w/ip18kj7IFbDbIoqi7YMg2MF7vxtWj6MrzwdBcGcQBHcuXLjwnhZ77YmIiFScEnXprWWBccB3KV9tOMJGzS/DprW36sh5by2NJcLHArt1cdxsbFrvBdgSgv76C3AkxZ8Hb1Ncmf8KbObEggo8nsQ455YHdoqiaBcsKe/JFPb/kEvKgyB4vL29/bEpU6bMqnasDeADrIjaPOz9pF7iFeKcWyuKol2xpTu7YPUPyn2H6MBmctzpvb89k8k84Jxriir8IiIitaJEXXpqIJZAnkX5asLzsKTvF9i0SOm7LYDvYFPPy42yvwtMBC6h762ovon1Py+nAzgdOKeP9y8Jzrn1oyjaOQiCXbz3O2EJT1fmYwUW7wce7ujoeGzy5Mkzqx5oY7qHwkmuNbATGlIFp5566goDBgzYBfgctpRn4y4O/8R7f3sQBLeEYXiz+rmLiIh0T4m69MShwK8oP/XxLeDXwMXYVHepnJWB/wG+RfkTJP+Xu/6+MteXsxHwBHYSJixx/UKscvZDwJvYKPtbqMp7r5x++ulrB0HwuTAMP+e935Puazm8771/AKsZ8EAmk3lSo5E99nvsvQDWqu22FGNpKePHj185k8nsFYbhXt77vYDVyxzqgaeBm733N7/44ov/nj59ei2KZYqIiDQUJerSlVHY9Oovl7n+XcAB0+j7iK70zFDgZGzZQake2B5LUn6MFXzrzmBslHZjyrfvymKfEckkfh6WuF8C/A4tbSjinFslm83umUvMPwes3cXhEXai5X7ggTAMH3DOqQZA330Pm9ED1qrtlynG0tKcc+tns9m9gf2DINiT8rVMPgqC4IYgCK4FbnPOaXmNiIgIStSlvNFYElaqiNVc4OfYdOhKrJGWnuuuRsD7wInA1d3czwVYpf6e8tjnxRzgj9h0+Zd7cfumlVtjvkcURXti04C7msreDjwSBME93vt/L1my5N9Tp06tZGHAVrcPhVH0C4FvW12qugAAIABJREFUpxiL5OQKJO7mvd/fe78/sEGZQxd6728Hru3o6LheLeBERKSVKVGXpBWAP2PTRpOWYCNUU9H057StCpwJHEfpaeuXYWvcS41OfQG4gZ69/6Pc/T+JjdhfXuY+W4Zzbiiwh/d+n9xU9s0o/7fMAk8GQXBXEAR3z5s3777zzjuvWi32xJbn5Nel30fXRRklJc65daIo2t97f2AQBJ+jdPHELPCA9/6qTCbzD61rFxGRVqNEXeK2BP4JrFXiuqeA47E1zVI/dgL+gK03T3oRqy/wYmzfqsCzWEu2rpLLDFbE7Argt9ia0paV6zO9N3BQEAR703UbvdeDILjDe39HGIZ3Oud0Uqt28rM+lgZmAcunG450xzk3NIqivbBZXAdhn01JEfAgMD0Mw+lK2kVEpBUoUZe8r2FTRZPTqRdiI7fnYQmc1J/B2Nr0U7HCcHGfYtX6r8ZGxu/FkvtS7/3k6PlfsPXoLcc5t1w2m907DMN9vff7Yic4ynkpCIK7vfd3tbe3/0vTdVP3CLBt7ucVKPRWlzrnnBsYRdHuQRAc4r0/mNIFTCPgfu/99Gw2e9XkyZPfr3GYIiIiNaFEXQZgFdvHlLjuHuAbgIpbNYYtsSnvmyb2R9jJlpHY2va4/NrzJViF919i1cZbyujRozObbLLJtlEU7QvshyV65YrsfRgEwe1RFN2ayWTu0Ohe3ZkGfD338270vhuC1IdgwoQJO4RheLj3/kuUriKfBe4Crli0aNE/zz333Lm1DVFERKR6lKi3toHA37Dp0UkXYcXG2msakfTXYKwI4LElrssn5XH/B/wGW3veUq31nHMrZrPZL4RhuL/3fm9guTKHdgAPee9vBW7NZDKPO+ei2kUqvTQOq6MB1qrtohRjkcoInHM7RlF0OPAlSo+0LwSuAa6YMWPGbRdddJH+7xIRkYamRL11DQL+Dhyc2D8POAG4suYRSSWNwWZKDChxXYR9oT0fawvWMpxzW0RRdAC2FnZbShfiA+sZf2sYhrcsXLjwznPOOaelTmI0uIOx2SFgrdrGphiLVJhzLsxmszvmRtqPAFYqcdiHQRBcGUXR5ZMmTXq41jGKiIhUghL11jQMuA7YM7H/NeAQ4LmaRyTVsDswHVunG/d34Cu0QM2BsWPHDhkxYsTO3vuDvPeHUnr6LFhbqAfyReAmTZr0BDYDQRrPZyi0DryN0h0spAk450JgJ+/917z3RwHDSxz2IjCto6Nj2uTJk2fWNkIREZG+U6LeegYCtwJ7JPa/AOyN9eGW5nEaMIHO7Y/+jK3jbbpk1Dm3WhRFB+RaP+1F6X7zAG8GQXCD9/6GuXPn3nv++ecvrGWcUjUZbGbQYKxV2xrphiO1cMoppwwbMmTIocDRQRDsQ+caE+3e+xuDILgkDMObnXMdKYQpIiLSY0rUW8+FdC4c9xTweUDVqpvP0lgRuTuBtRPXjQem1DyiCsuNqm2TGzU/ACuqV0oWa/F0QxiGNzrnNHOkeT2D9bf3wAhARcZayPjx41fKZDJHBUFwHLB5iUNmBEFwWTabveSss856qdbxiYiI9IQS9dZyMvCrxL4nsCR9Vu3D6bOvYNO6S/kQOL1Cj/Ntyid9rwHnVuhxamFNLFlfN7YvwpY6XJ9KRP2Qq9K+o/d+dK4idLn2afO993cD12cymeucczNqGKak50rg8NzP2wKPpRiLpGjChAmbBEHwNeB4YPkShzzuvb9o3rx5f9asGhERqSdK1FvH57D1mm2xfa8A2wOzU4mo734LfKeL6zej/+vshwDvAcuUuf7fwM79fIxaWwd4mOIvq58CO2JLH+qac25oFEV7AaOxYnDlnpvXc2vNbwjD8Fbn3JLaRSl1wgE/yf38daxtobSwk08+edCyyy57sPf+mCAI9qP4/0KAmcC0MAwvcs6pJamIiKROiXprGAU8S+cEbQesPVej6S5RPxdr0dQfX8XWcZfTiIk62AmbWymuBv8SsBWwIJWIujB+/PiV2traDvbeH5Jbb55caw82pf1+7/113vvrzzrrrFdqHKbUnyOw1pNgrdpOTTEWqTPOuTWiKDoO63CSbPUWYR0ffv/888/fOH369KYvuikiIvVJiXpr+CfFvdIj4IvADemE02/dJeozsS9f/SkWdAewVxfXN2qiDnAi1rotrm7aWDnn1oui6FDsNbojpVuoLcRmiFwbhuH1zrmPahmj1L3NgadzP1+LLfFICmjCYorSc6NHj85stNFGBwLfwroDJD9r3vbe/66jo+PiKVOmNNLyMBERaQJK1JvfkcBfE/tOB85OIZZKKZWoe4pfzwcAN/Xx/tcEXqfwpS1539DYiTrAH7A1m3kRsBM2Nb7mnHNbeO+/5L0/BNi0zGEfB0FwQxAE1wK3OOfqbgaA1I3BWOX3DNaqbYPE9YOwk5d/QwQ4/fTT185kMmO8998AVkxcvRD4i/f+gkmTJj1d4uYiIiIVp0S9uQ3BpravGdv3ILArjd1Du1Sifj+wS2z779j01744Azgztn0f9jeLa/REfRhW7X+92L6HsGS9FqOMgXNu21xy/iWKi9zFvY2Nml8D3KuWStILr2Cv7w5gKWBx7LrzgVeB36QQl9Qx59zA3Iye71CiaKn3/l+ZTOaC559//lpNixcRkWpSot7cTgUmx7aXYIXWXk4nnIoplagfB1wa214CrELvq9kH2Bf8eOJ4PPDHxHGNnqgD7A3cnth3BHaSo+JybdR2yiXnh1G+v/WzQRBcG0XRNZMmTXq8GrFIU9kWmIH1TI+7Dis6CMUFJvcHbsSWVjRcxwOpHefc5lEUfRfrNDIkcfVb3vvfZjKZi5xzc1IIT0REmpwS9eY1DHgDWCG276fAj9IJp6JKJerbYNO5t4jtOzF3bG/sDvwrtv0JVi09mfA3Q6IOcDXF63dfwJKaqBJ3nlsDuhvwJeAwYOUSh3msfdZVURT946yzznqtEo8tLWMp4Enss+3q2P5zKHzeHQ5MxwprPgeMBD6L9VsX6ZJzbvlsNntCEAT/A6yeuHpeEAR/6Ojo+MXZZ5/9VhrxiYhIc1Ki3ry+j03vzJsDrIUlno2uXKK+K8W/86PAdr2870uBY2PbF2JF1pLroZslUV8fS84zsX0H04+RxkSP88Ox5CgpwpKrG8IwvNw592pfH08EW6pyBoX360LsfZyfZfMT4CzgFmCf3L5laI7PQ6kR51wYRdEB3vvvBkGwd+LqyHt/UyaTOds591AqAYqISFNRot68ngc2jm1Pwr7INoNyifqbwLsUt/DaHGtN1xPDgPeB4bF9O2LVo5s1UQcrNnhkbPtG4MDe3MGYMWMGrLLKKvvkkvMvAsuWOCwL3AtcFYbh1c659/ocsUixFbB6BoOxdoOHY1OV8wnT34AnsNaNYIXmhiPSR865rbz3P/Tej6a43SXA3WEYnuecuxl1FhARkT5Sot6cdgIeiG23Y9P1ZqYTTsWVS9Qfp3Mrut5M9z8OuCS2/TKwIfblv5kT9W2BR2LbWWz2xTtd3SiXnO+VS84PAZYrcVg7cBdwVXt7+zVTpkz5sEIxiyT9CjgZS4zase4W51CoO7EW0Jbbfp7y3QVEesw5t4b3/nve+2/S+eTP8977c2fOnPnXiy66qD2N+EREpHEpUW9OP6e4J/ZVwJdTiqUaukrUvwhcE9vfm57q9wC7xbZ/jH3RH0JzJ+pgo41bxrZPpnOvdZxzbcCeueT8UGytb9IirEjdVUuWLLlu6tSps6sRsEjC6lhbxTYKLRUXYzNsPLbcIoOdiLoJW+IhUhHjxo0bMXDgwG8FQfBdYNXE1W9673+ayWQucc4tSiM+ERFpPErUm1O+LVHeaOAfKcVSDV0l6m1Y9ef4uugDsencXVkbeI3CeyLC2tq9Q2sk6v9LYVowwG3AvlBUrX209/4IYKUSt1/svb8dmL5kyZJrzznnHK39lTRcBhwNhF0ckwV+D5xUk4ikpYwZM2bAqFGjDsE+U7dNXP0h8NswDH+hSvEiItIdJerNZ3VsrWbeYmB5bE1ms+gqUQf4GfCD2HXTsTWrXZkITIht3wx8IfdzKyTqGwAv5jeCIFgwbty4AwYNGnRoFwXhstga4OlhGF7hnPuoRrGKlLMpVsm9u//bxlF8Ykqk4pxzu0RR5IC9Eld9GgTBtCAIpjjnZqQQmoiINAAl6s1nNMV9sO+jeDp3M+guUd+U4gJyS7CpiOUSyRCbMrtmbF+8n3grJOpgswdWDcOQk08+mREjRpQ6ph24w3v/9/b29ms1rV3q0PVYr/RMF8ccCVxZm3Ck1U2YMGF3YHwQBJ9PXDUPuLCjo+OnkydPbpYaMiIiUiFK1JvPWcBpse2fAz9MKZZq6S5RJ/fzVrHtk4DflLm/vbE11XmfYP2+F+a2WyVR/28hvsMPP5wNN9wwvz8+cv5X59wHKcUn0hPbU6j2Xs6OPThGpKImTJjw2SAIfgh8heITSYuDIPhTEARnqhuGiIjkKVFvPn8BjoptnwD8MaVYqqUnifrJWBXovMfovF4w73JsXWv8/k+MbbdKov7fkzwbb7wxe+yxxxsjR46c3NHRcfWUKVNmpRybSG/cD+xA+VH1VbBWjCI1d/rpp2+UyWR+7L3/ClZXJW9BEAS/b29vP1cj7CIiokS9+dxPcQK5F9Yeq5n0JFFfDniP4p7qn8XWr8YtjX1hHxrbtx3waGy7VRL144E/xLYvB76WUiwi/XEAcEOZ69qxlotR7cIR6ey0005bs62tbbz3/hsUJ+waYRcRkS4r40pjSi4sbtUCXx9ja1XjSiWdR1KcpL9AcZLeSpKvlZKL1EUawI3A09iyjaR3UZIudeDss89+68wzz/xWGIYbAJdSaCM6yHs/JoqiV37yk5/87NRTT10hxTBFRCQlStSbz9DEdnIkuJX8KbH9NWBAYt+xie1LqxZN/Ut2BlgqlShEKuOndJ767oE3UohFpCzn3OsTJ078RjabXS8IgosoJOxDvfc/GDBgwBtnnHHGVOfcMmnGKSIitaVEvfkkR4pa+Tm+heJ1qCuR6w2esz62jjWvA7iiBnHVq2RSU2o0UqRR/A14E0vO83xun0jdiY2wfyaRsA8DxkVR9NoZZ5zhfvSjHw1PMUwREamRVk7imtX8xPawVKKoDx3YOuu4r8d+Po7iOg3JxL7VJF8rydeSSCPJAj+j+D0eAm+nE45Izzjn3swl7BthJ4/zJ+CXA34yePDglyZMmHCSc25gelGKiEi1KVFvPsm+1qNSiaJ+TEtsHwwsj732j+7m2FaTfK18nEoUIpXzBzq/jt9KIxCR3nLOvTpx4sSvhmG4GTCdwuyQlYMguCCKolcmTJgwxjnX1sXdiIhIg1Ki3nzeTGyvnUYQdSRZHG4g1r5uX2D12P6PKV8lulUkXytvphGESAUtAn6R2NfdiPoQYFlgBWCd3GUrYOvcZcPcvlVzx6noolSVc+6FiRMnHh6G4Q7e+ztiV60RBMGFURQ9fcYZZxyUWoAiIlIVOgvbfF5LbG+eShT1ZRrFPdS/Tue/0xXA4loFVKc+m9hO/o1EGtGvgdOxk3QA2wM7YYn4SthMkvzPy/XjcRYCH2DLZz5M/Pwu8Crwcu44kV5zzj0C7DNhwoSdgyA4G9g9d9XGwHVnnHHGQ2EY/q9z7v70ohQRkUpRH/Xmsz9wU2z7KWDLlGKplp70UY9bFuupPji2r4PiE1VbA0+UuX0r9FEPgVlAvKrwhsBL6YQj0idtwKbYSaf1Y5eNqY8T0x4b0X85d3kR+D/gMeCTFOOSBpQbRT8b2Cy22wdBMD0IgtOcc6+mFJqIiFSAEvXmMxIbwck/txE2RXNGahFVXm8TdYArgcPLXPccxV90klohUd8BeDC2/TG2lt+XPlykLqyMvffz09J3ofhkUyN5HXgA+xx7HHgEWJJqRFL3nHNhFEVfAs4F1opd1R4EwaXt7e1nTJ48eWY60YmISH8oUW9Oj2JfXvO+iRVVahZ9SdSTMw3ifgCc38VtWyFRnwycGtu+EjgypVhEylkO2AfYD9gbWK0Kj7EQW9veAczN7ZtD4aTVMGwa/cDczyHVWac+H/ucuSV3eaEKjyFNYuzYsUOGDx/+fWAcxa/HT73358ybN+/8888/X8suREQaiBL15jQBmBjbvpfCWrZm0JdEPcSqPSe/2HdgReW6mnHQ7Il6iI3mrRnb91Vau6e81IcQGynfDzvZth2Q6eN9zcHe7zdi7/cZ2OyjGcBMbE35h/R9FskwbIR/JWzN+yrAirnL2tgU/DX7Ef/bFJL2OyicRBD5L+fcclEU/Qj4PjAodtW73vuJmUzmD865qMzNRUSkjihRb04bYOse88+vx9ZovphaRJXVl0Qd4DTgG4l995TYl9TsifoBFFe8X4AlGVozK2n5LHay6Chs6U5vvI9NG3+BwjrwV7AaDDtSvMSj1gYB62Gf0evn/t0G2IjeJfCLsRMOl2MzhVq9EKYkOOfW8N6f5r0/geIOP4+FYThWBedEROqfEvXmdS+wa2z7EuD4lGKptL4m6n3V7In6PcBuse1pwHHphCItbDXgS8AxWDu0nmgHnqF4bffzVYmuupYCtqCw1n5Xitcbd2UOcD1wGXAnqishMc657aIo+jnF/1954G9hGP7YOdddu0IREUmJEvXmdSTw19h2OzZ680Y64VSUEvXK2RP7ch+3LVaFWqTaAmxa+/exNedh14fjscQ8PwX8AeyzrRl9hsKU/92BoT24zRvAb7CaJJoRI/+VqxD/C2Cd2O6FwK/CMJzsnPs0nchERKQcJerNK4ONLG0Q2zed8pXPG4kS9coIgU+xtbV5D2CjeRqVk2oaio2cfw9rA9iVucDNWGJ+K9ZqsdUMxpL1/YCDgHW7OX4ucCnwK+C16oYmjcI5NziKorFY4dDhsave896flslkLtP6dRGR+qFEvbl9DZsOGbcXcFcKsVSSEvXK+B9s9C1pBjaifj+WuKtNlFTK8liXhTFYK8ly2oHbsIKG19L5/dfqdgC+AhyBFasrJ8KmxZ9DumvzpY4455b33k/w3p9IcW2Ex733J0+aNEmvFRGROqBEvbkFwENYpeS8N4HNaeyKwUrU+29VrNjW0j04dh7wMIXE/SEa+/UjtbcUMBY4ha5fcw9jBdKuxCqwS9fagM8DRwOHYp9V5VwLjEdt3iTHObdxNpv9WRAE+8V2e+Dyjo6O/1X/dRGRdClRb347YglWfO3nFVhF5UalRL1/BgC3U9yybxHwe2BTYBdsqm05WeAlCon7fTRH7QOpvAFYYUKHtS4rpR24Bjgfjfr2xwjgWOCHWMvJUiLgKqzXtt6zAoBz7rAoin5GcQHDOcAZYRj+zjnXkU5kIiKtTYl6a/g5NpoV9yPgpynEUgkTsRGkuC8Cz1Xp8QbTuZL0E8DoKj1etZU60XFibj9YcrU5lrDvjBWc62qaMlhLrHzifj/wJJYUSOs6CPvsWa/M9bOAC7HlF6247rxaBmK1SL6PVZAvZRFWWGwiVlBMWtzYsWOHDB8+/LvA6dgMmLyXwjD8rnPutpRCExFpWUrUW8MQ4FFgk9i+LPZF+uZUIpK0fIdCQp53O7AvXReQW4dC4r4L1ve5q8+Pudg05nzi/gBKCFrFisAvsc4TpXwEnI0l6XpNVNfnsL/1jmWufxWrF3B3zSKSuuacWy2KoslYjZv/8t7fEEXRSWefffZbKYUmItJylKi3jvWwxGm52L45WOKlNYutYU+scvaA2L7XgO2x0c3eGIW1ccsn7ttiI3nldABPU+h3fTfwn14+ptS/0dgI+QolrpsP/BqYglqH1drewHnAZ0tc57G6AN8HPq5lUFK/nHN7RlH0K4pP8C8Afppr56YCoyIiVaZEvbXshSVqbbF9HwD7YL2JpXntDtxA8ZTGucBOVGbJwDBgSwqJ+87Ast3cJjld/gnUFq5RrQRMw9qHJbVjfb0nYh0FJB0Z4OtYvYBSa9jfA76BtcATYcyYMQNGjRr1P9h7N14E8iXgxIkTJ96ZTmQiIq1BiXrr+R62NjFuNvYF+5HahyM1sB/wT4orQkdYlejrqvSYGaw/dj5x3wVYu5vbfIq9BvOJ+/3YWlqpb1tjr681Slz3NHAC1u5P6sNQ4AysAn8mcZ0HzsWqw6vGhAAwfvz4ldva2n5KcW0YHwTBpUEQjHPOfZRWbCIizUyJemu6ADgpsW8OltA9XPtwpIoOBv4ODErs/wFWZbuWVqF4xH1LirsRJLVjMz3yiftd9H6KvlTXGOzzJLnsYSGW8J2NPY9Sf3bEZjpsXOK6G7E1yrNrGpHUNefcblEU/Zbi6fCzgTPDMLzAOaeTOyIiFaREvTUFwM/oXAl+IVZs7E81j0iqYQy2Jji+Jt1jSXpyVkUahmPr4/OJ+8503Qca4HWKC9Qlq/FLbQzGihIeV+K6e7BR9FdrGpH0xSCsyvePKV4SBfAKNutG7zH5r9h0+LMoXkr1eBiG33bOafaMiEiFKFFvbWdiUyCTLsJG3DUS1piGYEnUsYn9HvgulrzXozas2FU+cd8dqyDelRnYtOp84v4IoCJH1TUUuBYrUBbnsdH1H2LFA6Vx7AZciRWJjPsE2B/1t5eE008/fe0wDC8ADojt7gB+uWDBgp+cd95581MKTUSkaShRl3HA1BL778N68ar4U2NZD1svvFlifxb4JnBpzSPqn+R0+a3o+nNrPvAUxUXqNH23coYB12Ntv+LmYoXI/lHziKRSVsWevx0S++djS2juqnlEUvfOOOOMg7ATdGvGdr8bhuFJzrlrUgpLRKQpKFEXgOOxUdbBif3vAd/GvphLfQuwROlnwIjEdbOx9aY31jqoKlga2I7inu7J121cFqtQnE/c7wXerG6ITWsZrGvE9on9L2JTpF+seURSaYOwpOubif1K1qWsU045ZdjQoUN/grX4++9SqyAI/hIEwVjn3AfpRSci0riUqEvelsBVlK7MPR2bCq//bOvT2sCFWJu9pGeAw7B+6c1oALA5hcR9T2BkN7dJtoV7ElW47s7SwN3YjIa4x4B9Uf/tZnMGtjQqbiFwEKCWXFLShAkTNguC4GKKT+bN8d6PmzRp0sWo/aaISK8oUZe4kcBfKZ3wzcYKDl1U04ikKyFWtOtnFBf1yfsrNjLWamsF16F4xL1UVeu4uVi3g3iRuoXVDLDBhMA1WJIW9yjWKUJJenM6BfhpYt9sLAl7pfbhSCNwzoXZbPaEIAjOwwqG5t0XhuEY55xm3oiI9JASdUnKAD/BkvIBJa6/GzgVtXFL277AFGwmRNI8rPbAb2saUf0aBWxLIXHfls7txOI6sP7f+cT9Hlp7NskU7PMg7j6siNTc2ofTZ6OBZctc9wL2XFfC0dha/lKewgoeNopvY58j8e8KL2Hr2OekEpE0BOfcKtls9oIgCA6L7V4EnDN79uwpF1xwweK0YhMRaRRK1KWczYA/YklNKXdg1Z2fqVlEArY+ewo2xbuUe7BRdI14lTcMO8ERL1JXLoHLS06Xf4LWmMZ5JPAXiv+veBh7/S1IJaK+e57ysyteBjak/8/pBnS9Vn8KML6fj1FrpUbWb8TWrGvJiHRpwoQJhwdB8EuKOwo8670/ftKkSY+mFZeISCNQoi5dacN6rZ9J6f7WHcA0rJ/qW7ULqyVtij0Ph1L6fTsb64/+J1ojgaykDJak5RP3XYG1urnNp9jIaD5xvx8bLWomm2JJ+dDYvvewk3fvpRJR/3SVqAPsRP/bkJ0L/G8X1zdiog72Of/1xL5JlG7vKVLEObeM9/5M7/1J2FIagI4gCH47f/788WrlJiJSmhJ16YnPAL/E+umW0gFcDfwC+HetgmoBATbFfSxWN6DU+zXC1qKfglrpVVKyLdyWFL5gltKOzS7JJ+53AbOqHGM1tWFJ6zaxfYuAPWjcZS/dJeoXYlO9+6oNeBtYuYtjGjVRH4wte4q3buvA3huNNJVfUjRhwoTdgyC4CFg/tvvlMAyPd85VaumJiEjTUKIuvbET1nN91y6OeQJL6v+KJS/Se4OAI4AfAZt0cdwd2Fr0J2oRVIsbjhXRyifuO1N6lknc6xQXqHu+SrEdAdxKZdcMn4bNlIk7Fpux0ai6S9Q/wU7Q9HVK/wHADd0c06iJOtgJiMewv1He89hJLH3WS4845wZHUeSwk8uZ3G4fBMHFCxcuPOXcc89tpLoXIiJVpURdeivA2n2dhU0XLuc9bG3r5VhhrnLWwRKaVhdgyd/RwOHAcl0c+zBW3Otf1Q9LymgDPkshcd8dWLGb28zAEp184v4IsKQCsWwN3AScDfyO/idNq2IFw+IF0f4KfKWf95u2ZKI+H/iQ4mUOR2OfW30xHfhybPs5bPlAXCMn6gB7A7dR/N3he8Cv0glHGpVzbosoii6huCDqm8CYiRMn3p5OVCIi9UWJuvRVG/BVbF30Zt0c+zxwRe7ydm5fgFWX/xE2rfuB6oRZ9zbGEqCj6X5d9H1YK7br0Dr0epScLr8VXX/GzseqgMeL1M3u42P/AkuYXsfWSP+zj/cDndcjf4C9Tht5Kj90TtTnAudTvM76Nmy5SW8th52cHBTbNw44J3FcoyfqAJdisyvyPgbWo++vXWlRY8aMGTBq1KgfABMp7sQxPQzDbzvn1PpRRFqaEnWphL2B72Nr2Ltax+uBh7AvwzsCn8/t/wgbeZpZxRjrRRu2hGA/4AvYqGxX2oG/Y4nYY9UNTSpsaaxKf7yn++Aujs9iI9n5xP1ebISpJ4bnbptfH/0odhKtt+s+18WqlrfF9o0BLu7l/dSjUon65tjJjfz/hRF2wuw/vbzvkykeVX4EcNhMh7hmSNRXxKrkj4jt+wmWbIn0mnNu89zo+tax3e+EYfhN59wtacUlIpI2JepSSRtgo3pfxRKHnroKK+L0UTWCqgOrYon5fthJjWV6cJsPgT8AvwHerV5oUkMDsMSw3hktAAAgAElEQVQwn7jvCYzs5jbJtnBPUr4l1mHYewks6Q+xkzynAm/0MMaLgRNi289jJ5OyPbx9PSuVqC+NLSHZPbb/NGByL+/7cWwGRd6J2N+8GRN1sN/h7Nj2LGBNbJaISK8559qiKDoRe+/FO01MX7JkybemTp2qGRsi0nKUqEs1DMams38N+CLFU9q6Ek9KHsdGkBut5VU8Gds6d+mqgFXcIqxA3GXAtVRm/bLUt3UoHnHv7rUyF6tREC9StzB2/XVYUbP8zJYsNpPlEiwB7epk2Ahs+nb8S/JXsPXpzaBcon4sNp077xXspGNPl5dsCjwb216CLYPYjuZN1JfGZnssG9t3PPY6E+kz59yGudH1HWO73wS+MXHixLvTiUpEJB1K1KXaVsCqUh8PbNHL2y7CRhCfxaZavohN730Taw2UpoHYuswNsFYz62NFcTajeNpwd7JYK68rsHXFqnjb2kZhfcrzifu2dH2iqwMr1phP3F/N/TuE4s93j/V+n4Ito1hc4r6+jRWjy3sbO5HQDKPpUD5RH4adJIzPAtqZnreaPB9b+pN3JXAkthSoWRN1sN/lx7HthyhOrkT6xDkXRlF0MlbjIV/3wQdBcHEQBD90zs1LMTwRkZpRoi61sCM20rd8bnsRlmAs1cf7W4KtK30ZW0v6IVbw6v3cvzOxCtt9nYa5NLbWd8XcZWXshMMoYHUsKV+LQmuZ3voAa6d1M3A7zTvlX/pvGHYCKF6kbtkub2HTkEtNqffYZ/67WLL4Z4pHje/G+qTnOeDMPsRcr8ol6tC5QNpFwLd6cJ9twDvASrF9X8De282eqK+NnRgKE/veTCUaaTrOuY2jKPoTsE1s9xve++MmTZp0T1pxiYjUihJ1qbZDsdHifM/p17C12m9SKKr2eWwdbG9GonuqHZiHJST5PtOfYF8uh2Pvgfya8aXpe/LdlcXYNP5bcpcnKL/OWKQrGawtYj5x35XuuwUkRdjr/0Gs4NxDWNXymRS/B9fD3q/NoqtEfXeK2x1+ip2g666n+iHA1bHt9/h/9s47XI6y+uOf2XtTCUnoISAQOihIC4j0Ir0JhF4CKCBVEGnmXoYbiCBRqogBJXQ0gFIEpChIEQKI9KIU6RBKCun3zvz++O78dnbubL27O1vO53nmSead2Zmzu3Nn57znnO+BFVAWQrM76qBOFJuH1k9AuhqGURHStes/IVsZPoiun+K6bqG/UcMwjIbFHHWjmpyI0myDiMvTwO4oAh6lL7Xd9cpCJCp1A41Xa280DuG2cFuj8oti7u1BhP1OlOFxZWjbyxRuu9ho5HPUHVSbvkpoezE91f+MdDgCLkDifdAajvpPgV+E1u9EkxeGUVE6OjrWdxznOrLvS6+mUqnDXde1jiiGYTQl5qgb1SDokX5OaOxOJExVyuz3SJT2uyaZOvA1UQp60vgo7f5NVDf/Bqqhfx64AtXlg9LxN0KRNsOoFg4Sb7wMOZ+l3tsDpz3gYhRtbybyOeqgfurhVP8HybSQjGNplPbeLzS2NvBa+v+t4Kivh+55AZ9SH/dnowkJ9V0fT+bvrhv45SeffNIxadKkhclZZxiGUXnMUTcqTX+k/HtwaOz3qN6zUgJwQ5GI28qoNnQp5NQvRXZN+aBcByjAbDJ17tPS/36a/v9HZOrjc006DEIpoUFP2CdRK644AS/D6CvroZrq0WTS2vvKIahkpZko5KiviP62g8/PQzXX7+U43qnAL0Pr/0TlPAGt4Ki3o5KiRUJjK1B6H3rDKJqOjo5N0tH1NULD//Q879Dzzjuvmcp1DMNoccxRNyrJEGAKqjsHRem6kChVUvRDdoVr0Yehh/BZ9K5dr1Tt+IrAM2jCAGAycESFjm0YoOv6XOBk4rUVgtZsTnopxYF/HU2u/aOPNtYThRx1gIfRpFrAOLL7hYd5AZXrBBwL/Da03gqOOsBUNEkUsDVgQl9GVXFdd6DneS4qvwjubXOBs7q6ui6j+PaKhmEYdYs56kalWBb4C0pVB0XPjwOuTsyi5NkcPfgHAjjHk10HbBjlsgdwOYpegpzO2Ug4cRbwZXp9dnp9Rmh9ZnqZHXrNk2ScfR9NAjSbSFMxjvqhwPWh9Vw91TdEApEB89A9cHporFUc9VvJlPqA1POvS8YUo9Xo7Oz8HurasFww5vv+/T09PUdOmDDh4+QsMwzD6DvmqBuVYC3UjmjF9PpsYD96P6S2IicghwokLrcjaoNlGOUyEJV6hB3tvjAc+Cq0PhNlnTQbxTjqg1B5y/DQ2OaoT32YK9DEW8DNZJf7QOs46pcCJ4XWT0EiooZRE1zXHe553hVk/w1OA37Y1dV1Z0JmGYZh9JlK1DIarc13UHps4KR/ilodmZMurkD1w6A0/Cmott4wymUeqqX+lL476ZBdXwxy/luVucBtkbHDI+v9yY4gQ2tHkKPX4JBErDBaFtd1p3d1dR2CAgTBpONSwJ87Ozuvd13XrknDMBoSc9SNvrAX8DdgyfT6W6iv83OJWVSfnECm1ncJ4A56O0eGkRTRzKpWr+2MOt37A4ND63uSuecBfIhKXFqVqK7H1sA2wKK1N8VoZbq6uqb09PSsT7ZGwqGe5z3f0dExOtfrDMMw6hVz1I1yOQpFhwNl9anApqim08hmIZrpD5SQv43qYK30xKgHorXog2P3ah0eR2J6AUOB74fWx0b2n4yE+1qVaLRyOzSB+xUqN7geCR5uiD1zGFXm/PPP/19XV9c2wI/JdFpZ1XGcJzs7O90xY8bECW8ahmHUJfajaZSKg1Tcr0GteQDuRhGUaQnZ1Ah8iiJxgVO0N3BWcuYYxv8zi+yo6FAyAoityo2R9SD9fQS9e6tH9201lswx3oY0AQ5FNevPIrG9x1Fd+xgyXTEMo5L4XV1dl/q+vwmaLAI9r5yz1lprPei67vIJ2mYYhlE05qgbpdCO6q3PCY1di5zOZlOIrgbPA0eH1scDuydki2EELATC6sgpMpoTrcpksqPk2yGF/cPJTFCCRObC0fdWpBTNjUWBzZD43B+Bz5B43x9R1H1zYEClDTRak/Hjx7+QSqU2chwn3K5tG8/zXu7o6DggSdsMwzCKwVJvjWIZgh6mdg6NXQicmYw5Dc1FwGnp/89CJQOv5N7dMKrOP5C+RMD3gT8nZEu1KEb1PcwDwPdC6+OQqvRaobEfouyiOFpB9d0BvgAWq+AxFwIvokmQ54DHgHcqeHyjBXFddwfP865DWTEBN8yZM+dHEydObGUBTcMw6hiLqBvFMAKJswROeg9wDOakl8sZZB7gF0XicsNz724YVef5yPrGiVhRX1wbWf8p2U76HDR52cqsSWWddFB3jA1R1P061OHgI1Ri5QLbk9FGMYyicF33gVQq9W3f98OTZ4cOHjz4Wdd1N0jMMMMwjDyYo24UYhUU0Qh+yGajWutJOV9hFMIDDiEjvLc6euA3kRsjKZ6OrG+biBX1xZ9QTXVAtLf8HajnfCtTq+tkWWA3VHb1IPrcXwF+CxwGfBPLEDQK4LruZ+PHj9/N9/2fAgvSw2t6nvdkR0fHqdg1ZBhGnWE3JSMfGwP3kBH8+QLYA3gyMYuaizWBp8g4AL9A0XbDqDUjgQ/I/CZ4wPJk1643OqWmvgNchbKH4tie/G3ZWiH1/T5gp9D6YygAsAG1j3pPQxNOT6WXZ7CJFCMHrut+y/O8m4F1QsMPd3d3HzphwoRmuu8ZhtHAWETdyMUewN/JOOlvA9/FnPRK8jpwABnRqtOBI5Izx2hhPkKOTUAKODAhW+qJaE/1gA+AR2poRz2yDBLZC3M0EoQbCnwLTXLcALxKRsyrWiyFou7nAQ8BM4C3sPZwRgyu6748a9asTdJCcwHbtbe3/7uzs3PHxAwzDMMIYRF1I44jUUphoG78DHoA+iwxi5qbDqAr/f95wJZkO02GUQtORwKRAa8iZ6vaDlatKCeiDvoc1oqMdZHd/SKOZo+on4neT8CrKAU9F8OA0ciR3xBN/C5eNevi+Rp4AYnUPY60V+x3rcXp7OzcB4lCBloxPcCE11577dwpU6b05H6lYRhGdTFH3QjjoIfP8APog8A+6KHWqA4OcAuwf3r9I/RA+1FiFhmtyDLA+0jMK2Bn4P5kzKk45Trq5dLMjvoAFK1eLjR2CuqXXgork3HcNwPWp/ZR74+R0x6ozD8DzK+xDUbCuK67QjoVfrPQ8NOe5x143nnnWdcBwzASwRx1I6AduBK1GwqYjFIZFyZhUIsxCNV3bphefxIJNdkDo1FLpgD7htafQI5UM2COeuU4BtXvB8xBfea/6ONxhwDrkXHctwKW7uMxSyXaHu45rH1mS+C6brvneeNQllswYTQjlUr9wHXd2xI0zTCMFsUcdQNgEaQ6vkto7ELgLJon7bURWBFFcwJdgMlYzbpRW9YD/kX2b8PeSAG90TFHvTIsCryBlNgDLgZOrdL5RiKnPYi8b4Qi+rXkYzJO++NoInVOjW0wakRnZ+d2SFshuMZ9x3Eu//LLL0+//PLLbfLcMIyaYY66sQzwFzKR3B7gBLKjJUbt2BwpSfdPrx+PMh0Mo1bcjTQpAt5GtepzkzGnYpijXhkuRHoGAXOB1YAPa3T+fsC6ZBz3Dcn+XmtBN/AmGcf9CWojmGfUCNd1l/Y873ogLCz3L8/zDjjvvPP+k+t1hmEYlcQc9dZmZVR/ulp6fQ6qk74nMYsMgBOBQIl2IbADpjBt1I41UepvuFa9mhHTWmGOet9ZH7VAC18bxQjrVZtlUaQ9SJn/LjC4xjbMBKaSSZl/kr6XAhgJ4rpuqqen52zHcc4hI647E/hBV1fXlARNMwyjRTBHvXUZjRzyoP7vS9SS7YnELDLCTCKjF/AF6mn/dnLmGC3GZWjCKMBD/bIfTMacimCOet9YFPUnD3+GH6CJndmJWJSbdmANMo775ki5v9bPPG+TXev+NKb50nB0dHRs4jjOLcCoYMxxnElffvnlSZYKbxhGNTFHvTXZAbgNPXgBvIMeKt9IzCIjSj/UC3jL9PoL6IGz3h6IjeZkEXTNrRIa+wpNGP03EYv6jjnq5eMAfwDGRMZ3Ae6rvTllMRRdv0HK/KbAEjW2Idoe7h/ApzW2wSiDs846a4n29vbJjuOEy4KeSaVS+7mu+25SdhmG0dyYo956jEXR2iB18VlUj2oPC/XHMuj7WT69fgdS5LY6SKMWbI0mi9pCY68iB2dmEgYZiTEeGBcZ+z1wVAK2VJJwe7gNkSPfL+8rKk8gVBfUuj8LzKuxDUZxOJ2dnScBF5G5TmamUqmjTBXeMIxqYI56a3EGcEFo/SHUI90euuuX9dEDXFBveTaKyhlGLTgdiYeFuRvYC6XDG83P/sAtZD8vPIuyfRpdYDBKtD3clmjCtJZYe7g6p6OjY4t0Kvxy6SHfcZzLHcf5qeu6C5K0zTCM5sIc9dagDfg16n0bcD3wA6xerhE4BLWKATlHe2KCf0bt+B1wZGTsVuBQpH5tNC+7AVOAgaGxT5DGyQeJWFR7RpJd674h2Z9HLbD2cHWG67pL9vT03OA4zk6hYUuFNwyjopij3vwsgh6qw3VVlwE/xlKoG4mJwE/S/5+F0o8tymLUgoGo68AmkfHbgIOwyb5mZT/gRrJTwecB2yBRuValHtrD9SBNmXDK/GtYlkutcTo7O08HzidTIvQ5cFhXV1ejaDcYhlHHmKPe3CwB3IVa1YB+3E8EfpOYRUa5pFDK8S7p9TeR4zQ9MYuMVmI5FMVbITJ+F3LoTPm4uTgEmEy2PoEHHI6cdyObcHu4DZETP7zGNswEXiLjuFt7uBrR2dm5HXATmTIJz3GcCa+++qo7ZcqUngRNMwyjwTFHvXkZhXqkr55en49SVa33Z+OyGGrvE/S9fwA57vYgYNSClYCHkQBXmL+jOuZptTbIqDgO8FOkg5EKjfcg4bjrkjCqAWlDbevCKfNrkv2Z1oJoe7ipgNVQVwHXdZfu6em5yXGc7YMx3/cf6enpOWDChAkm1msYRlmYo96cbAT8hewe6XuimXajsVkTpZ0OS69fCJyZnDlGizECOevRVN8PUEeCp2tukVEphiAl92gLth7gCDI6GUZ5DEUp84HjnkR7uNnAv7H2cFXBdd12z/O60G9y8Hz9fiqV2td13akJmmYYRoNijnrz8T3gdjI90t9F/XxfT8ogo+LsAfyJTHTmYODm5MwxWoxlgAeBdSLj81FpzdU1t8joK6uh9o/fiowvAA5MbzMqT7Q93Gigf41tsPZwFaazs3NXNLG1WHqoGxjX1dUV7aBhGIaRF3PUm4vD0UNyIP7zEnLSP0zMIqNadABd6f/PQ22EnknOHKPFWBL4IxIWizIZOAXTT2gUDgMuRxHfMF8gJ/3BmlvUuiyCWnIm2R6uG2mgBI67tYcrA9d1V/A87zY0+RJw05w5c46ZOHHi7KTsMgyjsTBHvXk4A9UVBt/pw8DeWI/0ZsVBav77pdc/Qg8EHyVmkdFqtCG149Pp/VvyCXACyu4x6pORwJWoLCrKv9Hvxzs1tciIox7aw32CIu1Brftj2ERcQVzXHej7/uW+7/8gNPya53n7nHfeea8lZphhGA2DOeqNTxtwBXBsaOwGJPxjbZOam0Eo6rFBev1JYFtMgduoLXsC19M7IgtwD7o3WVZP/eAAP0QtHxeN2X4TcDTWp7teaQfWINtxt/ZwybMHmszoNVne0dFxmOM4V6HfbIBZqVTqSNd1b6ulgYZhNB7mqDc2A5FTvm9o7DKUdtrKP5itxIoo5X2p9PpkJPxkGLXkWyjD45sx275EZRq/wRSnk2Yz4CIkZBZlDnAa1r6zERmBMqqSbA83C3iRjOP+T9RTvFUYiRz164ELiGQcdHR0rO84zu2oIw+A7zjO5R9//PFpkyZNsqCKYRixmKPeuCyOehhvll7vAU4Gfp2YRUZSbAE8REaE6DjsYduoPf2AU4FzgQEx298HzgN+h7UUrDVroe9lX+J/9x9DUfY3ammUUTXqpT3cx2TXujd7e7jjUYbjl8AvUOBkbrDxrLPOWqK9vf1Gx3F2Cr3m4VQqdYDruq00qWEYRpGYo96YjALuQ+lvoFTnw5C4k9GanARcmv7/QmAH4JHErDFamXWQMz46x/YXgbNRC0mjuqwEuMChxDtpM1Df9GsAv2ZWGUmwKPBtMo77d5AoZC0Jt4d7DrWHe7fGNlSTFPAo+nxBbSvHkz056XR2dp4OTCDzN/mB7/t7jx8/3gRhDcPIwhz1xmNd4F5gufT6V6hG9LHELDLqhUkoKgZSbN4YeDs5c4wWpg1l+LjE10EDvABcAtyC6SpUmo1RCdQ+ZLqARPkTaqdn+gGty0iya93roT3cc4Si0A3IWsDzZGcVvYwmJ+8OBjo7O3dGehBBC7e5wDFdXV031MhOwzAaAHPUG4vtUD/bQLTpQ2AXFKEyjH5I7X+L9PoL6CHMWsEYSbEEitieTG6l6s+Aa1GLMHMayycF7Iqya7bPs99TwJko8mcYYcLt4TZEvyUr1diGuPZwr9JYGR+dqNQkyuOoQ8+TAOPGjVsllUr9CWUhAeA4zqSPP/74BKtbNwwDzFFvJA5D6YlBdORl1CP9g8QsMuqREUhcbvn0+h2oLrWRHnKM5mMlJCh3MLnrZOcBU1CU6SGsjr1YVkOf61gkLpmLl4CfEYrqGUYRRNvDbUBGvbxWfIp+14KU+cdRNmG90h/Z+a0c2+9EEfZXTznllEFDhw69yvf9w0LbH0+lUmNc1/2k2oYahlHfmKPeGJwMXEzm+/o78H1UX2gYUTZApRCD0+tnIRVaw0iadYEOdP9qy7PfJ0hF/iakpGxkszSwP3LQNymw7ytI2OpGrBuI0Xfi2sOtRW2fJ8Pt4QLH/Xnq6/reGEXOc93nPOB2FGF/p6Oj42jHca4gE4z5MJVK7eu67lPVN7XiDEfXxibAquh+NSy97Ss08fIG8DTKmrCsP8PIgTnq9U0bUg09LjR2GxIGmpeIRUajcAhq3Qd6INgT9bQ2jHpgJVQf/QPi+6+HeR1lhtyH0ra7q2pZ/TIKZVHtjlLb2/Ps6wN/RRO8D2IZNUZ1GQ5sRMZx34xM7XWtiLaHewqYVmMbolyKSlHysQC1VR3X0dGxtuM4f0SOLcB83/dPGD9+/DVVtLGSbA+cgO5TxWodzAH+jD6rqVWyyzAaFnPU65cByNEaExqzHulGKfwStcsCPcRsiqJrhlEvLAociO5raxax/2yUUXQ3ctzfr55piTMQOT7bp5cNi3jNfNT940Lsb91IjnB7uMBxX5/k28M9Q22FKxdBkwcrF7HvLOBXY8eOvWmFFVa4iVDXDMdxJjmOc6LruvXa2m5t4Coy+jjlcidy9Bu1pHMq2df4ZNSuzzDKxhz1+mQxdMMKbno+So+6KDGLjEakDTk0O6fX30SpaNMTs8gw4kkBW6NU7n3IpEnmwwdeQ+mTwfIyjRtxXx6ly34n/e9oMuUr+fBQm6sbUSqt/X0b9Ui0PdwmwFI1tmEOSpEPUuYfA96p8jl3Rp16iuWDIUOGTDjllFM2dxznoND4E93d3ftMmDDh0wrb11d+iIRABxTasUhmoIzARswA7Ca71OECVHpoGGVjjnr9sRKKFAXRpflIJOjWhOwxGpvF0Czvqun1B1CnABPqMuqVgSi9+2BKS6EERdz/hZz211Da/JvA5xW2sS8MAFZPL2uglOGNybTcLJaXkHN+C82dWWA0L9H2cBtROYevWIL2cOEWcZVuD3cjup+Vwuv77LPPP775zW8eQXbd+l6u69aLbsf5SBQvjh7gETRJ8T76nB30na8I7IGy/OKyLDzgaNR/vpEwR92oOOao1xfroJtaoNg9HdUW/yMxi4xmYE1UrxdEKe3Hw6h39gdmIod75/SyA+VH4L5ADvvrwNuoDdw0JGr0cfr/lUiJddI2LgUsAyyb/v8o5JSvjh5Sy0kBXoCciPuBv2Cp7UbzMRiJoQYp85ujv51aUo32cEukj7F0oR2jrL766q+NGTNmZFtbW/D7PTuVSo11Xfe2PthTCc4Efh4z7qE0+HNR6818rIDKdA7IcZyDaawglTnqRsUxR71+2BYJJgU3449Q5POFxCwymok90fWVQg8cB6NInGHUGwej2j4P2AtlGIGu3Q2BnZDjvjH5leNL5SvgSzKp4zPSNsxGTvJCMpGt4ej3c1Ek6jY4PbYU+UXeSuU99P7vBx5GdayG0UoE7eGCWvfNqH17uBmovj1w3MtpDxcWeC2JoUOHcsghh8xacsklF00P+cAvUqnU2a7rJqFZtAsqq4tOOH6G7tn/LPF4uyGHfJHI+BzguzTOc7A56kbFMUe9+iyJHv7y3UwPQSk+QYrnK+hB1NIZjUrSiWa5QV0DtkQPH4ZRL/wI+DWZ36bJwBE59h0CrEfmIX4LVDrUqCxEwlOBM1CJSJ5hNBuN3B7uLlTWUzL9+/dnr7328tdcc83/f5++7982d+7csRMnTqxle7Nh6Bk1WqrzOQo4vVTmcTdHE5JRZ/1FVBKxsMzj1hJz1I2KY4569ZmEam1ycTLwKzIzk4+gHsMmCGRUGgfNWu+XXn8PCVYVSk8zjFpwPBIlCn6XbgYOpzRxuBWQSNW3ydSBr07tI3D58NDf3hsoxfZVpCPxEo3xMGoY9cYw9FtWD+3hAsf9EXq3h1sBOblDyj3JVlttxVZbbRUeejGVSu3puu675R6zRH6FunSE8dDE/xN9PPbBqJ4/ygloArfeMUfdqDjmqFeXbVB68YiYbQ7wC+C00Ngd6EZlPdKNajEE/Zium15/El2n9dr2xWgNzkAPNQGTUHS9EmmdDnpADsTbVkIp6kujGvKlqWzK+nTgEzQB9mn6/58gpzxY7B5vGNWjntvD/Qi4uC8HXXvttdlzzz3p10+VOL7vf9HW1rav67qP9M3cgoxAGh/Ric9fI2e6EtyHypvCfAisQn4dkRS9u4UEZUulMoRMmRPIAY8rO+pH9qTLNLId9UuArgLnynXsfLQhLYctUWnIEsBQNNH7OfqOXkTPd33NtlgEZUpsREZ7ZT56r5+gCamplC9QHP0MQfo00eOtB2wHfBN9z376/M+iLlnFlKK0o+fd7wKroev4a1Rq/DDwKJWfLB8EbIUmEgP9mnb0bPAp0iB7gsbtVtPQDAbeBf4bs20Aimz6oeVSav8jYrQmKyInIrj2fp+sOUaLcy7Z98IrSGYSeWn0MBg83G+D+pfvCYwBDkr/Owb4XnrbJmRSb5en9orVhmEUx6Io4n4y8EeyfwNrtSxAjsVHfT3WiBEj/JNPPtnv7Oz0Ozs7/XHjxnWfffbZP6jwZxblrBhbpiMnsVKsipy06Hn2y/ciJDoYfc2hZdpwb+Q4uWrud445Z6nL0yXYtSLwW1ROW8yx5wMPoQDgwBLOA5roug11QCh0nmlIFHB4iecA2DvmeBuGtm+L/mbynX8uEjbM9R7bgJPQxFm+43wA7FvGe4hjXdQudU6BcwZ/Q7+gvM/P6AMXoi/gucj4YmjWJviCPBRNMoxasgV6aAiuw2OTNcdoQRwUWQr/YF2Q9xWGYRiVYySaeLsURb7nUXvnvexlyJAh/lFHHfX/znpnZ6d/7LHHPjhmzJhSWlrmYj1619O/EmPHbypwrigPxpzn7gKvaWZHvR8qOZjfh/MU+4zXBkxEkeVSzzGNwhMqUXI56g4wAflIxZ7/SXo7u0sDj5X4Ps4r8T2EGQJcS/xkU6HlCySsaNSAdclc5A+Fxkci9crgS5kPHFhz6wxDnEzmWlwAbJ2oNUYr4aDIefhH6pxELTIMo9Xph5yEk4HrUQpx1R3uvixtbW3+nnvumeWsH3HEEdNHjhy5Z5mfwWJo4uILlFIdEOcI+yilt9IcEHOe2eSPCjero74E8PcKnKcYR30AiqL35Tw9KHpdLLkc9V+Uef77yWTkBW0RyznOMSW8h4ARFLJZmAYAACAASURBVI7+F1oWAkeFD1rJNjKGaEOpxA6aCZqZHv8W+uP/Rnp9FkqxeKDWBhpGmkvRdfkD9IByG/rRfSdJo4ympw11uTg8ve4DP6GPdZuGYRh9ZCEZJfeAemgPl5Oenh7uvPNOPvnkE3bYYQccx+Eb3/jGsL333vvPd9xxx2MfffTRiRTX3iyFOhBdhKKQnchZD9gy5jVfIMek0vwV/S6ES6AGozrpx6twvnJ5A/WTD5hAdgnrI8hxzMfHebYNRrXT347ZNg3pWj2Intm+Qp2jFk/v/x0UnV0i5rW5uAbYJ2b8XeA64G9pe/shrZc9UVp9WKk/hZ4tvyReGLAY9gF+Glp/HZiCNB6mpc+3HjAWPcOG2RE4DLVCvBWVpQU8hDov/Addu0uj7NIfos8tzIWo9v2TIm0ejiL3q8ZsewhlhExFGgIecuq3QF1t1gjt2440et5Bn7dRBU4he2bkWlTrOD00/hG6yAwjafohMYvg2nye3u1RDKNS9Ec/uMH15gEnJmqRYRhG8bQjQavDUL3wK5SWnlu1ZdVVV/VPP/30/4+sn3nmmf7qq6/uIadlpTzvaSMUNQ6ONY3edeeXxJzzr0V/aqXzn5jz5futSCKiHqU78rqfl3n+gMn0fk89qESsGF2AfshZ/weFI+pxWQw+cBn5J6ZGoZTz6OumU1y71LiIevA5LkDfeVuO17bTOzPPRxMoJ4TW30X6FLlYCk3ORY8zvgj7A8LPNcHyGoUzTtqBcfS+h3xIaZMsRpGsgNJzgg+8G7iHbDGGV9L7GUa9MAJ4n8w1ehvWEcKoPAOAP5H9Yzw2SYMMwzAqwDAkLumiyNkXJOSsL7HEEv5xxx33/856R0eHv9lmm/mo/v5iYMmQ3SNQpDTqJPwk5j3+JeZ855f5eRVDVHDZR+07c9Fsjvru9H4/C1EEuxwG59k2iHhxxYlFHntR4lO+byvitXGOejAhUUz5hoNU06OvDzSY/oeEXguxPPLfwsd4l+KehQ+JOf9U9LkUy0kxxzi3hNcbRXIXvQUEwjfAJ7EZEqM+2YBsdcoz8+9uGCUxGJX5hH9ExyRqkWEYRvVYGUXdL0VOTDniUmUtgwYN8g855JCsuvW9997bb29v94EZSA/kzPT/o6//gPgI6gsx+x5XiQ8qBxNjzndHnv2bzVGPi1J39OF4+Rgbc65nyR3JjmNVeosxdlM4MJnLUS92kgBglxzH8Igv2cjFr2OOsVqB1zjAS5HXfEl8W+5C3B05zmfUUZlNM7A/+W+ej6MLZi1gOUqbaTGMWnAo2bOZpj5pVIIhqNYquLbmA3slapFhGEZtWQFlWNYkTd5xHH+77bbLctaPPPJIf9CgQYVemytF+v2YfcuN7hZDR8z5HsmzfzM56t+l93t5kdIc51J4KuZ8pTi4ARfFHKeQgnqcoz6b3jXj+eiP+qFHj1OqBtgOMccopGK/U8xrTivxvAEbxxxrd+vbXRkWQ3USXp59NkNt2V5FM5Yz0/t/CVwFLFtlGw2jEDeQEfRKATejWjzDKJfhSOxmm/T6HJTS9+fELDIMw6gtY1CEcldqVFbm+z4PP/ww99xzDz09PcyaNYupU6cyd+7cfC97F4khxxGnuD6jr3bmYXrMWL707WZi15ixK1AApdIMpXcd9Zuorr1UrokZ266M49yFfKNiWYDqwaNcV+J540QXC0XU946sdyNtsnKYilL1w2xW5rGMCFdT+iyphxQFq9HawjDKpY3s2eQ36N2X0jCKYWng32Supa+BbRO1yDAMo3YsR7YuRyLL8OHD/X79+hWz7wV53ktc3f3O5X80BTku5nzP5Nm/mSLqj0eO001x4nHlsD29P7cL+3C8aBr4PPK31YuLqB9fxnnviDnOSmUcJ9qrvtBn8Vpk/3ImOMLcEjneoxZR7ztbop53xcySBs78PUhlc0/y33gMo9b0oHS2/6bXV0eiLtVKuTKakxFkt5WZjh4IrN2IYRjNThvwYzTRnXiZz/Tp01m4cGExu56BlNzXj9kWF4of1he7ChAXIMibDtAktNE7gPcqmVbPlWaDmLF/9eF40dcOoHcLtUK8VcZ5v46s99A7Ol0MsyLr+a7xJclurQYSDO8LH0XWR1of9b4xAEXTffI76oGDfjNSyXyj+qYZRtl8hdKTn0I3qR1Rm4qzkzTKaBhWQE560E/0S1THZZOShmE0O+ui58KNkzakTHZAk6p/QG2j3k6Pz0AZAmGq6ajHHbuaqfb1wmKo5jrMi1U831IxY6/34XivFnmOfMSVPRRiQWQ9EEosleiMVj4/eTl6+3570LdM6WgZ9OLmqPeNcSjimIsgZeVWoItMlNIw6p3XgcNROlEKKcS+hNJyDCMXo5CTPiq9/inwPXTtGIZhNCsD0TPh6ah/dSOTAg4E9gEmoYn6D4G1I/stU0Ub4o79YRXPVy/Eiah9VcXzxWUulOMo53ttqeWTRaV/FCDquFeDuO9qZHqpFMNb3VEfhGZ6RqJZrCHp8cXS/w5BN9yBZOoWZqMLYAniI4zBDM48NKv6C1rj5mI0H3eiCSYXzRr+DvgPEsUxjChrAg+Ribq8hyIz/0nMIsMwjOqzJXreyxe4aUT6Ayeg9l1xUda4tOlKEXfsclKiG404p7Zaae8QX/seTSMvhWjqOFQ38yJJSlGmL5dUszvqI9DD4+rpZRQSOFoapRdUo0Wag5SN/4dmHy9ACorhZXYVzmsY1aALXcf7oYmt21Faz2dJGmXUHeuhVihBitu7SO317VwvMAzDaHCGI7GpH1IjNfcqsBA5wNNzLF+l/90MaSuFia5XikXoHb0HiZM2O3H+QTXV7ufEjOUTfytEXN/vuHM0A3GR/6dRkKJiNIujPhjNvm2CRDDWQI55tVQSi7FnrfQSx/vIYX8d1W0+jerWy6mnMIxq4iOxxLWAdVD98R1IvbsWqUVG/bMRcD/KMgLd17bHMokMw2hevo9aZlUyzbVUeui70Ov3UOvgQrwOnBQZWw49a7/ZRxuibEnv97UQta+qBUmK58a1Jatm5524VPWhlP/7HRc9r2bqfpJ8HjP2BzJtjitCozrqawDfQY75Jki8o5HeyzfSS7i/4HTksD+NbkZP0rwXt9FYfI0EMqaiiOlmwCWofYrR2myBulgEk6KvIif948QsMgzDqB4jgMtQb/RKMA896xW7zECCb2ci1em+cB/FOekgnZEP6S0odwRwVh/tiHJEzNhT5BeT82LGynW4Fyu8S9X4kt4C1avm2LcSxPkZo4jvS14Mo2LGSumJ3kh8ETMW9/77RKM4t4uih7+d0ssKFTruAmAaEjz6lExdRnDhfo1m8eYhhXcH3ZxXTm97A6W4L04mpb7cG+dwpK69Y3q9BzlG96eXZ4m/ERlGLXgXicvcj+4bPwJeAH6boE1GsmwN3E1G2+M5dP+K+/EyDMNoZByU4n4hmQhnN7lTxYNlRsw+wVJqGeRGSNztO314HwE+0p8pZf9bgNMi44cBnVRGAAz0HL1nzPjNBV4XVxs9JGasGKKTEbVkPmrxFW5ptiHSy6rUZxzm5Zix9VAv+XKItvbziFeCbwbeRr5g+DrbLCFbEmF11NPx78ihjjayL2aZiRzcm4FzUH/orVDtyxKUzvfRBXcouSc5+qE/8vWBXYBjgF+iqNN/0B9aOe/lM+Cm9HuoRm29YRTDyWSuyQXo78loPXZFPW2Da+Exkis1MgzDqDYjgM2RA7U85TuB5bAEcCkK4JTz/Bi3TCnDjjVy2FDJiPo1McefQeHfl3YyrZCDZUIZ5x8Vc/5/FvnaQHQ6WH5ZxvlBJRVRG3Yu81iFGBlzrr+VeazFUWAzfKxCfcX3jjn/hmWcO3rdlJvV93HkONcU2P/eyP4eyU70VJ1lUA3M05R+03kTuAEpVG5DdeqGNkVtK/pCfyRwtxdwLko9ClJdil3moJnN3Wj8NiBG43E1mWvxc6qQ6mPUNfuRPXn6CLV9aDUMw2gF2oGTHccJekJXaukmXqytGG6JOd4cKpOevRW9nW0fidoWw0eR191Thg1nxJy/WEf988jrrivj/CD/IGrD3WUeqxjeipyrB5Xnlsrx9La7UNZlozvqp9Db/ollnrtuGYyixPdSfLR5NvBXlLazC+VFx+sJB81UHgb8GqUUx92s4pZpaPatEqlQhlEM/YB/kLkGn6e6qqRG/XAQ2ffpvxCv8moYhmGUzzapVOpVKuug99WBBE3Mfx1zzFdQ2nq5rE5vR9tH4svFZmvdE3ntnBJeCxkRtXId9f9GXldu+ngbvZ1nD/k71WAcvd/z70s8xhDgg5jjbFzgdY3uqA9HZSxRH3WNMs9fV4xAjvY0iruxvIVmZsbQGtGbpdF7vZ7iI+7/Ao6mb60VDKMYRqAf0ODau43GbU9jFMfRZKc93oW0OwzDMIzKsFz//v3/SHUcdB9lQ63SRxtPyHHslygv7Xct4p28HkpL+T475hjjinytQ3y2QCmO+t2R131I+c9Fx8XY8TGwWpnHy8cy9E5Z70GaYMXyW3rb+3QRr2t0Rx2U8RF9D6/T9yByNUUE87IRcCOF6857UJ3EscBKSRhaR7QhgYIJwDsUvhF/iG5YfVUENYx8bIBmrIPr7owC+5sjX79sWWD7cWRn+dxM44iRGoZh1Dv9hw4d2tnW1lauJlOxy5UVsveGHMefjlKgiykT7Q90kK13El7cEm1akd419HNRC7p8DCG3k16Ko35OzGtPL+kdZOiPlO6jx/sUlfcWy3A0WVGoS8H5MeeaTUbgOhftKNU7zocrxs5mcNQXRfpl0ffxLqVnO6fQ53YX+bscVIWtyU6XzbW8APwUiXUYvXGQoMlvkLpyvs9yDkqLH5GIpUYrcCjZN+bd8uy7X00sMkrlm+RPc4vW7E2i71odhmEYBjBy5MgjF1lkkWj6bDWWuVTu2bo/8Oc85/oAtbHbCdXDL4YijN8Cdke/I/kyai+jvMn9O2KOtTB9vA3ItGzrl7blDLKdsw9Qx6VyHPXViC9bfQNNBPwqfb7wclie461I7uf8u4B9iO9bviJyzG9BzraPgp756I+ycqPn8YDJSKMr/H0MAvYFnslhX7FCes3gqINKN74i/vO7Dz3/LhvzulT6tfujSbTw+WeW+R5K5tv0VsWLLl8AvwDWqZVRTUJ/1Mbiz+RXA/0aGI+pMhvV4WKybyy5RGrexMQP65G/A7/Lsc0l+15yJeakG4Zh9JnRo0dvNWLEiHeovoMeLBdV+C30I16hvC9LN+oRX24G3vL0rhmOOk5xDpWPnNpN6e2zFOuog5zaUt5vofTwTVAUPd8xPkeTAe+gNnVx+xRy1EGtp6N19uFlTnr7B+h7yrXfnyi+LK5ZHHXQtROnsxBepqO2bm+g7zXaKSC8VN1RXwHVLOT7Mv+D2j0tUm1jWoCVgQvIfQPy0YTIGVgNu1FZ2sj+YXudTJ/ZgGHpbRZVry/2Rd/LVZFxB83+h+8fF9TWNMMwjObj0EMPXWvdddd9qq2trZIObqFlFn0Te8vHXsTXmJe6vApsUQF7tiW3w5prmQFsn359Xxz1IfQWtcu3FFPHPQp4scT3E12KcdRB9eq5ouTFLL8hk7VQDM3kqIM6jj1G+Z9fePmizPdQkAGosD7XLIGHFNt3xmpWq8Ew4CeoNiLXl/82sENC9hnNyeJo4i24xu4n+2a9dnr8idqbZuRgEPAe+l4uD407KE0wfM8wJ90wDKMPuK678s4773z/sGHDKvEQX+riVvntDQZ+TPZzQLHLC8ARVDbjbgPia7zjlr+TLdLWF0cd9Bu6O3A7hQWzi3HUQbXgRwH/K/I9Bct01E1qpRLsbweOobfTmm95BrXXK5Vmc9Qh8/1HSyiKWbqBx5Eu0OLBwSrJpuhN5Up9fRKltDxW4fMavemHbnwu8XURAFPQxfB5jWwympu10A9jUGLxcyRqCJoY+mv6/6OBZ2trmhFDJ3Bu+v8XA6eiyZVrgLHpcR9phhRbb2YYhmGEcF137bfeeuvCRx99dNf33nsviQDVVyjrcnqNzjcaRbU3RsrVy6FMTh/VyX+AHPp/Ag+i1m7VwEFiaLsB30XR4uFpG95GPskf0/+GWZ9sxe4ZyBEtl2VQpHUIvVPCZyKHrljakUDZjkhkehkkHD0Epe5/gT7bl4GHkdM3v0y7ByLnexckBL400ryaB3wGfIImOe4jU99eKksD60bGplJ62vfa6DMOWIB00UplS1RWHPARyvQol7XQ38JWKDNiKfR9daOsj+koDf4N9L4foUoicoNRxCVXmvsrFFYcNKrDYJTunisl/hPyi1kYRinsSUYvwQMOTI8fTuaaK7U/p1F5lkf1Zl56uRA9AISVfD3gpKQMNAzDaGRc1/3WqaeeeuvGG2/sOY6TRBQ9WMpVHjcMowkYTe6WYR8Ch2DiQ/XAksAl5J5MuZXW6E9vVB+XzHU1B83E/iw0toDq1coZxfEHsgUof44EYMLpV0cmZp1hGEaD0tHRMXrcuHF37rbbbt7gwYOTdNB9FBEcXPU3bRhGXXIo2X2Uw5GY60nn1xt1xXrAc8Tf0F8nd9mCYRSLgxzB4Lr6H3At2ZNEZyVmnbEZvVvIhJVeu9G93TAMwygS13U37+jouPvII4/0ll122aQd9GA5vtrv2zCM+mMAcCnxN4W3yCgnGvVJO0qHn0fv728m6s1oGMXQnmN8CNkqpZ+TcdQ9lG2T67VG9WhD30uuzJr5SNzFMAzDKALXdXfq7Oz8x6mnnup/+9vfTtoxDy/vkF1vaxhGg1GOqMUI1Ld7k5htl6JI2dy+GGXUjLWBG5FwRhgfOA84J/1/w8jF9kifIoWc7w9Rql3QquVX9G7VFjAGuK0GNhoZjkXtU+LoQU78B2gSLxgLRF1mo+/zg2oaaBiG0QA4nZ2duwHjPM/beOrUqTz66KPMn1+ubldVOAL19TYMo0Ep1VEfgVQEo+nRXwM/QOmuRmMxELgCtX2I8lukCu/V1CKj0VgcOX+l9EjvQUqrW1bFouIZju5rSyH11BGofn4ZYDH09zEIRaKHhl4T3DtnovcyFzm3gZLnHOBTJNY4Lf1v8P9pJPM3tRjKeBpGbt2QICXeCe2zEN0LzkPvyTAMoyVxXTfled6uSItlg3feeYf777+fadOmlXO4hej5GTK/JQArUlov6jheB74VOqZhGA1IKY76CshJXzUy/l+UKvlSpYwyEuFo1Es5mib1u/Q2c9aNQhwCXInS3ou9t6yHeqhWkwHovrU2alGzMvBNYB0yznctWQi8j9rDvIq6YrwdWqrF5ahesZjvpgc56rehFnv/raJdhmEYdc3RRx/db5llljnQcZyzgTUAZs6cyX//+1/a29s/mzZt2r3PPPPMnQsWLPgaPS8FLZZmIwHVYBKX9L/dMacZBTxB7pa6pbA/aj1mGEYDU+zD9KrISV8hMn4vcBBV6vlm1JzvIuXnqCL3DSiFymZmjUKsAFwHbE0mMpsLD7Vq+2EFzz8AlXJsjMpzNgFWqeDxq8001Evz6fQylcr0vl0bTYgU0gXoQZGch4HTgH9X4NyGYRgNyYknnjhg+PDhhzuOMw74RmTzy77vX9TW1naz67pxjncpLIGc9DX6eBxQCdP6WIDFMBqeYhz1kehhcbnI+G3ISV9YaaOMRFkLPaRHZ3Svxdo1GcXhoP7bF6GobL4Uvvno3vJFmecaAmwHbIuc8vVpLvEcH3gDOe3/AO5HGgCl8gD6jHJ9F4GD/jQSmny0jHMYhmE0Ba7rDvE87yjUg3xkZPO/gQldXV23URkdn0HAQyhYUgl2RYE0wzAanEKO+kDgEXoLx92K2vf0dQbRqE9GIWd9VGT8J0hMyjCK4ZvALSjFPB9nAL8o4bjrADull83pm2M+h+z68Y9RHfY04DNUdz6X7LTF6WQezoYiB3cQul+2A4uml5Fkat+XDf2/r20rX0AO+/0oAlNosnRv4PYc2zw0mfIqqrmc0kfbDMMwGhbXdYd6nvcj5KBH79VPABd2dXXdXcFTtqHA114VOt6TqAWnYRhNQCFH/Xf0jqLeBBxOY6VBnwlskGPbm8C4Cp3nl/ROjQp4ErikQuepBSsiZz2cNuwBewB/ScQioxEZiBzA09H1E43o+ihCvBK5J/4cFGk4CF1/y5doQzdqU/NGenkztJQTne4rw4DVUYrjGun/B8vgEo81C0XLb0F/l/Mi2weg+vLoZxaUJbyPROKuwdIkDcNoUc4+++xl+/Xrd6rv+8egidYw96VSqQmu6z5e4dM6wNXEi/mWy7bA3yt4PMMwEiSfo34aSl0Ncz+wG43lpAPcg1KB4uhBTkJfWw6thpyAXJ/prcCBfTxHrRmFyh6WDI3NBDZFETjDKJZtkdbBssT/jeyN9BHCrAkcABxMbxHLfHwMPJdeHkdRkEZpGbkyyhLYML2MpviMgRnAXSgqfi+6t10OnBCz7yeord5VqPzAMAyj5Rg3btyotra2H/u+/0OUGRXg+77/l7a2tvGu606t0unPA35WYJ+P6J16n4sHgR36ZJFhGHVFLqdyM1SjGI5+vYoctJmxr6hv8jnqUHrqbRznI3XkXDSiow666d9L9rXwErARUjI1jGIZhhzHQ+ktNPcMEoBbBGXsHIGusWJ4E00iPoxqrJuphdgQ5LBvCeyMPqNi2va8D9yJ2mYODI3PRA+HV9A4kxeGYRgVxXXd9TzPOxPYl+x7arfjOLc6jvNz13WrGZA4Bk2U5mMGsA3wryKPuSnwVF+MMgyjvohz1AcDz6M0zICv0ANio7boKeSov4Gid+WSAt4ld9o7NK6jDnAKvWvTz6dyJQNGazEGpfsNJfse9Fv00LREgdfPQaUkD6EI8mtVsLFeGYIe3HYDdqH4MgAPOefnUBkVecMwjIbDdd3Ne3p6znAcZ1eyf38WAH9IpVLnua77ZpXN2B1lkOWbdJ2HAiWPofKtQhO0fwa+XxHrDMOoG+Ic9QtQhDnARw+E99fEouoQ56gHKscB30HRuHLYAfhrnmNDYzvqADeiFOSAbqSw/XIy5hgNzgrAzRQvejMXuBtdhw9g6dqg+/do9He5PxKqy8c/gYvRA6IJgRqG0TK4rru953nn0ltZfZbjONc6jnOh67q10Cz5LppkHpRnnx5gP+CO9Pps8uuXeEiH6YVKGGgYRv0Q7am7MnByZOxKGttJz8VDwI6h9cMp31E/IrL+IFKkbiaOR+m3QdZAO3Apao1lGKWyKIWF3DzkXF6PJroaseymmvhIQ2Iqynr5LiorOJDeYkigtMhNyQhoVqq1kGEYRt3hum7K87xdgXM8z9swsnkacOWCBQsuveCCC76qkUmroInSfE46wKlknHTQxHQ+R/0WzEk3jKYkGlG/meyo7weor/bXNbOoOsRF1A9EN7eAGUjoqtS6zWFIvCp84z0AORZhGj2iDkqruiMythPZ2QSGkY+VgfHobySVY59pKA3+KuDDGtnVTHwbtfoZg1rk5eIZ1BHjb7UwyjAMoxaceOKJA4YPH76/4zg/I7uME1SmeEkqlbradd05NTRrSSRsGrUnynlAR2TsI/R8GsdCYG0atzTVMIw8hCPqq6JUmzBn0vhOei7+BryHUnBBDvcewB9KPM6BZDvpz9O8M5t/QmnHYVXRczBH3SjMkuhaOZrcKuafowjvKJTqZ5THC+mlC/ge8GM0oRadmB2NBPj+isqdmvW+ZRhGC3D66acvOnDgwCNRO9CoUvrLvu9f1NbWdrPrurUu/RmExD0LOek3Ap0x4/mEe6/FnHTDaFrCjvoZZNdVv4Ai7M2Kh1Jqw4Joh1O6ox5Ne5/cB5sagbPJdtQ3RXXGTyRjjtEAHAxcQnabvzB/B36OylG2Rr3Fi1W5NXLjo4m1B1Bm1E/QPS5a8rQjKmH5JXAupgZvGEYD4bruCN/3T/Z9/zgkUhrmSeCCrq6ue0im1CcF3ETv2vgo96LnyTgbc2myzEMReMMwmpQgwjIEpdaE6xr3B/5Yc4uqQ1zq+1Lo/b5F5nPwgBUpvqf6GsDrofUFSIV5CXorUTdD6nvA/WTX91+PHADDCLMs8GtyK9G+ghzDKTWzyFgDlR7sS7yY6Fso68HS4Q3DqGtc11057aBHe6CDggcXdnV13Z2AaWEuobf2U5R/AVuRO4P1RWCdmPGLUT27YRhNShBZ2Y9sJ/0D4Pbam1Nz3gEeB7ZIr6dQ9O/CIl9/VGT9HlRfW6i9VKNzMdmO+hjgREzsyxAO8CPUQSJO1OxtVIN3K5ocM2rHG+h+vzH6fraJbF8FZTZchbKsZtXUOsMwjAK4rruB53k/9jzvILIzQT3f9+8FusaPH/9MQuaF+TGFnfR3USApX5lpXER9NsU/qxqG0aAEjvrekfHJqD1EKzCZjKMOcCTF3fzagUNijtUKPAj8D2UfgGayd6J5MjCM8hkKXIfEzKLMQxH0X5G/5s6oPlOBbVE/9ivJdHOAzETLtigbopX61BuGUZ84ruvu6nne6Z7nbRHZNs9xnOsdx7nIdd16qdfeHZhYYJ8ZSBvpkwL7xTnqFwOflmGXYRgNRDuwCL1bbJVap93I/BG1GRuSXl8d2ITCrdp2JFuF8zOas41dHB5KVz4tNLY75qi3OmugrgBrx2x7AvgB2aUiRvLcg9LcO4Gfkq3EvwZy6I/EyhMMw0gA13X79/T0HOA4zume50W7WMx0HGdyd3f3L84///x66hAyGnUVasuzz0JgH+ClIo43L7L+FdIUMQyjyWlHTunA0NjbwMvJmJMIXyPn4rDQ2FgKO+pREbkb0I23VbibbEd9y6QMMeqCPZBWwbDI+EzkBF6OpbnXK3NQh4+7gWuANUPbhqCJ283R33sr3eMMw0iIQMHd87yfOI7zjcjmj4FJ8+fPv/jCCy+ckYR9eVgZTYAukmcfH5VOPlzkMaMR9YuA6aWbZhhGoxE46mEeTcKQhJlMtqN+49ryXwAAIABJREFUIBLoyKV+vDhKGQ1zfeXNqmv+iT6fQMBlBWA5rO91K/IzJFAWFSd7FomW/a/mFhnl8ASwAXAFiqIHOMBJwGooAmSq8IZhVIWzzz57mfb29h+he85ikc0v+r7/y08//fSWSZMm1eOk4RJIvX3pAvv9DAV3iiVcKjYN3aMNw2gB2oF1I2P1IMBRax5BaserpNeHAXsisas4DgYGhNafRaqcrcRC1MLvO6GxdTFHvdVwUX/0KDcAx2BOXaMxF0V6HkWCcmEl5Z1Rec9umMicYRgVxHXdVX3fP9H3/aPJzvKEjIJ7Ui3WimEg6pW+RoH9fofakZZCOKJ+Hnb/NYyWoR0YFRlrxRpSH7iRbIfjcHI76tG09+uqYVQD8DrZjvpKCdlh1B4HCeVEW8PMR2rhl9bcIqOSXI9KoO4gIxoJKnG5D9gF6/JgGEYf6ejo2NBxnJNzKbi3tbWd77ruU0nZVyQOcsA3K7DffcCxZRw/cNTfA35bxusNw2hQ2slW+wW1imhFJqOWUYGY0g7os3k/st+3gPVD6wvI7dA3O+9E1ldIxAqj1jio5vz4yPh0FG19ouYWGdXgX6g06n5gvdD4ZsADqNOD1UkahlESY8aMaVtrrbX2An4CbBrZPMdxnMmO4/zSdd23EzCvHCYCBxXY5yVUVtldxvEDR72LeAV4wzCalHZ69zn+MglD6oB3gX8AW6fXg57qF0T2i/ZOvxP4vJqG1TFfRdbjemYbzcc4ejvpXyHHbWrtzTGqyKeo1/p9ZGfPbALchr7zch48DcNoMVzXHdLT03OQ4zin0jtF/HPgd93d3ZdOmDDh4wTMK5dj6J1ZFuVDlIVUrvDdfOA/tG72pmG0LO1k1yCCFIBblclkHHVQivuFZGqi2tGMaPQ1rcrXkfUhsXsZzcSeqC49zGfA92gsnYZhZLciCzOX3u1w6v081WQ6+n7vJvv+uB1qEXRyAjYZhtEguK470vf9Ez3PO8ZxnKhA3Nu+7188d+7caydOnDg7EQPLZ1fg1wX2mYmc9A/6cJ75qDTTJkUNo8VoB3rS/4bH6lFNsxbchlJ6g8hw0FM9qI/aHVgmtP+nKAW0VekXWV8Qu5fRLHwTicSFHc9pyHl7LQmD+sBL9C77CbgNGFOBc4xApTPtObb/DJhQgfPUgq/R/e8+1Kot4CQ0QfO7JIwyDKN+cV13Xd/3j/c87zB6C8Q95/v+ZW1tbTe7rtuIDuhGqHVlMb3S+zqJ/Vz6XIZhtBjtwGyyFcwXoXWVmmcDU8huTXQ4GUd9bGT/62jtGc5oBL2VszGancVQmUe4vGEhar/WaE56IfYAlqTvJS2Hk9tJb0S+BvZG5Q0rhcavRNfAkwnYZBhGneG67uY9PT1neJ63K9ltOwOBuEtd130oKfsqwCjgLxTXK70S7/NW6lft3jCMKtKOHkYXD40tR+vWXIOc77CjfiASPBmC2hOFubFWRtUpIyPrrXzdNDu/JtO+MOAkpOvQbPQHDqDvvWoPr4At9cY0lO75T2Boeqw/cBNqz2htgwyjBXFdt39PT88BjuOc5nneOo4T9s+Z7TjOTY7jXOy6bqN3FhoK3EXhXumdlNYrPR/mpBtGi9KORNRWD42NQv2xW5XHgP8Cq6bXg57qy5Kd6v00Sp9tZVaOrEdV4I3mYG96azNclV6albH0zVH/DrBWZUypO15Fk5lTyETLVkLCm1GRQcMwmpgzzjhj2IABA8Z6nnea4zjLRzZ/Cly1cOHCy3/+859/kYR9FaYfcDvq/pOP36N+54ZhGH2iHTmlO4TG1gf+nIw5dYGPegh3hcYOR456GFPfzG5TB7qWjOZiMHBJZOzfNJ+A2NcoMtw/vb4hsA7lT8aNjax/SXbmUqNzO3Ap8OPQ2LGoVv1fiVhkGEbNcF13Vc/zfozuddEU8BeBX6VSqVtc120m7ZrLge0L7PN34Ec1sMUwjBagHXgmMrZJEobUGdchZetwT/VwHtc8Wrd3esCyZNepLqC1MzGaldPIFl1bgCaumunhC6TM+zTw/dDY4ej9l8pAYP/I2M3ACeWZVrf8DKXBr5ZeT6FJnS0Ts8gwjKriuu7mnued5Hne3vQWUnsCuLCrq+semi9d+2zUii0fL6PfkGb7fTQMIyHa0cNpmC1RFK2VhcHeA/5GZubUiWz/M717iLcaO0bWn6cxWk0ZxTOM3v1hL6Gx2rCVwmSyHfVDgbMovQvG3sDw0Po7wOM0n6M+B0WOwmJJW6B7w18TscgwjIrjum5/z/P2BE7zPG/jyOYFwB98379o/PjxzVoOuB+FU9k/BHai/F7phmEYvWhHar3vkomODkIR5FZOfwdF1XOlOFnau5Sxw9iDefNxCnLWAz4Dzk/IllpwL/AJaqsGEgvaCfUPL4WxkfVrab7oUsDDSP1419DYOOx+YBgNj+u6I3zf/5HnecfSWzztc+C33d3dv54wYcLHCZhXK0aje3g0YBPma2A35KwbhmFUjKB10N3AiaHxsZijfjuqRxoeGf8QeLD25tQVS5P9YA6lOzNGfdMfOC4y9kuUIt6sdKMU9XAWweGUdm0vD2wbWvdRd4jRfbaufjmH7PvB5qjH8LPJmGMYRl9wXXe9tIN+KArehHkTuDKVSl3tum6zZ16OQvf/wXn28YBDkHaLYRhGRQkc9ZvJdtR3RW3aWnl2cC5wEbBNZPxOoKf25tQVR5AR3QJ4A3guIVuM6rAHsFRofTrwm4RsqSW/J9tR353SeqqPJbtu828o9b2ZHfXngAfIFiX9IeaoG0bD4LpuyvO8XX3fP8nzvO3IjiD7vu8/7DjOZU1afx5H0IZtmQL7/Rg9FxqGYVScwFF/CngF+GZo/FTUP7yVmZBejAwDUP/sMNfQGj/crcShkfWbaI0e2a8gx3PD9Hp/1Jru8iJff0hkfXJlzKp7fkO2o74/mvw1USXDqGNOP/30RQcMGHCg53mnAGtG+p/PA6b4vn/h+PHjX0nGwkQotg3bZRT/22AYhlEy7aH/X0F2xOwYFFH+pKYWGfXO0cDI0PrXqH7LaB4G01ufoZW+48lkHHVQ+nsxD2NbAGuE1mcCd1TOrLrmHtQzOYg+DQO2wsqEDKMuGTdu3KhUKnUM+k1fLLL5Y2BSE/U/L5Vi2rDdhwWzDMOoMmFH/VokArRcen0RJBx1VK2NMuqWxYDOyNhVQCv+kDcz25Bdk/c/Wqu04WZgIsoeATnt36Zw+8GxkfU/0jrdM7qRsx7+vdgZc9QNo55wXNfdpqen50THcfYg04I24CngklQqdbvrut0J2FcP/IzCbdieR0rwrfoZGYZRI8KO+nyU5v3r0NhYlNb8zxraZNQvE1C9bsAsJDBmNBebRtbvTcSK5PgSCQjtGxo7jPzRk0WAMZGxyZU1q+75C9mO+neTMsQwjAyu6w7s6enZz3Gc0zzPWyeS3u75vn8vcMH48eOfSMjEemEM0FVgn4+APVE2oWEYRlVpj6z/FvXFDepyUuhhcz0krma0LtvQe5Z5AlYa0YxE++S24kTdZLId9UOAM8ndU31fYNHQ+n+AJ6tiWf0SvU7WR1kJ8xOwxTBannHjxq3S1tZ2nOd5RzqOE+1g8yVwdSqVusJ13Q+SsK/OGI3u+9EsgzBfI7Hl92thkGEYRtRR70EKlg+SUfxcHUXZj6yhXUZ9MQK4nmwV2DeBS5Ixx6gya0TWWyntPeCvqE5z2fT60iiV+64c+4+NrE+m9QQWPwE+QC3qQEJ8o4DXE7PIMFoQ13U39zzvJGBv3/fbIpvfAH4zZ86cayZOnDg7AfPqkVGodMfasBmGUVdEHXWAh1Fk/djQ2BHAizSuY/ZvMvWmAbkiY5VgNvBQZOzlKp6vmvRHtbbLh8Y84AdIEdZoLvqR0akAOZtvJ2RLknSj/uc/DY0dTryjvhKwZWjdA26ommX1zdtk3ytWwhx1w6g6rusO6enpOchxnBM9z4uqlXu+79/b1tZ2qeu6D9N6k4j5CNqwLV1gP2vDZhhGzYlz1AFOR6nO4cjaRah1USOKA42r8fneB75X43NWi0lIzTrMhcBjCdhiVJ+lyO4DPo3WnZD5HdmO+m7o85kW2e8IstMlH6J1UyPfi6yPSMQKw2gRxo0bt0oqlfqh53lHO44TVW+f7jjO9d3d3b86//zz/5eIgfVNP9SZw9qwGYZRl+Ry1GchsYyngOGhfW8FNgdeq75pRh1wBooihrkX6EjAFqM2RFP/WqF3ei7eAKaSqdkPeqpfFtrHoXV7p8cRvV4WScQKw2huHNd1t/M872hgb7InVwGe933/qra2thtd122VzhPl8FtguwL73AOcWgNbDMMwepHLUQc9pO6PHLPgR2Bx4BEULX6xqpYZSfNT4ILI2BvAwUjLwGhOoiUiCxKxon6YTLa43uFkO+rbACuH1mfQ2umR0eyLgYlYYRhNiOu6w3t6esY6jnOc53mrRTYvBO5Ii8M9noR9DcbZKBsqH8+jyVl75jEMIxHyOeoADyCl44tCY0sjZ30nFG0ymo8z6O2kfwXsDkyvvTlGDYl2d2h1R+tm1IJwUHp9A7J7qo+N7H8rrdM7PY5oRkYrfxaGURFc193A9/1jPM872HGcaJbKZ8C1qVTqStd1o6UnRjz7AOML7GNt2AzDSJxCjjrARCQIdHxobDGkirwT8HTlzTIS5Hw00xxmNkqv+0/tzTFqTFQFeFgiVtQPM5DQ0P6hsaCn+hDg+5H9J9fGrLoler3YQ65hlEGo9/lxnudtErPLM77vXzF9+vQ/XH755dYCsXg2BK7D2rAZhtEAFOOoA5yIeuGG63SGo8j68cDvK2uWkQADgd/QO0I4G0XSH6mxPUYyTEN/60EK/OKoP3gr16pPJttRD3qqH4Cc9YA3sYnLlSLr1p/ZMEogJA53lOM4S0Y2zwfuSqVSk1zXjXaWMQozEpUm5dPOsDZshmHUDcU66j6KIH0NdIbGByJl5E2QM9/q9ayNyqpI+XSdyPgM1Dv6nzW3yEgKD0URVg2NrQb8Kxlz6oIH0GfyjfT60sAu9J7U+j3W9mjVyPq7SRhhGI2E67opYNs84nBvAVenUqnfua77ec0NbA4WBe4ju/1oHNaGzTCMuqFYRz3gHPQg70bGjwbWBvYDPu67WUYN2Q31fB4eGf8C2BF4ruYWGUnzCtkO12ha21H3UE/1s0JjncD6ofWe9D6tzMpAOAI4i97t2gzDSOO67nDP8w73PO9kYFRks+f7/t8cx5n02muv3TFlyhQTNCufFLo/r1tgv6uxNmyGYdQRpTrqAOcC7wBXkRFYArVt+zeKrP+x76YZVWZR4OfAj+hdq/UimtV/q9ZGGXXBVCSiE7A5amPTykxG6e5Oen2DyPYHgA9raVAdskVk/VlMLdkwetHR0bFhKpU62vO8Q8l+joKMONxVruu+W3vrmpKLgT0K7PNX4Lga2GIYhlE05TjqANcjZ+52slsTLQ38AYkt/QgT4qhXdkSO14ox224BfkhvUTGjdYi29tkZpWK2stP1JvAUsGmO7ZNrZ0rdsmtk3VpEGUaaM844Y9iAAQMOQRmI6/p+ryqZJ3zfv3L69Om3mzhcRfkBcFKBfV5FmiPd1TfHMAyjeMp11EHR89GofdGOkW27Ikf+p6iGvdXrNuuFxdHM8mEx27qB04BLa2qRUY88CXyJrheAJVC/8FYXL5pMvKM+A7i7tqbUHUNQF5Aw9yRhiGHUE0H03Pf9g8gWnwSYB0xJpVK/cl3XxMsqzw5IJDcfn6Nou7WeNQyj7uiLow56mN8V9d3uJKMUDap5vhrNHp8J/K2P5zLKZyBwAqqxXTxm+3+AI4AnammUUbd0IycrPKFzFOao34omuqK9wm+id//5VmN/VE4T8BFKfTeMlsN13aE9PT0HOY5zDLBeTPT8Fd/3r2pra7vedd2ZCZjYCqyFMjzzPefOQ066lfkZhlGX9NVRB6XDTkB16VcDW0e2jwYeRg/5Z2LiZLUkBewDXEhvoRqQQ3Yl6ptuqe5GmMlkO+rfR61tPkrEmvpgJhJfjLZMeioBW+oJB00EhrkOifAZRssQqj0/yHGcaPQ83FrtYSzTsJosibKcoiK5YXw0AW1dbQzDqFsq4agH/BfYDvVVn0DvFK/tgWfQDOf5wMsVPLeRTRty0F00qxzHC+hHyiZOjDgeQX/Tgfr7ANSi8SdJGVQn/D1pA+qQnYH1QuseKnkyjKbHdd2BnuftDpwMbBYTPX8T+L21VqsZA4G7gFUK7NeJSjcNwzDqlko66qAHtMvRTfIiYF8yKsmk/38ASpN8CKWR3o/NLFeKYUg45QRgpRz7TAcuAH4FLKyNWUYD4gO/JLu+71jgEkwk0siQAsZHxu7AUkmNJsd13bU9zzss3ft8schmi54ng/N/7N13mBvV1cfx7x3tuheMwdimmY7pLVQDoYVOSMCEEloAk7yJCRBC9SYTDBgICSEQQgsQCL2HXkLvvXcCptnGFBvbuK007x9Hm1WXdlfSVfl9nkePZWlWe7QqM2fuvecAl5C/6GeHjgEjEZGaVu5EvcNkrKf6BlhSuE3G/Q7YLnl5HzgP+3LV9OvuWR6rBXA4+ad6LcCmM7dh7V9EirkMmAAsmfx/P2wZxb7eIpJaczDpreoibEaVSMM59thjB/bp02dv4LBEIvG9HJu8A1wcBME/NXruRQjsV2Sbx4ED0ckTEakDrvgmZbEtlrCvX2CbmdhI/A3A3ahNRjGDsV7XY+lsn8Umm2zCkCFDePLJJ5kxYwbYLIebgGOBj/yEKnXsEOwkWqqdgbs8xFJOHwNLp/z/czpPSFTCXtgoTqqTqO+kdnHgjeS/Ha6m+IGySF1pa2vbzDl3CPY57p9x93zn3C1RFF108sknP4wSQF/2wgp+Fjqu/RDYGA1WiEidqFai3vG7fgQcBYwpsu2n2Bfuv7C11GJ6AzsBP03+2yf1zpaWFo444ggGDBhAFEVMmTJl8vTp04+67bbbbvERrDSEAKvevW7KbZ8Ba2FdH+qVEvWecdgU991TbpsLrIr9bUXq2vHHHz+ktbV1rHPul9j3Xab3gUuCILgsDEMlfn5thhUt7l1gm2+T26k+kojUjWom6qk2wBL2sUBrkW3fxUbY7wYepfnaIC2J9SfeAVsqMDjfhqNGjWK//faLYrFY6usaAf+OomjSxIkTn6lsqNKg1sMqm6d+Vh/AZnLU68wXJeo9cwLZsR+L1SYRqUtjx46NjR49+gfYTKJdgV4Zm8zDZqj9Q6PnNWMU8AwwrMA2C7HBjWZvMSoidcZXot5hSaxK/MHA8BK2n4tVo74buB9bD9ZoO8q+wEZ0Judrl/hzTwN/O/TQQx8eOXLkocCRZCf1TwBnnHzyyXfQeH83qazTsOQs1V+wE271aGfS+6HPxXrHV8rS2JTLVK8Db1Xwd1bKDtjfKpZy2zPYaFXcS0QiPRCG4VKJRGI/rGDmqBybvAlcocrtNWcQdlyzRpHtfgFcUPlwRETKy3ei3iEANgX2B/YBBpb4c7OAV7EWY48Dj1B/a4+Wx5YCrJ+8bEDh6VupJmNLBC7FZh78TxiGgxKJxC+A3wJDM37u5SiKzn777bevuuGGG3RgLaXohY1GbJ5x+8+wonPSHFbD+g4PSrntS2BDbP2nSF0YP3587yFDhuwWRdEBzrn/1XlJ8a1z7lrn3JVhGD7uI0YpqAWrlbJdke3Owo6DRETqTq0k6qn6Yese98OK0GVOPSvmv8BrWOL6Ljbq/g5+E/gWYDlgZWwN58rJy7oUmMqexzTgeqxo09PFNg7DcEA8Hh/nnDua7Km9bwFnTJ069eqLLrpIrdqkmGHAc8AyKbfNB35M/ReXk+JGYX3kR6XcthDYHvWXlzrR1ta2bhAEB0VRtB/ZJ7Ej4DHgklmzZt149tlnN9tSu3pyPjZSXsidWNFdDUiISF2qxUQ91QAsWe+YBr5sDx5rBpa4T025TAemYEn8F1gLsxnJ7WdiFdNz6YsVcmtNxtgfGAEsgVVAHoklNYtjB7XL0/UTDh3i2LTSu7Ge8y8WiCuvMAx7xePxvZ1zJwKrZNz9MfDnIAguDsPwu27GKc1hbWyqYWrl4wXYTJibvUQk1TAKeBA74Zjq/4C/Vz0akS5IKQx3OOntBDtMdc5d75y7OAxDFRurfUcDfyqyzUvYDDC1/RWRulXriXqm0VgBq+2wddxDqvA752CJCFhSXqz4XTm8hyVD92Br8ctWXTsMwyCRSOyM9VPP7AM7HTh/wYIF55x++unflOt3SsMZixVGS/3+WIgtXcksmCb1b1WsovLIjNsvxNb0itScsWPHxlZfffWtEonEOGxUNfNkeSKKogedcxdNnTr1Vs0qqxs7AreTvVQh1RTsGPGTqkQkIlIh9ZaoZxqJFTBKXePdp+BP1J5vsan6j2PJ+dNYwlxxYRiOSSQSIbBNxl2znHOXOefOCMPw82rEInXnMKw4T5ByWxxL3DL7rkv9Wh+bPrpExu3/Ag5CU0qlxoRhuFoikTgAe39mvm8B3gaui8fjl5166qmTqxqc9NQ62NKEAQW2mQt8H3i2GgGJiFRSvSfqmfpixY5WSbl0rAfvX+DnqmEqdoCQunb+LWxNvdcK7GEYjonH48c553Ym/T0xH7g+kUhMPOWUU97zFJ7Urn2AK7AaDKkuAsbTORNF6tNPsVHzfhm3XwUciJJ0qRFhGC4Sj8f3cs4dgJ28zzQT+HcQBFeEYfgf1PWkHo3AlgEuXWCbBLAHcGtVIhIRqbBGS9QLWRpbZzkM+8JfHDvbPjx5fRg2lWqR5PaDSR8tTNWO/e2+xirPz8VGwT9P/vsFNvWq47YPsAOFmhaG4VqJROIYLAFLTb4SwE1BEIRhGL7pJzqpUXtho6uZS0JewA6YNGJVf3oDZwJH5LjvIqyAU5frZIiUU8rU9gOAPbET9akSURQ9CFwZi8VuVP2VutYXa827YZHtjqH42nURkbrRTIl6d/Wnc23bbKxg1i7J/x+OHbg2lAkTJiwXi8WOjKJoHOlLCaIoiu6MxWKnhmFYtOK8NI3dsWQ9c9bKNGBfrAiZ1IdlsZaPmT3fwZL349FopHjU1ta2unNuf2xWx/Acm7wDXBsEweVhGH5U1eCkEhxwDfCTIttdChxS+XBERKpHiXrXbY0VVgKbwj6aBh1dOvHEE5doaWn5BXAk2W3kngDOOPnkk2+vfmRSg1bFTmKNzrg9wpL4IyljUUQpO4fVHTgLGJhx3zxsFP3yKsckAkAYhkslEok9sHXn6+TYRFPbG9cZwLFFtnkE+AFabiUiDUaJeve8QGeLl92wCqQN64QTThjaq1evI6Io+hWwaMbdzwGTgiC4LQzDhjxhISUbhCVzP8px3xTgV6iFWy1aAbgY2CrHfe8DP8YKXopUTXLd+Z7J0fPNyT5eiQMPRFF0xezZs29Rz/OGdDA2Ul7I28CmgDrViEjDUaLePftho4RgZ3K/7y+U6gnDcEAikTgEWwe2VMbdHwDnfvPNNxece+6586sfndSIAPgd1v4vV42Ha7DRkU+rGZTk1BfrRzyB3N0y7sBa7s2oZlDSvDLWne9BdiFDgDeBK9rb26847bTTplQ3QqmiLbD2tJlt9VJ9BWyCtbQVEWk4StS7pxUbaVom+f+NaKJWIGEY9orH43s7507Apjyn+hj4cxAEF6t4T1PbFGvTljkVHmx64uVYMv9FFWMSE2BJ0JlYgc1MM4HfA+fSoMt6pLa0tbWtHwTBAVEU7Y0Vds30mXPupkQicfnEiRNfqnZ8UnUrYK1qFyuwzUJge+ChqkQkIuKBEvXu+y12oAs2Srivx1i8GDt2bGz06NF7YgWmMtcNTgfOCYLgb2EYakSuObViI7Ynk3tUZBZwPnBq8rpU3rbAn4E189x/F/Bz4JOqRSRNacKECaNjsdjeURTtC6yYY5NvgZuAK4MgeERLq5rGosBTWFvdfCKsmOCVVYlIRMQTJerdNwgbPR6MrZVbCfjQa0QepfRi3yXjrtnOuUvb29vPPPXUUz/zEpz4tjbwD2D9PPd/DpyDrZPWOsPya8HaVx1D/tdgOtaO7dpqBSXNJ6Uo3Fhy9zuPR1H0EHDl3LlzbzrrrLPmVDdC8awVuJfc9TJSTcSWWImINDQl6j3zZ+ColOu/8RhLTWhra9ssOSV+J9LfX/Occ5c75/4YhuF/PYUn/sSwis0h2fUNOswB/okl7e9WJarGNgSr5P4rYOk828zFprifjk6SSAWccMIJi7e2tu4F7I0l57mOO54Brlq4cOG1kyZNml7VAKWWXIR9ZxVyI7AXquwvIk1AiXrPLAX8FzsLPAtbs65p3kBbW9uazrnfAvtgI3odElEU3QWEEydOfMFPdOJRXyxxPJ7sDgIdEtgU7POxYkLt1QmtYawNjMOmhmb2tu/QDlwG/AHQTBcpqzAM+yUSiZ2jKDrAObc9to/M9BZwfRAEV4dhqBNzciK2DKqQ54EtAdW/EZGmoES9567BRgrA1q2f5TGWmhOG4agoio6KougwLEn7nyiKHojFYr8Pw/BJT+GJP0OA47CkPV8yCTANuA64iiYq2NgNy2B1MvYD1iiwXQJrkTcBeKcKcUmTGD9+fO8hQ4b8AJvW/mNyf64/dc7d7Jy7IQzDx6sbodSwPYHrKXxM+hGwMbZPEBFpCkrUe2597Cwv2MjUclg1Uklx4oknjmhpaTkKK1Q1MOPuR4MgmBSG4b1oOluzWZTO6dn5psR3eBdL2G8GXq9wXPVgJLArlqDn6jOdSssKpOwykvMfYrVbMn0B3BBF0TUTJ058En3HS7r1gUfJ3YqvwyxgDPBqVSISEakRStTL42FsOhbAT7FkQnIIw3BQIpE4GBtNHZFx92tRFJ0Xi8WuCMNwnofwxJ8YVtfgeKy1WzHTgPuA27Hp8c2w5CSGdVfYFdgFWI/i3+FTgQuxdehfVTQ6aQoZyfkaH2nvAAAgAElEQVRuWEHVTN8BdwJXTp069Z6LLrpIJ68ll2Wx+gRLFNgmDuwO3FGViEREaogS9fLYFfh38vqr2MG0Rg0KCMOwXzweP9Q59xs6+9F3+CyKor8uWLDgwjPOOGOmj/jEq02Bg7HpkIuUsP1C4AngQWx6/DM0RuLeG1gX2BDYAmutlispyrQQq5x8JXALmuEjPTR+/PjeQ4cO3T6KorFRFO1K7vfh3CiK7ozFYtcAd+lkqxQxEHgcWKvIduOB8yofjohI7VGiXh4OeAMYnfz/1sBD/sKpH+PGjWtdYokl9nPOHQOsnnH3LOfcxc65c8Iw/NhHfOJVb2BnbJbKTsn/lyLC1l93JO3PYoWrarnVUyuwArABlphvhJ3wy9V/Pp+nsNk81wFfljtAaS6lJufAPcANQRDcHobh7OpGKXUqBtyKzQwq5K/ArysfjohIbVKiXj6HAxckr99J8R2QpHO/+93vdsBa3G2TcV+7c+5659yfwjB80UNs4t8QbIR9N6zHbqECdPl8gq3PfhdL5N/BujZ8RnWS+N7A4lhCvnLyskryshy5K2MXEgeew6aEXgt8ULZIpSmFYdgH+EEyOd+N3GvO/5ecz5s3744zzzxzVlWDlEbwd6xeTSF3Yd/38cqHIyJSm5Sol09vYDK21irCKi+/6TWiOtXW1rZ2ckr83mQnL08AZ5x88sl3oOUFzao3NhV8B2BHOmey9MR32Lr3qVjxq6lYX/GOKfQzsYrpc4AF2MFjDPsO7ZiePxgIsJMIi2DfBcOBYcnrQ8oQ5zRsWvvd2Np8rTuXHkm2UtuGwgXh5kVR9ABwQywWuzUMw2+rGqQ0kqOBPxXZ5g1gM+x7V0SkaSlRL68Q+H3y+iVYNWvppjAMl4mi6NdRFB1K9sHjG8C5QRBcGYaheqo2t1HYcpONkpc1sCS6EXyETd9/BquM/CI6QSU9FIbhYvF4fDfgR865bYE+OTabh50Q0si5lMtOWD2fQt/PU7A2bFruJiJNT4l6eS2Ojar3BeZjCcRUnwE1guOOO25w7969xwFHkN3C62vgH/F4/G+nnnrq5OpHJzWoP51rvTcG1sSmlrf4DKoEn2GzcFLX1qtnsJRFGIZLJRKJHwE/wmak5EqWvsOS85uUnEuZrYudbBxQYJu52NKmZ6oSkYhIjVOiXn4XAuOS1ycCv/MYS0MZN25c6/Dhw/fGps6tk3F3PIqi24C/Tpw48ZHqRyc1rhVYHpsmv3LKZRR2gi3XiGK5xbFp9VNIXyffcV2FuKSswjBcOZmc/xj4Hrn3+d8AdwRBcPPMmTPvPfvss+dWNUhpBiOBp4GlC2yTwOqQ3FKViERE6oAS9fJbGaswHWCjvctQ29Wm61JbW9uWwBHOuR+SPTL0ShRFf43FYlerRZCUaBAwAkvaO9aWDyT3+vNe2Hsujk1Dz7WOfRYwHVv3uxk2mj8MtUqTCgrDMIjH4xsFQbBbslJ7ZieNDlOcc7dFUXTz1KlTH1afc6mgfsDD2ImiQn4LnFXxaERE6ogS9cr4N9ZbHeD/sAqnUgFhGI5MJBLjgF8BQzPunuGcuyJZLV7r3cSHu7CCd2BTOh/2F4o0oqOOOqrvwIEDt3XO7RJF0S7Y6GUuk51ztznnbgceDsOwvYphSnMKgJuxIoWFXAocUvlwRETqixL1ytiSzgPy94BVsZE2qZBk5eL9gfFkjyItBG4NguCCMAwfQsW4pHr+D/hb8vpZ2KiRSI+EYbhUPB7fxTnX0a4w39KNV51ztzrnbgnD8OUqhigC8BeK90F/GNge66YhIiIplKhXzjNYMSuA3YHbPMbSVMIwHJNIJI7A1mVmTot/H7hk4cKFl06aNGl69aOTJrM0VmDSYevQV/UbjtSrtra21Z1zu2CztTYl9/47jq0Fvj2RSNx6yimnvFPNGEVSHAZcVGSbt7D38owi24mINCUl6pWzD3B18vpjWJVdqaIJEyasEATBL4GfYWuMU80Hbo6i6EIVn5MKexlYO3l9ZWyWjUhBxx577MA+ffpsiy2d2AlYMs+mX2OV2m+fP3/+PWeccYZ6T4tv2wN3ULjTxpfAJtjJcxERyUGJeuW0YDugZZP/3wQb6ZAqC8OwTyKR2DWKonHJnsGZ3gUuDYLgH2EYflnt+KThnQKclLx+NHC2x1ikhoVhuHzyu2oX59zmQO88m37onLs/iqI7pk6deo+KwUkNWQ14kuyT46nmAVsDT1UlIhGROqVEvbKOBv6UvH498BOPsQjQ1ta2rnPucGBfrKp3qnnADclR9ieqH500qI3pPCB9ENjGYyxSQ4455pj+AwYM2CaKoh2jKNqRzhO7meLAU1EU3R6Lxf4dhuHbVQxTpFTDsWV/yxTYJgL2B66qSkQiInVMiXplDQQ+xlo8xYFVgA+8RiTA/6aV7ov1vF8vxyZvApcHQXBVGIafVzc6aTAB8DnW9m0h1qZNazKbVBiGqyYSiR2xKe1bkH/U/Avn3D2JROLuWCx2XxiGX1cvSpEu64udiNy4yHYTgFMrH46ISP1Tol55fwSOSV4/BzjSYyySQ1tb2/pBEIyLomhfYEDG3QlsJOuKuXPnXnXWWWfN8RCi1L/LgIOS13+CzbCRJhCG4WKJRGIr59y2URRtT/5R8wTwEvBAEAR3AE+GYahuIVIPAuAGrIBrIf+k83tQRESKUKJeeUsCHwKtwBxsSphGRmrQcccdN7hXr177OefG0Vn8K9W32NT4f06cOPFx1OZNSrcHcGPy+pXAAR5jkQoaP3587yFDhowBtkte1sESmVy+BO6Nouiu5Ki5amRIPfoTttSvkMewz8P8yocjItIYlKhXx7+A/ZLXjwfO8BiLlCAMw3WiKDowiqJ9sCnLmf4LXBkEwRVhGP63yuFJ/RmAJWW9sRN1w7DlMFL/XFtb2xokE3Pn3BZAvzzbxoFngfuiKLo7Fos9p1FzqXOltGH7LzYlXi1RRUS6QIl6dayFtWhy2FrV5YAFXiOSkoRh2ALsEI/HD3TO7Ur2etIIeDyKoisWLlx40+mnn/5N9aOUOnEflswBjAFUsLBOJauzbwVshRUHHF5g8w+cc/c75+4HHgzDUPUJpFHsCPybwm3Yvsa63rxblYhERBqIEvXqeRA7qAOb9nqlx1ikG8IwXCQej+8G7O+c24bsz08ca8F3QxAE14Rh+EXVg5RadgRWpwLgdOAEj7FIF4RhODyRSGyeXGe+HXayNZ/ZURQ97Zx7IAiC28MwfLNacYpU0brAo2TXdUm1ENgBO/4REZEuUqJePTsDdySvv4atgdYa5zo1YcKEVYIg2B9rM5OrFc3CKIr+A9wQi8VuVcVmwZK7jmUSrwNreoxFCkgm5h0j5lsBKxbYfCF2gu7+IAjuf+ONN5674YYbtKxBGtlIrA3bUgW2ibDCcVdUIyARkUakRL16HHZwvlry/9sBD/gLR8ohDMMA+H4URftEUfQjYGiOzRYCD0ZRdGN7e/stkyZN+qq6UUoNeYPO74DlsUKT4tlJJ520dCwW2wLYDNiSztcolzjwonPuIefcQ8DjYRjOrkacIjVgIFYYLlfB1VS/AyZWPhwRkcalRL26DgUuTl6/B1vfJQ1i7NixsdVXX32TKIrGRlG0F7nXraZOj78uDMOp1Y1SPDsDODZ5fTxwnsdYmlYYhsvH4/ExQRBsFkXRGAon5gD/dc49EEXRAwsWLHhAtSikSbUAdwI/KLLdNVgBXc0aFBHpASXq1dUb+IjOBG4d4BVv0UjFdCFpfxm4I4qi2ydOnPgiOrBpdJtj6zoB7sXWb0oFhWHYEo/H13bOjcFGzLcCFivyY/9LzBcuXPigZsGIAHA+8Isi2zwCbI/asImI9JgS9eprA05OXr8M+JnHWKQKxo4dGxs9evT3nXN7RlH0Y6w1Vy6fOufucs7dDTyg6bQNKQZMw5ZIzAcWB2Z5jajBnHjiiSN69eq1URRFm0RRtDGwAfnbpQG0Ay85556IoujRIAieUCFIkSwnAKcV2eYt7GSYZpyIiJSBEvXqWxT4GOiPHagvB0zxGpFUTTJp3wIYC+xK/mI884FHoyi6K4qiO0855ZT3qhakVNq/sGmhAD8GbvEYS10Lw7AXsF4ikdjIObdxFEWbAMsW+bE5URQ945x7LAiCx4GndVJMpKCxwLVAUGCb6Vgbtg+qEpGISBNQou5H6vSx04CTPMYiHrW1ta3tnNsJ2Ak7yInl2fR94M4gCO6aOXPmY2efffbcqgUp5bY3toYT4FLgEI+x1JUJEyasEATBBsmkfCNgPWxJUSHTgSejKHoUeGLatGkvXnTRRQsrHqxIYxgD3A/0KbDNXGBrrP6KiIiUiRJ1P1YC3sbOTn+DtffSiE6TC8Nw0UQisU2yV/OuwIg8m7ZjtQ0eCILgga+++uqxc889V+sB68dgLHlsBb7AXueE14hqUBiGIxOJxPrA+lEUre+c2whbKlBIHHgHeCGKoseBJyZOnPgmqv0g0h3LA0+Rf7kW2HfXnmhmkIhI2SlR9+dW4IfJ66r+LGnCMAzi8fj3nHM7Y6Pt65H/8zoba5fzUBAEj33++ecvaMSw5j2IFTUD2BjrSdy0wjAc1ZGUY2vK18eWCRUzJYqiZ4CnYrHY08DzYRh+V8FQRZrFUOBJYOUi2/0a+GvlwxERaT5K1P1Jrf78ITbKHvcXjtSyMAyHx+PxnYIg2D6Kou9TeITjuyiKnu1Yg/vdd989deaZZ6pgWW35DXBW8vpErOdwwwvDsE88Hl/dObeOc26tKIrWxvoxL1LCj8/C+pe/kEgknkskEk+deuqpkysbsUhT6gM8gBWGK+RC4OeVD0dEpDkpUffrKWw0DWAP4GaPsUj9cG1tbWs457aOomhr59yW2HTqfOLAK865p6MoejaRSDx7yimnvI2mA/u0ItBRIPBlYF2PsVREGIYjgbUSiURHMr42NjrXUsKPzwZeAp4HXgiC4AXg3TAMtURApLIccCWdBS/zuRObFagBBhGRClGi7tdewHXJ688CG3mMRerU2LFjY6uuuup6wJbA5s65zbBpi4XMjKLoOefcM0EQPAu8GIbhpxUPVlK9Q+e00lFAXY4Oh2G4TCKRGA2sBnT8uxowpMSHmAm8CrwIvJBIJJ5vaWl5R0m5iBeTgOOLbPMCtr+ZU/lwRESalxJ1v2LYwfoKyf9vio2yi/SEC8NwdCKR2Nw5NyaKojFYIljMl1g/6ZeiKHo5CIKX3njjjfduuOEGjZhUxp+Ao5PXfwFc4DGWgsIw7NXe3r5cEAQrR1E0OgiC0VEUrQ6sCgws8WEi4L/Ay1EUveqceyWRSLx6yimnfFixwEWkKw4DLiqyzWRsJuDUyocjItLclKj792vgL8nrN2HVU0XKKjkNecNEIrEhsCHwPWBQCT86B3jTOfdGFEVvBkHwBvBmGIaT0dT5ntoKKyoHNo10F4+xMG7cuNZhw4aNCoJgRWykfyVsiv5KWG/yfK0Dc/kGeAt4LYqil2Ox2Kvffffda6qVIFKzdgJuo/DSlG+xdm2vVSUiEZEmp0Tdv/7Ax1iF4zg2QvW+14ik4YVhGACrJhKJ72FJ+7rAWsCAEh9iNvC2c+7NRCLxnnPuvSiK3l+wYMH7Z5xxxswKhd1oWrA2bYtgfYgXAypZsdyFYTgCGBWPx0c555Z1zo1KJBKjnHPLYbMuWrv4mNOjKHojCIK3E4nEG865t9rb29887bTTppQ9ehGplA2Ahyj8/b8Q2BnrqS4iIlWgRL02nA4cl7x+HtauTaSqwjAM2tvbVwqCYB1g3SiK1nXOrUPhCvO5fAG875x7N4qij6IomhyLxT4GPv7qq68+Uc/3NNcCP0le3w24vbsPFIbhYvF4fEQsFls6Ho8PB5Zyzi2FJeDLJi+9u/HQC4APoyh6zzn3HvBOEARvzZ8//41JkyZ91d14RaQmLI+1YVuiyHYHA5dXPBoREfkfJeq1YSTWoq0XNqK2DKADYKkJYRguBqyRLBi2ehRFo51za9D1BB5suvxUYLJz7uMoiqZgvbCnxGKxqcDn8+fPnzZp0qTp5XsGNe2nWIVlsLWhh6feeeyxxw7s1avXsJaWlsUTicTi2Kj7MOfc8CiKRgBLYt8fI7GWSt3VDnwURdH7ydkR7zrn3kskEu+3tLRMDsOwvQePLSK1aTHgCYr3Sj8Z+H3lwxERkVRK1GvHFcD+yesnAad5jEWkqBNOOGFo7969V0kkEivRuZ65Y01zKevfC1mAjcx/GUXR1865r5xzXwFfJxKJr4CvnXNfBUEwJx6Pz4yi6NuWlpY5wOwwDL/t4e8um6OOOqpv7969+/Tp02cw0C8ejw92zi2CTXdfZN68eSNeeumlk3r37h30799/3sorr/yoc24YdgC9GD1LvlO1A59GUfSRc24y8BHwYRAEHdc/VTIu0lT6YTUyinWbuRI4ENUkERGpOiXqtWNN4BXsNZmGTVedl7HN0kBAnbZxkuYRhuEwYMV4PD4KWMY5tww2U2RU8t9SK4V310ysEN48YD6da79nRFEUOedSb/sf59wsLKlNE0VRf2zGS8f/h6T8TC+s1kRfLLEejK317unJilItAKYAnwKfA59HUfQpMDUWi328cOHCya2trZ8pEReRpBhwI7B7ke0eAnbEvkNFRKTKlKjXlvuBbZPXc60Hm4D1W7+vijGJlN3xxx8/pLW1tWMN9bAoipYMgmCJRCIx0jk3HBgBDMcS4GY0Bys0Nw1rm/elc256IpGY1nE9iqJP4/H41NNOO22a10hFpN6cB/yyyDavA5sDMyofjoiI5KJEvbbsANydvP46VoW7Y7qZA94FzsF2siINb/z48b0HDhy4aK9evYYCiyYSiaHAolEUDXXOLeqcGxxFUf/kiPcg59xgLLkfgI3a96N7BdS6azY2wj0j+e+cjNu+xUb1ZwAznHMzP/jgg0GPPfbYefPmzaO9vf2VAw88cJOzzz57bhVjFpHmMQGYWGSbT4FNgU8qH46IiOSjRL32vAysnby+A3Bv8voWwCPAX7He6yJSorFjx8ZWWGGFQQCxWGxAa2tra3t7e6tzrtR2dCR/9n9T4xcuXLgwHo/PBujTp8/8MAx70lrtfWAF7MTcUtgUdhGRctoX+BeFj/1mYiPp6pUuIiKS4WDsYD0ifYr7Zcnb7vIRlIhU1F/p/Nwf4jkWEWk8W2NrzaMCl/nJ7URERCSH3thoWseOcx1sGu+c5P8/9BeaiFTID+j8zN/iORYRaSxrYktvCiXpcWBPXwGKiIjUmiOBo8juR30SnTvPf5I+yt5OSvVpEWkIvbD16xG2pr1cbdlEpLmNwrpCFErSI+x4RERERJJasfYnC4DbgN2Stw0BZmE7zwVYpfd2Oneoq/gIVkQq6iY6P+M7eI5FROrfUOAtiifpf/QVoIiISC1bFCsk1bHDnA78Cbia/DvVXbxEKiKVlDpzRp0dRKQn+gKPUTxJvw4IPMUoIiJS81Ymff1YIuPfzMtRfsIUkQoahq0TjYDJqCuHiHRPANxI8ST9EarbvlJERKQubQUsJD05z5WotwN/9xSjiFTWU3R+1tfyHIuI1KdzKJ6kv4EtsxMREZESHELxnWsc+I+vAEWkoibQ+Vk/0XMsIlJ/jqP4ccSnwNK+AhQREalXZ1LaTlZEGs/adH7On/Qci4jUl5+Sf8lcx2Um9j0jIiIiXRQAt1J4Z5vACsWISOP5iM7ZM0v4DUVE6sSu2PK5Qkn6AmA7XwGKiIg0gr7A83QWlsp1WcNbdCJSSefT+Tk/0HMsIlL7NgZmUzhJTwD7+wpQRESkkYwEPie9f3rq5Uf+QhORCtqJzs/5DZ5jEZHatgbwNcWXzP3WV4AiIiKNaF3gO3KPrB/nMS4RqZw+dI6OzQR6+Q1HRGrUctgJ/WJJ+jm+AhQREWlkPyZ7vXoCuMRnUCJSUbfR+Xnf1nMsIlJ7FgfepniSfjVW+0ZEREQq4Hiyd75PeY1IRCrpMDo/63/xHIuI1JZBwAsUT9IfAHp7ilFERKRpXEj6Dni233BEpIJG0DmT5gPPsYhI7egF3EfxJP0ZYICnGEVERJpKL6wSfOqOeKDXiESkklI/76M9xyIi/sWwApPFkvR3gWGeYhQREWlKi5JeOGZdv+GISAWFqGKziBgHXETxJP0TYFlPMYqISBk53wFIl60CPIutUdsbuC55e3+spdswrMhM6vURdI6+L4K97gOAVmz9Wm/g2+T9M7Cd/RxgQfIyHZgKTAO+SP47NXn7tIo8SxHZAHguef1RYEuPsYiIX6cCJxbZ5itgc+CtyocjIiIiqVqA5YFjsP7qTwL3Y2tYMyvDV+syP/n7bwdOB8YBY4DBFfobiDQLB3yKfc7agaF+wxERT/6P4vviOcCmvgIUERFpNssD+2BVn58C5uEnGe/u5X3gKuAIYGNUfVakqy6m8/O0r+dYRKT69gHiFN7XLgB28BWgiIhIo3PA2sBx2Oj0NPwn2pUYfX8KO/GwM9CvLH85kcb1Qzo/P1d7jkVEqmtbip+gTwAH+ApQREQqR2vU/RqC7Yh3SF5G9uCxFgJTsGJzHWvHp6Rc/zq53TfJf2cnf2Z+8jIoeftgIMDWvPcC+tK55n1xYAlszfviwJIpP9cd84DHgHuAu9G6OpFM/YAvsc/hTOxzt9BrRCJSDRsC/6F4i7WjsJPfIiIi0kOLA7/CEtR2uj4q/TnwIHAB8BtsZHolbP26D8Ow4jWHAmcCt2IJ9wK6/tw+As7GimiJiLmLzs/I9/2GIiJVsBp2gq7YPvMPvgIUERFpFH2AXYHr6VoCOwt4HDgHm9o2qspx90QrsD7wa+AK4A26VvDuLaw91YpVjluk1qQWkvqj51hEpLKWw1qsFdtHXuArQBERkUbwPeByrPVZqYn5rcDPgTWBWNUjrqzh2Jrb87BCc6X8TRLYyYqDUDE6aU5L03mS623PsYhI5SyJdVEptl+8lcY7PhAREam4ABs9v5/SEtEPsBHzbWm+RHR5rJ3b9ZR2MmMa1gJuSR/Binj0Mp2fg5U8xyIi5bc4NvOs2H7wQWyWnoiIiJRoEHA08CHFd7SvYdXdl/ESaW3qC4zFRgrmU/jvNxf4BzbrQKQZnELn+/8oz7GISHkNBp6n+LHDy8AinmIUERGpO72wUeFi7dQ+w0bOx/gJs64sgq3Jv5/i69pvB1b2E6ZI1WxM53v+P3m2Wbd64YhImfQDHqV4kv4e1nVFREREiogBPwM+pvD66nuBHbEp8dJ1ywN/AmaQ/++8ACusM8JTjCKVFgBT6Xy/Z46qDQOurHZQItIjfbGp7MWS9I+BZT3FKCIiUld2pfBasu+Ai7AWK1IeA7EK8oWK0M0BTk1uK9JoLqPzvb5Xxn1/x2aXiEh9aMU+s6XUZlnVU4wiIiJ1YwRwE/l3qN8AbcBivgJsAgGwO4XX803GZjGINJI96HyPX5Fy+8rAQqzfuojUvhhwLcWT9G/QkhYREZGixgJfkntnOh+4EJt+KtXhsNfkXfIf5FyPVdIVqScBsAvZ7ZcGAPOw9/ZXKfffmLzt3moFKCLd5oBLKJ6kz8RavIqIiEgeywEPkHtH2g5civU5Fj9agV8An5N/2uBPvEUn0j17Y8trdsu4/T4639ubARvRWXDxgWoGKCJd5oDzKZ6kzwG28BSjiIhIXdgFm3qWa0f6NGoPVkv6A2diU4BzvV4X0Hx96qW+3YG9d58ANk3edgSd7+lJwGPYCcMEVpRKRGrX6RRP0uejpVsiIiJ5OazPeZzcheKOI3taqtSGtcm/fv15VDlX6scywGw6378PYAfwHf//iPT39yNeohSRUvye4kl6O7akS0RERHIYBNxC7p3oI8BK/kKTErVgJ1Pmkv0aTge28ReaSJeMp/O9G8cO5L/JuK3j+uOeYhSRwlJnwuS7xIF9fQUoIiJS65YG3iF7B7oQOBIbaZf6sSbwHrlfzwM9xiVSqgB4BkvQO96/CXIf6D/tKUYRye9g8n9mUz/T43wFKCIiUuuWJXeP7i/QCGw9GwTcTO4Do195jEukVGuQv/ZC5tIOEakdPyX3ErrMyzG+AhQREal1KwOfoDXNjSpfzYEENlNCpNZNpPjB/kveohORTLtT2gm2E30FKCIiUutGk7u117VAL49xSfn9CKuoq9EMqTe9gbdInwKfeXnFW3Qikmoncu9rMi+n+QpQRESk1g0FPiB753k1VpBMGs8OWOX+zJF1FfGRWrc5hde6vuEvNBFJ+gG5C5lmXs71FaCIiEitawUeInvneTFWwKle9AKGFLiUqwBetX5PNWwJzCL9dZ8LbOgzKJESXED+ZP1tj3GJCGxHaUn65dTXcYaIiEhV/Z3snecF1FfCCbAnhQ8ItizT7zmzyO/pW6bfUy1bkn1A9Qkw3GdQIkUMAqaQewr8ex7jEml2WwCzKZ6k34hm7ImIiOR1ONk7z/upz51nsUT90jL8jhZyr+Ov50QdYH+yn8cTqDaB1LZdyP0Z/K/PoESa2OZkz9LKdbkb7V9ERETyWoXsNcofAov5DKoHiiXqs4EBPfwd+RKDek/UIfdMgT94jUikuFvIngI/2WtEIs1pS0obSb8H6OMpRhERqUP1Ns27pwLgcWCTlNtmJf9fr4WY9gRuKLLNQcA/e/A7bgT2KLJNP2wqeb2JAbcDO6bcthDYCLW7kto1AluTPijltqnAisCw5O0tWGLQN3l9YHI7hyUO32GVqRcAc5K3zQC+Tv4rIoVtho2SDyyy3QPAbtTnPlJERKQqDiH7LPf+XiPquVwj6l9k/P+hHjz+ULLbzEzP8TvrdUQdYFGyp/Y/QfOdyJLaNgQ7gXQAcDLwKOnv2UIV4bt6mYfVbHgWuANbQnMqcBg2grhEhZ+rSK3bFPiW4p+lR4H+nmIUERGpCwOxIkypO9BbvUZUHrkS9XNILzaVAJbv5uMfkfHYTwNP5vid9ZyogwomPAAAACAASURBVI12ZD6nvb1GJM1qILAV8FvgEuAxsk++1cJlBvAMcCVwEtY7emgF/h4itWYzSkvSH8Jmm4mIiEgBJ5K+A/0OWNZrROWRK1E/Frgv47bfd/PxX8x4nF/QmIk6wG2kP6f3sKnxIpUSA1bHRsnPwZbmLMB/Et6Ty+fYcpLjgDFoXa40ljGUlqQ/Ts/rw4iISBNrlqm9/bGCcYun3HYy3U9ea0muNerHAZ8CV6Xc9hE2qh514bHXBF5N+f88YCRwJ+nr/KF+16inWgmrVdCacts+wLV+wpEGtSSwPbAD1nd5kTI+9lxsaco32Kyaecnb2rF6HGDfAQ77zPbGqlD3T942BFvjXs6punOBR7C1vPcA75bxsUWqaXNs/1dsTfqT2Od7VpHtREREmt440s90z6S8B8c+5RtR74MdrKfevmUXH/vsjJ+/Jnl7o46og63FTX1eT/kNRxpADPg+cAbwCt0fqZ4HvA7cBEwCfoUVedwKeB/7XiuXfsBy2Am5H2ItLf+AfQe8QGmtqPJdPgD+BuyKRtulfmxBae/7J0kv8igiIiIFPE36jnSS33DKKl+iDnBhxu2XdeFxW8leF7t98r5GTtRXJrso15peI5J6tT7wZ7ILFRa7xLGZHZcBv8ZG5pan8DKMDbETc9W0FLA18HMs8X4B65jQlec6Azs5tjXWlUOkFm1JaS3YXsBmpYiIiEgJViL7IHg5rxGVV6FEfZOM22dTfMpehx9l/OyndCYKjZyog7XSSX1uZ/oNR+rIKKyw2puUnqxOB/4NTMCmwg/u5u+uhROQfbFCW0cD1wEfU/rf4RPss7ZW1aMWyW97rKZNsffv03T/sysiItKUjiF9Z/qg33DKrlCiDvBWxn0Hlfi4mYXVTku5r9ET9X1Jf25v+Q1H6sAY4HrSuy3ku7QDzwOnJ3+uXCPJtfoZXB5bfnQ7tl69lKT9eazAXmuOxxOplh0o7T37BJruLiIi0mUPkr5DHe83nLIrlqgfn3HfQyU85jCyK0+vmnJ/oyfqg8nuHd9IszCkPPoAhwCvUfxAfgZwObAXzT01th/Wxu08sttl5rp8jH2fNfPfTPzYBasLUew9+hilz1QTERGRpBjZxV9W8BpR+RVL1EeS3VO92N/gNxmP90TG/Y2eqEP2CZ79/IYjNWQg8DtgGoUP4OdjM1PGoqJpucSwacVXULxI12zgXGCEl0il2eyGknQREZGKWpP0neo0v+FURLFEHawtUur9YZHHzKxMPS7j/mZI1E8l/fn91W84UgN6AUeQXWQx8/IM8AtgqJ8w61I/rBXi3VgdkXx/2znYZ1NrgaVS9qW0ooiPoj7pIiIi3ZaZxN7tN5yKKCVR/0nG/R+Sf13sBhnbfkd2K7tmSNR/TPrzu8dvOOKRw0bF3yf/QXscW4O9racYG8mKwDkUrrL9FXAcjfe9I36No/CJoo7LIyhJFxER6ZHfkr5zPc9vOBVRSqLeB/g6Y5vv53m88zK2+1eObZohUV+H9Of3jt9wxJMNgZfIf8D+DVapfBlfATawRbEaG5+Q/+//MdbnXaSnfkl2a85cl3tpvP2diIhI1f2R9B3s8X7DqYhSEnWA8zO2uSzHNr2wVlGp222XY7tmSNSHkv78ZvgNR6qsH/An8ldx/xZbp65RtcprxXq1F+pHfx2whK8Ape79juIJesesPNWbEBERKYPM5PSXfsOpiFIT9Q0ztsnVU32vjG0+obN3eqpmSNR7kf78FvoNR6poc2wGRa4D9QXAhcBwb9E1r37YdPdvyD+7YRy2VEGkVCdTWpJ+O0rSRUREyuYS0ne0h/kNpyJKTdQBXs3Y7uCM++/KuH9insdphkQdsqdB5jppIY2jD3AB+ae/Xo3a9NWCocBZZLeQ7LjcCSzmLTqpFw74C6Ul6ddgMztERESkTM4lfWd7hN9wKqIrifqxGds9nHJfZhu3CFglz+M0Q6Lel/TnN89vOFJhSwFPk/sg/UPgB/5CkzzWwCrs53rNPsFmEYnk0gJcTmlJ+iXoJK2IiEjZTSJ9h9vmN5yK6EqivgTpbWdSe6ofn/EYjxX4nc2QqI8g/fl96TccqaAtgKlkv6cT2DR39UmuXQE23T1XH/Z5wKH+QpMa1Ru4kdKS9PPJ3yFFREREeuBXZJ8ZbzRdSdQB7sjYNkze/mbG7YcUeIxmSNQ3Jv35vew3HKmQY8hdMO5t7D0g9WFF7ORirmTr72jaspgBwH2UlqSf7ilGERGRprAzpY8S16uuJupjM7b9CNgs47Y5wKACj9EMifoBpD+/W/yGIxUwkdwH6LcDi3iMS7qnBUuucr2md6BCYM1uUeAplKSLiIjUhFGk73xnYwdzjaSriXpvbBp36vavZPz/iiK/sxkS9cx+8qf4DUfKKF8RqQR2gK6prvVtb+y7PvP1vZvG+56S0owEXqN4gp4AjvIUo4iISNPJXHv6Pb/hlF1XE3XILrKXedm6yM83Q6KeefJiF7/hSJkEwEVkv3+/Brb3GJeU15rAB2S/zvdjbd6keawKfEzxJL0d1TQQERGpqsyiMSf7DafsupOor5/jZ1KnwhcbUWz0RH0Zsg/ghnqNSMolV5L+BbCOz6CkIkaQXXsjAh4EenmMS6pnfezzXSxJnw/s5SlGERGRpnUg6TvkN/2GU3bdSdQhe8S44xKW8LONnqgfTfpza8TaBs3oKLLft1Ox0VdpTMOwQpCZr/tlPoOSqtgKmEnxJH0WsJ2nGEVERPJqtPXaudyJtSTrqPo7GtgUSzab2RXAWRm3RcCVHmKpNZnTH2/1EoWU07bAmRm3TUneXk8n78aQfxlGAvg99n3XUzsCW+a5bw5WiK8efIElbPeQ3lf9IOB54G8eYpLK+zFwFcULCH6DfZ6a/XhARETEm5tJP4N+td9wyqq7I+pDsArwqZedS/ydjTyivi3pz2sB1n9e6teKwFekv67fAKv4DKqbjqTw6OBuZfo9+WbcRMD0Mv2OaloUeI/sz/ZWPoOSivg1EKf4SPoUYC1PMYqIiEjSTmSvOV7Ja0Tl091EvScaOVF/kPTndaPfcKSHegOvkv3538FnUD1QLFG/qQy/Y4Miv6MeE3Ww2VSZU6GnY2vZpf45bOlWsQQ9Av6LncATERERzxzZB+vXeo2ofJSol882ZD+vTb1GJD11Btmv6W+8RtQzxRL1+cDiPfwdma0JGyVRB6vs307687nda0RSDr2wqe6lJOmvY+3aREREpEbsTfrOOoGt96x3StTLowU7gEt9Tvd5jUh6aj2yk7KrvEbUc7kS9UTG/3/dg8fvTfYygczHr+dEHeAksv+GP/EakfTEYOA/lJakPwYs4idMERERyScAniV9p/0W9Z9gKlEvjwmkP5848D2vEUlPPUL6a/opVpuhnuVK1J/I+P+LPXj8vTIe6yPs79ZIiXoAPEr6c/oE9VevRyOw93spSfqt1P9+SkREpGFtTvbOO7Pyeb1Rot5zawHzSH8+l3qNSHpqd7Lfozt5jag8ciXqvyT7/bt2Nx//rozH+QPwccZt9Z6oA6yKLRNIfV4neI1Iumo17ERSKUn6+UDMS5QiIiJSskvInta5t9eIekaJes/kqgY9Heu/LPXrOdJf00ZZh5wrUd8HK3qYetufu/HYI0lfKpAAVqAxE3Wwk7Spz+tLYIDXiKRUm2Hvw2IJegIrMCciIiJ1YBAwmfSd+XfU7zTnpchus1bptlNb5/id9Tha0ULutY17+gxKemwHspcxrOY1ovLJl6jvknHbNKC1i499XMZjPJy8vVET9UWwNn2pz+1IrxFJKQ4iezZErks7cJifEEVERKS7NiF7quhk1C+72eSqbn2e14ikHG4h/TW9wW84ZZUvUW8BPs+4/YddfOw3M37+oOTtjZqoA5xC+nN7G+sSIrWnK+3XZgM7e4lSREREeuwgsnfuz1L/xaakNCeQ/fo/SNdHIaW2LAEsIP11baQWe/kSdYA/Ztx+cxcedxOyE52ByfsaOVFfguzR2UboBtJoegNXUlqS/iX2fhYREZE69meyd/Iv0fM+xFLbMqf4RsCH6HVvBIeT/rq+5TecsiuUqK+WcfsCSn9PX5jxs5el3NfIiTpkr+//i99wJMOiZHdwyHf5AFjZT5giIiJSTjHgXrJ39q8Bwz3GJZUziezX+1tgTZ9BSdncQfpre5LfcMquUKIO2S0oS+mp3ofstdpbptzf6In6j8hO9qQ2rER2sc98lyfRyVYREZGG0h+b8py5038bq3gsjaEFOJfs13kmVkFY6l8MO+mS+vqu4TWi8iuWqP9fxn2l9FTfN+NnPiR9nXajJ+r9gbmkP8dlvUYkAFtg09hLSdKvp347j4iIiEgBvYF/kzuJ291jXFIeiwH3k/36fgNs7DEuKa+1SH99v/AbTkUUS9QHY10sUu9fp8hjZn42fpdxf6Mn6pA9tfonfsNpeodSWmX3CDgHCPyEKSIiItXQi+xq0RHWh/V0dCBQr9bDRggzX9evqd+WfJLbAaS/xnf4DaciiiXqANdl3H92gcdbiuze6ctnbNMMifqZpD/H0/2G07Ri2N++lAR9IfALP2GKiIhItfUCriL3QcG/sZFZqQ8OO4jLbMMXYa34tCa98YSkv85neI2mMkpJ1HfMuP9L7LstlwkZ2/4nxzbNkKgfSPZUaqmuQcDtlJakfwvs5CdMERER8Wkc2S2eOkZhx3mMS0qzHLmnukfYFNcl/IUmFXQp6a91I462lZKoB8AnGdvkW8LzdsZ2++fYphkS9S1If47P+A2n6ayEdWgoJUn/lOLLOURERKSBbQ5MIfeBwp3A0v5CkzwC7ETKLHIvYTgH9UlvZDeR/prv6TeciiglUYfs6cO35Nhm84xtZgEDcmzXDIl6Zmu7RmvrV8u2J7vrQL7LS9hyDREREWlySwFPk/uAYQbwS5T41YoNgKfIP02yEZM2SXc36a97I06NLTVRXxk7OdWxzUKyZ5JkzkC4JM/vbIZEfRnSn+MnfsNpGr8hvUZCocv1QD8/YYqIiEgtasF6Ec8h98HDR9gororN+TEKuBCIk/v1eRibVimN7y7SX/ud/YZTEaUm6pB94urIlPv6k93Kbkyex2mGRH0U6c9xstdoGl8f4HJKS9ATQBvpLQNFRERE/mcFcvdb77i8Boz1Fl3zWRyb3puvhc832AkUHdw1j+tJfw/s5TeciuhKon54xnavptx3YMZ975H/s9IMifrqpD/HN/2G09CWxmoAlJKkzwb28BOmiIiI1JMAm+6eORKVOYK7Gxphr5QVgXOBueR/DW4EhvsKULy5iPT3wRF+w6mIriTqg8meCdRRhOuhjNtPKvA7myFR34b05/iE33Aa1nZYF4JSkvSPgLW9RCkiIiJ1azFsNDdX66+Oy/vAccAinmJsNGOwEdNC6xmfww64pTmdSPr7oVD/8HrVlUQd4Gqy/yajSF8qEsfWaOfTDIn6oaQ/x3/5DafhOGx/WOp69CdQdw4RERHpgZWA60gv2pRrCvaZwGhPMdazQcDPgBcpfFD3BTbNWdPcm9tPSH9f5OoJXu+6mqj/IGPbL4FJGbfdV+R3NkOi/lfSn+PJfsNpKAOBGygtQY+Ai4FeXiIVERGRhrMBcC/FD0BeAI4CRvgJsy70wpYOXAd8R+kHd49iLZakea1I+ntiJo23BKWriXpAdqK9IOP/+xb5nc2QqD9L+nPczW84DWM0pfdHX0hjLlcRERGRGrAOVsm20JT4CJv+dx9wMJreB5acbw38HfiK0pPzzMt8YCLQt7rhS41wWBKZ+p7Y1GtE5dfVRB3g1Bw/k3oyo1jLq0ZP1IeSPSVb38s9tweF67lkzorayk+YIiIi0kyGYevxPqO0g5Q3sDXv29I8fdmHAwdg685nUPxvFKdwAbnUywfA9tV7KlJDriH9vXC633DKrjuJ+krkX55zYQm/s9ET9QNIf34v+w2n7rVgn7tCS8JSLy9gdRNEREREqqYPdhB4H6UX0fkam/Z9BLAR0LvqUVfG8lhC8RfsxESpo+QfYOtFx5O/V3q+y7+wkybSPPYh/T3wCRDzGlF5dSdRB3g8x89FwCYl/GyjJ+oPkP78tD69+0ZReuu1CFuP3ij7OBEREalTI4GjsdGDriSb84GngXOA/bA1f7VeaGdJbES7Dbgdm9bYlec8HfgbNm05tUDcelhl96481jfAr2m8tcqS2yCs93Lqe2BnrxGVV3cT9Z9iJ71SL4+U+DsbOVFfkeyR3zW9RlS/dqP0pUvzgHF+whQRERHJbzXgD1gBo66OEkdY0Z33gDuBPwGHY+v7lgf6VyH+Fqwg3vrA3sDvsTZQz1P6msTMy+fAP4BdKTz9vwVLvGd18fEfB1bv+VOXOnAZ6a/9w16jKa/uJuo90ciJ+gWkP7en/IZTl7o61f0TYGMvkYqIiNQRtbPyb3Fs9HlHrJXSYmV4zDnAVGAaNpo9BUugFyTviyf/D53rw/tjI/V9sGJsLVhbnd7JGEdgBZYWpzzTyduxXrn3JC+vJOMo1Shs1H2nLvzMAqyP9CnYqKs0pu9hJ8FSbYl1Bqh3R5LdH35fbG1+pXwMLJ3y/y+x74F6txTwPulTrw8ArvQTTl0ahS3V2rDE7R/C2ig20skeERERaQIBdsBzFHAt8BHdG52uxcu3wINYD+cfAoPL8ydjL2w0viuxfIYtI9CJqsaV2S7xRRpj+YNG1MvnKtKf1wfYCUopze5YTZVSvnMTwJno7ysiIiINZAls7d+pWFG6yZQ+xdDXZQY2onkRcAiwBpUt6DUYG13v6jKCx4B1KxiX+LMZ2Z+TQ71GVB5K1Mtjc7LfH4d4jah+9AXOpfT90FfALl4iFREREamy/liC+ROsYNtVWMG5Tyjew70clwQ2rf51rFjcWVhhoO9jbdd86U6xuThwBY0xlVfSXUf6az0TWMZrRD2nRL3n+gHvkv6cXqKxugNUyupY+7pSv1+fBZbzEqmIiEid0zS0+jQHO7B8Kc/9i2AJ8+LYiPxwOteg98emAA9O2dYlH3MBlujPxQrWzcbWkk/H1rp/nrw+PXl7rXkRazf1S2wd+oASfiYA9sdGfP4AnIcl71L/jsMqvncUWByEnZTZDnt/S3P6C9ZbvkOEFajU5z4/BxyG1UfoV8L2ETbq/ltsvyIiIiIiAliLuJvo+myBl7BpsdIYfkX2a3y+14h65mCy26ztWuHf+XjG73u+wr+vknK9H87zGlHtGwbcQdeWPu3pJVIRERERqRt7Yuv6uzq1/yos2Zf65oD7yX6Nf+4zKPFia2wmRer74F1Km3nTrHbFZlOV+t35JPW/vEREREREqqQfcDI2pb8rCfss4HjS2zdJ/RmKjQSnvrYLsJoK0hxWwtrKZdYsGO0zqBrWF5t5UmrBuARWq6TVR7AiIiIiUt+WA26h69Ph3wP28BCvlM9obEpu6us6B9jWZ1BSFatghTYzi0iqEnluGwNvUfr34xRgey+RioiIiEhD2Rp4ja4n7M+g9ev17Idkt/CbR+XXeIs/o4HPyP4sH+szqBrVBzgdKxRa6nfiPfjt9CEiIiIiDaYVq/ScOcpayuV2YMXqhyxl8BuyX08l641pHaxDRebrfbHPoGrU2lghzVK/A+di35/OR7AiIiIi0viGAueQPdJa7LIAuBBrfyf15Uiy194uAI7wGZSU1W7kPgl3AdaSUUwr1sZwAaV/970OrOUjWBERERFpPhtgFYu7Oro+Cwix4ktSP8aR++TM1XT2XZf647DEM9drez4aAU61FvAipX/XJbCTmiquKSIiIiJVFQCH0bV2RB2XycD+aLSunuxHdruuCHgFWMFjXNI9iwJ3k/vz+UePcdWaXlgXjFzv/XyXz4DtfAQrIiIiItJhCPBnYD5dT9hfBLapfsjSTbsC35D9On4DHOQvLOmi7YCPyL1EZby/sGrOGOANuvad9i/sJIiIiIiISE1YHriW0nsJp17uwgo0Se1bEXiV3K/jw1gPbqlNg7FaEbk+o18AW/kLraYsgq3P78p32RRgdx/BioiIiIiU4ntYwtbVZD2BVYhXwl77+gKXk/t1nIOte475Ck5y2oXs/ugdl8eBEf5Cqym7kv/vlO9yPbCYj2BFRERERLpqW7rXfz2OHfiuXP2QpYvGY4l5rtfxeew9IH6tBdxJ7teoHVuP3uotutoxEriJrn1XTQP28BGsiIiIiEhPtGAVw6fQ/YRdhcpq23LAfeR/He/HugRIdS2DTXNvJ/fr8hqwkbfoakeAfUd9S9dH0Rf3EK+IiIiISNkMBE4h/+hroct84DxsxEtqkwMOIXehuY5lDdcBo30F2ERGAH8lf3HH+cDvsWrmzW5jbOZHV0fR9/IRrIiIiIhIpSxJ4VG+Ygn7hcDwqkctpRoBXEP+IlwJbBr2tqhHd7mtB1xB4e4LDwNreIqvlgzFepzn6h9fbBRda9FFREREpGGtBdxD15P1CJgFnIZaINWyNbCkptDr+A7wa6CfpxgbQYCd9Lidwn/r14GxnmKsJa3A0cBMuvad8y6qiC8iIiIiTWQL4BG6l7DPAE5GI1y1bFuKTy2ejk3V3thTjPVoVWAi8CGF/7YfAPthCX2z2xZ4k67P4pkI9PEQr4iIiIiId2PofsI+G5vGunTVo5ZSOKwy9tMUfy3fA0LUiz2XEcBRlLam+m3gcLQOHWApbElAV79XngBW9xCviIiIiEjN2QV4ke4l7POBS1CSV8s2wYrKLaT46/kc8AdspL1Ze7KvAfwWeJDS6jrcD+yMRtDBClhOBL6j6zN1fo7+hiIiIiIiaTpGYF+newl7HLgW/r+9e4+Rq6oDOP7tbrstobTdlnYXim0p7VYeVeymKYESEm0VIU2IER9EGv5QgpqAiTFAiKRijGDEhGjUYoymgURBMcQAMUQUuqSJ9uGrpYWw1b62FPoKtNvnjn/87mSHptWZO3fuvL6f5OTObrd7z5neu93fPef8flydd8dVtllE/e4DlPdv+g6RpG4l0FOH/uZlEvAp4HFgB+W9N0eBnwML69DfRjQW+DKwl8p+bowATxArFyRJkiSdQweRAGsr6QL2AjAArMi74yrbeOLfZw2Vle7bQyRQu5fYNnFe3h3PQCextHolsXVjADhB+Q+jBoj635Py7ngDW0bUh6/058Qm4Po69FeSJElqWsWA/Q2qD9gtCda4phC12P9E5WWzjhN74H9KZPW+GZhHzK7W2xhgNrAc+CrwQ6JM2rtUfh1vJMZ3cZ4DaAKLife00vfzAFF5oF23VkiS1DL8JV+qny4ikLuPWDqdxibgUeBpYvZSjakHuDFpy4m612mcAAaJVRm7gH3Ekui3ktdDyXE45ffvAmYAvUmfpxNLp3uIYHo+sID0M/5HiAcXLxDlDAdTfp9WNY8o1fhpKvv/eQT4GfAAsL8G/ZIkSZLazjjgDqpbEr+b+CXd0m6Nr5NIQvcQMWte7vLwStshYoZ1B1HW7F9EhvX1wN+Tz21PvuYAEexl3YfTyXkfJR5QjM/g/WtFHwB+QqykqPQ9fhVYlH+XJUmSpPbQQcykbSB9YHSUmFm7Kue+K71xQD+xZHkNsJnaBM15tINEpvZVxNaMtCsH2sVM4EfAMSp/r3cAX8CVcZIkSVJulhLJxaoJmgaIvfDuV20+vcBNwNeB1cRy8SHqH4gX235gHfAL4H6iqsGcWrwRLWo68DCVl1orELkAVtGcCQclSVKZfBIvNbbriD3sN5P+fn2TmGVfTSyHVvOaDPQRe5l7ktZLBH7FfeUziL3maZwm9ri/zeh+99K974PANqK8nCo3nXj4cjeVB9oniQcjDxI5CSRJkiTV2YeJutunSD8Legj4Ps58tovJQDex/3kuUTqtP2kLk8/NSb6mGx/c1lIvce9VUrKv2EaAp4iHM5IkSZIa0DxidjzNntZiOw08B9xCY5T7klpVH/A46e/XtcA1ufdakiRJUio9xD7VvVS313gP8B1idlVSNpYAvyUeiqW5L/9BPEiTJEmS1IS6iIRx66guYD9NZOteiUmqpLSqTQK5mbgHTQApSZIktYgbgGeobh97gUhU9QgwP9/uS02pC7idmAVPe89tAT5HlGiUJEmS1IIuBX5AJI+rJmAfAV4CPg9MyHUEUuObCXyL6srmbQVuwwBdkiRJahsTgTuJ2bpqAvYCEfSvAVZgAjq1t37iXjhB+vtpkLg3vZckSZKkNtVBBNjPkz65VWnbCXwP+FCeg5DqaCJwF/BPqrt3NgCfxT3okiRJkkrMBh4CdlF9wF4gApf7gFl5DkLKyVXAY1S/jeRFYHnOfZckSZLUZDqAZcBTwEmyCdrXA/cA03Mch5S1KcSy9AGqux9OExngl+TbfUmSJEmt4BLgQeA/ZBOwHwOeBe4ApuU3DCm1TuAm4NfE9VvN9X8U+DFwWa4jkCRJktSSOoBPAr8ju1n2U0Tm+Ltxebwaz+XAw8Buqr/WdxMPvGbkOgJJkiRJbeMi4H5gM9kE7AWi3NtfgQeAK/IbivQ+lwLfAP5CNtf1n4FbgXE5jkGSJElSm7sSWAVsJ7ugvUCUqHoMWIp1pFVbs4n8CQPEA6Nqr91hokSblQ8kSZIk1VUHEVSvBg6TbdC+j0hsdyexZ16qVtbBeQF4HbgXmJrjOCRJkiSpLOcBnyGSxh0n26B9BNgEPAJ8FBif05jU3DqBa4BvAxvJ7no8CjwBfAwYk9toJEmSJKkK04C7gLVkN3NZ2t4DniNmRz+Y05jUHC4EbgOeBN4m2+vuVeBLwOTcRiNJkiRJNTAT+ArwInCC7IP2AlFG7pfAF4ms3c5yto8xQD/wTWAdUVUgy2trF/BdYEFeA5IkSZKkPHUDtwPPAEeoTdBeIGZSnyUyeV8LdOUxOOWiE1gEfI0oG5j1rHlxafuvgBuT80mSJElSW5gArCAS0e2ldkF7gZjJX09klL+VWJqv5jCW64+cswAAAsZJREFUmDG/h0guuJ/aXCPDwO+BlcAFuYxMkiSpAi4ZlZS3scANwC1E8D67xucrEKXgNhJJ6ortrRqfV//fbCIw7weWEMngzq/RuY4SuQ5+kxyP1Og8kiRJVTNQl1Rvc4FlSfsEMCmn8x4EtgAbStoWIrBX9i5mNCjvBxYDPTU+5zDwR+BpYun8uzU+nyRJUiYM1CU1kvHAdcByImi/mnx/Th0CNgPbiJrZb5Qcj+fYj2bWS2TnX5C0K4l95hfmdP43geeT9jIRrEuSJDUVA3VJjWwGEbR/PGm9derHCJFtvhi0b02Og0Sm8GN16le9TCSWrS8oaZcDfcCUnPtyDHgFeIEIzl/P+fySJEmZM1CX1CzGELOzS4lZ96XAnHp2qMQ7wB5gZ3LcXfK6eDxYt95V5nxgFnAJUW7vbK/zDsbPtB34AxGYv4T7zSVJUosxUJfUzGYyGrhfDyykcUtsHSeW1p+rHSw5DjO6ZPsEo4HoKUb3WY8Ah4k9/cUxdyfHTkb3+ncRwfeE5M+nJseztanE9oNGs42YNX+FWM6+s77dkSRJqi0DdUmt5AKilvq1ROC+mFimreYxQuQJeBlYSwTne+vaI0mSpJwZqEtqZR3AfOAjSVuUHK2t3jiGGM26vx5YR9RPlyRJalsG6pLaUbFU2BXEvvd+IhmaPxNrqzQoLwbmQ3XtkSRJUgPyl1JJCtOIoL2PmIXvI7KZX0bs81b59gGvERnYtxH16f+GQbkkSVJZDNQl6X/rJLLL9zEavM8H5hEz8+0axA8D/yZK1RUD8teSY7NkuJckSWpIBuqSVJ1uImC/qOQ494zX3ef8243pOFFibogoLTd4ltdDQKFeHZQkSWplBuqSVHsTiaX1U0pa9xkfFz83mdFM9R3Jx0WTk89BlFzr4v3l24aBY8nr94CTyevDRGm3g2W2w1WPWJIkSan9FxagEXniubi2AAAAAElFTkSuQmCC" width="65%" style="display: block; margin: auto;" /> ??? You could stop here if you were interested in linkages with the network and an outcome. --- ## 8. (optional) Combine both estimations into joint model <img src="data:image/png;base64,iVBORw0KGgoAAAANSUhEUgAABaAAAAJTCAYAAAD+AbHxAAAABHNCSVQICAgIfAhkiAAAAAlwSFlzAAASdAAAEnQB3mYfeAAAABl0RVh0U29mdHdhcmUAd3d3Lmlua3NjYXBlLm9yZ5vuPBoAACAASURBVHic7N15nBx1mfjxT3XnIoQQjnAE5T4NIIJyCB4oKip4EldE8Y6uisguP04DBQkQWVxEd1VQWWXx4lARUBFU0IAXgsp9hBsC4Uog1yTprt8fT89OT0/P3d3VM/N5v171Sld1dfVTXdWTmaeeer4JkiRJkjQ2bQJsAWwOTAc2BTarerwhUASmVtZfHygA6wITgNXAciADllTWeQEoAc8DTwGLK/8+CTxd+fexynJJkqRRL8k7AEmSJElqoonAzsCOlWmnyrQjMC3HuJYA91amu6se3wOsyjEuSZKkhjIBLUmSJGk0mQHsDxwA7FWZJuUa0eCsJRLRfwMWADcCdwHlPIOSJEkaKhPQkiRJkkay3YCDgdcD+wAb5RpNczwH/AX4HfAr4J/5hiNJkjRwJqAlSZIkjSTrAwcRSeeDgZcMY1vLgQeJvszVPZo7ezc/TVQkv1BZfylRibyc6P88gegHndDVzmM9YBzRX7qzl3Rtj+ltK68bqieIRPSvgGvp6j8tSZLUdkxAS5IkSWp304D3Ah8AXkskeAfjIaLP8j2VqbPf8qONC3HQXkLPvtQ7A9sMcjtriVYdPwAuIwY/lCRJkiRJkiT1YQJwKHARUXGcDXB6gUjIngfMIqqOR5LOCu8UuJKowh7ovq+qvOZIYHKL45YkSZIkSZKktrcT8F/Aswws6boM+DnwGeBlQKH1ITdVAuwCfBq4AniRgX0uzwFfr7xWkiRJkiRJksa0A4BLiJYS/SVXFwLnExXSE/MINkfjiM9qPnAz0ZO6v89rAfFZ2YJRkiRJkiRJ0pgxEfg4cBv9J1HvI9pS7JBHoG1se+BUoqd1f5/h7cAngUm5RCpJkiRJkiRJLVAg+jMvpP82EucTFb9W7/ZvJlEZvYi+P9dHgdlAMZ8wJUmSJEmSJKk53g78k74TpH8C3g+MzynGkW4ckeC/if4rot+RU4ySJEmSJEmS1DCvBG6g92ToGqIH9H55BThK7QP8iPh8++oRvXdeAUqSJEmSJEnSUE0GzqH3wQU7gPOArfIKcIx4KXAusIr6x2Ft5fl18wpQkiRJkiRJkgbjNcDd1E94lomK5+1yi25seinRV7u3CwIPAm/KLTpJkiRJkiRJ6sdU4FtEkrlekvNq4OW5RSeAXYGf0/vFge8A6+cWnSRJkiRJkiTVsQtwF71X174lv9BUx0HAQuofr3uAmfmFJkmSJEmSJEldDgWWUL+i9nxgvfxCUx/WAeZTvy3Hi8C/5BeaJEmSJEmSpLGuCKRAiZ4JzPuA1+cVmAblFcAt1K+GPh8Yn19okiRJkiRJksaiicAV1E9aXghMyi80DcFEon93veN5FR5PSZIkSZIkSS0ykfoD2a0Cjs4xLg3fkcAKeh7b64Ep+YUlSZIkSZIkaSyYDFxLzwTlY8B+OcalxtmTGDiy9hj/AZiaY1ySJEmSJEmSRrEpRBKyNjF5O7B5jnGp8TYF/kHPY30TJqElSZIkSZIkNVgCXELPhOStwPQc41LzbAD8mZ7H/FfEAJSSJEmSJEmS1BBz6ZmIvBnYKM+gBuk04M4+puuBQgPe54R+3ucvDXiPVlmfqHquPfZn5RmUJEmSJEmSpNHjMKBMz+Tz+nkGNQQX0TORWj2VgIOG+R7jgGf7eZ/Vw3yPVluPnpXQZeDwPIOSJEmSJEmSNPLtDiyje/JxEfCSPIMaov4S0KuBHwzzPQ4F1g7gfUaaGcRAk9X7sYIYsFCSJEmSJEmSBm0CPQeiWw28Js+ghqG/BHQGdADThvEePwfW9PMeIzEBDZFsXk73fbkTmJRnUJIkSZIkSZJGptPpmTz9aK4RDc9AE9CfGOL2N6L/5PNITkADfJCe+3NmrhFJkiRJkiRJGnG2BVbRPdF4Ua4RDV+9BPTKmvkyQx8k8GgigV29vXoJ6ZGcgAa4kJ5J+x1zjUiSJEmSJEnSiHIZ3ZOMjwMb5BrR8NVLQK+gZ9I4A3YawvbvrNlGqc52R0MCen169oP+Wa4RSZIkSZIkSRoxXk5UAlcnGI/INaLG6C0BXbtsFTB3kNverc52VtdZNhoS0ACH03O/XpVrRJIkSZIkSZJGhEvonlj8M5DkGlFj9JaAXkPPVhlPAoVBbPs8erYsWVvn/UZLAjoB/kj3/fpJrhFJkiRJkiRJantb0DNx+rZcI2qc3hLQV9KzXUYJOGiA250ALKFnkvmqOu83WhLQAAfT8zPbKteIJElS2xjMlXxJkiRJY8dHgWLV/D+AX+YUS6v8Eni+ZlkJ+PgAX38IMKVm2ThisL7R7FfArVXzBQb+mUmSpFHOBLQkSZKkemp7PX+LqG4dzUrA/xCDEXYaD7wHmDaA13+yzrLHgBuHH1rbO79mfjT0CpckSZIkSZLUBDvSvaVCB7BBrhE1Vm8tOD4FzKzzXAfwiX62uSk9W5asAk4GNqmzzdHUggNgKrCS7vs3M9eIJElSW7ACWpIkSVKtt9bM/56erSlGqzuAf9K92ns8MLuf132ISEBXmwD8b+NCa2svAL+rWXZwHoFIkqT2YgJakiRJUq19a+Z/kUsU+bkAWFM1nwCvAnbq4zWfBCZWzZeJxP0jDY+ufdX2CK89jyRJ0hhkAlqSJElSrVfWzP8plyjy80Mi6VytA/hgL+u/imhbUq1EJLLHkj/WzO+dSxSSJEmSJEnSKDQ57wAaZDzdexmXgXVyjajx+uoB3elyogq6ep2ngGKd7X2TSFBXr7ucrnNiLPSAhjh3qj+z0XjuSJKkQbICWpIkSWqMlxAVoAuBK4H5RN/gg4BtgXH5hTYoW9I9ybqIGFxurPkOPf9e2hg4sGbZJOAIot9zp9XAD4ik9liyBnisaj4Bts4nFEmS1C5Gyi/BkiRJUru7l0hOngt8Gjik5vnVwMNEgnoh8EDN43ZJVk6vmX88lyjydw3wLN0/jxLwceC6qmXvJJLQ1cYDFzY1uvb1ON2TztOBu/IJRZIktQMT0JIkSVLjrAL+Fbie6P87teq5CcAOlameRdRPTC8EFjcn3LrWrZlf1sL3biclIon8BboGFxwPvAeYBiypLPskPftFP8LY65vdqfZ8qT2fJEnSGGMCWpIkSWq8HxMJyB8D+wzwNZtXpgPqPNdBVJY+UGe6k8a2yBhfM7+mgdseaS4Ejq+z/DDg28AWRNV7dauODuB8ogfyWFTb13pC3bUkSdKYYQJakiRJao6HgdcBXwY+O8xtTST6SG9b57m1RMVtb609Xhzke9W2AhnLg8jdC/wdeDldVc7jid7e3waOJD7/6iTrBODiFsbYbmornpfnEoUkSWobJqAlSZKk5ukAPke05Pg2sH4T3mMcXcnpN9V5fjH123osJNp+1KptobBBwyIdmc4HzqMryZwArwJ2JtpvVCefy8SxfrSF8bWbaTXzY7WFiyRJkiRJktRSWwF/JFoztMu0ikhEX0skWo8nBtmrXueFZnwYObuInp/FCuBTddadRrSVqP3criMSztXL1wDvr7ONTeq8X0bPdhWjwXN038cZ+YYjSZIkSZIkjR0Tga/SM3HZ7tPFwAnALGBPela5jjSDSUADXELPJHRGVLhXzy+jfsuSsZKAnk7PRH2hz1dIkqRRzxYckiRJUut0AJ8HfkcMcDdSErlH1Fn2LN3bedwC/IxIro823wHeW7Mso3v7jdXA92nsgJAjzb4187cxOs8HSZI0CF6NliRJklrvp0Ql8V/yDmQYNiJ6Ie8DPE208RitycZrgWdqliU18+OJiwpj2QE18yP5/JYkSQ1iAlqSJEnKx4PAa4CvENW0I82fiJYcOxCD9L2YbzhNVSaqoFf1sc6DwJ9bE07bOrRm/ve5RCFJkiRJkiSpm3fSc/C2dpzWApcBr27Ox9Ayg+0BDZFo76139yrguD5eOxZ6QO9Kz30bKS1mJEmSJEmSpFFtA+A9wP3kn2Tubbod2KlZH0CLDSUBDfBX6iehS8CMPl43FhLQ59F9367JNxxJktQuHIRQkiRJap3JwO7AbkTF6K6Vx9PzDGqAXgoszjuInH0VmE/P/s83AE+0Ppy2sT7w4ZplY70ftiRJkiRJktRU04ADgX8HLgbuIFpY5F3JPJDpbiKhuKhm+SkN/YTyM9QK6KEa7RXQJ9N9v54CJuYakSRJkiRJkjSKbAAcABxNJDfvoPd+we06lYGrgDdU7dfxNeu8AGw27E8rfyagG2dD4Fm679eJuUYkSZIkSZIkjWAzgEOBFLiSaL2Qd/J4ONMK4Hxg5zr7uh49k4sXDOlTay8moBvnG3Tfp+eJlhySJEmSJEmS+rE98AHgHOC3RHIt74Rxo6bFRD/jvgbPA/hCzevKwJsG+Pm1KxPQjfFqeraV+fdcI5IkSZIkSZLa1FSijcbxRGXzU+SfJG7GdB/RKmTyAD+X8URP6OptPED0uB6pTEAP33rEuVS9P/cCE/IMSpIkSZIkSWoH44GZwGzap2fz8zS3wnoB0TokGcLnVa/S9RqgOIRttYN6CejVmIAeqAJwBd33ZTRUxkuSJEmSJEmDlgC7AB8G/hv4K5H4yyvR3AHcDFxItCt4M9EG4/NNeq9LgL2H/SnCuXW2f3YDtpuHc4je3dXTY8DhTXq/jYCH67znfU16v2Y7g57nwn/nGpEkSZIkSZLUItOBdwDzgGuBJeSXbH4B+APwVeCjwB5E9XWtVwKrGvi+S4HzgJcO4fPrzXjg93Xe62MNfA+1v/fS826BWxh4SxdJkiRJkiRpRJkBzCISrjeTXyuNJUSbi/OAI4kWH4UBxD8NWNigGBYSPazXH9AnN3ibA4/WvOdK4O1Nej+1l4OJXtnVx/8JYIs8g5IkSZIkSZIaZRywFzGI3iXkN1DgE8RAhfPpSjYPpbcywGUNiOfmShzjhhjDYLwCWF7z/h3Ae1rw3srP24mLDbUXH/bNMyhJkiRJkiRpOKYBbyN6zt5Az+rLVkyPEEni44lB1jZu4P4dNYy4SkQS/NUNjGegPkDPSvM1wPtziEXNdxg9+6aXgQ/lGZQkSZIkSZI0WLXtNEq0Ntn8It3baGzTxH19OT0rSgca4/nAjk2MbSA+Qc/jsxb4dJ5BqeE+QVxcqL344XGWJEmSJElS29sRmA38AHiM1iab1xCDp30d+AjwMgbWs7kR1gfuH2S8jwDH0rz+zkPxfnomJzPgImCdHOPS8E0kLsTUHtu1OPCkJEmSJEmS2lRnhfP5wEO0NuHc2bc5BQ4CJjd1T/v2AwYe961ENfb4XCLt3/vo2Z4hI5L7zawgV/NsAfyR+snnI3OMS5IkSZIkSepmG+CjREXsI7Qu2fwC8GvgdOAQYJNm7+ggfI7+4y8B1wKH5hTjYL2D+Mxr92Mx8NYc49LgvZn6g3suA96dY1ySJEmSJEkSm9NV4fwAra9uPh44AJjQ7B0doj2BVfS+H8uBb5B/f+eh2BG4g/r7dQmNHbxRjTeN+N7WDi6ZAfcBu+UXmiRJkiRJksaqzehKOPeWfGzGtJCoqp4NzGz6XjZGX32fFwFfZOQnadcDLqP+Pj6F7Rva1SHAo9Q/blcBG+QXmiRJkiRJksaSdYn+yfOBm2lNsnk5sIAYEG0WsFHT97I5LqHnvt1GDOg2Mce4Gi0Bvkb9StoM+CmwfW7Rqdo29H7BYC0whziekiRJkiRJUlMUgb2I1hbXAh00P+H8JN3baYyG5Oxn6b6PC4j+zqMxuXcksIK+j/Fqomp+85xiHOs2Ji4i9dYO5jZgn9yikyRJkiRJ0qi2LfAp4HLgeZqfcF4E/IBop7FTC/av1XYnErIdRNuQ0dpLdxyR1Kw+tvcAX6b3ROfyymum5RDvWDSFuLCzlN4vDMxndFz0kSRJkiRJUpuYQmvbajxFV4XzXozOKuBOU4A/E+1DXpJzLM20BXATPVttTK08vwtwI72fE88CZwAzWhr12LE5MA94ht6PwZ+AXfMKUJIkSZIkSaNHbVuN1TQ34byYsZNwrrUNMDnvIJrstUQVe+fxXkMc69rjnBA9vO+l93NlNdEre99WBD4G7EG0OllJ75/5w8TdB8WcYpQkSZIkSdIosBkx2N1l9H77faOmZ4j2HUcRFZVjKeE8liTA0XS/gLEYeGM/rxsPfIbuSet60w3ABxj9CfxGWwd4P/A7+r8T4XPAhHzClCRJkiRJ0khWIAYRO51oq1GmeQnn54CfEcnI3SvvrdFtClGpXH0eLGBwLTTWBb5ItN/o6/x6keid/Ras0u1NEXgT8F3gBfr/vp5KHENJkiRJkiRpwNYFDiVuuX+c5iWc1xJJ7flE7+jxrdg5tY0dgdvpfk6cz9AraScDnwbupv9zbxHwFeD1eN6NA14H/CfwBP1/dvcAnyV+TkiSJEmSJEkDsi1ReXwt0EHzks4LiSTjLGBaS/ZM7egdwBK6zouVwEcbtO2EuKBxJQOr2F9GnPdHA1s2KIZ2twlwJFF9/jwD++4uIL63Vo9LkiRJkiSpX+sQSbrzgIdoXsL5GSLJNRvYugX7pfZWJCreqxPD9xEtV5phJnA28CgDP2dvA74KfBDYoUlxtdr2wBHE9/0fDPyzeAw4h+jBLkmSJEmSJPXppcSgbb8kKk6bkXDuIAYtOwl4FfZxVpeNiUrj6vPlKmCDFrx3ATgQ+A7dK68HMj0L/ILod/w24m6Bdq0CLgDbEHGeClxNXAQazP4uBS4kBoH0+ytJktqGI5JLkiS1p92AdwLvAvakOb+33QX8mkguXg8sb8J7aGTbC7gc2KoynxGVyScR1dCtNAl4O9Hn/C3AZkPYRgdRuX1v1XQf8DTRX/qFhkRa33rEII3Tge2AnYh+2p3TxCFs8yngGuKCwFXEBSpJkqS2YgJakiSpPRSB/YBDiKTzTk14j5XAjcB1wM+IQcmk3swGvkbX4ILPEu0grsktoi4J8Arg4Mq0HzEo33CtAhYTg/o9XXlcAl4kBt9cQSSxVxMXbCYTieOJlcdFYGrl302IZPPmlcfrNCC+EvAn4m6IXwG3EBcFJEmSJEmSpB7WIao5zyeqL5vRWuNBugYPXK81u6URbhLwbbqfR7cQLSLa1TTiuzSXqOgfbLuOdp2WEheMziAGgGxF2xNJkiRJkiSNYBsBRxID/L1I4xNWa4GbgZRon+AdbxqMLYG/0P2cuoio7h1JCsAuwEeAbwC/J6qZ804o9zUtBv4AfBP4KDEQo72cJUmSJEmS1K9tgKOJysw1ND5x9TSR0D6SqASVhuJtwHN0nVeriPN2NNkA2If4rpwBXArcBDxAtNRoZoJ5BfBY1fwDRIJ8X2DDJu6zJElSrqyIkSRJao7dgMMq08savO0yUeV8NfAL4G9EQksaigQ4DjiTrorbx4i2LX/KK6icTCEGN9yU6N88nfh81iN6THf2fJ4ArEtXT+iOyuMSXQMZLqarj/QiYBnRG/pFov3OY8BLW7BPkiRJkiRJGiX2AOYBd9P46snlwE+JW/M3adUOadSbSpxX1efa9UQCVs3xD+JzLhOfvyRJkiRJktSrmUS/5WYknZ8h+u/OIiozpUbaA1hI1/lWBs4jKn3VPD+i6zPfO+dYJEmSJEmS1IY6k8730Pik8wNEEvAgTASqeY6ge8/jF4h2MWq+U+n63D+ccyySJEmSJElqE51J5/tofNL5jsq292rRvmjsGgfMp/v5dzeN71Ou3r2Prs9+fs6xSJIkSZIkKScFIiGcAvfT2ITzGmABcDSwRYv2R9oCuInu5+JPsQ9xq+1G1+d/Rc6xSJIkSZIkqcX2Bv4TeIzGJp2fBy4m+jmv17K9kcJrgUV0vwhyPJDkGdQYNZH4/DPg3pxjkSRJkiRJUgvsQnN6Oj9HDCJ4KDChVTsjVUmISvvVdJ2Xi4E35hmUuJc4FmuBSTnHIkmSJEmSpCZ4KZGYW0Bjk87PYtJZ7WEKcAndz88FwIw8gxIQrTc6j8luOcciSZIkSZKkBtkQOBK4FijTuKTzM3Qlnce3bG+k3u0I3E738/R8vCjSLqoHgnxfzrFIkiRJkiRpGNYnks5X0r0NwXCnpzHprPb0DmAJXefqSuCjuUakWh+m6/icmnMskiRJkiRJGqRJRGL4ImAZzUk6j2vZ3kgDUyQqa6ur++8Dds8zKNW1N13H6Ec5xyJJkiRJkqQBKBADq30PWErjks6PAucC+1feQ2pHGxOtZarP3auADfIMSr1aj64LBf/IORZJkiRJkiT1YWcgBR6gcUnn5+iqdC62bE+kodkLeIiu87dMVEJ7waS9PUpXixR/zkiSJEmSJLWRDYDZwAIal3ReAVxCJJ0dqE0jxWygg+4DYr4l14g0UL+m67htn3MskiRJkiRJY95EIjl8Cd0TbsOZVhGDEx4JTGndrkjDNgn4Nt3P51uAbfIMSoNyHl3H7tCcY5EkSZIkSRqz9iISNYtpTNK5RFROH030zZVGmi2Bv9D9vL4ImJxnUBq0T9N1/I7LORZJkiRJkqQx5SXA8cA9NK7Fxs1E0nmzFu6H1GhvBZ6lexX/0blGpKF6HV3H8X9yjkWSJEmSJGnUmwp8DLieGEStEUnnfwAnAFu3bC+k5kiIizIlus7vR4F98wxKwzKdrmP5p5xjkSRJkiRJGrX2As4HXqQxSecniJYde7ZyJ6Qmmgr8lO7n+fXApjnGpMZ4mjieS4mLDJIkSZIkSWqAzYi2Af+kMUnnlcTghIcC41q4H1Kz7QEspOtcLxMXWDzPR4c/0HVst8g5FkmSJEmSpBGtCBxEJIpX07jBBGcTFaLSaHMEsJyuc/4F4LBcI1KjXUDX8T0o51gkSZIkSZJGpB2BFHiExlQ7PwTMB7Zr3S5ILTWOOMerz/u7gZflGZSa4hi6jvFROcciSZIkSZI0YkwCZgHX0pgBBZ8HLiIqBO2TqtFsC+Amup//P8Uq/9HqYLqO89dzjkWSJEmSJKntdQ4o+ALDTzqvJRLYRwLrtHInpJy8FlhE13dgDXA8XnQZzbak63j/LudYJEmSJEmS2tLGwLHAnTSmxcYtxK3oG7dyJ6SczaZ7b/TFwBtzjUitkAAvEsf8yZxjkSRJkiRJaiud1c4rGH7SeUllWwe0dA+k/E0hBuas/j4sAGbkGZRa6ma6jv1GOcciSZIkSZKUq/WJSs1/MPykc4lItM0GJrdyJ6Q2sSNwO92/F+cDE/IMSi33v3Qd//1zjkWSJEmSJCkXndXOyxh+4vlxYD6wXUv3QGov7yAq/zu/FyuBj+YakfJyEl3nwSdyjkWSJEmSJKllphLVybcy/KRzB3AlMAsY18qdkNpMkbgAU6br+3EfsHueQSlX76LrXPhyzrFIkiRJkiQ1XWe1c+fAWMOZ7gSOB6a3dA+k9rQxcC3dvyNXARvkGZRytxNd58Mvco5FkiRJkiSpKSYR1ckLGH7SeSlwEXBQS/dAam97AQ/R9T0pE5XQhRxjUnsYB6wizouH8g1FkiRJkiSpsXYFvk4kjYebeP4j8BEcUFCqNZtoQ9P5XXkGODjXiNRu7qDrwsSUnGORJEmSJEkalgJwKNEKoLoP7VCmF4h2Ha9o6R5II8Mk4Nt0/87cAmyTZ1BqS5fRdY7smXMskiRJkiRJQ7IJ0Y/5IYZf7XwzUdVppZ5U35bAX+j+vbkI7xBQfafTdZ4ckXMskiRJkiRJg9I5qOAKhpd0Xglcgr2dpf68FXiWru/OKuDoXCNSuzucrvNlXs6xSJIkSZIk9WsijRtU8C6icnrDlu6BNPIkxHelRNf351Fg3zyD0oiwB13nzOU5xyJJkiRJktSrGUAKLGZ4SedVWO0sDcZU4Kd0/x5dD2yaY0waOdah68LFXTnHIkmSJEmS1MMBRMJ4DcNLPN9DVHBu3NrwpRHt5cD9dH2PysB5wLg8g9KI8wBx/qwGJuQciyRJkiRJElOAzwB3Mryk8xrgMuDA1oYvjQpHAMvp+j69AByWa0Qaqa6m6zzaJedYJEmSJEnSGLY50WajepCzoUxPAfOBrVoavTQ6jCO+P9XfqbuBl+UZlEa0/6DrXHpPzrFIkiRJkqQxaC/gIobfZuNmYDbRc1TS4G0B3ET379XPgPXzDEoj3sfoOp9OzjkWSZIkSZI0RhSAQ4Ebacyggvu3Nnxp1HktsIjuLWyOB5I8g9KosB9d59XFOcciSZIkSZJGuanA0cDDDC/x/DjRrmN6S6OXRqfZxABxnd+vxcAbc41Io8k0us6tW3KORZIkSZIkjVLbA+cByxhe4nkBMIvoUytpeKYQdxBUf8f+CmyZZ1AalTqr65cTd8BIkiSNeP5RKklSezgA+Dwx8FRxiNt4Efgh8DXg9gbFJY11OwI/AWZWLbsAOIqohpYaplgs3jV+/PjNisXi5P333/+AV7/61Y8Bk0ql0jqV59cFJlRWn1oul4sAWZZtUGdz5SRJllYvKBQKS4FyqVR6EVhbLBaXA6s7OjpeXLVq1bJzzz13ZdN2TpIkjVn2qpMkKT+TgA8SrTZ2HcZ27ge+CnyXSEJLaox3EAN/dg4uuAr4DPA/uUWktjdr1qzizJkzNy2VSpsmSTKNaK0xLcuyDarnq6YNqh5PySvuihLwArCUuBPnRWBJkiTPZln2LPBslmXPJknyLPBMoVB4GlicpulTROW2JElSDyagJUlqvenAZ4lE1nB6M/+OaNdxJVBuQFxqP5OJc2RD4nb8ycBEYDyRqEqICxkr6UocASyh6zb+54lexSaHBq4InAEcR9fvy/cD7wX+mVdQyt+JJ544fdy4cZsVi8WXlsvlTbMse2mSJJsBLwE2B2YAmzL0O1lGqrXEz5kngEVJkizKsmxRlmWPFIvFR9auXfvw0qVLH/na177WkXOckiQpByagJUlqnW2JaudPEInEoVgNXAF8Gfhzg+JSPtYj2jt0TptVpk0q0+bAug16rxLwNJEgehJ4qjL/IHBvZXoEL2QAbEy0sjmoatnVwIeIZL5GsZNOOmnzCRMmbFcqlbYvFArbHh5dtAAAIABJREFUZVm2HdHrewviOzmxSW/9ArCko6Nj7ZNPPrltqVRi8uTJT2y66aY3JUmSJUmSEReWyLLs/y42JUmyJsuyZf1tPMuypFJ9TZIk6xMXtKZkWTY+y7J1gMmV5esTF7c6p0ZbRAyu+3CSJA9mWXZ/lmX3F4vF+9I0faIJ7ydJktqACWhJkppvT+ALwOEMffyFp4ELgf8CHmtQXGqNScQ58CpgFyLZvBNRKdlOVtGVjL4X+DvwJ+DRPINqsb2Ay4GtKvMZcDZwEibnR4VKe4yXAtuXy+XtgO2IwV87Hw/3os+LwGNZli0qFApPZln2PLAky7IlwJJisbgEWFIqlZ4vFovPE0nlJWmadp5fmxNVxAA3AfsPM57hSE444YRpxWJxaqFQ2DhJko2zLNsoSZKNgI2ADZMkmZ5lWWfV90sY3ue3HLgvy7L7kyS5D7gjy7I7ly1bdre9qSVJGtlMQEuS1BwF4O3EwIIH9bNuX+4Fvg58C1jRgLjUfDOIpNEBREJzLyIJPVItAv5WNS1g5FUCvx64vp91ZhMDeHYO8PYccATwq6ZFpaZJ03QCsHO5XJ4J7Eb02d8J2JquYzwYHURiuHp6PMuyRcVi8THgyWXLlj16zjnnLG9A+M8RfaGXVP4dMY499th1p0yZskWpVNqUqBrfMkmSLbMs2ypJkq2JizvrDXKzJeDBLMtuT5LkrizLbi8Wi/8E7k7TdG1Dd0CSJDWFCWhJkhprAvB+4HjgZUPcRgb8hhhY8Crs3dvuNgTeBBxcmTZr4LY7iLYZzxAVuCsqy9YQA4RlROXyOkTP2amV13XeYr8u0UN6owbGtJaojP4lkZy9lfY+Rw8kejd/rpfnJxF3Fny8atnfK695oLmhabjSNC0A25bL5d2yLJtZKBR2y7JsV2AHolf6YCwC7k+SZGGlNcRCYGGxWHwwTdNnGh17H24C9qs83oxomTNqnHDCCRtMmjRpK2D7SquTHbIs2544ZpsPYlOrgNuSJLm1XC7fCtxaLBZvS9PUi7WSJLUZE9CSJDXGVOCjwP8jqr6GYhVwKTAfuLNBcak5Xgm8tTLtzdAGHFtD9GC+pzI9QiSanqSrV/OSRgRLXBiZTlef6enEebodsDPRFmSoSeqngGuIZPQvaVzMjbAJ8A/gK8CX6jy/JXAZ0R6l08XAp/COg7ZzwgknbDBhwoRXZln28kKhMLOSaH4ZA++pXwIeybJsYaFQuL9cLi/sTDYXi8WFbZS4/A7wscrjA+m/en/USNN0CtEeZackSXbNsmwXooJ9WwbWwqoE3A38BfhzlmV/KhaLd1gpLUlSvkxAS5I0PNsQ/Z0/ztB7Xz5O3Pp/ASOvtcFYsgPRkuEIomfsYDxAVA3fSlfC+UEiCd0uNiIS0TsDM4F9iN7VgxkwcxUxYN/FRDK6o8ExDkYC/IKoSj8c+FHN828l4tywMt9B3LlwXqsCVO/SNJ1aLpf3Ii72dP673QBfnhHfr9uSJLmjXC7fBtxRLBbvSdN0dXMibqhjgf+oPP4s0YZpTDvqqKMmTps2beckSV4G7JZl2cuTJHkFA6uYXg78LUmSvyRJ8mfgJgc8lCSptUxAS5I0NHsR1c7vZegDC94GnEMkxkZCUmQsmk60VDmCSMgOxItEsvnPlekvREXzSDSOqD7cl9j/fYgE9UA8T1T0X0z0jW51m45/J75fAK8G/lh5nADHAWcSbUogBvacRRw3tViaplNKpdIrkiSpTjbvyMD+VlkE3F6Z7siy7J8rV668s0G9mPPydqL9EkR7mKNyjKWtpWm6GbBHuVx+BdA5bUf/587CJEn+UC6X/1AsFhekaXpvs2OVJGksMwEtSdLgHEBUSb6dof8/eiPRDsD+zu3rFcCngSMZ2ACCDxDH80rg94zuCwqbAm8BDiF6X08bwGvuJxJp3yaqEZttTyKZ3NkDeAti0LipwPeAd1WtewPwL4yyPrvtatasWcWZM2fuViqVDigUCq/Ksmwv4qJGf21s1hKtif6WZdktxWLx9o6OjtvOOuusZ5sedOttCyysPP4NwxvIdsyptGrZO8uyfYB9kiTZG9i4n5c9mWXZAuAPwG/mzp17R9MDlSRpDDEBLUlS/xIi2XYiXQNDDVaZaAcwj6iKVfspAu8kWqq8pp91VxHH82qi9/FYvZ17HLA/0ebiMPpvTfIc8C0iGf1Yk2JalxhEcBvimK4hLiLsBlxOVxuHjGh98+9EclNNkKbplHK5vE+WZfsD+ydJsh+wXj8vKxN9fP+WZdnNxWLxZuDvbdSjudkKxJ0Uk4mfLUMdV0AVaZpuXy6X90mSZN8sy15D/Dwo9PGSJ4nk/29KpdJ1Z5xxxqMtCVSSpFHKBLQkSb0rEC02TiV64g7FMuAHRCuA+xoUlxprAvBJIhG5TR/rlYnq5ouJRGY7DbbXLvYFPkC0LZnex3priMH/5gJ3NTiG7xKV652/5z4IzCF6rHf2s36RGOTtsga/95iXpumMcrm8f5IkB1SSzi+n7zZFGfGz8f+SzStWrLj17LPPfrElAbevW4E9Ko83wJ83DXX88cevv8466+yfZdlrKgnpVwIT+3jJvURC+tpVq1Zd5/kpSdLgmICWJKmnicQt+V8kBp4bisXAN4CvElWfaj8JkSidS9+Dm90D/A9xIcEquIEZB7wZ+BDwHiLJX89aImF8Go2piH4f8OOq+TJxzLaqWnZPJaY7G/B+Y12SpunMSjuN/SsJ574u4gAsBW7KsuymYrF408qVK//2pS99aWkLYh1pfkAMnglx5439yZsoTdNJwN7lcvlA4I3ExbTxvay+huhr/8tCofDLNE1vb1GYkiSNWCagJUnqsh5RFXkcMGOI2+jsdXsBsLJBcanxDiL6cO/Zxzo3AucBPwFKrQhqlNoU+Ffgs/Teh3U1kYj+IvD0EN9nW+AfRJVz5631Gd1/370C+DCRBNUQpGm6Y6lUekOhUDgwy7LXA5v085KHiWTdjVmWLSgWi3ekaVpueqAj3xzg9MrjjxEXwdQiaZpOAV6bZdkbsyx7I7A7vf/t/AjwS+CXK1asuG6ED4ApSVJTmICWJClaBXwW+Dxxq/NQmKwcGbYlKtPf3Mvzq4DvE8fytlYFNUasQ1REHw28rJd1ngdOBr7J4AboHA/cRAweWW8wuwz4I/BTIiHaOTnwYD9OPvnkrYrF4huAA4E30Hc/4hJxEeBG4MZSqbTgjDPOeLwFYY5GhwGXVh7/B3FhVDlJ03STyoWXt2ZZ9hbiwlo9K4HrgCsKhcKVaZoubl2UkiS1LxPQkqSxbCvgWODjRHJssMrAz4H5OLBguysSic+5dPUBrraGGBxvLjH4lJonIVpgnAHs1Ms6vyf6ct87wG3OB47v4/nOitvaQcc6iB7R11fiadbAiCPGSSedtHmxWDwwSZLOpPO2fay+ikjs31AoFG4E/pSm6bJWxDkGzAQ6WztcBRyaYyyqkqZpAdizVCodnCTJ24C9qX/hq0x8P64ol8s/mzdvnuNASJLGLBPQkqSxaBfgBKK/Zm89HvuyBvgh0cLBPrLtb1fg28A+dZ7LiIHoTsZBIlutc5DPs4Gt6zy/iviOnUm06OjNQcCvGfzvtSXgaqIi/td0JanHlOOOO269SZMmvQF4E1HhvEsfq68F/pIkyW+TJPnd0qVL/3juuefaaqg5JgDLiX7qC4Ht8w1HvTnxxBM3Gjdu3JuSJDkEeBu930l1Z5Ikl5XL5Uvmzp17RwtDlCQpdyagJUljya7EbcwfoH61Un86gEuIKlmTle0vIS40pNQfBO+3xPnwtxbGpJ7WIdrfnARMrfP834jBIu+v89x0okp0Y3pWN9cqEd/7xUQ/3W8QbTjGnDlz5sxMkuSQLMsOSpLkNcTAq/WUgbuTJFmQZdl1hULh2jRNl7Qw1LHubuIugTIwBccVaHuzZs0qzpw5c79yuXwIcYGttwsHDyRJclWSJJemabqghSFKkpQLE9CSpLFgd6LVxhH0n6Sq50UiYfUl4IkGxqXmWY84Zu+t89wSol3Dtxhcn2E11+bAfwPvrvPcC8BHiB7OnRKiBc7b6f132urj+1vgfOBnxF0MY8aJJ544vVKheTDR/7y3/rUZcEdnhfOqVatumD9//vOti1Q1fgq8q/J4D6K/tkaOZM6cOXsmSfJO4jju1st6dwKXlMvlS+bNm3dX68KTJKl1TEBLkkazPYiqysMY2v95TwNfJwakMwkzcuxEDAZZb6C7q4BPAw6M1r5mEYno6TXLM6Jdx0lERei/A+f0so3OauelwPeI7/ADzQi2HaVpOq5UKu2XJMlbgLcAe9L7xbcngWuAawqFwm8cNK2tnAmcWHl8OPCjHGPRMKVpumO5XJ4FvI+4MF7PrVmWXVwsFn+UpqkXvCVJo4YJaEnSaPRq4o/2vioj+/IwcC5RIbuigXGp+d4LfJe4Xb3a08Cn6F5Bq/Y1nUhCz6rz3NXEoIM30DOpWq4suxX4JvC/jJG2BWmaziiXy2/PsuzgJEneCKzfy6qrgRuJhPM1aZr+A+8EaFcfAi6qPD4dODXHWNRAX/ziF3dKkmRWkiTvo35ldIm4a+P7q1at+snZZ5/9YmsjlCSpsUxAS5JGkwOI1gqHDPH19wP/QbRuGFO36I8SRxDVrrX9vW8hEtMPtTogDdts4L/oOVhoB9HXu/p32ReAC4k2G3e3JLp8JWma7lXpNXsIUeXc2+/2C5MkuSZJkmuA36ZpuqxlUWo4Xgn8tfL4UqJyVqNMmqY7l0ql9yVJ8gHiDp5aK4nWQd+/6667fnXppZeWWhuhJEnDZwJakjQaHEAMNPfGIb7+70TF8/eJqiONPLOJQeVqK2L/l6h8HhNVsKPUAcTgn5v38vy9wFnAjxnlx/mYY45ZZ/31198/y7JDsyx7L7BFL6uuyLLspiRJrsuy7Lq5c+c60ObINIW4sJIQg2321kNYo0RlgNAPAR8GNquzyqIkSS4tl8vfnjt37m0tDk+SpCEzAS1JGqkS4G3AHGCfIW7jD8Bc4NpGBaVcHAN8me6/16wBPgdckEtEarQtgMvp+V1fSCSon2x5RC2SpunWldYahyRJ8npgUi+r3p8kyVVZll1VKBT+kKbp6haGqeZ5GNiSaJ2yLrA233DUCrNmzSrOnDnzTVmWHZFl2buJY19rAXBhoVC41LsaJEntzgS0JGmkSYB3EonnPYe4jd8QiecbGhWUcnM08JWaZR1E7+ArWx+OmmhXonfx1Jrl9xB9359reURNkKZpAdi3UuX8dnqvel2bZdkC4Oosy66cN2/ePa2LUi30K2IgSYj2DPfmGItycOyxx667zjrrvBv4aOUiVO2dPsuSJLmkXC5fOHfu3BtbH6EkSf0zAS1JGkkOIm61f+UQX38jkbj+XcMiUp7eBPyS7j2fVwDvBn6dS0SDtzs9+xtXexJ4vAHvswM9E7fVlhI90NvZNKLFxiXAO2qe+wPRgmdE9m4/6qijJm600UavqSSdDwNm9LLqc8Bvsiy7as2aNVfOnz//+RaGqXycC3yh8vhdwBU5xqKcpWn6knK5fATRdmrbOqvcA/xPoVD4Vpqmo+KinCRpdDABLUkaCQ4CzgReNcTXXwd8EfhzwyJS3nYkjue0qmXLier43+QS0dA8BWzSx/N/BvYd5ntMBBbTdwL6V8Bbh/k+rTIB+BFxoaHaV4mK+BHh2GOPXXfy5MlvIKr130nvx+eBJEmuSpLkyieeeOKGCy64YEQm2TVks4mBNQFOBObnGIvaROVOiVdnWfahLMs+CEyuWWUVcGWWZedZFS1JagcmoCVJ7ewg4Axg7yG8NgOuBk4Dbm5kUMrd+sAfgV2qlpWInuAjpfK50zPARn08nxHJ9uFUJ7+PGGBzXB/rXAu8eRjv0WrjiZhfV7P8k8C3Wx/OwKRpOqNUKr0DeHflVvoJdVZbS7QHuqJcLl81b968B1sZo9rOa4DfVx5fRAxOJ/2fE044YYMJEyZ8gBhwt17Lnr9mWfbNYrH4ozRNV7Q4PEmSABPQkqT2dADRo/n1Q3htGfgFcApwawNjUvu4HHhPzbJ/I25VH2n6S0B3AGcT5/NQXQscSPdWJfXWGUkJaIDpwF+ArauWrQb2A27JI6B60jTduVQqvStJkncRF9Pq/f69nKhCv6JQKFztrfOqshHxcwLgrwztgqzGiDRNX10ulz9N3FlRO2DpEuA7hULh62maPtD66CRJY5kJaElSOzkAOJ1Ilg1WmUhMngLc3cig1FY+RFQBVvtf4MgcYmmE/hLQEO0zZhBV3oM1A3iUnoNW1RqJCWiAlxHV8NXtK+4E9iJuQc/FnDlz9kqS5L1Em5Cde1ntmSRJfp4kyRVLly699txzz13ZwhA1siwmLrgsI871rOb5pM4yjWEnnnjiRuPHj/8I0cJlx5qny8QgvV87/fTTf4vnjiSpBUxAS5Lawf5Eq4w3DuG1a4h+sPOAexsZlNrOxsQASxtWLfsrcYt6Ry4RDV+9BHRG99/RykRyeCi9rY8nvlsT+9g+jNwENMBhwKU1y04h7qJolWTOnDn7VpLO76V7VXa1B4GfZVl2xd13373g0ksvHcpFBY09NwCvrTzekrioVO1DxIU4qVZyyimnvCHLsk9X7sKobcV0B/C1FStWXHzOOecszyE+SdIYYQJakpSn1xDJsaFUPK8Bvkv0iH64gTGpfX0d+Neq+VVEpeud+YTTEAOpgF4DXAZ8YAjbfwDYpmq+TFRSj69ZbyQnoCGq4j9UNb+SqDx+pFlvOGvWrOLOO+98QFWl80t6WfVW4Iosy342d+7cfzQrHo1q3yT6+wK8he697t9CVLm+t9VBaWQ5+eSTXzpu3LjPZFn2SXr+v/N8kiTfSZLka2maNu3npiRp7DIBLUnKw37AScAhQ3htZ6uNkxjewGwaWXYCbqd79dbxRH/kkay3BHQH3auWVwObAEsHse39gJtqlq0lEtATa5aP9AT0BkTrnU2qln0P+Egj3yRN03HA68vl8mHAu4BN66yWAX8CLi+Xyz9xEEENwnRge6KtTLWjga9UHn8BOK/yeDPgn8APK+tI/TrmmGPWmTJlyhFJkhwF7F7z9BrgkizLvjx37lzH0ZAkNYwJaElSK+1K3Bo/awiv7Uw8fxFbbYxF36N7n+eFwExGbuuNTgOpgIZIQH8G+M4gtv0t4jObULO8NrkNIz8BDVEFen7VfImogh7WhapZs2YVZ86cuV+WZbOyLPsX6iedy0TS8NJCoXB5mqaPDec9NaZdRVzAOIuuvu9vBq6pPD4f+DTxd9yvKs/9O/CfrQ1To0GlX/3RwOH0bM9xI/Cl008//SrsEy1JGiYT0JKkVtiZqFg+gv4HQ6vVmXieQ/T/1dizFZFErP7j+H307Ps7EvWWgH62ZnkG3AzsPcDtrlPZ9uSqZauBJcD6jM4EdBG4Ddilatk3iMT9oKRpOgF4c7lcfh/wDuIzq7U2y7LrgctLpdJPzzzzzKcGH7LUw4HAb4E/EP9nPkq0d+ns+/x74HVE0vmcyrJZRJseaUjSNN06y7Kjsyz7BDCl5unbgC8XCoUfpmm6OofwJEmjgAloSVIzbQmcDHyMnpU1/elMPJ9C3Fqvset04gJEp7uIavpyPuE0VG8J6K8QidPa6uWdGdiFmCOIHum137tzK9sdjQloiIrv71XNLwNmAC/298JK0vmgStL5ncC0OqutzrLsOuDytWvXXnHWWWc924CYpVp/AV5FtNz5GPBT4uLRVOJnxsFExX1nL/e9iQFZpWE54YQTNhg/fvynkiT5PLB5zdOPZ1n2lWKx+M00TZflEZ8kaeQyAS1JaobpRHXWF+iZ6OpPBlxNJBz/3uC4NPIUgIeAl1Ytm020lxgNektA7030b65OIHcQPa9PGcB2rycG+ay+4+B+4FTgQkZvAnocMfBgdeLkE/TSumT27NnjZ8yY8cZK0vldRC/pWiuzLLsGuHz16tVXfulLXxpMH25pKN4N/KRq/gJiwNW9KvMPEhd4i5X5zQAr8NUwaZpOKJfLRwD/RlzwrfYc8NVCofC1NE2fa310kqSRyAS0JKmRNgL+H/B5ogXAYHQmnk8BHPhGnV5N9KHstJyoaH0hn3AarrcE9PbE7fWH0D0JvZjY/1Kd13R6CfAw3ZPPHUT/9CcY3QlogDOBE6vmfwm8rXOmMpDgG6qSzvU+/1XANVmWXdLR0XHl2Wef3W8FtdRACXAnsCPxPc6IKv6pddbtIP6/tUevmiE55ZRTDs6y7NgkSd5Q89yyJEm+uWbNmv8888wzF+USnSRpxDABLUlqhCnAZ4mkT71eqf25DjgB+Fsjg9KocBZxbnT6MfD+nGJphr4S0DsDP6d7IrlMJIp/08c2v1iZqpPMZSIxfSCjPwE9E7i9an7VhhtuuOnnP//53fsZSLAjy7JrgUtXr159hZXOytlHie9qp4z6f7stJH5eSE2Vpuke5XL534AP0FV9DzG+wI8LhcK8NE0dJFqSVJcJaEnScEwGPgccR/0kWn9+QbTauKWRQWlU+TPdB947HPhRTrE0Q18J6IeBJ2ueX0P0Rj+8l+0llddVtywpEQnmtxKJg9GegAa4F9ihc+a44467b9KkSTvUWW81cG2WZZeYdFabmUC02tiM3gfvLRMXo0bTd1dtbs6cOTMLhcIJWZa9n+536KxNkuQHpVJp3rx58+7LKz5JUnsyAS1JGorxRHXWqUQ7gMG6iRic8PoGxqTRZyIxCFd1snS09TrtKwG9kOj5/Hm6fwariQreJXVe9xrgBrr/jreWqBq/nLGTgL4A+GTnzOGHH75ghx12OKAyWwL+BFy6Zs2aH5x11llP5xGgNADHAP/Zx/MlYrDRT7QkGqlKmqZbZ1l2TJZln6R727UycHm5XJ4zb968gQyaK0kaA0xAS5IGowB8CEiBrYfw+r8SrQF+3biQNIrtQfd+4A8ztPOunfWXgH4ZcEfNc6uJljffrvO67xLV0ROqlr0AbEL0ih0rCeiPU/X5bLvttr/74Ac/uDrLskuKxeLPHDhLI8S6wKPANHr/u+0UYG7LIpJqpGk6o9Ka41NES7ZOpSRJvm9FtCQJTEBLkgbuIOA/iKTgYN1JJK0vw4GSNHDvJc6ZTr8A3p5TLM3SXwIa4O/A7nT93pYBN9O9NQlEsuppuleirQbOJ6qoYewkoPcF/lg1/3fgFTnFIg1HStxt1JsPAxe1JhSpdyeeeOJG48ePPwr4At3HA7EiWpLUaz8xSZI67UO0yriWwSefHyIqYnYHLsXkswZny5r5h/IIog1cQPR+7pQArwJ2qlnvMKI9TrUJRFX0WPNQzfzWOcQgNcJXgRV9PP9IqwKR+nLWWWc9e/rpp6dr1qzZIUmSs4FllacKwKxCoXD7qaee+r00TbfNMUxJUk5MQEuSerMLcAlRRfi6Qb72MaICZmcieVZqbGgaI9avmX8mlyjy9/06yzqAD9Ysm03P3+3uY2wO8ll7rkzFO/80Mj0HfIveL+A+3MJYpH6dddZZT5922mnHFwqFbYDTiDZQAOOyLDuyXC7ffeqpp55/8sknb5FjmJKkFjMBLUmq9RLilv3bgFkMLmnzDHACsCNwHpEkk4Zqcs18X1WAo9lS4EpiMMFOE4mEc7EyvzWwH91/t+sAvtGC+NrRWmBV1XyB7q1JpJHkHOpfyC0Dj7c4FmlA0jR95vTTT08LhcJ2dE9Ej8+ybHaxWHzg1FNPPf+kk07aNMcwJUktYgJaktRpQ2A+cC/dE1sD8SLwJWC7yr8rGx6dxqLair+xXMH6HXr+3rYxcGDl8UeIfs/VxgM/bG5Yba328yrnEoU0fI8Rd0LUnsOL6fm9l9pKVSJ6hyRJvkLXxcEJWZbNHjdu3L2nnHJKmqbp1DzjlCQ1lwloSdJk4CTgAeB4BlcluBw4E9iKqHx+oe/VpUFZXjO/bi5RtIdr6NlWogR8nEjMf4LuAwuWgF8BT7YkuvYznuh/3am2IloaaebT8yLcg3kEIg1FmqaLTzvttGMqiegL6LqrZypwarlcfvCUU045/phjjvFuFUkahUxAS9LYNY6odL4POIOe/Xb7sobo7bwDcDLwfMOjk3qeV2P5Nt0ycCHdk6jjgfcA7wI2r1k/I/rGjlWb1cwvySUKqXHuJlrxVHMAQo04aZo+dtppp32qKhHd2V5mQ2D+euutd9+cOXNmz5o1azB34kmS2pwJaEkamw4F7iB6Pc8YxOsy4FJgJvApYFHjQ5P+T2113za5RNE+vgdMqrP8W/TsD7scuLrpEbWv2nPFSlGNBmfUzC8bwGsmAhsQLXu2rUyvAPaomt+wss6EXrYhNVyapg+ddtppn8qy7OXE75adbbe2SJLk/F122eW2U045ZVaOIUqSGmgs91KUpLFoX2Iwo/2H8NrriDYbf2toRFLvXkZcKOm0mNFXBf0MsFGd5dsDC+ss/xuRPOr8HS6j5+9zq4nBB79Q5/UfICqpJ9YsvxZ488BCHhGOAr5aNf8j4PCcYpEa6U5gZ+J7fzFwM7AJcRfEdOJn5GaVZbXf84FaSfy8XQQ8XfN4EfGz6V4GlgCXBmTOnDn7JUlyFvC6mqd+VygUjkvT9OY84pIkNYYJaEkaG3YC5gJDqSS5HTgRuKqhEUn9KxKtE6ZULduW0VXNOtgE9Gzga/RfqfgK4O91lo+VBPTFwBFV8/+PuPgmjSRbAHsSCecdK9PuwLQ8g6ryOHAPkYy+F/j/7J1nmBRl1obv7plBkqgIKJgQI35mBAOYUHfNcTG7u65pzTmgDpYzKOq6Zl3TumbXxbymVYxgzqIouqLCKigISoaZ6fp+PF12dU3n6dznvq65oN6uqj7T01391vOe85zPgPeRYG0YOeE4zs6RSOSv6L3u4QIPRSKRkaNHj0703WgYhmGUOSZAG4ZhVDe9gEbgBOT5nA3fogaDtyP/WcMoBS+C1pPyAAAgAElEQVQBO/i2/4ysY6qFbAXo5ZC4k0qAnoRschJRCwJ0GPie+Gz5bYEJpQnHMDKiAQluw4BB0Z8NShpR7kxH1RoTgNei/19U0oiMimLEiBF166+//pGhUOhi4q3iloZCoZuWLl06esyYMT+VKj7DMAwje0yANgzDqE66ohL081F38WyYhTIFrwGW5Dkuw8iW84n3PX0a2KNEsRSCbAVogAeB/ZBgFWQJcA7x9hN+akGA3gp4w7f9C7IjWFqacAwjIfXANsCu6LO3CdkvFGfCYiT+tgFzo2O/oPtAb36wfHS7G4XxgV6KROj/AM8g2xBb2DbS4jhO10gkcjKqxPM3y54DXD5nzpxrrr/+epurGoZhVAAmQBuGYVQXYeBw4DLkB5kN84EbUdbz3DT7Gkax2BCY6NtuBVanehpg5iJA/waJOImaSbeibLGZSY6tBQH6JuB43/aDwMElisUw/KyKBOddgZ2JF9SyYSkSlJ8DpqHP+w/ADFQhMSO6vTDH83dDc4iVkK90P7SI0xtdf9dFjT4TLYJlwix0zXkGidJm2WGkZOTIkSs2NDScjXob+L+/vgQuaGpqGluayAzDMIxMMQHaMAyjetgZCHrmZcJS4E5gFLphNYxy4xPiLSXOAy4vUSz5JhcB2rOYWAGJUB71wLPA3imer9oF6G7Il9Yv7B0APFKacAyD3sBByJN8qyyPXYQ8lScir2XPb/kbYGtKayvTgERovz/1ptGfbITpCPAK8m1/GGVnG0ZCLrzwwnXC4fBfgH0CDz3vuu6Zzc3NExMdZxiGYZQeE6ANwzAqny2AK4AdszwuAtwPXIj8ng2jXDkDLa54zEDNCKvBUzQXARokqg5OMP4w8E6K46pdgA6+V34AVgNaShOOUaN0BnYBjgD2JXNBdjrxvsnvUHlWWA1IjB5KzM96IJnddy5B16K7gccx2xwjCaNGjdoRuAoteHhEgPtaW1vPvvTSSy2hwjAMo8wwAdowDKNyWQ2Jx0eTuBQ/FS8gn9j38x2UYRSAFVHGX3ff2EnIMqbSyVWAzpVqFqCXBT4nvmFVM6ruMIxisC5wKhKel81g/x+QBcWz6DM4q3ChlZS+wG+B3VC1Vs8MjpkN/B24AZhauNCMSsXXqHA08U1nfwEuCYfD1zqOY4sYhmEYZYIJ0IZhGJXHCqgx28m0F5HS8QkSnp/Jd1CGUSgcx6l3HOcy4Ezf8Ewk9vxckqDyhwnQ+eMFYLhvexGyCLBMOKPQ7Iy8aXcj9YJwBDXIfAaJzu8DbsGjKy/qgC3Ra7UbypBORSuy0LkaeLOwoRmVyFlnndWta9euZwPnouoDj/8CpzU1NT1VmsgMwzAMPyZAG4ZhVA71wJ9QRl+fLI/9HrgYZRO1pdnXMEqO4zgD2tradg6FQjsDu7z00kvbjx8//nXk8etxHco2rGRMgM4P/ZHYUBcYnwe8hSwNPGuDarBuMUpPPcp0Ph3YKM2+nwD3Idsry+aNZ13kj30YsFaafd9EFjsPU3vCvZGGCy64YLW6urpLUDPuX3UO13WfrKurO9VxnCmli84wDMMwAdowDKMy2Bll/2yY5XELUPnqJUiIMYyyxHGcfpFIZGgoFNrZdd09gFX8j7uue1Bzc/N6QJNvOIIyXl8pYqj5xgTojhNC2aSZxN8CvAu8jgTp14EfCxeaUYWEgP3Q9+r6Kfb7HgnO9wIfFSGuamBrdI06GOiVYr/3gJHoumUYcTQ2Nm4bCoWuBzbxDS8OhUJXhkKhMY7jLCxVbIZhGLWMCdCGYRjlzUDgSmD3LI9rRSLTRahhm2GUFSNHjuzd0NAwHNgp+jMgxe5zgfObmpruAD5Ftgoe36BmfJXqnWoCdMc5GzVi9fMw+h2HItuiVPgbv02gNm0RjMzYGrgc2DbFPu8D1wIPYM0vc2UZ4CDgLFJnl48DzkOCtGH8iuM44ba2tsNDodCVQG/fQ9+5rnt+c3Pz3aWKzTAMo1YxAdowDKM8WRE1zjqR9iXl6RgHnAFMzHdQhpErjuN0BbaJRCI7o4z+zUjuldqKMgbHhcPhccCrvkZCw1DGs//YCUjErsRmQzOBHsSscULIw7KQAvTdxL9WDcCLqElYpfEb4Gnir5MvoveYGx1fHwnRw5Bw2D/NOecCbxMTpCcAi/MZtFFxrIUsf5ItBrchn+JrUFa9kR9C6LN8OrArie9dXeBB1CPg++KFZlQCjuP0cl33Etd1gw27XwiHw6c4jjOpVLEZhmHUGiZAG4ZhlBedgOORX/NyWR77DsoErGQ7AqNKiHan3zRqqbFzKBTaluRNMyPA56FQaILruuPC4fB/HMeZm+L0V6OGX35uBE7KQ+jF5nwkQAf5C/BTAZ5vU1TeHuRz4M4CPF8hWQ95wi7vG5sNbA58m+K4fsQE6aGkXgwBZbF+TEyQfpHC/G2M8qMOXWuagK4JHm9Dn5vRqBrDKBwD0dzodyS+h/0ZZUzfgVUwGAEaGxs3i9pyDPUNt4ZCoZsWLVp04RVXXGE2dYZhGAXGBGjDMIzyIITKTS8D1sjy2CnIC3EsdtNllJBA48DfkHoRZUooFBrnuu64lpaWF8eMGZONoNcA/AfYMTB+PHBzdlEbFUpP4A3UwMyjFWWoZusLuyywJTFBeijQJc0xU4hvbPhpls9plD8bosa9Q5I8Pg5l3X5ctIgMgC3QXGmnJI+PB44GvihaREalEGpsbDwiFApdDqzsG//edd2RZsthGIZRWEyANgzDKD2DgauQ+JEN81E3+Muw8nCjBJx//vl96+vrhyVrHBjgR+AV13XH1dXVPec4zjcdfPqeyCZhLd+YC5yMsqGN6qU3Epk3CYyfhrx3O0p99NyeIL090CfNMTNQc0NPkH6byrSEMZT13IiqExoSPP4q8h1+o5hBGe3YBc1/Nk/w2EK0MH9dUSMyKgLHcXq4rnux67onoes9AK7rPllXV3dyHuYnhmEYRgJMgDYMwygdq6Kb3KAvXTq8BoOjgB8KEJdhJMRxnF6RSGTHqOA8DNggxe4LXNd9w8tybm5uLkRjt4HIgsFvYeEiv9B8CJFG+bESEp+DjcnuAv5YwOcdgATpQUiU3pzU8+gFwIfEBOnxyCLAKG9WBO4ncUPOn4FzgduwaqNyIQQcg5o1L5vg8ceB3yNfd8OIo7GxcZNQKHQTsI1veBFwxZw5c8Zcf/31S0oUmmEYRlViArRhGEbx6Y4yc04nfZl3kKeRx+Fn+Q7KMILksXFgIdkTeBj5p3u4yA/9r0V4fqN4rAq8QLztBsgKYQ+Km3HcA1kzeFnSw1DzyGS0AZOJCdKvYp7B5camqJHgmgkeexL4M/BdUSMyMqU/cAuJFw4mA/th8yYjMZ4tx1+BXr7xL4ETm5qasrV0MgzDMJJgArRhGEbxCKHmOVcCq2d57OdIeH4q30EZhkeWjQPB5+OcQePAQrIH8BDtBcDr0OempegRGflmCPobrxYYfwY4AGWtlZIGYGNigvRwlE2biunEBOkJwAeoIadRfP6A/OOD15DpyFv+8aJHZOTCH1GT2uUD478ARwD/LnZARmUwcuTI3p06dbrCdd0/ENNIXODelpaWM8eMGTOzhOEZhmFUBSZAG4ZhFIfBwDXEl/llwk9AM/K0bc13UIYRaBy4C+1v3P10pHFgodkeZSl2D4xPAA5EQpJRmfweiYPBipGn0KJeuXrge7YdXoZ0KssagHnAW8Q3Nyy1sF4LnIu8hIPYtaMyWR1VxWwRGHeBk4Cbih6RUTE4jjMsEon8DTUh9Zjjuu55zc3NZr9jGIbRAUyANgzDKCz9gIvI3ud5KRJcRqHMHcPICyVuHFhotkWiZNAL9HtgBPB60SMyOsIyKIv92ASPPQQcSmVlt6+MFiM9QXow8dYxQTxbG0+QfhmwLLz8kkx8vhWJlZX0/jJidEYL938KjLuoWak1JzSScuyxxzastNJKp4dCoVFAN2/cdd1xruv+efTo0V+VMDzDMIyKxQRowzCMwtAFOAW4gMSNcVLxJHAqMCXfQRm1h69x4DDXdYeiJmrJKEbjwEKzGfJx7R8YX4qqCS7HRKVKYBPgH+jv6cdF3t7nIV/lSqYb+v08QXoYqSsQQN8L/gzpSVTeZ7RcuAJ5xftZiJra3V/8cIwCcAqyPWvwjbnImumqkkRkVAyO4/Rva2v7WygU2tU3vAi4IhwOX1qkPheGYRhVgwnQhmEY+Wcv4FoSNzJKxbvAGcD4vEdk1AwV0jiw0PREAtJvEzw2ETgKeKeoERmZ0oCug020zw6ej/52/yp2UEWiDlifmCC9Le0XUoL8CLxNTJB+B1hSuBCrhqtQI2A/vwC7Y5US1caewFja+3ufjcRpw0jJqFGjRgA3AH18wx+7rntsc3PzWyUKyzAMo+IwAdowDCN/bIZ8nrfL8rhpKJvvASyTzciSCm4cWGhCwDnApbQX4FuRD+j5wIIix2UkZ2vgdhJ7JX8J7A98UtSISk8/YoL0UFIvKIGy+z8mliX9IuolYMQ4nvY+wD8DuyIPbqP62AE1IPT3CHCBg6neBS0jjziOs7zrupe7rnsMMQ0lEgqFbl+0aNFZV1xxxbxSxmcYhlEJmABtGIbRcVZEXs0nogy2TFkIXA+MRpl9hpERVdQ4sBjsg/xc+yR4bArQCPwTiBQzKCOONQAHNRtMJK7ej0TDal0syYZlgS2JCdJDad+cMUjQtuPTQgZY5gwDXiA+u34muo5+VJKIsmdz4L00+xyHrnsdYWPSvyYnUjlN/RL1CJiPPkMflyQio+JwHGd4JBK5BVjbNzw1HA4f5zjOs6WKyzAMoxIwAdowDCN3GoATgIuB5bI89kngZOCbPMdkVCGBxoG7A6um2L3SGgcWg+WR97M/c8nPp+hzPLaYQRmsiMrgT6V9eTzAdNQI7pFiBlVh1CO/bE+Q3gHoneaY6UjA9ATpt5FHerXTH/2u/tdnHnrdJpYioBxJJ0C7wAek9vvPhGvQwk+yRplLkF1OpQjQoM/Hf4j/nb4GhgCzShGQUXk4jtM5EomcB4wk/r00tqWl5fgaXOg3DMPICBOgDcMwcmNP5CG5TpbHvYs6sL+W94iMqqEGGwcWi12BW4DVkzz+Mmocah6whWU51BzsLKBHgsdd4DZkofJLEeOqFgYQE6SHAQNJPedfAHxITJAejywpqollgDeBTX1jEWA/4ImSRJQ7mWRAg/7un+f4HPVoMXOFFPssQp/hShKgAY5F3wN+XgB+g1XCGFngOM7GkUjkNrSA4TEdOL6pqenxEoVlGIZRtpgAbRiGkR1rA1cjATobZiGrjRuAtnwHZVQ21jiwqCyLGtydQPLMvjfR5/wR9Hob+WEAEp7/RHwZvJ9PUHXIy0WKqRbog2w7PEF6C1L7xLcBk4kJ0q8A3xY4xkJzGXBuYOwC5BFfaWQiQC8B/op+x1zYF3iI1LZilSpAg+ZiJwbGTkG2aIaRMY7jhNva2o4OhUJXAd18D41taWk5ccyYMTNLFZthGEa5YQK0YRhGZnRF2XjnkfrGPUgL8DfkM2v+pQZgjQPLhNWROHMUyUWW6chH9TpgdpHiqkYGIZuNQ1BmZSKmoUW6v2OLdIWmAfn7elnSOwE90xwznZggPQFZPBQiW3Q74Dvgqzyecwu0qOT/nI8FDqIyq0UyzYD+ETWxzOXz9CTwW5J/XqGyBegGZMWxo29sAbARsuQwjKxwHGdAJBK5nfj31BzXdc9rbm7uqB+7YRhGVWACtGEYRnr2Qtkyycr2k/EkstvI5420UaFY48CyZVNgDLLnSMZ8lA19HyrVTiborIaEVANWQYLzEUjsTMYslIV6E8raNEpD0LZjgzT7zwPeIr654aI8xNEdidv/RgsSHV34CQGvot/J4zskNM7p4LlLRTIB2iX+3i4C7I6E1mzoA3xPvGC/lPYVI5UsQIPE+U+Itxl5CBhRmnCMKiDU2Nh4TCgUupL4Kp+nwuHwnx3H+V+pAjMMwygHTIA2DMNIziaoHHPbLI/7HDXmeSbvERkVQ5aNA2cCL7uuO8513edHjx5tGVjFZ3tUor8rqedH04EHgXuJF4GGI5H6ciRo1yLLAQcAh6FmX8lsZECv443oGmsZ/eVHX5Q57AnSQ1DWaDI8ayBPkH4ZXddyYW/gcfS+uBgtAOdqM3Qg+rz62Q14NsfzlQOZZkC3omvSQVme/wx0DUtmUeRR6QI0qALm9sDYdsgH3TBywnGc/m1tbbdFEw485riue0pzc/O9JQvMMAyjxJgAbRiG0Z4VAAf5A6byPwwyG3nL3oj5xtYc1jiwalgf2UX8HlnvpGIyqnRYgoSYTujveADwaAFjLCf6I0FvN5TZ3znN/h8A1wD/JHdR0Sg+3VG1gCdIDyN1FQfAFOIzpCeR+XXuMWCf6P+/A84H7snieNB9zkco29l/3v2yOEc5kkqAXkK8nVMLymjOpqnk58B6vu0IiReTqkGADiF7Fn8TuXHoWmYYHWLUqFEjgJvxWRy5rvt0XV3dMY7jfF+6yAzDMEqDCdCGYRgx6oCjgUuAFbM4rhXdgDlUbkmvkSXWOLDqWRE4FjUrTJW9HuQLVML9cSGCKgM6o6oQT3ReP4Nj2pClwjWooZ1R+dShv70nSG8HrJHmmB+Ad4gJ0u+Q3HZlVSSEdiMmgL6NsnNfyzDGfYlfCGpF1iJfZnh8uZJMgF4EdAmMLUXN9W7J8NyDgHcDYy3R83QLjFeDAA26nr0aGNsKWcwYRoe44IILVqmrq7sF2MM3PAs4oampaWyJwjIMwygJJkAbhmGIwagUfMssj3sZ3dxNzHdARnlhjQNrljCwDRKVDyPzxanpSCTyfiZQmQtU/ZAo5QmNg0if5ewxCbgbuAuYUZDojHKiH7H3yVBSL8oBLEQZ8Z4g/Rrxns+nAVf7ttuQ8P1U9LH/ponnOeIzWe9FnuSVTjIB+luU6bwxsXs8F73GqSpy/NyAFuL9320LUVbw3oF9q0WABngJ2QZ53AX8sSSRGFVJNBv6FuI9x8eGw+ETHMeZVaKwDMMwiooJ0IZh1Dp9Ueby0aS+UQ6Sa0mwUUHk0jgwEom8VldXN87KK6uSLsCeyJ5jd7K7ZrQBnyLhaDLKlJ6MRLRyyIbvAwwE1o3+bIgW5rKpBgE1Xb0PuB/9fkbtsixa1PUE6aG0z9D1E0FZz96CzRtICNyYeDusNvS9ewfQCPyY4Fz90XvR/xndhOqoTEglQF+OKg2C/s0D0Wubik7It7uHb2wpWkTqDhwc2L+aBOjdgKd92wtRI9VsrEsMIyXR3iC3EZ8N/UO0QeFjpYrLMAyjWJgAbRhGrdKAMqgaie9UnY7FwJWoQc/CAsRllBBrHGhkwPKoQdp2vrG5xIs22dAGfIME6a+RmPYjyqCeiWwLpgMLcjx/HdAbCcx9o//2RtmqKxETnNP5+SajBWWuPosar1aDwGcUhnokAnuC9A7ovZiKWWgRJNE9iwvMR7ZZ16LvZ49zgct8228hW4VqIJUAvQm6fvgF6CXAVWjRPBUj0MJRvW/MRX+rU6huATqMPMv9NjK/R0kGhpFPQo2NjceEQqG/ooUdj7FRIXp2sgMNwzAqHROgDcOoRbZDN0z/l+VxT6LmZFPyHpFREqKNA7dGN9g7Y40DjdT0Q1lym0S3l6Iy7QdQluau0Z+tSJ3pmSu/oCzRBdHnXoxEINCczhOReyDhuRvtMyHzwdeoJP/Z6L9mL2PkygBigvQwlKmbzf2JG91/KhKdH4yOTYie0+NkZC9RDaQSoPsDDyG7jAbfYzPRAlRbivM+BwwnPtv8W2BNJExXswANMBq4wLc9FjiwRLEYVc6FF164Vjgc/gfyIPeYBhzV1NT0fInCMgzDKCgmQBuGUUusDFwBHE52178vgdOR76RRwVjjQKMD/B/K8F0tuj0P+B0SbYLUA+sR7528PtlZdpQj9wBno6xswygEKwFD0Odm2+j/61MeITwh+mP0Hn2amJDqImF2ap5jLRXpBOhd0XzFf71pQ2X//0lyzpWRtZj/mCXAxaji6wGqX4AeQnzjwTko+94Wmo1C4WVDX0WsyacbCoVuW7BgwRlXXnllrpVPhmEYZYkJ0IZh1AJh5PF8BbBcFsctIGa3saQAcRkFxhoHGnliB+BRYhnG05EH9IdZnGN55Km8PhKn10PWF6tRHvOx+cgGxPOmnowWYAYDd0b3iQB7Ee+VahiFYBBwKxJbPXE5Vz4DNshHUGVCOgE6jK5RfXyPtQKPAAclOee5SGz2fz+6yJJiGrUhQIeR5Yu/SVwm3tmG0SEcx1k/EoncSXwj9EnhcPgIx3HeL1FYhmEYeaccbngMwzAKyRbo5mhwlsc9CZxI9WRM1QzWONDIM/ujpnqdo9uTUMOqfF0buiIhem1k8dGb9l7NfXzPnwszae8t7f1/KhKdp6U4/jpkYQDKCtwSVYYYRr5ZHi36HocWPOpS754R/wQOycN5yoV0AjToNTydeEG5BV1LEjXW+wrZoXhEgJeBnaLbtSBAg2yFfuvbPhxd/w2joDiOUx+JRM4EmohZZ7UCl3z22WfNY8eOTWWfYxiGURGYAG0YRrXSE7gIOInsyt4/RM12xhciKCP/WONAo4Ccipp3edeQN1AG8E8lime5aCyet3NnYl7TLjFhaS4qufe8ojtKPSrdHx7d/hz5XP+Sh3MbhschqJlgosaELhJFvWzoMJnfxyxCn+U7SO2BXClkIkCviz6n/tdoKZrf3BI4bit0bfPTChyBxHuoHQH6cuAc37aDMsMNoyg0NjYODoVC96AqKY83o9nQ/y1VXIZhGPnABGjDMKoNz25jDBKhM2U2cCEq+a2GG9SqZeTIkSs2NDRsgzUONApHCF1DzvWNPQocRqzpX63RE3gbWCu6/QSwHxIFDaMjrA3cCPwmuh1BljDzfP/+jBZWFkR/fomOL4ju49++DNjGd/79gMcK/UsUkUwEaIB3o/t693su8AHtvzNvBf5AfMPSBWghwLve1YoA/Wfgb77tO4EjSxOKUaucfvrpXXr06HGZ67onE/v8znNd96zm5uZbSxmbYRhGRzAB2jCMamJTdCO0dRbHuMC9wJkoO9YoM6xxoFFkOiHRwV+yfz1wGia2bgy8TqxZUjMwqnThGFXCesgeYi4Skxd38HxvEu+lug3tM3wrmUwF6OPQtashsJ/f17gz8j3u5nt8KfAPJMZ61IoAvS9abPR4AtinRLEYNY7jOL+JRCJ3AKv4hp9pbW096tJLL51eqrgMwzByxQRowzCqgeVRieSJZOcX+RFwAhJUjDKhI40DlyxZ8tzll19utgBGriwLPEQsE9NFfoxOqQIqQ/YDHkZzSBcJ9Q+WNCLDiGcisKFvexPg4xLFUggyFaCXQ17v/szmpchWaGR0+xDgbmSz4+Gihfy3fGO1IkDvAjzn254J3IUWMN4ErE+EUVQcx+nV1tZ2SygU2t83PCMcDv/BcZznkh5oGIZRhpgAbRhGpTMCZfislMUxvyB/6BtRxqxRYrJtHAi85rruhLq6uietcaCRJ/oCT6NKCpBQcyRwf8kiKl+akWURSITalsSCmGGUgk+BDXzbGwGflCiWQpCpAA3ycN6f+CzoWcDKyG7sJfT59S/efwOsGThPrQjQw4EXUjw+Hb327wETgNeoXVsmo4iMGjVqBPJvXyE65IZCoetnz559zvXXX7+khKEZhmFkjAnQhmFUKusBNyBLhmx4Ejge+F/eIzIyxnGclSORyLbRLOfdgNVS7G6NA41CswHwDLB6dHs+Wtx6tmQRlTch4F/A76Lb3wKDMRsjozx4B9jCt70l8i+vFrIRoH+Drm1+26o2YE+UKT418NgSZKtzReA8tSJA7w08nsX+rcAXxMTo94BJWK8JowBceOGFa4bD4XtQDxSPDyORyKGjR4/+rFRxGYZhZIoJ0IZhVBpdUYfykcSXlabjC+Ak4PlCBGWk5pxzzlm2a9euW/p8nK1xoFEubI18PntFt6cDe6BmXUZylkX2RZ7VwQRgJ5Q5bhilZBx6L3rsgaobqoVsBOgwso3wV4m1oqaMHwKNxFtcRdCCcLCyqFYE6D8Bf+/gOX5BiyCeIP0aanRtGB1mxIgRdQMHDjwLVSJ5lQ2LgfOampquLV1khmEY6TEB2jCMSmIvlPW8erodfSwE/gKMQZk9RhE4/fTTuyy33HJDrXGgUebsB9wHdIlufwbshoQcIz1rosxST7y/Dji1dOEYBgB3IPscjxOpLpE0GwEaYDQSiv1CcwuqWOjnG4sALyIbrCC1IkD77YXyyRRigvQEtMBZ601tjQ7Q2Ni4ZSgUug9YyxtzXfex1tbWo8eMGfNTCUMzDMNIignQhmFUAmsjn+ddszzuSZT1bGJSgUnQOHAY0DnFIdY40Cg1JwPXEFsYeRMtcs0qWUSVyS4ou9RrYnYscFvpwjEMLkRCoseNaC5QLWQrQK8DTCb+vm8pyp70j7UChwJjE5yjVgToh5FndqGZjxbePUH6FdQw0jAyxnGcHq7r3ui67uG+4R/C4fAfHccxCzHDMMoOE6ANwyhnugDnRn9SiZlBvgJOobpKbsuOXBsHRiKRpy655JLvihOlYbQjhJqQXuQbexw4BGsmlStnoUoTUGblTsD40oVj1Dh7oAVoj3eRR3m1kK0ADapU2ILYvZ+LvKDrffvMA3qTuFqsVgTo74jPCi8m04n3kn4Hq9wzMqCxsfH3oVDoBmSNBcqu/8uMGTMab7311pYShmYYhhGHCdCGYZQrewHX0r4TeyoWocY5lyE/NCOPWONAowrohMrzD/ON/R34M8r+M3Ln78g/FWAGEvys2atRCnoTn03aijyQq8WHNxcB+mgkFDckeXwp+gyfkOTxWhCg1wM+920vAS5Hwv2WwIpFjmcR+ju/BbyBqnRs8d5IiOM4/SORyL3ENyh8NxKJHDx69OivShWXYRiGHxOgDcMoNyOB4P0AACAASURBVFYFLgWOyPK4F5DP4+S8R1SjZNk4cKHruq97thp1dXUfOI5j/oZGOdEdlZZ7Vj4u0AQ4pQqoyuiMysiHRLffB7ZFPvyGUWwmEmuQCZpT3FuiWPJNLgJ0DyTKL5PkcdBn950kj9WCAO2v5AB4Dvitb7sfEveGofnQFqR+PQvBdPS396w7XseusUYUx3HqI5HIhciGqC46PNd13eOam5v/WcLQDMMwABOgDcMoHxpQ5s1oJBRlynfA+cDdhQiqluhI48Cffvpp/PXXX2+loka50hd4Cr2nQe/f44HbSxZRddIX2R14Jez3kv1iomHkg0vQ3MDjKWDPEsWSb3IRoAHuB35H4izo/yKv6GTUggD9HnptPU5Gja+T0QBsTEyQHkZ2VXv5oBX4gpgg/RowCS2wGjWK4zg7RCKR+/DZyYRCoZvnzp17xtVXX21WY4ZhlAwToA3DKAd2QE2CNsjimBbgb8AFqJmLkSXWONCoEQYCzwBrRLfnAwdGx4z8szXwErHMwDOBq0oXjlGjBEXaNiTOVoMtTK4C9E4oqze4sLwUzaWuTHFstQvQWxCf/Z3r+6UfEqMHoWzpoaifSTH5Bf0unpf0a1SP/YyRIY7j9IpEInciT3yPSa7rHtzc3DyxRGEZhlHjmABtGEYp6QdcjcSgbDC7jRwJNA7cGVghxe7TgQmu646zxoFGhbIV8G+gV3R7BsqCTCTeGPnjD8Cd0f+3IU9/E/yNYvM+saoHkL3COSWKJZ/kKkCH0eeyR2C8Dc2pZqQ4ttoF6PtRI1qPfGXM1yNvab91x0CKfw8+hZggPQH4ADWqM6qb0KhRo05B/XE6RccWASObmpquLV1YhmHUKiZAG4ZRCkKoLPtqoGcWx00HzgPuwcoLMyLXxoF1dXXjHMeZUqQwDaMQ7INEEy/77Cvk//zfkkVUW9yARC2AOchf1l57o5gcB9zs256PLBJmlSacvJGrAN0RqlmAXgf4jJhnLmjR7MkCPd9yqEmrJ0hvQ3Zz4XwwH9moeYL0K8Q37jSqCMdxtohEIg8Aa3tjrus+VFdXd4zjOD+XMDTDMGoME6ANwyg2GwG3oDLtTGlFNziNwNxCBFUtWONAwwDgKCQ81Ue330bZbDNLFlHtUY/K/XeMbn+OMtLNuscoFsugzM9+vrEbgZNKE07eMAE6vzwK7OvbnoTmqsWaD9UB6xNv3ZGqB0ehmE7MR/o9ZONhvT2qBMdxekQikZuJz/T/JmrJ8Vap4jIMo7YwAdowjGLRBTgXGEmsDCwTJqAsuo8LEVSlE2gcOAzYkpjoFqQN+BBrHGhULyHgouiPxxPohmthSSKqbVZE4v+A6PbjwP5Y6bdRPE4n3oO8FQm4leyBagJ0/tgZeD4wdjDwYAli8bMssAkxQXoHoHeRY2hBc29PkH4P+LTIMRh5prGx8fehUOgmoFt0aAlwrllyGIZRDEyANgyjGOyIshHXzeKY2UATcD0mVvyKNQ40jKTUo8akR/vG7kBl+K0licgAiSivEbvZvRhwShaNUWt0Aj5BNgseH6DF2paSRNRxTIDOD92QDcVavrE3kSVGOdq89SPeS3oLYs1ei8V0YmL0BOB1bHG34rjwwgsHhsPhB1Gmv8ejS5YsOdLuEwzDKCQmQBuGUUhWQk1/jsjimAhwH3AGle/TmBescaBhpKU78C9gt+i2ixawnFIFZMSxP/AQmne6SMj6V0kjMmqJfZHNgp/RyNarEjEBOj/cRvyCZQSJz5ViR9AAbExMkB6GPM6LSSvwBTFB+jVkYVKOAr7hw3Gcrq7r3ui67h99w5Nd1x3R3NxcyRUihmGUMSZAG4ZRCELAMcBlpBZLg7wH/Bl4txBBVQrWONAwsmJl1CzK8ztvA04Abi1ZREYiRgMXRP8/Hwk9dpNrFIsHgQN92y6y5im11UIulIsA3QqcSmUK0CehCjs/1yDLlkqmH/Fe0kOJNeItFnOR9ZJn3fE68FORYzAyJIElx2LgPLPkMAyjEJgAbRhGvlkX2W3smG5HHwtRtuKVSDyqKaxxoGHkzFrAM8TK6xcABwFPlSwiIxlh4DFgr+j2N8AQrDGkURx6oQWPlX1jC5BA91FJIsqdNdF8qS4w/i0ShAvBKcDwwFgEzfeeK9BzFoodgf+gDGKPL1Djv2qzk6gH1iPeumMgxdcAphDvJf0WlWuBU3VELTnGAv/nG75n4cKFx1955ZULShWXYRjVhwnQhmHki87AedGfbDzpnkRNBqcWIqhyxBoHGkZeGIKuH15jpp+QuPlGySIy0rEs+vt4N7kvALtiHt1GcRgKvEh8I+RvsIWQWmIAys5d0Tc2F1Vk1EqDveWAwcQE6W2AnkWOYT5a+PGsO14BfixyDIYPx3G6RyKRW1FliMfn4XB4hOM4n5QqLsMwqgsToA3DyAfboyyY9bM4ZjoSq+8uSERlhDUONIy8szcqCe8a3Z6ChMwvSxaRkSnrouy35aPb1wKnlS4co8b4A3BnYGwSqj6aXvRojGKyBlr08jcdjCCP8H+XJKLyYQAxQXooygYPFzkGr8Gh5yX9LrKDMIpI1JLjZmLWLfNd1z2uubn5/lLGZRhGdWACtGEYHaEnMAb5PWd6PYkAtwNno6yTqsQaBxpGwTgS+Tt7VQPvAHti2VOVxC7IOsWzEDgGfS8YRjG4Djg5MPY5+q6279/qZB2U/b5qYHwk6ldixNMd2JSYIL090KfIMbQAHxNv3VErWeolpbGxcVAoFPoXWpgAIBQK3Tx79uzTrArTMIyOYAK0YRi5EAKOAP6KfBUz5WPgWCqnw3jGZNk4cBbwkuu644DXmpubbUJtGOkJARdFfzyeBw4A5pUkIqMjnANcHv3/UiT+jS9dOEYNUY/se34bGP8CvQ+nFT0io5AMRJnPfQPj96CMeLfoEVUm/Yj3kt6C7Cz38oGXJe1lSr9O9fl2lwWO4/SIRCK3AyN8w+9HIpHfjR49+utSxWUYRmVjArRhGNmyNrLb2CmLYxYCfwEuRUJDxeM4TndgK1/jwM1Jfk21xoGG0THqgJvQApbHndFta2RUudyBMtoBZiBf0v+VLhyjhugMPALsFhj/BtgP9V4wKp/tgbHEegV43A38iRpsfJ1HGoCNiQnSg4ANihxDGzCZeOuOSdiiQr4INTY2nh4KhS4j1rRzVjgcPsxxnEprPmoYRhlgArRhGJnSAJwBXEx2GQ9PoyaD3xQgpqLhaxw4zHXdoaFQaHviO6j7scaBhpE/ugEPAnv4xi5HHvJGZdMZeBUJzwDvIzFjUckiMmqJTshLfv/A+GI0b7mj6BEZ+eRY4Abaz9VuB45DlnBGfulHTIweihocdk15RP6ZC0wkJki/jpoUGzniOM6wSCTyIPr7ggT+K8Lh8PmWUGMYRjaYAG0YRiZsh7KeB2ZxzAzgXCq0yaA1DjSMsmBF1Bxq6+h2G3ASuh4Z1UE/5OPt3djeA/y+dOEYNUYdqqY4PMFjtyKv6Kqo3KohugB/Q/YaQW4BTsDE52JRD6xHTJAehu4liq1BTCHeS/pt7HOdFSNHjuxdX19/f7S3DQCu6z5dV1d3hOM4s0sZm2EYlYMJ0IZhpGIF1JwlmyaDLnAvcBpQURMSaxxoGGXFANSobt3o9kLgIOTdalQX26AGYV51zenANaULx6gx6oCrad+YEJQ9+Ufgy2IGZOTMZijxYcPAeARV8DVj9gylpgcwhJh1xzaoqXkxWYAqFT3rjleBH4ocQ8UxYsSIuoEDBzYCo4jdF04Nh8MjHMd5u4ShGYZRIZgAbRhGMkYAN9LeNy8VE1FZ4xsFiSjPBBoH7gqsnmJ3axxoGMVjMBKa+0S3ZwN7owwmozr5I/CP6P/bgL3QAoRhFItDgdtobxmwGNn+XIJ5zpcrnk1cE7JW8TMXZUM/VuygjIwZQEyQHgpsihaGionX4NCz7ngXffZrlb2Qv/YXwQccx9k7EoncBSwfHVoMnNLU1HRbEeMzDKMCMQHaMIwga6HSxV2yOGYRcAVl3mTQGgcaRkWwC/AwsGx0+2vUKGxyySIyisVNwPHR/88GtgT+W7pwjBpkE9SccECCxz4CjkIilVE+bIN8nRPZxH2MPL6/KmpERkfpjkRoT5DentiCdLFoQe8fv3VHLSWf9ES//1OoeuB7/4MXXnjhOuFw+GFgI9/wPfPmzTvu6quvtj4OhmEkxARowzA8vOwRh9Rex0FeAv5MghXyUmONAw2j4vgj8l31PqfvAntipbG1QgPwHLBDdPszYCuUwWgYxaIncBe69gRpAa4DxmCNzUpNPzRnPQoIJ3j8LuT3vLCIMRmFw2tw6HlJDyK7+5V8MAPNSzxBejzwc5FjKCYHo0atC4HrkS3jr7/vWWed1a1r1663AYf4jnm3ra1t/0suuWRaUSM1DKMiCAHPF/D8e2OdzA2jEtgWNWbJpsngD8in84GCRJQDHWkcGA6Hn3ccp5onkYZR7pyLRB1vcXwccAAmPtYaK6IGUV4G6mPofWAVKEaxSWVFNj/62CXAvGIGZbAsEpYvIFYp42c6cCLwaDGDMopOA7AxMTF6ELBBkWNoQ9VZfuuOz6iu76vHgH2i/5+NKl6vxWdP0tjYeGwoFLqemP3NrHA4fIjjOOOKGqlhGGVPiMI2YuiBTcoMo5zpgRqynETi7JFEeE0GT6cMsn9ybRxYV1f3tOM4/ytOlIZhpKAOuAFVUnjcDRyN+a3WKpugG/lu0W0HlQAbRrHpg8SWg5M8Ph35Dv8du14Vmq5ovnoeied6LkqmOBdbuKxV+gJbEMuU3ob2nu6FZi7qieMJ0m+gPjKVymrIesS/2PMt0AjcR1Rsdxxnm0gkMhZlqgO0uq57dnNzszUUNgzjV0yANozaZXfk9Zyq8V6QL5FI9GJBIsqAXBoHAq+5rjuhubnZfBsNo7zoCjxIfKn7dcBpFHZ+YpQ/+wMPEZurHgSMLWlERi2zF1ooSzbn+F/08VuBOcUKqkZYGWU0H0fyxtiTo4+/UqygjIqgDlifeOuOgRTfhnQK8V7Sb1PGPXMScCK6vgX5GBgJPA0wcuTI3g0NDQ8CO3o7hEKhf4ZCoaMcxzErHMMwTIA2jBqkD3AlcEQWx3gd4McARfVFtsaBhlG19ASeQDeFoFLWk9HCmGGAGtuOjP5/Pspmm1i6cIwapxPyqW8meUO0xWih5DJgUnHCqlo2QVYbRwBdkuzzHcpAvwNoLVJcRmXTAxhCzLpja2T9VEwWoL4zniD9CsoqLlfCwKvE5mtBXkGVCW86jlMfiURGo0oEj4/C4fD+juNMKXCchmGUOYkE6IWotD4f/AMrRzOMcmIEcBPQK4tjXkFZz58XJKIA0caBgyKRyNCoj7M1DjSM6mNN4Blgvej2EiQyWIar4ScMPE4sQ/4bYDCVXc5sVD7LAecApxKziQkSAf6D7IQex3riZEoPVP3wB2LNSBPxM0qMuA5rMmh0nAHEe0kPIfm9R6GYTryX9Lv4fJbLgPXQPVeq3jrjUEP7iY2NjYeHQqFbiS0ezY76Qj9X4DgLQR9ge/S+WAdYBVgm+thitBD2JfAWum+2OYphJCGRAD0HZSUZhlE99AduBn6bxTGzUebZbRSwUsIaBxpGzbExKtdcJbo9GzW4mVCyiIxyZlngTWLNpV4AdsWyHY3S0xc4HziS5EI0qBr0EeSX+iJaPDdidEKf6cOQ1UmybGeQ8Hwz8Bf03WEYhaA7sCkx647tgJWKHEMr8AUxQfo9VFVRSnuyi1BPhlREgIeBcxobG1cIhUKPoPtQ0LXvgqampisof5u1ELAbqszbBdm5ZEIL8BzqHfB8YUIzjMrFBGjDqG7C6ItzNJpMZYIL3AmcTYGaDFrjQMOoWXZGNyY9otvfoAl+USosjIplXZRZtHx0+2qUZWUY5cAKwLGoQd6qafadjrzNn0aZcrWaGd0dfR/sBhxAeguEL5Ggcxey4zGMYtOPeC/pQaROmCkEM1BmtGfdMR4tyhSLTsD7wP9lsO9C4PrddtvttsGDB98C7OQ94LruE3V1dUc4jlOuzUI3Rg1Nt+rgeV5B3vSTOxxRafiE+AXBO4BLShSLUSWYAG0Y1cv/AbeT3Zfn1+iLMq8rtueff/5K9fX121njQMOoaX6PrkleWetEJD58V7KIjEriN0i087KQjgb+XrpwDKMdYWAP5IW6TQb7L0KZjeOiP9U+3xmAROe9UEbhMql3B/T6XIsyyC1z3CgnGpBQ6bfu2CDlEfmnDYmbfuuOz1AWcqHYKvo84Qz3/6m+vv7y8847b41wOHyib3xiJBLZZ/To0V/nP8QO8Wd0zemUp/MtQguU9+bpfMVkAWoW7nEVcGaJYjGqBBOgU9MVdc7tiS7wM4Cp6MNYKLqg7Ine0eefG33e6RTGTzuEJoQrooy0EPrSmh19zhkFeM5yYo3oTyf0+k5FImw2rIA6dPdCk+mZwI/o9SsFDSgz7GIym9yD/ua3oy+VDmeWWONAwzACnIqyVr3rwAvI57Ncs1+M8uQ81AwX5Lu4A8qMNoxyYzNkKXEIypzMhK+B14G30fv6Q4rc+DmPdEVzvy2jP9sQs11KxxTgfmRZYtUxRiXRF9iCWKb0NsQLeMVgHvAxMUH6DfLvSXwdqrDNhv/tuuuuzw4ZMuRQYq/JT8CIpqaml/IaXe74Gx8HWYoWCp9CeoF3n98XaQl7oizvZN7h5yDroErCBGgj75RKgD4bNZHx8xK5db4fim5s/fyCyuASTdp6AzcGxm5BN8OgzJqDgWOi564P7NuCskPvBf5JfvyLVgCOAnZHq6iJLlzzos/7OPAAHROju6FmdIeiSWGPFPsuRBPgt9EF92VS+y7WRePz8wDwaA5xHovES4+fgOMzOO5+4v9uTxC/6jgIfWnujt4PQZZBXzKp2Aq9hrsBA5Ps8w1qsnUHKpcqBlsjz+ZMSqM8JqJMsrdzfVJrHGgYRhLq0I3KCb6xe4E/YU2KjewJIVHqkOj2dDSftCx6o1ypA3ZEYvT+pJ5zB1kKfIDE6Iko03EySnIoJ1ZBNjnrIt/cLYGNaH8PlYqfgAfRHP51yt8f1jAyoQ4ls/mtO9Yn8+zhfDGdeC/pt0l/r5uKbuiatGa2B6622mpfH3744T0aGho8250W13VPam5uvrUD8eSDc4HLEoy3Ie3oYtL7zvcCmpGGkehvfDzysK8UTIA28k6pBOg1kX/Q8r6xNrRq9EoW5+mFJmZBv7WDgH8lOWYNJAz6ORaJduugic8WGT7/q0i4+zLD/YPUA6cAF5Ddaz4Zifj/zuE59wWuJ71HXTImANumeLye9qLCBWhFMVtuQX8bj/8Bq2Vw3BLiy2b+glYduwM3oDLwZNm4ID+vZILoBij7au8M4vBw0aT6TOD7LI7Lhq7AKOAssmuScFX0uKwmIdY40DCMDFgGuAct1nlcB5xOYctDjeqmC5oreokMb6JMaFvINMqVtVF23C3ISmY31HRvjRzP9zNqTjY5+u//kCg9A/gBVeJ1RFzyswzQB2X59UGJG2sQE5zXRY1Cc+ELlKjxDGrOaIuSRi3QA1l3eIL01qT3QM83C1AykOcl/Srt9ZF07IZssbKma9euHHbYYXP79u3764JcKBS6dfr06SfdeuutpbgO7IoS7YKi8Q/onj/bJK1tUdJgsMdRC5qvvJ59iCXBBGgj75TSgmN/1ITDLwR+j1bOZ2ZwfAgJsHsExm8mdZZsMgH6dZTd2yuD5/YzG2U2fJzlcV2QSL5nlsf5uQhoymL/09CFI5X4mo730SpuMspVgL4UZblvnsHxXVBpb5DfITEl14YTU9GX9aQcj0/Gruh9n82NzOto8eSzTA8INA7cidTXCWscaBi1TU80+R4W3W5D1UrBCiTDyIXVgXeQIAZwN/CH0oVjGElZH5VtrwKcT8xCBlRB54nR25G5bVom/ITEk0VIRFiK5scLUSXjvOh+yyHRpRuaO3dG8+BuyF5uefLHQiQ0P4tE5yl5PLdhVDIDiPeSHkz+PIgzZTrxXtLvkb5J6n2oojpr6urq2GOPPdh00039w+NbWloOGDNmTCZaUL7oAXxK++S8WcBwlOmdC5sh7SEoQn+B9K5KaEBrArSRd0rtAZ3IP+hZJCqny446i/Y+Oh8ha4RE4qFHIgH6LGTZ0d83NjMayzfRWFYHfktiH7cf0Srmf9PE7NEJTUYTZRJH0IX/M5ThsBL6ItooybmuRNnQ6dge2ZwExecWlD30DfqdZ6HJZw+UEb4x8aJvJQrQV6Lf4zeB/WYgv70F6HVeE2VJd0OTZD8novdronKaKUjQ9TJOeiPPr/UT7DsbTSryMeleAZUKHUPmiwoL0aLFX0jzGbPGgYZhJKEXqf0E+yNxwbsGLkGVJ8kqkwwjF7ZB3zve9/2p6HvaMMqFTYHniNm9fRUdS9RrI1FDs4F0LGmk1ATL/t8l9T2aYRiiGxIwPeuO7dC9ajFpRWKp/zM8iXjtqFd0LJGlZUZsvvnm7L777oTDv95if+W67j7Nzc2f5nrOLLkcVUr7iaBM5fEdPPcewJMJxkeS2O6j3DAB2sg7pRagl0EXtKCgme5DuRUqFfF7zM5H1hmT0zxnIgF6JrEL51wk6N5Be6/jMPIdvI72r9F4JPJm4ll2GfIZCnJfdDyRl+EglOUatAdxUWlIooubn3cCx7ro9xhN+sYEa6OO1UchIaHSBOipxMRTF73Of0WlR346A/sBDxNfurgDWjAIWls8hsTcD5LEMQS4BpVW+Xkb3WB0pMRoBLIT6ZNuRx9Po+qAqYketMaBhmFkwK7oOp/sO2cjdK3xMknmAPvQ8Um8YSTiz8T6h7SiRIEXSxeOYfzKIOA/xErrJ6PqsWz8yldCfsqbowU9z/KiW/7CzAtzidmBfIaSVd4ivV+qYRiZ0494L+lB5F6Vmysz0EKSZ90xAWkEd3XkpGuttRYHHHAAnTv/+uvMBw5vamp6vCPnzYDeKBkteE29Bc0v8sE/kT2sn5+QJrUgxXFhVKHiZyG52Y11J143ayNxE/B64i2VvkMVMR43ARemea5k505FA9JLtkN2T71Rot1CVK3zOcpSfxW9dh1heZSUOBjpKL3Q/HEW+n1fjf7kamPVgF5vP/OI1xVDSM/cGX23944+/gP6fD1CrMllKjqj32UbpNcth3rhfYfmws+R/0z7HtG4B6M5Si/0Xp0ZjflVZJOX9HlLLUADrIUuYP4PWCsS/V5LsP8KSPAL2g0cQXyjuWQkEqA9fkEi8kdpzrFuNLagXcdxQDoD/a3RxTqYSXsWEkVTUY9sS/YJjP8IrIcyphMxAGVd+LmE9BeQICGUnZHq9SlHAdo/fhAqC8+UXuj39We+R1BGdCZNBDoh244DA+NnolXEbOmLStj3y+KY2WhRJ+696ThOfVtb2yY+H2drHGgYRiq6olLEP5JYUB6OGs56nn7foWav2VpUGUY23IzmX6DvuyG0n/MYRjEZhvxEvWvhJHTDlskNZSasRkyMXgvdBHo+zb2jP5n2A0lHK7qx/BFZJc5EItRXSHD+PLptGEZxqUf3/35BeoMix9CGFp+WJ3GVeMb07NmTgw8+mF69fpVX3NbW1ksvvfTSbPWKbDiD9vrLfHSNzVe/on5IewreYx+FEh6T0Zf2vaO8vmXZ8ijqA+bxAYltSYcj25CO8CHK3s+EtZEedQDtRdtERFAi353AA2QndG+M9C8vkSYV89GiShPZN/3dByUp+tmSmI/47ijrfsMU52hFyRXnk7xi6gyUuZ9Kt50Z3ecuOt7YdwhqxLkTyfUij0XovZ3w9UskQHtCU0c5ncwznkbQvjR3Gnrz+lc5QmhFYN/AvnegD3EmpBKg9yVzcXJH2mfZTEG2FakyQR+hvXiYzSpbF5RVELTkSCVo7o+yej1cJKwWIjOhnAXoPyCfyGwYjeL3czay9ciUzsjmZBPf2FR00xDMsk9GCFlt/IXsOqiPRWL5TGscaBhGB/krmvBsSvuFyMOBvxO79n6CvE3N/90oNA3A8yiBAJSBuRXZZ+AYRj7YAfWo8W6m30OZ+R3N2sqGMDEhehm0eLiM7//+LLdf0H2L5xO9GN08LiYmPBuGURmsjDITPRufYeTXx72gLLPMMuy3336su+66v47NnDnzne+++26nJ554Yl6KQzNhELLbfMg39gGa0/rJRlfKlET6z0tI8E1GNQvQy6B7iuNILwYn4xTg+gz2qweuBk4gsZVqKuaiHmr/yOKYZAL0u0jHOSOLc32AMpz9bgV9kV45OOERibkGabO5sBxKYhxB9pZgc9Fnyf+ZS/gHryO1xUKmBEsGUjEWqfz+5oGrodWNvYmJ5CfTXnz+lPY+0rnwLNllxr6ERHN/ZusAlOHwXJJjVke/j58fgfOyeN5FSFB8NTB+InpzJRK/gzYNP1N7ZXHjyF587kb7hpbvkj5TPchiJFr73xerowaUwQtUItZGH/wds3jO74ETHMeZ2NbWtl+wcWAolPD6YY0DDcNIxibIYxckWPg5FS2AehO7l9BEO7ifYRSCFpQ98w66uRyIsj32p+MZH4aRDbujhA9vcX8C8gAt9mJIBJXy/lDk5zUMo7TMQAtg/45u16ESf791x/pkL8QVhSVLlvDggw8yfPhwhg4dCkDv3r0Ht7W1/bD66qufNnXq1NvI/nu9J3ARSkRb1zfej/jkMI9UWcm5cgftBehhaCGwo8J6pdEbCfLD0u2YB7ogC5Sg/pYpPVByTT+UPd0RriJ2H5UpmyHxdjj6Xu+D7C3WyfI8pwHfIq0wG1ZFtorJetGlowfwIPq9b/AGc11xKARnIHsK/yrUnsRKI7YArggcsxBZKgQbxuXCDel3acd1tLdWOIjkAvR+tC+Ju53sSzzGo4zarXxjA9CXyzsJ9g9aJfRAnnTFzMYoNTfmcMwBtC9ruIzcbmifR+WKa/nGdiC1AF2PMtsdMvT4+ZejEgAAIABJREFU6tatm7vxxhu/MHz48Gl1dXXXRSKR1ZOIzQCzXNd9MxQKTXBdd5w1DjQMIwn1aPXf+/7yhOUQ+l4+y7fvwygb2hpNGcXkJyQ4v4YyPPdFZZ3NpQzKqCn2Rokpy0S3X0HeqLUmLhiGUT60oWS9T4klYvVAdgSeIL0V7W1FS4brurzwwgv8+OOP7LXXXtTX17Pyyit3OeCAA2559NFHT/zmm29ORIt76Qij+ehf0e8XtAPYlvYZnfOAN/LwawR5AVU9+7U3z/c4mW5UCj4nZmkGyjD2V5WPQ4mjqUilL3VDr0UiQXM6Eiv/g8TSWej7tFd0/63Q9+yqCY5Nxt9JLD5/iRYFnkdJew0ogWEfZDO4gm/fEKqIn0l6q91kHEi8+DwRWYi8jX7Pbugz+SfaZzZvDxwd/V3+RUx8dlE/nseRDc68aNzbomzvlQPnaUb3aNMyjLkn0hz7B8Yj0ed9AiVmzoyOrYQ8vI8ifmEnjDTT/6KE37ISoBejP857xJufj0F/pJuITeo8TkYX1I4yh9w+/K+jP6LfFmKrJPsme+y+HJ7XOy54vq1JLEB/E9iuQ0LqsdRGdtBitHqTLdsHthcCz3QgjpeIF6BTrfxtihYnUlYjdOrUiVVXXZU111yTddZZZ0mfPn06oSz8RFjjQMMwcuE0YuV0LprkLINuZvyLsNehEi+7rhil4EPg9+jmKIS86j4jUPpnGAXgEHQ99O6rnkFJDPlu/mMYhtFR5iIBdwLyogUlsnk+0oOQCJbIzrJoTJw4kZ9//pkDDzyQbt26seyyy3LooYdu/OSTT47/+OOPn0D+uMl0oCFIPB0S3Z5D++zPRDYR71OYOewiFGsw43ozykuA/p54kfVq4t8HH5O7CAtyPAiKzxEk8F5B4qaM/0Pzu3uQ9rcd8jVOp2Edir6bg/wV2asGEzSnooXjK6PPFbRHuQp4GfU+yBbP/mIp+h1up/377HX02o5Bv5+f85BI7WlTX6HfLZHu9zK6H3saNSf06I4cEzJxXgihxKP+gfGPgcOQzWKQGcie8UZ03/gXYpUWIVSZuBHwYyIBegFwZAaBpSOXbMov0arL/b6xBjSRC5aK3Ef+SiTep71vcSa4KBPZL0Cvj1YWE5XbbR3Y/hndHOVCopW/rdEbLsgb0Xj83sFHo4vgGFSmk6kXcSXyEbl1Mt02sP0JHcu2/zKwvV6CfbqgMqGzSNBAJhwOs9JKKzFgwADWXHNN+vfvTzj860cjuEBjjQMNw+goayAhz0UTiEVokfgxNAkk+ti5aLJhGKXkYXQTcy6xCfTnJJ4sG0Y+OAY1wvQmY/9GXok23zIMo1KYEv3xsqS7IXHUE6S3Q/PBojJt2jRuu+02DjroIPr27Ut9fT377rsvvXv33vvFF1/cw3Xdu9F9s5fV2Q/NRQ8hPrv5ciRC++mf4CnfzfOv4Oc92gvQ2VopVDJ7AUcExlpQlnqwF1wyIkhgfZnUCySdSWw34c0PU/E9cmF4jvhkwW5IhN4zs1DjCKPY90cNipMRicY3mHjr1TWJ3WN9HY0rVfPfX9Ai+BfEJ/YejhZu0i2y/IH2meOvIT/qdFpYBL1O84hfrOiDhOnzEwnQS0mfWl9IHkAv+DG+saD4/AWZN+3LhI87eOwI33YYvUmCDZrCtG+g9xG5ZyBPQh9afxfKZF8Mi5EwHewiOxh58MxFDRWfQ97Sn1FdGWz/zeGYZWj/pRAmvilitgS7nXaPPo//JmUQWvUKgbyaV1555V8F59VXX536+pSFC9Y40DCMfHIjmsh5E/mFaOK3cXR7CSpX+2exAzOMJJwP/B+6SeiOygSHEN/ExTDywQnIws+7Pv4T3WBXc1KHYRjVzwJiWdIe/Yj3kh5EhhaRHWHu3Lnceeed7LvvvgwcOBCAoUOH0qdPn7pHHnnkyCVLlhyCMmvnIuvK7oFTzCBxs7oVEowFm/7lk+8SjK1SwOcrNxIJvxeTufgcJFVy4UHIa9rPu2h+mAmLUAb1Z0h49tgN9ebKRVu6ktTis58xtO/9VYd0w8NJLT57zEBJGKf4xlZBCZCpEmBDtM/AnoWs7bJJxLwNvV5+7/PjgEvKyYLDz6nIXiKRP4xn1TE/j883swPHJuoOnajbbA/aC+kdaQ6yFHns+P1dEl1IPZrRl8UOSWLbl1iDxznI8+UFlE2U6IJZSeTS/CXo/QzyId+ig7Ekep7pvu0JwM2bbbbZ8eussw79+/enc+fk3+1LliyZ3dLS8kqXLl0edl133KWXXmoNZwzDyBeHoQZafpYn5hX4M/reeKWYQRlGGiLoxuFNYAOUFPAAmgibMGjki3ORnZ3H7ejmqpoSOAzDMDy+j/54DQ77IovUfZMekSdaWloYO3YsQ4cOZfjw4YRCIdZZZx2OPPJI7rnnns4LFiw4PcXhl5BYOEt0r1/I5tmJzp1Ku6kmtkILF34+If47NJ8kSlQ9E1WHZ8o0ZEHiT+AMo+/5s7OMZyHZ/a4v0d69AJQs+noW53mCeAEaVNmQSoDeHTXz9nMJuSVxjCZegO4J7FKuAvQilLF7W4LHLqN9dnFH6UiDkETiZqKLSSJRuqNdsX8hcwF6KbArKj04gdT+3yugtPu9UQr980jAzuYNX04k8hNKR6IvpUIQtM0AOHvDDTc8es0112wIPrBo0SKmTZvGtGnTmDJlCtOnT++JPtg7Ia/0CahE4g0s28swjNxZEbgWiSn+xVPvu+Mn5GP2PWo88QvWeNAoH+ahUsc30fxrZzR/PCvVQYaRIUHx+WbkrWjis2EYtcAIVCEXzDItKK+99hpz5sxhn332YdGiRYwfP54FC1Le5n9LYj0JEusmxRaguxbw+cqJ3ROM3UB2gnCmLEv7Rn5foir/bPk78ov227kEvaEz4d+0t4BJRStyOwj2e7sny+dNpJmulWDMT3BBqYWYNU+2vI8sQ9b0jQ0tVwF6FeDSJI/9Hnm65PMC0ZFGfImODXZUTTbW0QaAweMTPYefJSi7/EbgeOSPtFKaY+qQcL0r+hCeTOU1VcnldS54SVEyzj777OOJ+j+3trYydepUvv76a6ZMmcKMGTNw3YS/Tg+0suhfXZyCxOj3oj9vk5sXtmEYtceVaCEu2ffKisT3awBNUn5BjVYuJLMu5YZRKCYDB6OSxzqU/TKJ/PUPMWqPELo2nuEby8RT0jAMoxpYE9ld/LZUAUyaNIlvv/2WxYsX09aWVr98muT3vsHKdOi4NpOKRAuU6bSbamG7wHYEeLBAzzWY9n20Hs/xXN8AHwCb+8Y2RrYc2SQ45nI/9L8EY69leY5ZKDnIr2stl+aYYA+0N4DZWT6vnzeJF6C3KkcBug41GEy2ojYArWQdmMfnXDb9LkkJpsZD4hWORF68iY7NhuAbKNOVlS9QN84z0YdoR/Rm25ZYaXUijkLlNvtQnDLWUl6UE33QrkKlBPmk3UKK67oNoVAovHjxYj799FOeeipTu6B2DIj+eIb/C1BTQk+QfhVdWA3DMPwMRw0osr0Gf4/KtO7CFruM8uA/wCj0vgQtwH+CFmQNIxtCKAHGX856OZl1lDcMw6hk6lASWzNlkLWbJuvZz/HIcuBc2mfAziFeGIP04lxHSHTubDx1K5UQsGVg7DMSa2P5YPMEY+934HzvB85Zj/SzN7I4x9c5PG/QoSECTM3xPH4BOpX+2AtYNzA2KYfn9BP0VV+5HAXoi4Dt0+wzAnm73Jyn51yxA8cmEsoTfaB+oX0pc0fKVuppbxORTWo/0Xg+jP5cjWLbDJWq7oZWq4ICxO7A0eTvtU9Ft/S7FIyfEoytTPavcda8+OKLj2277baXNjQ0sN566zF+/Hjmzu2oWwug1zOYJT0didGedcd7VF6Gu2EY+WMZdH2P0D6DIEhbdJ+vUTn6HZjHrlF+jEE9RQ5Gk/DHUIZMpfe3MIpHHUp+OdI31kj+kxIMwzDKjU2Qx32++yAVi61Qr5KngJHIshISJ5sVUoBOZMdaSMuPcmFZ2leWf1LA50uk603uwPk+TzCWKmEzEbmI7S2BbU9L7Oh5Uum/q9Be+zsI+E0Oz+sRtLpZMVHpQSnZCfms+JmFmiAFvSWvRqsP+SBRs8NM2TCw7SK/oSAR2q8AdCT+9WnvHzytA+cDxfgeyujYAf1uiUoGTktxjlbav9FzXSnN9sOdT+bS/mIR/FsXhPfffz/y1VdffQ0QDofZbLPNCvl0fYE9kXg0Hv3ObyLv10NRBrVhGLXDRcA6pBafvZrHz1Gm9DrArZj4bJQnLvAn1AEd9L03lsQ9GAwjSB3qJO+Jzy6aB5v4bBhGNdMFWaK+Q+WKz372QEl3dwFrkDipLJ01aUdYOcFYLSyEJxKEC5nQl8jbuyPZ1omOzbZ5ZFAby4V8nCMdiXqgrUCsqj6Xn+BrtVw5CdArAfcSnyHsAn9E/j3B7qadkXdM9zw89+akz/RKxpDA9pck/1C9GdjuRXoj8GQETckTnb+jTEIrHl8GxtdDKyTJCK7mJVrxS0cI2DSH4/LJK4HtjYBVi/C8kzfccMPhRMWcQYMGLerevfs1KEO50E2+OqEymVOQFc5X6ML7POAAe1G8Bo2GYRSXjUjd2dkTnt9BTWo3Qo0pCtFExDDyySLgAP6fvfuOk6I+Hzj+mbmjF8GKomJv2LGLGo2xoTFRscTYkogVEW4PETkcDxXk7kBEMWKvUbHErlFjLGhssSModkWsIJy0u535/fHs/nZ2drbe7s6W5/167Qt2tj3Xdmeeeb7PA99Hru8JXBdcOKpMdEZOVkRbmYWRVYDTAotIKaUK7zfIALMLgU7BhpJXJjJPbB7+RVZ+7Rvyxe+5Pyng65UKv6ry1gK+nl9r3Y68nrcVBnS8jW6pyjaxnguzVFpwmMhUR++ZoSnIcgmQJcEHIO03orZC+vmd0sHXXwt5o302y8cNIjGB/GqK+/8XOMaz7TiSD1xM5U8+27LpRZOp5ciZQm+lx/okP2v3M/HVy9vk8Lo7AGvn8Lh8+jfS7zrKAP5M/OTzgpg0adLndXV19xuGcZxpmt3OOuusb5qamkYiOwHbA4OR379B5Pb9zcZqSFuWA13b3AMOX0Ia9Ov0d6WC1xup8FwLOVHbC1lu1RWpZqkhtuO0HDmp1Y7snE3Ef2lWtNXGa5H7PFK48JUqmC+Bo5DP9s7IvuMbyCR2pby6Aw8QG7YVRirpc50Gr5RSpa4P0ITMfSrXAXmrkCK6xWkuuyHH0W6DkK8738MIu+C/8v2tPL9OKfJL/nYr4Ov59dXuSN9yv8dmM4CwnPjN73kN/+4OOSuVBPSFwO88216LbHc7HXljcJ+xOhk5mLi1gzGcSfYJ6LN9tt2X4v4PIdOy3VXepyOJ9mwqW3cksU/210hVWiH4JZpTVYy/R3wD812RP95sGu2fl/4uBXc/0o7E3bcohEz/LXjPppqammbbto8DcBznfMuyrrIsaxWxIYJR6yJLo6IJ6cHkVnWeDe+Aw1bkTHk0If08sUozpVR+dEZOem6JvMduRizZvB6xpHO+hZE+z0uRk8C7Iu03Popc8tKkXqkimI2sqLsmcn0q8AHwXGARqVLUA9ln/23k+irgBCQhrZRSlehoYDqyXxmUVci+bkcciuSG0nkPGOvZtjoyD6sjQ+v8DCbx6wpTmOJBP0Hm/Px6bRcyT+HXiaAjFct+FdwFnwkWkB99tt0LtOTzRUohAb0PsrTfbTEyLMav+fbxSILL/Ud8DZKw/rADcRwdieXFDO+/E4mV198g7UKSmY9MZD/UtW0jZIK2leHr1iDVOt72KddSuP6bfm1CUvUsehP5fkb1BI4Fbsnw9fZDTiwE7RvgRuAc17Y1kOqXP1Lgit/Jkye/EQqFXkCGQfZfunTpUKQthte3SEVitCqxBlkdMAgZODg4cr2QLXd6EhtwGD154B1w+AaFbyGiVKXYGGmHswuwNZJw3ohgPrc7I8nuzZLcvpBYQvptZIf6fbQntCpNM5BVVsOQv6f7kEqoalgKq9Lrg+zL7xm5vhLZh304sIiUUqpw+iGJZ+8q7VysQJJz0ctyn23eyy/A/sA4pJCiI54hs+QzyGreeUhRh9tfyH8C+i8+217DPzkb5ZdnyLVlbDFaKySzGDkecB+/bJHkvvnglxzeBHg3x+fza9WS6udWzn7y2bZxIV7I8VyK+Q1dExmc543h6FQPQqpXvI95l8zK+Qf4PDZ6WUBmQ9fWQ6rBvI8flcFj90feUNyPa0dacaRjIFNova+7iNRv2I3ACHLrl70W8n1xv95npF6Ws5VPjAvwb77vNQhJZvj9fDIdsrjS87jJGT7Oz/rI99cby51IhUwu9gL+SQa9lEOh0BGhUMiJXN4i9+VQvZFE9AVIovoHkv8dFOqyCklCT0NOMAzM8WtRqtL0IjaE92GSvweW0+VX4AVkKefRpJ4boFSxdQL+Q+z39R1y/0xXlaMv0kov+nvRSnz7MaWUqhQGcAaSIHS/532NFBG8BDyKzOi6GmnHGUL64B+DvDdG25GuQW7J0V2Jf8/t6MVvRlYq432e42c61rLBaw0kEe99nXPTPK67z2NSzWlJ5QvP82SaYP/V87hcK2Ff8zzPSgqzYhPgcBK/b1YHnu8Fz3PZpM7hHOnz+t6ZcZm4zvMc3+XwHCAFle7nuSnFfWuRE0Lu+7+T4+umfBGvrkiSKh+ilY9+DKQi1jvUbQbS+iCVK5FE7hGubdshSynPzDpKWT4c7Zs5O/IcDyW574FIVeyGnu3/A67K4LWeQ36h3HHWIG/uOyPJYr++MgMijzvY57ZzkIRiMgOABuBipIz+XqRFQrrBUXsiX6t3Kc4tyC9kMnORn/1g17Z1I695GvCyz2N6IknyC4kdBH7r89rF9jXS9/lh4iuI/4RUJl4O3EX6yaSbIkmYY4n1m0qbTO7Zs+djra2tc5Gk/o6hUOjw5ubmXPqvLkF+Ji8hbUVATqREK6QHITsBHV32lEonYm1CoqJV0tHLi3RsUq1S5cBAesEdErnsTf6GvCxDEtgLI/9fipzkXIHsAIeJtczohvSQPxDZofocOVkYRnrwr0PuQ0e7I6uK9nFtmwM8ATyJ/K2vzPG5leqoNuTE/+vABsjf4+3I53Sq/RtVudZBBi5vF7n+C3AY/vusSilV7tZGjr12IdYXuVgr19ZFjkf/TP56TT+EzNvKxk1I8Yf7+LcvkjPJVz7schKTrUuR3E8qy5B9FffxwXo5vP76JOatMuX9fcg1T/ACkmdwP89hFKat1X+R/Tj379VB5JaEXp3E5PE8KrcCuh3J17lzrNsjucS89oEuZAXU+BSvG/K5/1tkfjZkDWSgjPc50lUSD/B5zFgSK84+RCpnhyNnqCYiZwD8vs5fkOWcmeqR4rmWAvcgfyQjkYF3zyK/EH73vzmD17vV53FLkF+wq5AWIGcgy1HPR1qavJfk9eaQ2c9o9xQxvw3MjHxt10a+vhWe+9xB4pmfICqgo0aRWLnu/l4+EXmdC5AWFOOREyJPkLyacY1MXriuru4UVxX0mxRuKEQPJBk9Amkz8hn+cRfy0o7047wN+X0cSGFbhyhVLN2R/sk3kng2OtOLjewAPI2crB0B/AFJYG9G9hWcfYCPkfeqZBXKnZGd112AIUjly2RkFceHJL7fZnppRVZjnJ3itZUqtJ2Ir/C5KNhwVEA2QFoIRX8Pfkb2Y5VSSuVPJ2CEYRhLyO/xY5jscjFuf/d5vjZk5lZHDcY/f3BZho//xPO453KIwS/nlmkFtPd45e4cXh8kAeyNIdvZa9mY43ktm+RtBFM5h8S4Z6Z5TDlXQIPkPr3xZ1Jkm5VCJpOSJaD3QJbju++7lOz7wQxG3iDcz/ML/j2Lowb4xHk6Uu3bmsPX2Ep8pW+m+iIVYB35/k4ns+ScXwI6l8s85PuXqYYcX+cpZFpsKSWgQfo+5/MDM6ME9NChQ2tCodA8VxJ6SJ6+nkysh5wFs5Ck1zIK+57hd/kFqdyeFIllzUJ+wUrlUQ2y03Ub8hmXze/9UmRHcyKycmJH8rskEKTSr6P99mqRnbohyMnMB0ls25TuEkb69p1KxwaFKJWLE4n/XTwi9d1VhdkImdES/R34Dqn4UUoplT8HmKY5l8IcK/rNSMpUP6T3rfc5Pye7vIfXlvjvD3+N/2A7P/d6HruK7Pbbe0dezxtDpgno9z2PeyaL13YzkAIzbxxDc3y+dOp8Xivb5HlP/L936ZLJ5Z6A7kXi38NyYNscX99XIRNHfgnovvhXVp6YY/wX+jzX6yRfIjDA5/6nR27bBWlIn+nX9w7SNiNX3ZBlGdkm9RYgrSwy1YB/76FML2HgejJMmHqMJvFkQ7KLjVRfR392pZaABtgcabmRrLo7k8snSEI342Us9fX1p7oS0G9QuCrodGqRquRhSFLtA5JXhhfysgD5UB6B/2RhpYK0M1JVnE0i9kOkOvp0JPmR66CRUrEhsmPZjJxA8p4sTnZZhqwCOoL8tSVRKp3JxH4Hl6AzCqrFlsTPolmA/uyVUiqf1u/cubM3kZrPSzuJgwSz5T4R7b58ihz7Z2sg/slLG2k9kanzfZ7jipSPiDGBf/g83iHzBPQsz+N+JPdh6Kf6xPETsbZX+dSXxP7VDunnzEUZyDGZ9/GZtHgp9wQ0yEBQv/xVJvPcUtkW5Jt7bwefKJV7kenibqcglVJu7yO9j3NhAlNI7IlzK/CYz/0HIGe03IYhCVaQJcznINNKk72ZvY60iLiW9L1/M7EB0ubjMJKfXWgHXkH6G/0d/z7RqXRH+mbvjywr3JnU1XQOUhHyAPK9/DDL13PbEjkZMQT/M34rkFYVTcjXGDWM+OEvPwFnZfB6dxH/5vgw6fssZWtL5HfkAGQJb6pk0VLkjf5Z4F/EGvFnbOjQoTUDBgyYQ2SVgOM4h7W0tDyRQ9yFsBrS1ynaS3ovcu8bm6tfkdYu7l7SnxU5BlXdTOQ97jwyG1q1DOkt+ijyvv55wSIrDT2QlUZHIG1DMulHtxDZAZuO/2RmpfLFRPYVovunnyGfa/p7V7m2QVZ2RY8fvkCGwX4SWERKKVU5uq222mrjW1tb68LhcCELCm4gVkzYEdOQfXiv5cgAxmakqC6VLkhx5JjI/70akf7SmVobSWS7v39twAmknpnWG8ltHZvk9rfIrIhyNIkJ74lIgtLO4PFuNchn7v6e7YuR/ODDGT7PWkgB2jxkdkcyFyE/N7eVSN9xb37SrTPysx7u2W4jsb+QJr4jkTaFbrsj+Z9sXIfkwqK+R2ZVZOsb4vOkNyM5rFS6I8eo3rY2C4CTgH9n8fqdkNXA5yPfh95BVVEGKV0C2m1DZAd1dSRZ/yVyUPJ1AeNbD1kOuBaS2PsJGTA4D2lFkC8G0ndz48hr9UT+sJYgZ7feJzaoKl9qkWXk/SKv2YpUnrxH9gn1UtIH2AT5mtZEvretwCLkQCYvvy+hUOg0Ymet3mxubt6VLBPZRbQJsYT03kiSvti9nN0DDqMDUZcXOQZV+foiO77nIicTU/kaqe59DPl9TLcjW8m2QwYwHousPkrlV2TFxTTks1CpQuiNnATfJnL9aeBQkg9sNijdz+BqtyfxBQ1eOyPt3qItvT5CThxmutJOKaVUEuutt95fFy9ePGXZsmWFbqu2CtiK/BQdmUjV66lJbv8ZKRh5DDlhuSCyvT+SXzocST72SfL4q4hVNGfjFiRB62Yj+8U3I8e5vyK5nK2RgsZziLXq+Ab5/rhbxmaagN4g8lhvod1XSOHX1yTmi75F9tf99Iu8tl8l7X+Qr+dfSAFKlIkU4O0GHIUcO3RBThZMTxF7DZIs3svntvuRgs7/EBu0uBrwe6R9h18/8eisr3QqIQENko98A/8OCC8gv5fPI6sE3DohKwB2AvZFvqfRwsSlVGmrxQEklpTn46yZUgUV6QX9UbQVR319/SFBx5SFnsQGHN6LLCMp1FKsZJc2pGXIdcDJyJtjNZ6EU/mxNnAl6WcHLEZOHB2ADtRMZiukKsQ7bMVv6eKjdKz1lVKpbIn8zUZ/51K18DqqKBGpbHUmdYVStLI9+jOeQ+IqSqWUUlnaY489ftOvX7/PKd6x3ZV5/hJMZH80n+0l25FK4lytSfrj5nCS7b8is9ce9GzPtAUHwNVpXtt7eSvN822PFHWmeo7lkft8R/JWrt4KZT/9Sexj7f2+LcC/B7j7ci+Zt/ushBYcUTsjJ1tSfW9WIicdviT9rLR8F7eWjQEkfjM0Aa3KQigU+qsrAZ2quqccRAccTkIqlDvSpzzXy2Kkys2KxFLs1iGq/PREls+l+pC1gceRHshdgwmzLBlIpcK1pE7s20hfu1wmWiuVzsHEz3lINnPjXmSVjyotIZKvPNuX+Pfu/6GDjZVSqkNOOumkbXfcccc3ampqinkM10puFaGZOABZGdPRGN9HVgJ31G6kT5J6L4uItbvoSAK6K9LeNNPXTZeABskBvJLl1+O9ZJKABjm2fz7H17CRkxzZFBBVUgIapJL+X+T2/fNevs/xayh7A0j8ZmgCWpWFYcOGdQqFQp+WaRV0Op2Qg/kRxAYcFnNHJnr5JPL6IyLxlPswuI4Ygiz/qebvQVQn5HvxLcl/d1Ygvzs6xKrjeiN/g6nOvK9CdtDWDShGVbncA1iW43/w8CLSe1KVjnWQJZ5f+tx2KPFDv19HTzorpVTOGhoaBh500EHP9+7dO4jjtcsK/OV1QloVvJVDbK8jvYZzHdjnZyvgmQxf/19IO8yojiSgo36DHON8lua1M0lAgxSdHIe01svme/sjMv8tXdtDNxMZNJmumtd9mY1Uj2er0hLQUQeSWyJ/BVLsdyrQC6pz+fkAMu8BrVTJCYVCfyV20Pt2z549B1mWle0ggHLRD1kuOyhyGUzy3lqF0gq8Q6yX9Avk/iFQbgxkZ2ddpOp3VrDhBOYAZEcgWcXtN8A1kfs5vVrRAAAgAElEQVT8XKygqkQtcAwwkuQ7cK3I7+c0kvfrVSobBlJlf1zk+rdIr/IFrvt8ibTi6Y8OKywVNyEV658Cm7q2H458fkVXpLwQ2ba0qNEppVQFsCxr2/nz5096/vnnD/vqq6+CyCf9giRYi7XPvRWSgNsN2Bw52Rk9Hl2EHBd+DLyKJNs+LmAsg5FVu3tH4lgD2Q/+HPgvsjrrDc9jdiDWExrks+/VDsTQJ/J8vZHeye4K4Wyf20D6BR+MfE39kH2rnsjPeQmSpH4feDby3O2+z5ReLTIj4jAkv7BO5OtoR6pzFyBJ1seRQrhcrEViD+nXyL79xNbI/mXUKtIPQPSzD/EDMRcgrcdytRFyXLx/5P9rICvJHORn/wNSzPch8nXPRk7+V7UBJGbmtQJalY1IL+g50SroUCj056BjKqIapLr0ZCTZ9wHJ+14V8rIAeAQZRjCYym6zsAWx9iivAPsFG05RrUYsqZnsLPwFVPbPv5QciJwISlV1of2hVb50Qw7i3NUw0Z14A+l755DZUBpVeLsQ69s517X9eOJ7SD6B/GyVUkplwbKs7UeNGnX3brvtZhuGUexjL/floiJ8uUoplReagFZlr66u7ihXAvrz4cOHd0n/qIrVC0kCX4AkhX+g+DtCq5BExTRiAw4ryVjiv96n8Z8QXEkOR6Y8+/28W5He5asFFl31MpDe2h+T/G9xEvFn+5XK1QCkKib6+3VzZPuarm1fk99ltip7BnKCINq7+73I9j8jA4ijP6tH0BOGSimVlYaGhj3HjRv32JAhQ+xu3boFmXh2kGrjngX/opVSKk80Aa0qQigUeimahK6rqxsZdDwlZj0kSTUNaZuxguLvIH2LHOxayFKpYrcOyadOwNvEf33twI3A+gHGVQg9gDvw/5mGgRkUbuiJylxn4Bxk+aXfz+pdYMvAolOVZDCxamcHOBs5Aef+fTsqsOgUwEnE/zzeBM4kfvXKPchnmVJKqQxYljW4oaHhkdNOO83p169f0Inn6EWPeZUqY9oDWmgPaFV2Ro0atYdpmi8jf8eLwuHwZlOnTtX+s/66I0vz3b2kNy5yDGGkh1W0l/RspD9SufTv3gXpLeYdSLgcuBqpOi3337/NgAeA7Xxum4+crPxPMQNSaa0DNCEJKK+lSD/Y+4sakapE5wLTI/9vA8YDEyPXw8j7+X4BxKWkEm4+0ncx2ofyS2BD133uRAbg5No3UimlqoUxfvz4w4CLWltb93zmmWd49913g44p6mukB/OKoANRSuVGE9BCE9CqLNXX1z/kOM7vARzHmdTS0nJh0DGVkfWIJaT3jlyK3RdyKVKpGU1I/xdpIVKqmoG6JLctAq4ArkKS0uXmMKTyua9nezvQggy5W1nsoFTGDgeuJbEi3wEmI/0CdUCh6ojrkP1FkDY83iXAOyDv56q4JgJjUtx+HVK1Xi4ne5VSqugsyzJt2x4CNNi2veurr77KCy+8wMqVJbXrewYwM+gglFK5q8YEtEli385laGJBlaHRo0dvadv2+0j/yRW2bW8xZcqUr4KOq0zVIkv2ownpwcgE2mK/T35LLCH9JvA6pfP+1B1JsGya4j7fAI3ATZRPtdl4JMFsera/i/QQfS/hEaoU9UWqVE/0ue0p4DhkorZSyWyMnGgbgAyb/Rp5T/4aOTloIZ8RXjbSkmiYz22qcDZFprl3TnL7fGQ4KcjffjQJvQI5Ufoo8nmrlFJVKZJ4Phq4BNj6008/5cknn+THH3/M9qmWI++tNrF9rV+RlZNbkriCMlvzgW2QVUhKqTJVjQlopSpKKBS6Afhr5OqNzc3NfwsyngqzGrArkoweBOwFrF7kGNqQRGg0If0i8FmRY3DbH3iW9J8f84AG4D6kCrUUGcAU4Hyf2/6BtNz4tagRqXwYhiSivUmp/wEHA1kfVamqUgtciJyYyma44AqgP8G2IjKRVhRrAf0il+j/VydWgNEbSQZ0RwZ2dkKSBmFiRRltSKW3AyxGhjH+EPn3W9f/g/x7+icyY8F78hDk63GQ93mb+J/lS8jPWJPPSqmqZFlW53A4fLxhGOOQtha0trby5Zdf0t7e/t2iRYvumT179uPt7e3Lie0LL0beV6Mn8UBWQCazPvAK+ZkX82eknZJSqoxpAlqpMjdy5Mj+NTU1HyEHkmFg5+bmZl0GXDibEEtIDwJ2o/iDjb5FktHuftLFbHtxM9JPMxOvIcujnytYNLkxkHYh53q2twPjkHYiqnztgvR+3tCzfQ5wIPI3pFQquyIHu5tn8ZjRSE/yQusLDESqwTaJXAaSnyqzbLUBXwGfIn9fH0T+H70Uym+BZzK8r40kqeciJxZmFSoopZQqZaFQqEf37t3/BoRITAy/6zhOy9y5c++cNWtWR9uWrYEco2zVwecB+VzZHm2lpFTZ0wS0UhWgrq7uMsMwxkauPt/c3Lw/pVt1Wml6AjsSa92xLzIYrZjagY+IT0jPoXC/A6tHnj+br/MxYCyl0SO1Blkuf4pn+4/A0cALRY9IFUI/JAm9l2f7B0gSemHRI1LlphsyYHU4cuCbKrnrIC2INiK//cY3AHYH9kBOeO5EYv/pUrUIaSP1auTyGvmZc1CLfJZsQfqfiYG0UJmAvO9rL3ilVNUZPXp0r65du/4FuABY13Pzy8CkxsbGR8nPsUM34GnkuCgf/oiseFFKlTlNQCtVAc4+++ye3bt3n4ss/8UwjOObmpruCTisahYdcBjtJT0I6FrkGH5BDvyjrTteBn7K4/MfB9yd5WNsJCF4IfBJHmPJ1o3AXzzbFiJJyQ+KH44qoB7Aw8ABnu1zkcR0qqWjSkUdBNwGrE36feeOHCgbSJXXQcjv527I50kl+QRJRr+M9Gafn8NznA9MTXF7NPH8MzKE9EpKZ5aCUkoVzYUXXrhGp06dhgPnkThoezZwRWNj4yN5fMkaZJXJH/P0fG8gn4VaWKVUBdAEtFIVoq6u7kTDMO6IXP162bJlW8+YMaM10KBUVHTAoTshHcSAw0+JJaTfRJIAHRnm8SDwhxwe14a08RgPfNeB18/FKKDFs20hspx7TpFjydWBaW7/GPgiD6+zF9LaJ5lvgA/z8DqF1gW4F/i9Z/vTwGGUz7BMFay1gJnIe160pYNXGFlB4T3hkUpf4HfAIZGLtzItWz8j76vfI+9t3yFVxwuJDYaKDuRz93s2SewLHa20XhP5+teOxLe26/+9OhjvfOBJ4AngP5GYUlkDSWL3xv8z1Ik8x9XAZcDSDsanlFJlx7Ks9W3bHoXMxejhuslxHOexmpqayyzL+m8BXvpq4Jw8Pt9ByP6aUqoCaAJaqcphhEKh54D9ABzHubylpeWigGNSyfVDeoxGe0nvTWJlQqG1Au8Qa93xAtklhNdDKob7dOD1rwEuB5bk+BzZ+B3wOPHDqL5Eks+5VOEFJV0VyFNIIqsj+iPfG78kW9QNyKDGctAZqdj3VuRMAeqKH44qYycDM5BVLcnaP2wPvJfiOdYFjkda/uyR4nmSWYy0XZoXuXzkuhRzHgDIYMMtIpctXf/fgvikRyZWIJ9D9yCrZX7xuY/fChaQhPoqYBrSw19XNyilqo5lWZs4jjPCcZxhxK++tB3HebympuYSy7LeKNDLNyIDyFNZgBwDpdq/jHoJ2KejQSmlSocmoJWqIPX19Ts6jvMGcjC7yrbt7aZMmfJR0HGpjNQggzrcCemdyGwHLZ+iAw6jvaTfQJICyZyFJGM64kegmcIuk94Y6T+6pmvbIqS36scFes1CSZeAtpEEckd6HI8BLkESt8liuJHySUCDVHQ+TeQkncvpSDJdqUxtBNyBvE9H2z1EOcB1yHujWzfgcCSBfQjxJ8JSaUP6HbtXr5RLq6D1iF/5k007qpXI3+ttwENIcnl3wFux5yCV5zch71kLOhy1UkqVmYaGhkHAGMMwjiL+2KENuMs0zUmWZc0tYAjDkM++VJYgs3L+S2afBb8Bnu9YWEqpUqIJaKUqTH19/bWO45wZufpIc3Ozd9m5Kh+9gB2IHcDvjiyDLqbogMNoQvpN4gccmsiy6XxUKHyBVEPfQH4nXXdBks/bu7aFgSFItXC5SZeAXglcRGKrkWx8iiTtU8VQbglokMGZrwEburatQFYjvB9IRKpc1QAh4FLkfdB7wN8PaYfxG+TA/EhSt7SJWgo8CzyD9El+j8ppE9MV2Blp73Mw8rnRJYPH/Yy00dkdGfobPX5xgH8g7ZyCnCuglFKBsCxrcDgcvsAwjCHE53ZWAveaptloWVahV/kdATxA6hOrq5C2Z88iq3hWS/OcjyP76UqpCqIJaKUqzMiRI1evqamZR6TS0zCMIU1NTY8HHJbKH29F2S5kdgCfTwuRyuhoNd63wIvkb9DiG0gF7rN5er7LkcGHbnVI+4VylMkglvnA5jk+/17IyYZ0MZRjAhokgfUS8e0B3kKSWx3pia6q0y7AXST+vT2IvF/vnsFzfAo8CjyCvJdWy8C87sj7zYFIj/ats3js/5D3n/8VIC6llCplxvjx4w9H9m339Ny21DCMm9va2iZdfvnl3xYhlj2R/fVuKe5jI8PL74tc/57UBTUO8tmq7+9KVRhNQCtVgerr689yHCfaFuHjlStXbjd9+vRqOaCtNp2Qyt5oQnpvYJMixxAGfkKGUuXTM0gi+s0OPMf2kce7qzLuAk7swHMGzS8B7W0DALLznsv37nqkTYC7/YZ36Fo5J6AB/gTc6dkWomNV46p6dUfaCHnbbqTyFtLG417g60IEVYa2Rnpjnwhsmua+HyP9nm9F5gkopVRFGzZsWKd11lnnBMMwRgMDPTd/D1xrmuaVlmUtLlJImwCvkH7//3zk/TrqK2D9FPefBRzbsdCUUqVIE9BKVSDLsszW1tZXkQQUwCXNzc1WgCGp4lqP+F7Se5HZ0u9S5CAVE2PJbVDgv5Dhg1ELgG0p7wFVmVRArwJmAsOzfO5uwA/EVwevQhLQ7gr3ck9AgyT+hrquL0UGp3Wkd7aqTj2Bc4CLSV0F9jWyTPkWJAGtkhuEnAg7Dmmdk8zPwGQkuZFqXoFSSpWl4cOHd+nTp89xhmE0AJt5bv4MmLZ06dKZU6dOLeYQ2jWQNlFbpLnf5UhbOLePSfw6osLAdsCHHYpOKVWSNAGtVIUaNWrUXqZpvohULa50HGfnlpaWOUHHpQJRC2xJLCE9GKk0K6fPgDbgZiTBk2mC8FCkh5zbkcDDeYwrCMkS0KuIr1peiixxzGb1w4lIcszbx2858Ym1SkhArw3MA/q4tl0DnBtMOKoMdUV+Xy4EVk9ynzDynnMVMkwpkxNIKqYXkrzYHemnncwXSC/oO8jvDAGllArE6NGje3Xt2vUvwAXAup6b33Mcp7mmpuYuy7KKPSegK7JKce8097sTOInEz70PgG2SPOYW4LSOBKeUKl3llHxQSmUpFArNILYk+NWePXvuZVmWHpgpgN7AbsRad+yJVDOUulakTUILkmBN5SXid46fRXqNlrtkCSxvG452ZDn7/Vk89/PI74Tp2V6JCWiAkcT3Al+JLCldEEw4qkyYwCnAJcAGSe6zHBlUeASyEkN13E7IUu7jiT/Z5vYe0rpJZ18opcrS2LFj162trR0JnImchHN7yTTNiZZlPUEwJzRN4B7gmDT3ewIp+vCbrfE/5P3cqw3YCpmLoJSqQN4DTKVUBVm5cuUYYr0ld//111/PCDIeVVKWINULFpIgWRPpuXkKUqk3G6moLTU9kSro+Uh7iWRJiH2JTz47SI/fSva957pBdgniDYF9iN83WEll96e9GqmcjOoCnBdQLKo8bIWcqLkJ/+Tzt8AooB+y0iTVoCWVnbeQz6iNgMvwPwm5HfAY0kO0X9EiU0qpDho3btymF1988bTa2tpPgHrik8+zTdP8XWNj4z6WZT1OcKtpmkiffP4f0sM52WDnZCvzrkeTz0pVNK2AVqrChUKhIcCjkatLTNMcOHny5EpOKKn86YFUKERbd+xL6l6cQfgUaADuJn7Z9R3EDxp8BPh9EeMqpGQHHZcDdUgSNcpGBr1kMgm9AVnq7n38FKS/bSVWQIN8bVe7rv+IfM90cKtyq0X+vizi+6FHtSItXJIlRlX+rYEkaUbg/zNZjCxdvx5tfaKUKlENDQ2DDMMYgQxIrnHdZDuO83hNTc0llmW9EVB4bmcAf09zn2+APUhdvPAfYD/PthXA5mkep5Qqc5qAVqoKhEKh+4GjIlcfa25uPjzIeFRZiw44jPaSHoT/gX+xfYAsh5+F9PT9lvi4fgv8O4C4CiFZImVLZGiLt4J5HNCc5jkN4Evip5KHgaeA54BGKjcB3R05YHL3gj6G7FqXqMq2M/L7vqPPbSuAGcgJoJ+KGZT6fxsh71En4r+6819I4uTz4oWklFLJWZZl2rZ9BHISzdtLeTlwi2mazZZllUpF8BDgIeIT5F5LkJV076Z5rqeAgzzbmoDROUenlCoLmoBWqgrU19f3cxxnDtAXwHGco1taWh4IOCxV/jZDKst+E3Acbi8jrUXGu7Z9isRaKRVwyb6OHkjPPW8P5/lIVUkq+yIVKd4e0scirVkqOQENUgF9juv6XcRX0KvqdQ6yCsCv3c8jyBDCL4sakUpmB2AmMt/AazHwZ6Q9h1JKBcKyrM7hcPh4wzAuIHEQ3y+GYdxqGMYVlmWV0iyKXZB9xB4p7tMGHIbsg6fzEPGrEpcg+5o/5hifUqpMaA9opapAU1PTQuDC6HXDMK4ZM2ZM3wBDUuWtFlnW/B6llXwG2Iv45DPAvVRO8jmdmcS3IgFJvu+S5nF/IbFX3zKqZ5DX3Z7rhyC/56p6dUVOslxNYvL5Z6Si9vdo8rmUvIMM1D0DaYni1gc5YTAJPf5RShXZ6NGje40fP36EbdufGIZxK/HJ58+B85ctW9b/kksuGVFiyeeNkVaOqZLPDlKQkEnyGRJbnE1Fk89KVQXdAVOqSjQ3N88EXoxc7dfW1pZuSb5SfnYCXkUO4kuh9UYmXg46gCJ6gMQd+1XAqSke0wOpdHYn2VYBN/s8V6V6hfiDn9WB7QOKRQVvA+AF5MSM1yxkEOHMokakMmUjP5vtSUyGGMjJ00eJrAhTSqlCGjt27Drjx4+3unbt+iVwJfGtzt5xHOcU0zQ3b2xsnNbc3PxrQGEmszpSiJBu/st44NYsntc95PwnZJWRUqoKaHWPUtXDcRxnmGEYbwFdDcP4y6hRox6aMmXKw0EHpspCT+BSZLl5qv5vpegfwDRgMvBLwLEU2nKkmvckYgnlzsDJyAA1v4TyUKCTZ1tn4JbChFiSwkjC8SjXtt2RSe6quuyOtGlYw7N9CXIi58FiB6Ry8hnSY3QE0lvUfcxzKHIi9SC0L7RSqgAsy9rMcZzhjuMMI7FgYzZwRWNj46OU7gq9zsROuKZyE3J8kA33vugVyOerUqoKaAW0UlWkpaVlruM446LXTdO8YcSIEenOait1CNJuYwTll3wGqfAdi/SCvoDyqdzO1U0kJpS7AUckuf8wEvcHPgLeznNcpe5Vz/UdAolCBWkwMrDOm3yeh7R20ORzeXGQisP9kcG0bpsjJ53S9cdXSqmMWZY1ePz48ffatj3PcZzziO1z2o7jPGqa5u6NjY2DGxsbH6F0k88G0oLqgDT3ew44K4fnjyagvwWuyeHxSqkypRXQSlWZXr16TW1tbT0c6d27VqdOna4D/hBsVKpErYn0Zftz0IHkyepI65CzkeWCt5PYL7kSvIxU9W3s2mYi/fnu89x3Y2AP4ocPrgRmFDC+UjXXc31j33upSrUv0pqhl2f7w8gKgkpfPVHJXkL64M9C5gRERVut/A54P4C4lFIVwLIs07btIcAY27b38ty8ErjXNM1LLcv6KIDwcjGJ9Pv+7yOrxlaluZ+faAL6UmTeiFKqSmgFtFJVxrIsu7a29jRiy52OrKurOynImFRJOhGYQ+Ukn902RNpLvAMcGGwoBXMD8QcFJvK1ruu536kkHjzUAncVLLLS9bnn+oAgglCBOAR4ksTkcyNyglaTz+VvAVIJfadnez+kV/S2RY9IKVXWLMvq2dDQMMy27Q+Rk5Xu5POPwCVtbW39GxsbTy6j5PPpwOg091kAHAYszvE1VgJfIFXWSqkqYqS/i1KqEoVCob8iSSqAX2pqara/4oorvgwyJlUSBgDXIj0yy0E70Bq5LEQSrP1dt/8TWVK/2OeyCFhRzGDzJNmSzR7EKkn6A18Sf6J5JTAOiA4gNYCviP9+hYEniG/XEUIScd08MdyIHKhUiv7A167r3wLrBRSLKp4dkX6c3T3bxwMTih+OKjATuJ7EAZPfArsC3xQ9IqVUWbEsa31Xf+c+npvnO44ztaam5hbLssqtuvdQJJGeapX8UmAfpIgjV5cgbfGyGVyolKoA2oJDqSrV3Nx8YygUOgxZPrWabds3IctQS7UfmSosExiOLIfrWeTXXoR/ctgvaey9rdXzXHcg1dtRDyCtNqrNN8B/gP2I9e3uApxJLAF9AIkV0SDJmWrknT7fI5AoVDGtgxxsu5PPDjAK6R2sKo8N/A357DjPtX1d4CEksbI8gLiUUiWuoaFhB9M0z7Zt+2QS54m86TjOVTU1NXdZltUeRHwdtDNwL6nzQ23AMXQs+QwyV+afHXwOpVQZ0gS0UlWsU6dOZ7a1tQ0G1nYc57f19fXnNjU1TQ86LlV02yJJxz1yfPwKJCnsvixPst17+R6pYs6XsOd6OQ5NzJcbkL62bpsiVX6vA39Fvl/uKumlSAV0NfLuE5XjAaTKXGekJ/oGnu3nUn490P8OnJHktp+Q6v6VSW7PxqvAbkluewE54VUOHOB8ZBXIcNf2Qcj38pQgglJKlSbLsgaHw+ELDMMY4jiOewW57TjO4zU1NRMty3o5sAA7bgDwGKkLUBzkc+ZfeXi9+9GCJ6WqkiaglapiEydO/KG+vv5sx3HuA3AcZ1JdXd2zLS0tc4KOTRVFF+AipApsGdLzOVnVsbvy2Fud7E36BslbEV3NVawPIkkn92f9KuA0YB6y+qGT67aVwM1IhUs18h54eSuiVWW5Ghjs2TaF8ks+p7MGMARZDdIRW5I8+VyOoknoTZDvT9TJSHXflCCCUkqVBsuyOofD4eMNw6i3bXtbw4jrXNpqGMZd4XB4yqWXXjovqBjzpBeyEqhfmvs1IvuI+aDJZ6WqlCaglapyTU1N94dCoTuQYXPdDcO4Z+TIkbtNnTpVl6BWPge4DOl1Wil+9lz3azFRLVYgwwRPQao9ifz7Z+ADEgcRd6G6+/F5f1d+CiQKVQxHkti//GngggBiKYZT6XgC+q95iKPU2MAJwCvAQNf2K4DngTeDCEopFZwLLrhgtS5dupxq23a9YRj9PTcvBK4zTfMqy7K8+5vlqAbZT9w+zf3+gfRtVkqpDtEEtFKKZcuWndW9e/fdgC2AbWtra68k+XJeVTlWBR1AAXzmub5JIFGUjpuQfqdu3YBJJO4DfEDH+/qVs409172/S6oyrAXM9Gz7GDiOym27cihS3bYwx8fXIieuKtFS4I9Ie5G+kW21SAuj3ajeFSFKVRXLsrZwHGek4zgnkziU9m1gimma91iWVUn7zlcBh6e5z3+QlXNatayU6jBNQCulmDFjRuvo0aNPtG17NtDZcZxhoVDo+ebm5ruCjk2pLM33XN8hkChKx3+RRKo7EW+S2G5iFXBdsYIqUTt5rnt/l1RluAxY23V9FTJUaVEw4RTMYqBP5P+1yHDWlhyf62DiVwi4n7sSfAz8BWlbFLUjcA46jFKpimZZ1mDbts+zbfsoEueGzAauaGxsfJTKS8DWA2enuc8c5ARdPmYIKKVUwvJbpVSVmjx58huO44xxbbpu1KhRWwQWkFK5eYv4ntRbUVmJklzMJP7gwe+z30SWYVYz7xDONwKJQhXSDkii0e0S4N0AYim0ezzXT+vAc3kfe3cHnqtU/RO407PtYmDNAGJRShXQ8OHDuzQ0NJw8fvz4d23bfhEYSiz5vAq43TTN7RobGwc3NjY+QuUlnw8HJqa5z0Jk9cziwoejlKoWmoBWSv2/lpaWK4GHIld7mqZ5p2VZnVM9RqkSsxSp2Igygd8FFEupuJ34YYNe7cj082ruedwH2N2z7ZUgAlEFNYH4Cre5QFNAsRTaPwD3LIeBwKAcnmcNEpdo355rUCVuFPCL63ofIBRQLEqpPBs7duy6F1988YS+fft+ZRjGrcB2rpt/ACa0t7dv2NjYeLJlWe8HFGahDUJOInqrvd2WA38AvixKREqpqqEtOJRSbk5tbe1p7e3tbwEDgF1aW1snAnUBx6VUNp4i/qDiSGBWQLGUggXAv4H98T/gMJB+p9XsUOKT9B8CXwUUiyqMHUhMpNZRuT1+f0FOKB/v2nYK2Q/WOxEZUBr1GvBex0IrWd8jLVomu7adHbleCQPHlKpKDQ0NO5mmeabjOCc7jtPVc/NHwAzTNK+3LGtZEPEVUX/kc6FHivvYyPv+q0WJSClVVYygA1BKlZ76+vp9HMf5N3KSygH+2Nzc/FCahylVKvYFnnddXw6sR+UsI0y2FLQHkOzg6TjgDvxPPC8C1iF5Ii4ENCLDC90x3Aicni7YMvEUcJDr+hXAmCT3VeXpZuBU1/VXgL2CCSXv/k7i4OCdkL/rJ13bfkbeC7Pp5/k/4vujn428lyzx3O8FYL8snrdUdUP65q/j2nYB8UlppVSJsyzLtG17iOM45xmGcaDnZttxnH8bhnFVhfZ39tMLeAnYPs396oAphQ9HKVWNtAWHUipBU1PTi47jXBK5agA3jhkzZqMAQ1IqG7OJXzbYDTghoFhKxUNIIr4NST5FL+1IIrlSq0AzsQngPTitxB631aw30uPT7fIgAimyp4mv5F+dxCrwVLYjPvm8Crg3D3GVsuUkDh78G1q0o1RZGD16dK+GhoZhtm3PAR72JGg6+/EAACAASURBVJ9bDcOYaZrmdhMmTPhdhfZ39lODzPlIl3y+AU0+K6UKSFtwKKV89erV6/LW1tb9gQOANdrb2x8YOXLk3lOnTl2e7rFKBSyMVDte7NoWAq5HEq7VaAWSgBvgc9sTRY6l1NQTf0L+NeDtgGJRhXEM8UuOvwQeDyiWYrKRauULXdtOAe7P8PHegY0PIr3ie3U8tJJ2IzKcMjoDY3NgT+DlwCJSSqVkWdYmtm0PQ1aDeIdPfwvMNE3zKsuyqrGdzlWkP/n4FHBWEWJRSlUxTUArpXxZlmWHQqETkH6R6wM71dbWXg/8OdjIlMrI9UgLhWjv0k2Qnna3BhZR8J4KOoAStAHxbRkArg4gDlVYv/dcvxVJzlaDW5D3wmgF76FAP2Bhmsd1Qt4zvc9VDX4AHkZOXEQdgSaglSo1hmVZv7Vt+zzbtoeQuLp7NjDNNM0HLcuq1gKEeqR1UipzkHkB1fo9UkoViS4nU0qlNGrUqJ1M05xNpP+rYRjnNTU1TQ84LKUycR0wzHX9G2BL4NdgwsmbXHpAd0Ql94D+B/FD2j4DtkAPwipJZ6TPeXfXtp2orCr3ZD2go1/jbOL7XYeAljTP+UfgAdf1BcCGyAqTXlRuD+ioE5Al61HvIoMslVIBsyyrZzgc/pNhGMOBbT03twH/NE3zSsuyqv2k0eHAP/EfQB21ENgD+KIoESmlqpr2gFZKpTRlypS3DMM4M3rdcZwp9fX1lXSQqSrXROKHbfUHGgKKRZWe/ZDhjG6XocnnSrMj8cnnr4F3AoolKN6VH6dl8JhTPddvQ5LP1eJJ4qvkt0V6iSulAmJZ1mYXX3zxVNu2vzYM4zrik88/ARPD4fDGjY2Nx2rymUHIPItUyeflwB/Q5LNSqkg0Aa2USqupqek2pKUBSOuee0aPHr1+gCEplYnPgamebfXA3sUPRZWY7siwHfdKsLepnhYD1WRXz/VXqI6hU253E786YiCSnEhmbaRVh9tt+Q6qxC1ClqVHmST+LimlisCyrMHjx4+/17btuY7jnA+s5rr5bcdxzjBNc8PGxsaxl1122TdBxVlC+iPDp3ukuI+NtFl6tSgRKaUU2gNaKZWhJUuWnNO7d++tgcGO46zjOM59w4cP32/69Okr0z5YqeBcjvQtj54wMZEk4+5ANQ6iUeJaYDPXdRsYTnVVeFaLrTzX3wwkimAtQQYIuns6n0Ly78VJSA/oqFeADwsTWkl7g/gKy62AZwOKRamqMnr06F5dunQ5wTCM82zbHui52XYc5/GamppplmU9E0iApasXMmS3f5r71SOfC0opVTSagFZKZWTmzJlto0ePPsFxnDccx1kH2L1r165Xkdh3UqlSshRZbv4vYtWumwH3AodQvu0W2ogtDzeJTxYVQhfi25mU8/5DPXCyZ9vVwEsBxKIKbxPP9fmBRBG8W4lPQJ+I/C34nUT2/n1U6/DWTzzXNw4kCqWqiGVZm9m2/Tdkhkdfz82LDcO4zTCMFsuyvgwgvFJXg/Su3z7N/W4EphQ+HKWUiqdDCJVSWRk1atRepmk+hwx2Ahje3Nx8dZAxKZWBa0icAj4NOD+AWDpqWJLtN1GYhPr2yIAar7nI4LFycijwCPE9ET9EltaX+3BK5e8N4ttN7EHlLTlON4QQ5ETVZ8ggwahjgPs9j9sFeN11fQWwLrDYta0ahhCCJOLdyfe7iE/iK6XywLIsEzjAtu1hwFEk9i1+y3Gcv9fU1NxhWVYhhi1XCu/wbT9PIcMJy7UAQylVxjQBrZTKWl1d3QjDMK6MXG03DOOIpqamJwMNSqnUOgPPAPt4tg8j1t9cVbaBSJVzH9e2RUg7lo8DiUgVw1xgS9f1bai8dhKZJKABJgDjXNcfAX7vuc/VwDmu635J12pJQP8ReMB1/SFkYJdSKg/GjBnTt3PnzqcCZwGbe25uMwzjfsMwputAwYzUAc1p7jMHmYOyOM39lFKqIDQBrZTKSSgUmoHsMIK0ORjc3Nz8boAhKZXOGsBrxC/Jd5Dev9cEEpEqlm2QExDruraFgSOAJwKJSBXLp8S3Ttg0sq2SZJqA3gz4iNj+fzuwAbAwcr0z8A2wpusxByMtjNyqJQF9KNJLNeoppHWTUqoDLMva2XGcMxzHOZHEQXnfAzeHw+FrLrvssq8CCK8cHQY8TGLluNtCZAXQF0WJSCmlfJRzD0elVICWLFkyonfv3psDByIHow+PGDFi92nTpn0XcGhKJfMTsrRzNrEDHgOYjnweTgsoLlVYOyOJozU92+vQ5HM18PY47hpIFKVhPvL+NzhyvRapbm6JXP8D8X8n31DdQ/e6ea4vDyQKpSqAZVldbds+Djjbtu3dfO7yOjB90aJF9+qA86zsBNxD6uTzcmRFhyaflVKB0gS0UionM2fObBs+fPjRXbp0mY1MiR/QqVOnRy3L2k/7s6kS9g7wJ2AWsT7mBjAV+UxsSfI4VZ72QJLMfTzbr0JPOFSLVs/13oFEUTpuIZaABhnSGn3fO9XnvuGCR1S6vL8r3t8lpVQa48aN29Q0zdNt2/4riSeCVwIPm6Y507KsZwIIr9yti7QG6pniPjZyovG/RYlIKaVS0AS0Uipn06dPXxIKhX6P7NSsDezS2tp6C3Ac0tpAqVL0MFLpdz+xCjcD6Z23BdKSY1Uwoak8OgHp7+1d3tsC1Bc/HBWQBZ7rG1LdB+L3AFcSS1gMRIY0fgsc5LnvHUWMqxQN8Fz3/i4ppXxkMFTwE+B60zRvtCzrx6IHWBl6IH38N0hzv9HAg4UPRyml0tMEtFKqQ5qbmz+rq6s72jCMZ4AuwND6+voPm5qaLg46NqVSeALp7/ko8ZUjw5BkzNHoUsVyVQtcClzgc9sVwJjihqMC9pnnunfQVbVpRZIRJ7m2nYq023AniWYjAxyr2Wae697fJaWUi2VZa9u2fZpt22cQ33sfwHYc59+GYcz88MMPH5g1a1Y1r67oKBO4HdlfTeUmdGWfUqqE6BBCpVRe1NXVnWQYxq3I+4rjOM4pLS0ttwcdl1Jp7AM8hvQxd/sBqaCt5v6n5Wgt4G7gAJ/bNPlcnYYB17muP4SsgKgkmQ4hjNof+Lfr+k+RyxaubacDNyR5fLUMIZxH/PfkN8DzwYSiVOlqaGgYZJrmMMdxTiKxd/r3wM2maf7dsqzPix9dRZpM+pVczyOrWnRFn1KqZGgCWimVN/X19RMcxxkXudrmOM6RLS0tOuRLlbrtgQeATT3b25Cd/AkkDjJTpecY4GpgHc/25cCZwG1Fj0iVgh2Bt1zXv0f6ZtrBhFMQ2SagDWQg4SZJbl8OrAcsTnJ7NSSg10HakkSPldqRXvK/BhaRUiXEsqze4XD4eMMwzkLeZ73edBxnZmtr6+1Tp07VAZ75cxpS2ZzKXGAvYFHhw1FKqcxpAloplU9GfX39nY7jnBC53mqa5m8nT578WqBRKZVeb+BW/Csj5yPVgP8pZkAqY+sgiedjfG77Cmmn8npRI1KlpBb4EVjNtW1PKqsPdLYJaIBLgPFJbruD+BYdXtWQgPYmed4Adg0oFqVKhqva+UQSZyy0GoZxl23bMyZMmPBOEPFVuH2Bp4kN0fbzE/IZ93FRIlJKqSxoD2ilVD45v/zyyym9e/deHTgY6Gnb9mN1dXX7tLS0VHsvSVXaliCDckYDlyP99aI2Q5arXw+EgKVFj04lMxS4FljD57bngOORildVvdqBJ5HhuFFHU1kJ6FzcDDTgX4xyS3FDKUlHea4/FkgUSpUAy7J627Z9ItLSaEfHSZgz/j7wd9M0b7csy3tySuXHpsjw7FTJ5zZkv0iTz0qpkqQV0EqpvLMsq3tra+szyBl4gK9t295rypQpXwUZl1IZOhRJNvf3ue0boBGpjGsvZlAqzr7ARGSJqddKpG3KJECHHCmQfu53ua5/B2yAHKxXglwqoEFWdXirlr8GNiL1306lV0D3R4bQuocy7kx8KxelKp6r2vlPxA9sBvmsfdg0zZmWZT0LJGSlVd6sjpw0TTVE1wFOQYYTKqVUSdIEtFKqIOrq6tY0DOMlYMvIpvdra2v3nTRpkvYjU+WgN5LEPJf4auioj4BxwH3oQVcxbQNYSIWPn/8CfwXmFCsgVRa6IyeP+ri2/Qn4RzDh5F2uCeidgV082z4ifbuhSk9AT0De36Pewb/HrVIVZ+TIkd169uw51DCMEch7hNdHwE2mad5oWdaPRQ6vGnUCnkKGx6YygeRtlZRSqiRoAlopVTChUGhjYDYy8AnghZ49ex5sWdaKAMNSKhu/QaqhN0ty++vIDv9TaCK6kLZE2gWcgP8JgV+RhNFVVNZwOZU/1wBnu66/B+xAZfzd5pqAzlUlJ6BXAz4n/mTFOcCMQKJRqkgsy9rGtu2TkTYbfT03a7VzcGYic0hSuQ84Fv25KKVKnCaglVIFNWrUqB1M03ye2ACoB7/44ouhs2bN0qXxqlx0Q5Kb5yOVlH4+AK4E7gR02nv+HIh83w8j+T7LP4FRwGfFCkqVpc2BD4lvq1ApVdCagM4fb/Xzj0hLkl8DiUapArIsq3c4HD7eMIy/4T9kc57jODPb29tvnThx4k/Fjk8xFrgszX3eQN57lxU+HKWU6hhNQCulCq6+vn4/x3GeBLpGNt3Ws2fP0yzL0kpFVU7WQhITZ5N8iO8PSH/oq5Feqip7nZHhgXXA9gMHDmS77bbjpZde4uuv476lrwBjkMSXUpm4C6mij/oS2IryP2mkCej8GICcpOjm2tYAXBpMOEoVRprezquAh7TaOXBHA/fiv+or6gtgd2SugVJKlTxNQCuliqK+vv5ox3HuIVZ9dl1zc/NZ6I6tKj9bIgmJo0n+OboKeBKpiH6E8k9wFZqBDBQ8EVlGukb0htNPP51115UuPl999RWvvfba5x988MG5wGNBBKrK2qbIaoUurm3NQH0w4eSNJqDz43FkCG3Ud8AWJH6tSpWdMWPG9O3UqdNQwzDOAbb3uct84AbTNG+2LOv7Ioen4g0Cngd6pLjPUmAw8G5RIlJKqTzQBLRSqmjq6upOMQzjJmJn869qbm4eEWRMSnXALkAISUQnq4gGSV48ANwBPIf2KHbbGmmDcCKwsffGPn36cPrpp9OtWzfvTa+ZpjkReFhXUqgsNSPV9VE2kjh9KZhw8kIT0B33F+BGn203BxCLUnkxdOjQmq233vogZDjv75GBdm4rDMO4zzCMGy3Leh4tCikF/YFXI/8mEwb+iBQ4KKVU2dAEtFKqqOrr6//iOM4NRN5/DMOY2tTUNCrgsJTqiA2Ac5EhMd7BPV4LgScil2eARYUNreR0AvZGqgwPBbZLc//2bt26PXL88cd/ucEGGxwNrO+5fb7jOE3ffffdzTNnzmwrQLyq8vRGKsYGuLYtQPqfLggkoo7TBHTH7Ay8SHyP/xeRIbR6gkuVHcuy1rdt+0TgTKSHudcc4DbTNG+0LOvHoganUumFvPfskOZ+w5FWb0opVVY0Aa2UKrpQKHQeMC163XGci1taWhoDDEmpfOgJnAKcg1T2ptMO/BdJRj+FJIsqcTjnRsDvkITzb5EEYDo/IZWHVyM9DrEsq3NkWNJYpA2K2xfAVNM0r7csSwfxqHQOQE4AufeD/wfsQ3kOctqHxPec+5G/o0LoBJzm2fYt5VmNtw7wOnIiMWoZsCPwcSARKZWD4cOHd+nbt+/vHcc52TCMQ4kfuAqwxDCMuw3DuN2yrHJe8VGpTOBBpFI9lenAeYUPRyml8k8T0EqpQIRCoVFAS/S6YRgXNjU1TQowJKXyaSBwEnAysG6Gj/kVSUK/6bp8UJDoCqcnkrgZFLnsg3/1lZ8VSFLwNuAhpI92gsiS4mORAYTePpbfOY5zZU1NzQzLsrRvq0rlcuBCz7bbkb9ZVR26AP9G+s+7nQrcWvRolMpBQ0PDTqZpnuo4zom45idEOEhF7Q1Lly69b+rUqTqPonRNI31i+SngcKSAQSmlyo4moJVSgamvr5/gOM646HXDMOqbmpqag4xJqTyrQap+TwT+QGbVv27fAG8B84CPIpe5SCuPIPVGhnNtAWwV+XdbpAoz1cR2rzDSF/tOpE92Vkljy7IG27ZtId9jt6WGYdxsGMYVlmWVa1sFVVgmcqLjcM/2S4GG4oejiqwWed851rO9Bentr1TJcg0UPANpIeO10DCMew3DuN6yrPeLHZ/K2t+A69PcZw5ysuyXwoejlFKFoQlopVSgQqFQE7GDPccwjOFNTU3XBBmTUgXSGZlYfkjkkq7/cSpLkGT0V0jf2h+A75Fl8N8D3wE/R+67mMwGC/VCkjLdgX6Ry1pIBffakcv6wOZkXtXtZyHwZOTytCvOnFmWtW84HL7QMIxDPDctA24Mh8Mtl1122RcdfR1VcXoBLyMnT9yagfrih6OKpDPwD+Aoz/Z/AYdRma2QVJkbNmxYp379+h3mOM6phmEMIXGgYNhxnCcMw7jBNM3HLMvSKtnycBDwGKmHWf8I7AF8UpSIlFKqQDQBrZQKXH19fbPjOHWRqw4Qam5unhJkTEoVwfrEktF7IwnfQlqKLNtcDrQRq8buQ2H3B35Feqw+jfS7fpvMEuJZsyxre9u2Q8CfiO9/2QbcbZrmJMuy5hTitVXZ2gx4FVjds/0q4HwK9LuqAtMVmEVi5fs8JMGzuOgRKZWCZVnb2LZ9MtIaZh2fu8wF7gmHwzfridayszVyErRPivusAPZHZoYopVRZ0wS0UqoUGKFQ6BrgrP/foD2hVfVZj1jv5L2RpZbdA40oN58Cs4n1sX6NJP2cC8WyrE0cxxnhOM4wJOEUZTuO8zhw6YQJE14tZkyqpO2K9Nbs69l+PTJUtK3oEalC6A3chwxFdZuPtPH5sugRKeXDsqw+4XD4WMMwTkb2B7x+AR42TfM2y7KeRU+UlaM1kaTypinu4wB/Bu4qSkRKKVVgmoBWSpUKo76+vsVxnJGubVc0NzePCSwipYLVCRlmGO2xHP13C6R1QJDakETzXGK9qecB75BlH+dCuuiiizaoqampQ/or9vDc/DQwsbGx8bniR6ZK0I5IC4a1PNtfQvoEf1v0iFQ+bYn0md/Gs30eknz+pugRKeViWVYtcIht26cCRyCtYtzCjuM8DdxaU1PzT8uyVhQ7RpU3XZEBqHumud844LLCh6OUUsWhCWilVEnxDiZ0HGdyS0vLBUHGpFQJ6o9UzayH9GZeK/L/tSLX1wFWi9w30xYby4CVkX9/ItZP+gdifaa/QxLPn1FGU9gty1rTtu1zgeEktlp4y3GcK+fOnXvnrFmztPdrddsKeBb5W3JbAAxFlkqr8nMkcCux98SoD5Hks55cUIFpaGgYaBjGScAp+Lfimoe0kLrFsqzPixqcKgQDuB0ZTp3KPcAJaHW7UqqCaAJaKVVyQqHQBcD/t98wDKOlqampHt0JU6ojokMGuyHV1UuQA6DpkdsfI7EvakWxLKunbdt/RQbM9ffc/IHjOJO/++67f8ycOVNbLlSvLZEBmRt5tq8C6oBr0M+ictEZqR6sI/GY5zVgCDLcS6misixrfdu2j0b6Ou/ocxdtsVG5LgHGp7nPS8CBSFGAUkpVDE1AK6VKkk8S+u89evQ4x7IsO8CwlKo0XYHPkYppB9gWqPghfZZldQ2Hw6cahlEPbOK5+TPHcZpbW1tvnjp16vIg4lOBWwO4EzjY57YXkZYuHxU1IpWtHYEbgZ19brsdOAMZyKpUUViW1ce27aFIT999SDwODwPPOI5zW2tr64P6+VORjgXuJnUO5jNgd2TVmVJKVRRNQCulSlZ9ff1ox3GucG268YsvvjhDl8krlVcWcHHk/zcApwcXSnFZlmXatj0EqUbaxXPzD8CMVatWTZs0adKi4kenAmYAo4HLAdNz23Kkiq0ZSRqp0tENeT8LATWe21YCFwDTih2Uqk5Dhw6tGThw4P62bZ8MHI3/YOE5wG3t7e23XX755doOpnLthbR46priPksi9/ugKBEppVSRaQJaKVXS6urqzjQMYwaR9yvDMB5ub28/XitDlMqbtYAvkMTNSqT1wMIgAwqCZVkH2rZ9CXLw57bUMIybDcO4wrKsBUHEpgL1R+AWoLfPba8CZwFvFTMgldTvgBnAZj63fQUcg7TeUKqgGhoaBpmmebLjOMcjcxm8vjEM437btm+ZMGGCvn9Uvo2Qzwu/34WoNuAw4JliBKSUUkHQBLRSquTV1dUNiySho9VM/1m5cuWR06dPXxJkXEpVkOuAYZH/TyB9f8KKZVnWvuFw+ELDMA7x3LTcMIwbDcNo0UFQVWcj5G/kIJ/bHOA+YCwwv4gxqZhdgIlIz1Q/s5ATBT8VLSJVdcaNG7e1aZonIIPj/E6CLAHuB243TfN5bSlXNXoDs5EWZ6mcBfy98OEopVRwNAGtlCoLoVDoSKRvWnTp2vumaR48efJkrUhUquO2AD5EWg38DGwI/BpoRAGzLGt727ZDSDKh1nWTDdzvOM4lEyZM0GWy1WUoUmG7ps9tbcDNSEsbXUZfHFsAlyKVzX7HNJ8jvZ7/VcSYVBVxDRMcCuztc5ew4zjPAbcvX778/ubm5qr+XK1CtcDjyOqMVJqR4chKKVXRNAGtlCoboVDoN8BDxJZCfxYOhw+aOnWqVp0p1XGPAIdH/n82cG2AsZSMcePGbV5TUzPacZyTgc6um2zHcR4GJk2YMOHVgMJTxdcPmI4kPf20IlVsVyOtbVT+bQ+MRIa51frcbiPf/4uQn4dSeWNZ1tqRYYInIC2b/I6nX3cc546ampq7Lcv6vrgRqhJyDbI/lcpjwJHoPAGlVBXQBLRSqqzU1dUNMgzjcWJ91BYahnFoU1PT20HGpVQF+A3wXOT/HwNbIYkcBYwdO3ad2tras4DzgdU8N88Grmj8P/buO06usvrj+Oe5s+kNCDUBQ5OqgCAlgDSpUkQwCigoIlHRQMpuAgT8XYmUJLsEjKAExNCRUFSaAiIopAChCZFOqAk9pJed+/z+OLPu7OzszG52Z+6U7/v1mlfu3vvM7JnM7uzcc5/nnAsuuAcrySCV7zCs7MNX2jjeCNwFTAFmFSuoChYAR2CJ56/nGPcgcA4wtxhBSXUIw7B3FEVHeu9Pcc4dBnTLMuxN4LYoiv7461//+uUihyilZxRwaZ4xzwBfo8pXnIlI9VACWkTKzqhRo7ZMJBIPAFuldi1yzh0zefLkf8cZl0gFmAPskdo+FltxIGnCMOwfRdHPsOWyAzMOP+u9n/LSSy/dNGPGDM1mqnwB8F2sbvpWOcbNAX6DJaTVQLdj1sVmmp4JbJtj3FNY4lkNvKRLjBgxose66657KFZe4zigT5Zh7zrn7nTOzQjD8LHiRigl7Gjs/T6RY8x7wJ6pf0VEqoIS0CJSlkaOHLlJTU3N/cDOqV2rvPenNjQ03BJnXCJl7kTg5tT2v4H9YoylpIVh2DeZTA53zo0GBmcc/q/3flIikbg5DMPVccQnRdUda+J5HrBRjnGLsaTETcDDaMl1W3oCRwLfA74B9Mgx9hXgfKzRoFYfSKdkJJ2/SXPJt3QfAbcFQXBrGIaPo587aelL2KqobD87TVZgq86eKEZAIiKlQgloESlbI0eOXKempuZuYN/ULg9cUF9fH8YXlUhZqwFeA4akvh4KzI4vnNIXhmH3ZDJ5gnPuHKxsSbqFwFVBEPwmDMNPYwhPiqsnljQ9C/hynrHvY411/4TN3q32cjfdsaXoJwLHA+vkGf8ocDm2SqPa/++kEzKSzsfQusQSwHKsVu8NCxcu/Nu0adPWFDNGKRubYCteNssxJsLe4/5clIhEREqIEtAiUtbCMOy9bNmym7z3x6btvqpv376/CMOwMbbARMrXaKAhtX0bVmJA8hg2bFhiu+22O945dzat6wIvds5d45y7PAzDt+OIT4puX6xsxHHkXoYN8Ak2I/oh4B4sOV0NNsJqaR8FHEr2xF+61VjC+VJ0YUw6oZ1J5xXAfUEQ3AzcF4bhymLGKGWnF/AIzWXM2lIH1Bc8GhGREqQEtIhUAldbW/t/wP/9b4dzDwZBMGzixImfxxiXSDnqB7yDnZAngW2AN2KNqMyEYXh4Mpmsc84dlHFojXNuRhRF9RMmTHgmluCk2LbFynOcAAxqx/gImxH9ANa88Ang44JFV1ybYjVP98YSzzu2834vY2VLrgEWFCY0qXQdSDr/DZgRBMHdYRguLWaMUrYC4A6sd0Yu1wKnFT4cEZHSpAS0iFSM2tra04ErsTICAC8kEokjJ06cqBmHIh1TD4xJbV+GdXOXDgrDcNcoimqxhEdNxuF/AA0XXHDB31AN0WqQAA7ESnQcR+76oJlewxLRc1L//gdY1tUBdrF1gF2whHPTrT0J+CYfYCVKbgKe7PLopCqEYdgziqJDyJ10XgncD8xYuXLlPZMmTVpSzBilIjRgq8dy+QdwBKDyLSJStZSAFpGKUldXd6j3fgbNJ/cLvPdHNzQ0zI0zLpEysyk267kbsAT4ArAo1ojK2LnnnrtJTU3NT7DawJm1bV8FrliyZMm0KVOmrCh+dBKDXlgy7HjgEPLXO87mHWxm8Cupf5u2F2AJtWLojdU63TZ12yZ12w7YcC0ebwGpRCDwIGrSKGshDMPeURR9ndyNBFd67x8CZiQSiT+HYbi4qEFKJTkNW52Ry0vYyo/PCh+OiEjpUgJaRCpOXV3drt77u2mebbXUOXfi5MmT74kzLpEycxNwUmp7HDApxlgqwrhx4wZ07979J865M4HBGYcXAL8LguCqMAw/jCE8iUcNsBc2M+5wrH54Zz+ff479PH2ENcJcmNpemToGdkHJA0tpnpHXA0sq12CleByWHO+FJZQ3ATbAajdvAvTpZJyNwEys5MH9wHNoNYCshXPOOWdgt27djvHeH+ecOxhrCJpppff+b8CMVatW3a2ZztIFDsWaU2aucEr3CfYe/1pRUMUv1gAAIABJREFUIhIRKWFKQItIRRo7duygKIruobkZmAcuqK+v/xU6wRVpj52BZ1Pb7wFbYk3ApJPCMOyeTCZPcM6NAXbKOLwKuDUIgt+EYfh0DOFJvDbCkhp7YWUrdsJWIlSCFcBcrIzI49iSdPVpkLUyfvz4wUEQfAv4lnNuf7I3+1yeSjrfoaSzdLEdsPexXCtYVgJfxy60iYhUPSWgRaRijRgxon+PHj1uw5odAeCcu6WxsfE0LXUXaZeHsbq1AKcAN8QYSyVyYRgekkwmxzjnDqH157LHgN8EQXBXGIaNMcQn8esF7ArsgSWkdwM2J/eMu1KwCisvMxerXT0HeB6b9SyyVsIw3DqZTB7nnDsO+53Idi77GXAPcFcQBH8Pw3B5UYOUarA+MBvYKscYj31uurEoEYmIlAEloEWkog0bNiwxZMiQC7ESAk2e9d4f29DQ8FZccYmUiSOxE3mwxmc7oxUEBXHeeedtn0gkzvTen0zr0gbveO+vbGxsvPriiy/+JI74pKR0x1YkbEdz3eVtsWTIBhQvOb0Kaxb4GlZ/+hWs1ukrwHxUw1k6KQzDANgjiqJjgKOBL7UxdCHwlyAI7nz//ff/OW3aNDV6k0Lpha3eGJpn3C+BCYUPR0SkfCgBLSJVoba29iSsSUiv1K6PnXPfnjx58qMxhiVS6hzwArbUFKxh2kPxhVP5wjDsH0XRqVjDwi0yDq8CbvPeN0yYMOG54kcnZWJDLBGdXrd5Q6yUx4DUmHWw3+++NJf4WAUsx2YpL8EuNi3CSu98hCW4D8ES34djjQJFutSoUaN6DRgwYB/v/dHe+2/T3M8j01vOub845+4GHtEqESkCh/XHODHPuD+lxuiCvYhIGiWgRaRqjB49eq8gCO7ETsjBTqrPqK+v/0OMYYmUutOBaantv2HN0qTAhg0blth+++2PAUbQXAblf7z3jwBXJRKJO8MwVG1uKYZRwKWp7QuB82KMRSrI+PHjBycSiaO898c45w4iexNBsAuid3nv75owYcIzRQxRBOBi4Ow8Yx4DDsYu6ImISBoloEWkqqSaE96J1dIEwDk37fPPP/+FlmyKZNUDW06/cerrnbFarlIk55133raJROIM7/2Pgd4Zhz9zzs1wzl0ehuG8OOKTqrEVVm4D7D1g5xhjkTJ3/vnn7+icOworrbE32c9Lk1it3buDIPhLGIYvFTNGkTQ/BP6YZ8wbWAPZjwoejYhIGVICWkSqThiGPZcuXToNOLlpn/f+n42NjSdefvnlH8QYmkip+iXwq9T2tcBpMcZStc4555yBNTU1pzvnzgA2yzjssfIoVy1cuPCvuqAmBfJfrAQHWImY+fGFIuVk7Nix/Xr27HkwtormCGDTNoZ+5py733v/11QTwUXFi1Ikq/2AB7AL8m35FLuQ8nJRIhIRKUNKQItI1RozZsxw59wVNDds+hA4sb6+/uEYwxIpResBb2PN8VZhiacFsUZUxVKNuQ6Komg48C1aN51bCFwXBMG0MAzfKHqAUskmA7Wp7Z8DV8YYi5S4MAy3TCaTBwNHO+cOoe0E3pvOuQe99/ekks4qKySlYltgFrBujjFrsIsq/yhKRCIiZUoJaBGpamPGjDnMOXcTMDC1aw1wdn19/RTUPEQk3e+An6a2Vf+1RIwfP35ITU3Nj733p9Fc375JBPwduCoIgnvVpEu6wAHAP1Pb9wFHxheKlJra2to+vXv3Psg59w3v/RHAkDaG/q+0RhRFf/31r3/93+JFKdJuA7Hk8xfzjPsJzb0yRESkDUpAi0jVGz169GaJROI27/1eTfucc39ds2bNDy677DIt/RQxXwReAgJsqekXgGWxRiT/M3z48G6DBg06OplM/sQ5dzD2OqV7D7gxCILrVStaOiEBfIAlZlYB6wNLY41IYhWG4XbJZPJw59w3sFIFbc1y/hD4u/f+vkQi8UAYhp8WL0qRDuuONV5u1QQ4gy7Ii4i0kxLQIiLAiBEjevTo0WMScGba7leBb9fX16vhmoj5C3BMavsXwBUxxiJtCMNw0yiKfgz8DNgwy5B5wPVBEPwxDMMPixudVICbgRNT28di7wtSJcIwXD+KogOdcwd77w+j7VnOEfAM8FAQBPcAM8MwjIoWqMjac8ANwPfyjLsNOAGtmBQRaRcloEVE0tTW1p6ELaPrk9q10ns/oqGh4ZoYwxIpFfsBj6a238RmRSfjC0dyCcOwexRF3/LeD3fOHUDrWdGrU7MRrwPuU91VaafvATemtq8GhscYixTYiBEjegwcOHAf7/0h3vtDgK/Q+r2kycfOuQe89/elajl/XMRQRbrKBPLPan4K2B9YXvhwREQqgxLQIiIZxo0bt2MymbwDazwCgHNuWmNj48gpU6asiDE0kVIwG9gztX0ccFeMsUg7jR8/fkgikTgFOJns9Sw/Bm5Jleh4qrjRSZlZDyvDUYM1Ix2MZgBWlPPPP//LwCHAIc65/YDebQxNAk865x5wzt3/4osvPjljxgxdlJRydipwbZ4x84G9sPdBERFpJyWgRUSyOOOMM/r26tXrGufcd9N2/xc4QSU5pMp9F7g1tf04sG+MschaCMNw7yiKTsFey3WyDJnnvb8uiqJbLrzwwneKHJ6Uh0exFREAu2OzAaVMhWG4ZRRFB2L1bg+idUPTdK875x50zj0IPByGoXplSKXYH3gAq//clsXY557/FCUiEZEKogS0iEgOtbW1I4FJQLfUrhXAqPr6+qvii0okVgngFWDL1Nd7Y13ipcyMGDGix7rrrnsoNiv6WJrf59LNA2YEQXBjGIavFTVAKWVjgYmp7V8BYXyhSEeFYbhxFEVfS9VxPgTYIsfwpd772c65h4IguFtNTKVC7YhdVB+QY0wjcCSWpBYRkQ5SAlpEJI+xY8d+NYqim0lbtu6c+3NjY+NpU6ZMURd3qUYjgSmp7duBYTHGIl0gDMONk8nkSc65HwA7ZRnigSeAGclk8vYLL7zwreJGKCVme+ziBMBc4KsxxiJ5pBLOTTOcDyB7GZ4mjcBs7/2DiUTiAZXVkCqwMVZerK2Gmk1+DlxZ+HBERCqTEtAiIu0wduzYflEUXQl8P233O9777zc0NPwrrrhEYtIPeBsr35DE6qW/HmtE0mXCMNwliqKTgG+TfWakB54Ebg+CYEYYhvOLGZ+UjFeBrbGfh82A9+INR5qMHz9+s0QisR+wD1ZWYIccw5PA0865fzrn/gk8Fobh0mLEKVICegEPYzWdc7kEOKfw4YiIVC4loEVEOqCuru4U7/0VQN/UriTw67feemuCZghJlZmILcMHmAqcGWMsUiDnn3/+js65YcCJwDZtDGsq03FzGIavFC86idllwFmp7eHA1THGUtXCMNwymUzuGwTBPt77fcmdcAZ4wzn3kPf+odWrVz90ySWXfFaMOEVKTADcgZWgymUGcAIQFTwiEZEKpgS0iEgHjR49epsgCG4Bdk3b/WgURSdfeumlatgl1WIw8AbWrGc58AXgk1gjkoJKS0afgM16z+YN59w9zrm7gX+FYbi6eBFKkR0MPJja/ivwzRhjqRphGNYkk8mdnXP7YjOcDwTWz3O3/yWc16xZ8/DFF1+s92oRuJz8F8+fwH7Hlhc+HBGRyqYEtIjIWhg+fHi3/v37jwfOx2ZQAHzuvR/b0NAwLcbQRIrpBprL0pyDLVGVKhCG4a5RFH0bq/+9dRvDFjnnHvDe3xcEwf1hGH5YxBCl8LoBH2FNu1ZgSVAlabrYueeeu0lNTc1ezrm9vPd7YfW2e+e4SyPwDPAY8O8gCB7X755IK2dhqzhyeQMYCuj3R0SkCygBLSLSCbW1tQcD1wObpO2+w3v/04aGho9jCkukWHYCnsU+T7yP1QvWjNcqc/75538lCIJve++/CezYxrAIa1Z3r/f+vkQiMTcMQy1nLn8zsFrhAEcC98UYS9kbMWJEj4EDB34lmUzuFQTBnt77oeRvjLbMez/HOffvIAgeA2arhrNITkcCfwESOcZ8gq0weLkoEYmIVAEloEVEOqmurm7jKIquds4dlbZ7IfDj+vr6e+OKS6RIHgK+ntr+IXBdfKFI3MIw3DyKoiOwE/wDaXum5ofA/alk9ENhGH5atCClK/0AmJ7a/h1wRnyhlB0XhuFWyWTyq6lk855Yaa8eee73ETDTe/8v4PHUxZzGgkcrUhl2Ax4F+uQYsxo4HPhnUSISEakSSkCLiHSRVIPC3wL90nbfEATBzydNmrQkrrhECuwImmc9voDNivbxhSOlIgzDnsC+URQdDBwDbN/G0Ah4yTn3mPf+oSAIHgzDcFHRApXO2ABYgM0kfAebravf/yzCMBwURdFuwG7e+92cc3uRv3ZzEpuBOdd7/xjw+IQJE+ah/2ORtTEYmJP6ty0eOAW4sSgRiYhUESWgRUS60Nlnn715Y2PjH4EDmvZ57+cHQfDDyZMnPxpfZCIF47AyHDulvj4MeCC+cKRUhWG4XRRFR2IXLb6GNbDMphF4wjn3sPf+XytXrpyti3glbSZWJxVgF+C5GGMpCeedd94WNTU1u3nvd/Pe74bVbV63HXddAMwGZnnvZ6dmN6uutkjn9Qf+TfNnlbacC1xc+HBERKqPEtAiIl0sDMNg6dKlY4AJNC+lTXrvG1avXv3LqVOnrooxPJFCOA24JrX9AJaEFmnT2LFj+/Xu3fuQKIoOBw4CtsoxvBFLaqY3VVtYjDilXc4FLkxtn5e2XfFGjRrVa8CAATsmk8ldnHM7ATtjCa512nH3xVizwKe8909FUTTrwgsvfKuQ8YpUqW7AvcAhecZdi32eERGRAlACWkSkQMaNG7djMpm8Hqvp2OQ159yPNRtaKkwP4E2am3FqFqR0yPjx44cEQXCgc+4gLCGda4k0wKvAbO/9E4lE4gng2TAM1QAzHjvR/Ps+m+bZ0BVl/PjxgxOJxE7e+52dc7tgz3sbcjcya7IESzbP9d4/5b2fW1NT86oacYoUxe+Bn+QZ8wh28Vx/R0RECkQJaBGRAho+fHi3/v37j8dmhTWdpHrn3NXLli0bc+WVV6pTvVSK87BZ/2BNyU6NLxQpd+edd962QRAc4Jzb13v/Nay2cC6rgGecc094758IgmDuiy+++OqMGTOSRQhX7ALU5lg970HAB7FGs/ZcGIZDgO2iKNoRq1u+Q+rf9sxqBlgEPOeceyaKormpZPPLSjaLxOJ84II8Y+YB+2C/uyIiUiBKQIuIFMHo0aP3CoLgD9iJLADOuTe896fX19c/HGNoIl1lPeBtrLP8GmBL4N1YI5KKEYbhplEUfQ1LEuwH7AgEee62DHgeq1H+TBAEzwAvhGG4sqDBVqcrgDNS26diF6FK1ogRI3oMHDhwK2CbKIq2dc7t4L3fAdgO6NvOh4mAN7z3zzrnng+C4Dng+TAM5xcobBHpmO8BN5A757EA2Av7/CIiIgWkBLSISJGkZkOPxmZiNDXf8sCNyWRy5JQpUz6NLzqRLpGehLoYqw0r0uXCMOwPfDWKoj2993s45/bAZt7m0wi8ArwIzPPez0skEvOAV1TCo1OOAO5Lbd8BfDvGWAD7mzto0KAtoij6IlYq44vA1ql/v0D+CxjpPgXmOede8N4/671/PpFI/CcMQ61iEilNBwB/o7kXSzbLgQOBJ4oRkIhItVMCWkSkyEaNGvXlRCLxB2D3tN0LgTPq6+vviikska6wJZbcSwCfYUkeJWikKMIw3BTYI4qiPbz3X0nV6d2wnXdfA7yGJRlfjaLo1UQi8drq1atfveiiixYULOjK0QP4GJs9vBRYHyuLUjBhGAbYRYfNoyjawns/xDm3eerfLbCyLTUdfNgPsJ+B/0ZR9GIikXhp9erVL1500UXlWlJEpBp9Cfg3ucvmJIHjgb8UJSIREVECWkQkDmEY1ixdurQO+CXQM+3Qzd26dRt58cUXfxRTaCKddSfwrdT2mcDUGGORKpdqHLdLWkL6K8AWdOwz8FIsOf2qc+5N7/3bQRC8lUwm31q9evXbEydO/LwQsZehPwPfTG0fCjzYmQc799xzN+revfsmwKbJZHKQc26Qc26zKIo2d84NATajeTVRR6zGSme84px71Xv/SmoW/LwwDLUSSaS8DQZmYe8PuejziYhIkSkBLSISo7q6uq2899dgSwWbLPLeh/369ZuqpkVShvYBHkttv4ktd1cjOCkZtbW1fXr16rUdsGOq9u+OzrkdsCZ6HSnL0ORz4G3v/VtBELzvvV/gvV+YSCTeTyaTHyQSifeAD6ugxMePgatT278BzsocEIbheo2NjRs459ZPJBIbRFG0AbChc25D7/1gYBNgU2Bj1i653GQN8Kb3/rUgCF5pmtHe2Nj46ssvv/y2mlOKVKQBwL+AnfKMmwKMLnw4IiKSTgloEZH4uTFjxpzunGugZfOjud77nzQ0NMyNKzCRtTQLa+oDVgv2jhhjEWmXMAx7Y7WCt46i6IvOua2991un9m3cBd/iQ6yW8Cepfz91zn3ivf8U+NR7/0kikVgELE0mk8sSicQSYNEnn3yybOrUqQUtZ9FeZ5999rpBEHTv3r17H+zv1TrAOslkcp0lS5Zs9vTTT0/o2bOn69u375Idd9zxQefculg5jqZbty4KZTXwTirpPz+KordSs9PnJxKJt1588cX3lGQWqSo9gL8D++cZdycwDGsiKiIiRaQEtIhIiRg1atTgRCJxGS2bNzUCVy5fvnz8lVdeqVq6Ui6GAbeltp8E9ogxFpFOC8OwL5aYHoLNlB6C1Thvum1U4BDWYKVAFmHNa1dhDbTAVs1451z6vlwiUjO9vfcJoH/TAedcL5rLQq2LzULuk7p1ZkZyR6wE3gPeB97FeiS8471fkEgk3gXmA+9rhZCIpDjgeuD7ecbNAQ6ife+TIiLSxZSAFhEpMbW1tUdjdemGpO1+z3s/sqGh4faYwhLpiATwMrBV6ut9gJlZxu2OJahFytqoUaN69enT5ws1NTUbRVE02Hu/kXNuEDZzemOsWd6GwEDWrsxHuVuMNfj7yHv/cRAEH3vvPwA+Aj4OguAj4N1Vq1YtuPjiiz+JNVIRKTeXAqPyjHkN2Bt7zxERkRgoAS0iUoLCMOy9dOnSscA5pM06897fA/yioaHhrdiCE2mfM4HLU9t3Yt3m020B1AFnFDMokbidffbZ6/bs2XNgMpkc6Jxbz3s/EFgvCIKBWFmLvsA63vs+6V+n/u0O9MaWmxdaEkscr8BmJS8GVnvvFzvnVmCzCBdhM7AXAYveeOONdefMmXPRypUrSSaTc0899dSjlyxZ8mmplBARkYpzBnBFnjELsOTz/IJHIyIibVICWkSkhNXV1e3ivf89sGfa7iXOuQv69OnzmypoaiXlqw/wFjbjMwK2xWYgNfk1Vif64OKHJlL+hg0blthqq636A/Ts2bMfUJM6VJNMJvvlu38ikficVB3UNWvWrEomk/9bln7JJZc0lfroKAe8AwzGEtgbYTWvRUS62jDgVnKvKlmC1YV+pigRiYhIm5SAFhEpfa6uru5k7/2lWDKvyavAqPr6+ntjikskn4uBs1PbVwC/SG0HWJIKLFElIpVjGnB6avv7wE0xxiIilWk/rOlgzxxj1gBHp8aJiEjMlIAWESkTZ5111kbdu3dv8N6fRMv377udc6MmT578elyxibRhEPAmVjZgOdas7RPgMOBv2AzLPtgSfxGpDMcAf0lt3wqcGGMsIlJ5dgAewxqltsUDPwKmFyMgERHJTwloEZEyM3bs2D2SyeRU59weabvXAL8LguC8SZMmLYkrNpEsrgNOSW2fB1yIJaW+m9q3E/CfGOISkcLoBXyM1ar+HNgA+xslItJZg7CmxkPyjBsPXFT4cEREpL2qsQu3iEhZmzRp0hP9+vUb6pz7Ac3dvLsBZ0ZR9FJdXd0p6AKjFN9QYAKwdcb+eppryZ4JbAx8K+34FwsfmogU0QrgkdT2AGCf+EIRkQrSH7iX/Mnnq1HyWUSk5CTiDkBERDrukUce8TNnznxuzz33/EMQBP2AXbGLiv2Ab+29994H7bXXXs/OmjVrYbyRShV5F0s03QkcgiWdX0vbvxVWbmNj4Ctp93sWW0orIpVjAHBkavsT4IEYYxGR8tcNK+2T74LWvcDJpBqsiohI6dAMORGRCjBmzJjtnHOXA4em7Y6AO7z3dQ0NDW/FFJpUF4eV3Dg59fUK4DbgBWBy2r7u2EXwZGr8acUNU0QKbDPgLew94WVgu3jDEZEy5rBazqfkGfckcCCwrNABiYhIxykBLSJSQWpra48GptJyeeJyYOqqVasumjp16uJ4IpMq0gP4B82zlCJsdv5qLPGczgOPA18rWnQiUizPAjuntrcFXokxFhEpXxOBsXnGvI597vig8OGIiMjaUAkOEZEKMnPmzFf23nvvq7GZpbtjSxa7AfvW1NT8cOjQoZ9vuummz82bN8/nfCCRtZcE7ga+g5WEafqsEdD6wrfDGpZNRkQqzWBgv9T2fGBWfKGISJn6Oda8OJcPsZnP7xQ+HBERWVuaAS0iUqFGjRo1uKam5pfe+9NoecHxv865/5s8efKMuGKTqrADMBvoTf4L3gMAzc4XqSx70Zx0fhj4eoyxiEj5ORa4ndyfIZYDB6MLXCIiJU8JaBGRCjdmzJgdnHOTgW9kHHooiqLaSy+99Lk44pKqcChwH9lnP6fbDXi6KBGJSLEEwPvARsAaYENgUawRiUi5OAj7/NAjx5gkcDzWnFBEREqcSnCIiFS4WbNmfTRz5syb995776eAXYANUoe2dM6dPnTo0CH77bffs48//vjnMYYplel14D3gm3nG/RN4sfDhiEgReWBH7O9OAngG/Z6LSH5fBe7HVlDlMgK4sfDhiIhIV1ACWkSkSsycOfPVQw899Pdr1qx5AxgK9AUC59xXvPc/32effTbbfffdn5wzZ466h0tXegZYD9gzx5jngX8VJxwRKaIarB48wErgrhhjEZHStzXwEPa5IZdfof4RIiJlRSU4RESq0NixY/slk8mznXMjaTnDZAnQEATBpZMmTVoSU3hSeRLAn4Ejaf3ZIwncDJxS7KBEpOD6Ah9jy+g/xcpwJGONSERK1WDgcWBInnF/AE7HVlmIiEiZUAJaRKSKnXPOORusWbNmDDCSlnX2PgEm9+3b9/IwDFfGE51UmH7ATKw5YZBx7Alyz5AWkfL1AHBIantfLMEkIpJuAPAosHOecX/F6j43FjwiERHpUkpAi4gIY8aMGRIEwbne+9NoWZ7pXe/9hH79+l0bhqE+7EtnDQGewpbWpiehPwfWiSUiESm0M4HLU9uXAOfEGIuIlJ5e2IWqffOMexQ4HCvnIyIiZUYJaBER+Z/Ro0fv7Jz7tXPuqIxDLwH/17dv39vDMIziiE0qxleBfwM9M/YPxJboi0hl2QJ4I7X9AvDlGGMRkdLSDSvR9Y084/4D7A98VvCIRESkIJSAFhGRVkaPHr2Xc+4i59yBGYfmOecmzp8//6YZM2aojqesrWHAbRn79gLmxBCLiBTei1j5HYAtgTdjjEVESoMDrgV+mGfcG8A+wMJCByQiIoWTyD9ERESqzaxZs96dNWvWdXvvvffjwJeATVKHNgC+NWDAgOP33XffpYMHD35h3rx5agIjHTUPK8Gxf9q+2cAz8YQjIgX2BZqX17+O1X0XkerWAPwsz5iPgK8DbxU+HBERKSTNgBYRkXxcbW3tUUAI7Jp+wHv/YhAEk/r06XOjSnNIBzngb8Chqa8nAL+MLxwRKaCvAf9Kbf8dq+MqItXrXODCPGMWAwegi9MiIhVBCWgREWmvpkT0r4CvZBx7wTl3weTJk28HNCNa2qsn8AiwJ3ArcGLasf7YzPsNgA3TtjfC6kU7mhsX9gNqsEZGPbEGRSuAJHYCC9boMMLqTH+AzapamNr+MLX9eZc/QxEBW3W5EFgfWIX9Li+JNSIRicspwHRy5yJWA0cCDxUjIBERKTwloEVEpEPCMAyWLl36HeB8mmt6NnkGmNC3b9+/aEa0tEMPYA/gLiwpdQ+wI9akrH8M8azCygO8iNWcfAMrF/I8zYlsEVk7NwDfT20fD9wZYywiEo9jgDuwi8ZtSQLfTY0TEZEKoQS0iIislTAMg2XLlh3vvb8A2C7j8Gve+8n9+vW7NgzDxjjik5LTE5s5v2fabYtYI+qYt7AmiU23p7FZ1iLSPicAt6S2rwVOizEWESm+/bHSWz1zjPHAcOCaokQkIiJFowS0iIh0Sq5EtPd+PnBZFEXTpkyZomRddekPHIzVb9wTSz53izOgLrYGmxk9Gysj8hCwKM6ARErcAKz0TTes7M0mWFkcEal8XwYeBdbNM+4c4JLChyMiIsWmBLSIiHSJYcOGJYYMGfJd59zZ3vsvZxxe4L1vWLFixVVXXnnl0lgClEJzwM5Yc7HDgb3pXMJ5OZakWkBzveaFqe0PscRVU8J3CdCIzUheic2u6oXVnW0q5TEg9XVTTemNgI1TX2+cuvXuRLyNWDL6fmyG1zOoHrpIpoeBA1Pbe2GrCUSksm0JPI79nc3lCuAXhQ9HRETioAS0iIh0udra2oOBC4ChGYcWA79LJpOTpkyZ8mnxI5MuFgD7AScBR2EzGjuiEXgTeDl1eyV1exlLPBfbRtgs/m1St21Tty3oeDL9A+Be4Gbgn2impwjAGKA+tT0B+GWMsYhI4Q0G/oUloXO5EWtOqAu3IiIVSgloEREpmNra2oOcc+d677+ecWgJcFUQBJdPmjTp3Thik07ZAfgOcDL5TyrTLQDmAo9hs6HmUh51lGuwRPQ+wL7AbsD2tP9z1PvA7cAM7LmLVKutgVdT289ipXlEpDKtj5XdyGxYnekh4EhgdcEjEhGR2CgBLSIiBVdXV7eL9340NlM2kXZoDfBn7/3khoaGJ+OJTtppAPAj4IfATu28z3+xkhSPYEvtPywMVsBvAAAgAElEQVREYDFZH6ttfQBWcuRL7bzfC8B1wB+AzwoSmUhpexlbYQCwOdbgU0QqywDgH9gF21zmAF8HlhU8IhERiZUS0CIiUjTjxo3bMZlMjgNOxGaVpnscmFhfX38PWoJZSrbEOtL/BFgnz9jlwEzgHuDPVFdiaSPgMKwUySHk/79aic2IvgSYV9jQREpKAzA6tf0z4PcxxiIiXa831gvha3nGvYiV8VJJNhGRKqAEtIiIFN2oUaO2TCQSo4BTgT4Zh5/13tcvWbLktmnTpq2JITwxBwAjgaOxWs9tWQbcBdyEzXbSa2b1og8Evgd8C+iXY2wE3Adchv3/iVS6g2j+Wb8He48RkcrQC+t/cGCece9iZa3eLnhEIiJSEpSAFhGR2IwYMaJ/9+7dT3XO1WGNatItBK5qbGy87LLLLlsUQ3jVak9gIrB/jjFJYDZwPXALVtNbsuuJzYg+Gfgm0D3H2CeAs7GmhSKVqhtWjmcdrAb8+tjqiXQ7YrMjRaR8dAPuxFYC5fIh9hnjpYJHJCIiJUMJaBERid2IESN6dO/e/WTn3GisuVu6z4Gra2pqrrjkkkvmFz+6qrEDcBGWJG3LAuAK4Brgg2IEVWE2wOpo/wLYNMe4+4BzgOeLEZRIDG4FvpvaPga4O+P43WhmtEg5SQA3AifkGbcIWwXxTMEjEhGRkqIEtIiIlBJXW1v7de/9Wc65zBk0EfAw8BvVie5Sg4ELgB/QskFkuqexEhF/Ql3qu0I34NvAKGD3NsZEWFmT89ASZak8J2MrKACuAn6aduwIrIZ8j2IHJSJrxQHTgB/nGbcYOBhQ02kRkSqkBLSIiJSkMWPG7O6cqwWOo3XDwueAqX379r0pDMOVxY+uIjisseBEoH8bY+5LHf9XsYKqQvsAY7FZoNksBcYDv8WS0iKVYD1sFUUN8D62IsBj9eafxVZkZL7vi0jpcdjfpzPyjFsOfAN4tOARiYhISVICWkREStrIkSM3qamp+Qnwc6xWaLrPgesSiUTDxIkTNUu0/bbCZisd1MbxOVgt4keKFZCwJ3AJ1vwxm1nY7LJ5xQpIpIvsgyWVl2Xs/zewb2p7N2ylRfrMaJ2niJS+S4BxecasBo4F7i98OCIiUqraWmorIiJSEmbPnr105syZjxx66KFXrFq16m1gS+fchqnDPYE9vfe/GDp06I5Dhw59f9asWe/EF23JqwHqsFIa22Q5/l9gOFALzC9eWAK8B1wHzAW+DGyUcXwzmpc3z0KzoaV8JLAGmz2xpffJ1P6NsOX4YLOgZwF3Yc0JASagUksipeyXWJmoXJLA94C/FD4cEREpZZpZICIiZWfMmDH7OufOxMpzZF5M/a/3/qoVK1b84corr1waQ3ilagMs8XxglmNLsKZ3v6c5OSTxCYDTgEk0J+PS/Rv4DrCwmEGJdEId9vP8GjZb8i6szMYLqeNPYo0JG9Lu0wPVnBcpVWcCl+cZE2GrGm4ufDgiIlLqlIAWEZGyNWrUqC0TicRw4HSspmi6xc65W733V9TX1z8fQ3ilZFfgDmDzLMfuxxqAqYRJ6dkYq615fJZj7wPDgJlFjUhk7dRgSeZdUl8/ga20+CNWEshjF8L60Xx+0htYUdwwRaQdTgX+QO5cggd+hjUZFRERUQJaRETK39ixY/tFUfQDrE70dlmGPOqc+12fPn3uCsOw2mbUnYLNbO6Vsf8zrM7ztKJHJB11NPYaDsrY34gtf55Y9IhEOm5XLPGcwFZaJLBSP5u3Mb4f1oRTRErHt7HVCvlKedYB9YUPR0REyoUS0CIiUlHGjBmzm3PuLOAEoFv6MefcB9776cBV9fX1b8YRXxE5YApwVpZj/8T+fz4sakTSGesDNwGHZjn2e+zii+pCS6mbhCWmmkRYyZls1sEazYpIafgmcDu2oiGX84FfFz4cEREpJ0pAi4hIRRo5cuQmiUTiNOfccKyBW7okcJ9zbtr8+fPvnzFjRqXVPXbAVCwpmWka8AtgTVEjkq7ggLHARbRO2t2CzXZvLHZQIh3QG5gHbEr+GZTrYSs1RCR+BwN3Y81Ec7kMGFX4cEREpNwoAS0iIhUtDMNg6dKlBwHDyd60cAFwvXPu6smTJ79e9AC7XgK4Bvhhxv6lWGO724odkHS5o4AbaN2g8Dbg++jigpS2g4EHyH8esgHwceHDEZE8DgDuxS4g5XIFdoFbRESkFSWgRUSkatTV1W3lvT8d+BGW3EjngZne++v79et3YxiGy4sfYafVYInJEzL2L8RKN/yn6BFJoWyPJfE2zdh/J3AiUG21zqW8TMdm7Oc6F9kIlQkSids+wN+AvnnGXY81J1QpKBERyUoJaBERqTphGPZcsmTJ8cBpzrkDaP338BPghkQicc3EiRNfLHqAa+8K4IyMfQuwGYfzih+OFNgQ4B/AVhn7r6P1DHiRUrIe8AqwLm3XgB6EvX+JSDz2xpLP/fKMuxP4LioBJSIiOSgBLSIiVW3UqFFbJxKJH2EJu00yjzvnZnvvrwmC4LZJkyYtKXqA7fdD4I8Z+94Cvg6US2mRnwOH5Dj+MfDjLvg+p2DlWNqyCjuZLgebYUnoL2bsPxOrAy5Sqk7CGmu2ZTPg3SLFIiItDQX+Tv7k8wPAMdjfTRERkTYpAS0iIkKrWtHHAt0yhqzEGvDc8NZbb91XYo0L98WSkN3T9s0H9gfejiOgtTQd+EGO4x6bkTW7E9/DYf8nmaUr0iWxciblYhDwKLB12r5G4DDg4VgiEmmfu4FvkH0W9ObYRTQRKa5dgYewFQq5PAwciX0+EhERyUkJaBERkQwjR47cpKam5hTgdFqXNwB4D7gxiqJrL7300leKG10rmwFPYvVSmyzFErXlVvN5OrkT0KuxWd4/7cT32B/4J7k/A5VbAhqsJvQsYEDavk+APYA3YolIJL8vAP8FetH6d3Ir9LMrUmy7Aw/S8m9JNo8BR2CfN0RERPJSAlpERKQNYRgGixcvPjCRSJzqvT8OS5Jkmum9vy6ZTN522WWXLSpyiA6rz3ho2j6PNSG8rcixdIXp5E5AAyzDGkiuWMvvcR32/9M9x5hyTECDzXi+F0ik7ZsFfA17TiKlaCQwJcv+bYBXixyLSDXbBVtNtV6ecbOwvzelXJZMRERKjBLQIiIi7TBixIj+PXv2PNZ7fzJWVznzb+gq7/2DQRBc36dPn7vCMCxGM54zsMaD6ULgV0X43oUwnfwJ6EbgZODWtXj8PsBHZL+QkK5cE9AA44BLMvaNJnuCT6QUBMBMbLZ++vvqdthqk/Ww38ee2O9uAuifGtMbWI5deGu6ALgEe59YAXyGygOItMfOWPJ5YJ5xc7FeDZ8VPCIREakoSkCLiIh0UF1d3VbAKVEUneKc2zzLkAXOuVudczdPmjTpqQKFsQE2OzB9meyD2KwkX6DvWWjTaZ2AXgX0SPs6wkpoHLwWj/9DYBot63uvoXW973JOQIPV1T0q7eul2GzSBfGEI9KKw8oHbZO67YyVPEo/N1mJJZ0763NgIfBh6rYAuxD1DvAS8ApWrkakWn0F+/yQL/n8BLbi6vOCRyQiIhVHCWgREZFOGDNmzG5BEAz33p9Ilm7x3vv5zrk/FaBe9O9oWQv5c2AnyqvpYKbptE5Ar8BmPKaXzPDAECyB1BGzsFmW2RqepSv3BPQg4AVaNpCaDpwaSzRS7bbEfu92pDnhvA02e7lUfAK8nLq9AjyPJds+jjMokSJob/L5GezC76cFj0hERCqSEtAiIiJdoLa2to9z7njv/Q+AA8ie5JwD3LxmzZo/XX755R904tttgSVJ0pOko4DLOvGYpWA62RPQmSUzVmGlRjJLTeSyBfA6LT/7rCZ7LehyT0AD/Bz4bdrXEbADlmATKZR+2Gzm3YB9sKafG8YaUecswEoOPAY8DjyFSnpI5dgVSz7nq/n8LJZ81koBERFZa0pAi4iIdLGxY8cOSiaTw5xzw7AkTKYImOW9v3716tW3Tp06dXEHv8XvgZ+kff0K8CWsnEQ5m072BHRP7P8svbnem9jMyva6ABhLy3IenuyfhSohAV0DPIclnZtch5UhEekqNcDewOGp287kX2HQEcuxWciNWOJ3RWp7Sdrx3tjv8Tqpff1ScfUC1qd1iZ3OWAn8G2v++jdgXhc+tkgx7QY8QP7k83NY3wsln0VEpFOUgBYRESmgsWPHbu+9Pwk4yXufLWG6ArjXe/+nKIrunTJlyoo8D7ke1pgrvTbqycCNXRNxrKaTPQH9BJbIT08KeyzxNbsdj+uAd7HSFE0asZIcX8syvhIS0ADfpWWzxjVY3d3OzL4XGQwcgSWcD6ZlHfqO+Ay7ePYydkGpqT7zqcDR2EqSRzsZK1gSesPUbRBWP38TYCuay4Gsba3pt2lORj+I1VsXKXV7A/fT3MyzLc9jyWeVohERkU5TAlpERKRIxo0bt2MymTwZS7JunGXICu/9P4IgmOG9v6O+vn5ZljEjgN+kff0mlkBp7PqIi2462RPQ5wITgL5p+1enxv+E/A4G/k7LmZlJ4LTUY2SqlAR0Avgv8MW0feOASfGEI2VsHeDbwPexizYdmeW8BCtd8STNNZZfxhoBZjMAeBEYhl0kKrQA+ALNyegdsZrVO9Gx94HlwF+Am7D3m0p4T5bKsy9wH1l6VmR4DvvbqeSziIh0CSWgRUREiiwMw5olS5Yc4pw7CTiWlonVJouBvwK3rVq16oGpU6euSu1/Evhq2rjxwEUFDbh4ppM9AT0aa5T0Q1rWbF6OzW7MN2v8FuB4Wi7F/wxLNL2fZXylJKABxgD1aV+/iJVrEcmnB/ANLOl8JC3L17QliZWlmIOtTpiDXQRJdvB7D8N+Nx/v4P26Ui+sTMGeqdte2AqC9vgI+BOWjG7PKg2RYmhv8vlZ4BCUfBYRkS6kBLSIiEiMwjDsuXTp0kOwhMuxZD8xXATc/cEHHzx88803X5tMJpv+fkfYzL33ihNtwU2n7QT0s7SeDdmIlR+5lbb1x5JB6Ynr1VjDxnps2X+mSkpAb4Al8tKfz1bAG/GEI2VgENbEcjh2gSefD7AZv3/Dasp2Va3YfjTXei4VW2PlR47AGiz2bsd95gGXAzeQ/2KZSKEcCNwN9Mkzbi5wKPBpwSMSEZGqogS0iIhIiQjDsPfSpUuPAr6DzTzslTnmtdde489//nPTl7OwWo6VYjptJ6B/D7xOy8aDEfAIVqOyLacDV9C6EdlOwEIqPwEN8DCWfGgyAvhtTLFI6foqMBK7GNY9z9g52AqNvwNPYzXZq01PLAl9OPAtYEie8R8DVwFXkn3lhUihHA7cSZbPFBmexmY+K/ksIiJdTgloERGREjRq1KheiUTiYCwZdBypWUsPPPAAzz//fNOw84AL44mwIKaTOwE9DvgVLUsBeCzx804bj/kUsCstP/M8i5X02IDqSEBnluG4BTgpplik9HwDOJvsDTnTvYqVlLgJeK3QQZUZh5U3+D72nr1ujrGrgduw9+6XCh+aVLmjsZ+3fI02NfNZRERERESkmo0YMaL/mDFjTh42bNji3r17eyzp6mk5q7USTKf5uTXdlgM/TR3fGEsOpx9fCZzTxuNtg82STh+/CvhZ6vgGWb6fp/Kahw2l5fN7Pd5wpETsBTxK9t+BptsnWNPTPWOKsRz1wMop3Y4lm9v6v10DXA0MjidMqQInkPtnsOn2OFauSkRERERERKpcAkuepp80VtoJ43RyJ6ABHsQSxOlj5pN9VdfFWII6fexqmmcnVksCuict/8+StK+hnFSmbbEZkZkXZ9JvrwJnkb9erOS2MRBi5Tfa+r9ehdWIzjVrWqSjTqf1Bdtst8eovM8SIiIiIiIispa+QMuTxmylI8rddPInoL+DzRxMHxNhs3zTBVhztMwZhzPSxlRLAhrgbVo+x23iDUdiMACrP5z5+5P+e/QgVpJDZfq6Vh9s5cXL5J5t/nPsvUukM0aS+wJT0+1RoG9MMYqIiIiIiEgJ2p2WJ45z4w2nIKaTPwHdHVicMWYVMC3jsQ6n9eyvxtT+JtWUgH6Mls9xv3jDkSI7BniXthNR9wG7xBZd9QiAb2O1n9t6Lf4NbBdXgFL2xpE/8eyBf6Lks4iIiIiIiGQ4gJYnj4/EGUyBTCd/AhpsFmdmOZJlQO+0MbfTeqbnh1gpkybVlIC+n5bP8RvxhiNFsiFwPW0noeYAB8UWXfUKsGaFb5L9dVkNXIJdcBNpr1/RvuTzPeRvSigiIiIiIiJV6DBankA+EG84BTGd9iWgM2eDNyVsTkwdH0DrBPVK4KKMx6mmBPRdtHyOx8UbjhTBcbRde/jl1HGV2ohXb+BsYBHZX6dn0GxoyS8AptK+5POtQLd4whQREREREZFStz8tTyL/FW84BTGd9iWgoXUt1STwcOrYGbROQHtaJ3KqKQH9d1o+x8NzD5cylsAa32WrAbsGm1mr2Y+lZRPgDrK/Hy3GynaIZJMArqV9yecbgZp4whQREREREZFysBstTySfjTecgphO+xPQtdis5vSxEbAZ8Bwtk28R2WtmV1MCehYtn+O+8YYjBTKQ1hcb0t8zdosvNGmHYbRuntr0HnY5Sh5KSz2AO2lf8vl3qMGliIiIiIiI5LEJLU8mF8UbTkFMp/0J6I1p3WRwJTbDK/MxVgHDszxGNSWgF9DyOW4RbzhSALsC82n987wSqEPJy3KxPnAz2d+b/g6sF19oUkIGYCuh2pN8nhhTjCIiIiIiIlJmHJaMTT+pXD/WiLredNqfgAZrrNeY5T6ZM6NXA+tkuX+1JKD70XJG+BpUA7TS7At8Tuuf5XeBoTHGJWtvONlLCc3DLkhK9doIeJr2JZ8viSlGERERERERKVNzaXli+Y14w+ly0+lYAvp4siegM5PPt7Zx/2pJQB9E6wSWVI79gSW0/jn+F7ZSQMrXV8k+q/0lYHB8YUmMNgdeIX/iOQJGxROiiIiIiIiIlLPf0fIE88J4w+ly0+lYAro72Wd9pt+SwCFt3L9aEtC/pOXz+2O84UgXOgJYQeuf4Smo5Eal2JjspRZeBb4QY1xSfNsD75A/+dwI/DimGEVERERERKTMfY+WJ5nPxxtOl5tOxxLQAFNpXXIj/baQthsvVUsCeg4tn99p8YYjXeRwspdoqIszKCmInsA9tH6t3wQGxRiXFM+ewMfkTz6vAI6NKUYRERERERGpAAOx+r3pJ5vbxBpR15pOxxPQu2a5T9NtJfDrHPethgT0EFo2a0yi+rGVYFvgM1ovuR8ZZ1BSUN2BO2j9fjUX6B1jXFJ4hwCLyZ98XkrbK35ERERipaV5IiIi5eMT4DHggLR9PwLOjiWa0vA08DKWkMvUHbiuuOGUnB/Rcgb4E8CCmGKRrtEfuJOWjTU9cCbw21giWjsDyV0mYAbwRhd8n82Ak3Icvw5bKVHqVgPfwUronJy2f1fgqox9Ujl+CEwjf+PYT4EjgdmFDkhEREREREQq3w9oOePpAypn9tt0Oj4DGqzRUrbZYHPy3K/SZ0D3AN6j5XP7SawRSWclgPtp/TN7VpxBraXtyT2b89Iu+j4X5fk+u3XR9ymWBHA3rZ9HbZxBSUGMw1Y25Jv5/D7w5ZhiFBERERERkQrUG1hEy5PPM2ONqOtMZ+0S0Gur0hPQP6Xl81qCzZ6V8jWB1j+vf4g1orWXLwH9AflnfeYTAG/n+T7lloAG+z2eR+v3rQNijEm6ToLWTYfbur0ObBlPmCIiIiIiIlLJMmf0LQD6xRpR15iOEtBdpTfwDi2fV0OsEUln7YqVYUh/TWdiM93LUb4EtAe+2cnvcVg7vkc5JqABtgA+ouVzeZPK+FtQzXoDf6V9yee5wIbxhCkiIiIiIiKVbkNgGS1PRC+KNaKuMR0loLtKSMvntAIYFGdA0indgBdp+Zq+C2wcZ1Cd1J4E9F2d/B63tON7lGsCGqzhXHqTUQ9cHmtE0hnrYX0e2pN8/gda0SIiIiIiIiIFdgEtT0ZXYzMky9l0lIDuCtthCef053RJrBFJZ51F65/TI2KNqPOyJaCfzPh6DbDRWj7+AOz9I9fjl3sCGmAKrd+/VA+4/GwBvET7ks83YE12RURERERERAqqL60bzD1D+S7HByWgu0I3YDYtn89CNFOunA0APqHla3ptrBF1jWwJ6OnAExn7Rq7l4/8s43HmAtdk+Z7lnoDujdUBTn9O98YakXTUHtj7dHuSz5djtc1FREREREREiuKbtD45vS7WiDpnOq2fzyqUgO6IK2j9fE6INSLprPNo+XouprxLbzRpKwF9Rsa+59fy8edkPM4IKjMBDXAcrZ/X7rFGJO31LVqX1Mp2i4C6mGIUERERERGRKnczrU9Uz4o1orU3newn3sVOQK8p0PcrtFNp/Vz+EmtE0lm9aN1obnysEXWdthLQ2Upn7NLBx94h4/6rgPWp3AQ0tK4dfEe84Ug7nEXrGt7ZbivRhUQRERERERGJ0TrAy7ROoB4cZ1BSdPthdcDTfw7ewJJuUr5OoeVrugj7na8EbSWgAf6UsX9KBx97csb9b0/tr+QE9OG0/jugxqOlKQH8lvaV3PgM2D+eMEVERERERESabYclptJPWj+lchIrktvOtJ4luwQ1IqsEj9DydZ0UazRdK1cC+oiM/R/T/qZrNcD7Gfc/KnWskhPQYOVK0p/bufGGI1n0Be6hfcnn97D3dxEREREREZGScChWuzhztuTQOIOSgtuV1snnCBgWZ1DSJTai9fL87WONqGvlSkAHwDsZx45t5+MelXG/hVhzTqj8BPSZtHxuz8QbjmTYDHiO9iWfnwU2jSdMERERERERkbbV0fokdjHwtTiDkoLZC1uenfmanxdnUNJlfkzL1/XJeMPpcrkS0ACXZBy7q52Pe3vG/SanHav0BPQGtLwQGWFJT4nfbtiM5vYkn+8D+sUTpoiIiIiIiEh+F9L6ZHYptqRdKsfXsYsLma91PeBijEu6zi20fG3PjzecLpcvAb0NlkBtOrYGmxWey0CsYVv6Y6aXoqn0BDS0bkb4g3jDEWxFyjLal3y+muYZ+yIiIiIiIiIlaxytT2ojbEZhEGNc0jWG07rhoMdeX6kcr9Py9a20cjr5EtAAszKOj8zzmJklKJ7IOF4NCeiQls/vd7FGU90c9vc4s5ROtluEvXYiIiIiIiIiZaOW7Ce5dwPrxBiXrL2+wJ/I/roq+VxZ1qHl67sK6BlrRF2vPQnon2Yc/0+ex3w6Y/wZGcerIQF9OC2f35x4w6lafYA7aN+s5xXAd+MJU0RERERERKRzfk7rxoQeeImWy9Kl9G0PvEDr1zIJjI4xLimMXWn5Or8YbzgF0Z4E9ABaly74ShuP9+WMcauwkhzpqiEBvRktn99H8YZTlTYFnqJ9yecFwO7xhCkiIiIiIiLSNfYDFtL6pHc1Nmu2R3yhSTvUYEu4V9D6NfwEOCy+0KSAjqPla31vvOEURHsS0AA3Z4y5rI3Hm5Ix7k9ZxlRDAjrAku/pz7FvrBFVl73J/jc32+0FYPNYohQRERERERHpYpsCs8l+AvwfYM/4QpMcdgKeJPvr9gywZXyhSYGdSsvX+4/xhlMQ7U1AH5Yx5uP/b+/Oo+QqyzyOf9OdfSEJZIEYCKIkICAEEAUiikRcENfDzLghDg46ogLH0SgzCB71HJxRZ0RFcEMz6CCOAuIQFHAUAZGBCLIEgsSYQEIWspB9654/nqqTqurq6rpV1ffW8v2c855UV3dVPbdu1T2pX733een7xdkwYHXJ35VbeLUTAmiAFRRv4wuyLadjfIC+4X9/4xZihr8kSZIkSW1jJOXDl16iTceX8cNwsxhHzE7fRfn9NR8YlVl1SsNHKN7n38i2nEFRbQDdBSwr+bu3lfzN20p+v5I4e6BUpwTQiynexlnZltP2hgFXUF3w3At8CejOpFJJkiRJklJwJrCc8h+KnyPaPRhuZmMYcB4RnpXbP6uAszOrTmkygC72hZK/u7Hk9zeV/L6/RTk7JYB+kuJtnJltOW1tMnAH1QXP24mzGyRJkiRJansTgKuBHsp/SF5OBKHlZhCq8bqAi4E/039wcT0wKasClbrSFhzfz7SawZEkgD6U4uPVLmD/3O+mED3tC+/n8H7up1MCaFtwpOMk4BmqC59XAidmU6YkSZIkSdl5LbCE/j8wPwacS7TvUOONAM6h8j5YBrwxo/qUndKWErdkW86gSBJAA9xd8rcX5a7/eMn1v69wH50QQHfTtw/xmEwrak/nUX2/5weBGdmUKUmSJElS9kYTs2830P+H51XAZ4GpGdXYbiYDl9B/q41e4HngM8DYjGpUtmZT/HpYlG05gyJpAP0PJX/7cO76h0qu/1CF++iEAPogirdvdbbltJ1K6yn0d/bK6EwqlSRJkiSpyexHLIy0jcr9K68BXp5Rja3ueODbVH6O8+Nx4JXZlKkmMJ7ilhM7ab/e7EkD6H2ALSV/XxpKbwMmVriPTgigz6B4++7Ntpy2cjDwANUFzz3EF41DsihUkiRJkqRmdiDwPaLHaqUP14uBy4jerOrfIUQI8TjVz5grDDC+A+ybetVqBqU9wU/OtpyGSxpAA1xb8vdbS37+rwFu3wkB9Oco3r52XMAyC6cDa6nu2L2BWPBXkiRJkiRVMAP4N2A9A3/Yvhf4KBFeKxb8Op/oWdvfQo9JxnNEv1Fn0nWWH1L8Org023IarpYAem6Z2xSO0we4fScE0PdQvH1nZ1tOyxsCzAN2U93xehH9L4IpSZIkSZLKGEuEn49R3Yfvp4CridlfIzKoNwvdRIh1GXA/1YXOzzDwLPPScRvOOO8k76d4/y/MtpyGqyWA7gL+WuZ2vcDTxHuxknYPoKcCe9i7bT3EF2Kqzf7AHVR/jL4O+/ZLkiRJklSzLiJU/inRC7qaD+ObgBuBC4ETicWb2sEI4BXAx4CfARup7vnYDtwAvJUIyl4E/HyHdHYAABFlSURBVLLK2+bHNiLo7pRwv5NNoe+syyMyraixagmgoW+Lifz4QhW3bfcA+iKKt+3/si2npb0aWEF1x+XdxCxpz1KRJEmSJKlBJgAfAH5D8Wy7gcZO4D7ga8B7gcOAoemWnthQYBbwHuAKot3IDqrf5h7gTuCD9N/L+SxgVYL77CV6cJ/WyA1VUyqdffmVbMtpqFoD6BcD68qMmVXctp0D6CHAoxRv27xMK2pNSVturCFaw0iSJEmSpEFyIPBJImRN2lIiH0o/TsyU/lci2D4FmEZ64fRQ4ABgDnAu8EVipvIikoXNhbPh7gY+RfTSrsZ+wHdJ1jO6B7iGOO1e7eldFO/zjcDETCtqnFoD6Hq0cwD9Joq3axdxXFP1krbcuBeYnkmlkiS1GE8TkiRJjTKBmAn2euANRIhcr9XEDLPVwMqCn/cQYVwPsJkIW7YTLSpGAqOAYUQ/ziG52rqItgaTiWBmau7yZOr/P9GzwK3AAqJX8/oa7+cU4CqSLWK1Efgs8HXieVD7GAkspfhLhs8SbVha3eFEb/lCPwDOGcTH/A7xJVOh44EHBvEx0zCEWHzwFQXXXQe8M5tyWtJpxMKf1X6hdxXRXmrHoFUkSZIkSZIGdDRwAfAjYnHCpDOJm3ksJQKei4DZNPZL/eHAJUSYnqSmR/FU8HY0j+L9vIU486DVOQO6cf6OvmdHHJtpRa1jGNE/vNqWG88Tz7ckSZIkSWpCk4nTxD9HLL73DNkHydWMlcSs5s8Dbya9lhe1LFLYC9wMvDClGjX4xhEz/wv38bWZVtQYBtCNMZb4Qqxwm27MsqAW8kKiXVKSL/naaSFQSZIkSZI6wjgiAHoX0VrgOmAhsShfkn7I9Y5VwB+BHxMB+buBlwH7DN6mV62WRQq3ApcT4ZRa3z/Sdx+/PdOK6mcA3RhXUrw9O6huQcZOdzawieqPqfOB0ZlUKklSG7AHtCRJalbdFPdszl+ekvv9eKKv81jiNOp87+d8L+hdRH/oXmBD7jZridmkpT2ldw/61tRnItH39yPENlfraeCfifBEraub6FN8dMF1q4jw9JlMKqqfPaDrdwZxxkPhZ7ovAZ/IppyWMB74JtX3x94EfIhoJSVJkiRJktT2TgEeIvkM79tItrChms8xwE6K9+tCWndW5kxgXcm4cpAf84oyj3l0xVs0r1nEF2uFr4en8KyHSk4FllP9cfNBnE0uSZIkSZI60FBiJvQ6koXQO4Ev0xxtRVSbz9B3v7ZDP2glMxFYTPHrYBcwJ8uimthwoiXRHqo/Xl5FnFUjSZIkSZLUsSYBV5MsVOkFngXOI4JstZYu4Cb67tN5WRalVA0DfkXf18BHsyyqib2UOFOg2uPjWlq/v7okSZIkSVJDzQbuInlbjseJBQ7VWsYCD1O8L3uAi7IsSqkYDvyMvu/la7IsqkkNJb6Y2U71x8Q7gOlZFCtJkiRJktTshhBh8jKSB9H3ACelX7LqcDCxeGbpvrwkw5o0uEZQfvb73bnfaa8jgPuo/hi4i1jkNckCr5IkSZIkSR1pDBGkJJn1l59Bez1wSOoVq1anAlvpuy8vzbIoDYrRxEKipfv6CWBKhnU1m26Sz3peBBybRbGSJEmSJEmt7FDgf0g+G3oH0Vd6cvolqwavATbTdz9+k2jXoNY3HfgDfffxo8ABGdbVbF4E3Emy49184ks7SZIkSZIk1egtwFMkD6LXAhdiiNkKTgY20ncf3g/MyLAu1e8UYCWGz5V0A/8EbKP649sa4tgoSZIkSZKkBhgJXEz5kHKg8RTwN0SPaTWv44gvDcoFbadlWJdqdx6wk777dCEwKcO6msls4ouWJMe0G4CpWRQrSZIkSZLU7qYAX6d8qDXQuBd4VfolK4EjgSfpu+92AZ8BhmVXmhLYH/gZ5d+HPwfGZ1da0xgDfIl4bVd7DHsOeHcWxUqSJEmSJHWamcSCgz0kD6LvwiC6me1D/+Hlw8AJ2ZWmKpxFzFov3Xc9wOVAV3alNY03AH8h2XFrAdFLW5IkSZIkSSl6OfA7kofQvcBtwPHpl6wqDAHmAXvou9/2EItMjs2sOpUzDbiJ8u+1jdivGKJtxnySHac2EK1MJEmSJEmSlKEzKd+6odogenb6JasKb6T8Ana9RG/vd+KM2qyNJdqjbKL/1jeHZFZdcxgCnE35meGVxq3AgRnUK0mSJEmSpDKGAxdSfiG7gcYe4EfArNSr1kAmEDOe+2u38gjR9kHpGkbMzO3vC4KtxCz27qwKbBJHA3eSfNbzubhwqiRJkiRJUlMaC1wGbKO2IPp64MVpF60BnU7lvrm/Bl6RWXWdoxt4L7CE/vfF7TjreQLwVZItMtgL3IyzniVJkiRJklrCDOBayvcRHmjsBK7CRb+azVjg34Ed9L/vfge8A2feNtp44ONU/hJgNc7c7SJmhidtt7EMeHMG9UqSJEmSJKlORwE30H8Lh0pjG/A1IsxW8ziIaMuxm/733RKiBcTEjGpsFy8ELgfW0/9zvTn3N+MzqrFZnAjcT7JjzG5ipvS4DOqVJEmSJElSAx1FtNdIGkLnZ0TPB16SetWqZDaxUFulfbcR+DbwKlywsFqjiQUef0HlMwh2AFcAU7Ips2nsT3whkvRsi4ewbYwkSZIkSVLbOQm4g9qC6D3AT4HjU69alZwK/JKBZ7kvI2bqHpVNmU2tG3gd8APgeSo/j1uBbxGzozvZSGKW/UaSHUe25G43NP2SJUmSJEmSlJZTgbuoLYjuJQLPV6ddtCo6gpjtXM0ClA8BnwdOpnP7RY8F3kr0O1/JwM/ZCuBfgElZFNtEuoCzgb+S/Ljxc1ygUZIkSZIkqaPMBe6j9iB6IXAWnb3wWrOZDFxCBKbV7MN1wI+B9xPtFNrZkcAniLMAKi3mWDgeAN4LDM+g3mZzGvGeT3qceBI4I4N6JUmSJEmS1CTmEkFbrUH0w8SsSE+rbx7dwByiP+9AbSVKZ/peD1yQu/3ItAtvkHFE/RcQ2/Ms1T8HTxOL481JvermdDi19ZDfAlwGjEi9YkmSJEmSJDWdLuBvgUeoPYj+M/BhYEzKtauyMcB7gAXALpLt023APcDXiH07FziI5pn1Phw4jGin8Ung+8CjJF8UbwPwPeA1uGBj3nTiOUn6XPYA1wIvSL9kSZI0GJrlP36SJElqD13Am4FPAyfUeB8biH7E3yB6xap5TCYW3HsDcDq19zTeCizOjb8QM4zXEP2UV+fGGiKQrNUwYAowlWgPUnj5UGAmsRhgrTPvFxOh/ALgt8D2OmptJ1OIMP/DwKiEt30Q+Bjwu0YXJUmSsmMALUmSpMFyGnAxMSu0FruBG4l2Bnc1qig1TBdwPBFGvz53uZFtVHYDm3JjNzGbejuwk2jPkDeB+FwzlgidR+XGhAbWArAZ+A0RON8KLGnw/be6SUSP7PNJfhbDaqLdxreIGdOSJEmSJElS1WYD84kQsdb2HAuB82jdnsKdYBhwHNE3eT7RyqLW/Z312J2rfz57+1m7kGB5+xLh8QZq6/N8OTA+7aIlSZIkSZLUfo4gAr2kfYQLx7NEYDUt5dpVm2lEf+VPE/2V7wXWkX3AXDiWAbcDVxJh86nEIoSqbBwwD1hP8ud8D7Ew4YzUq5YkSamzBYckSZLSdghxqv451D6jeTvwQ+AK4E+NKUspmgzMInoxT8/9nO/PPDn378Q6H6OHvb2knwVWsbfP9HLgCaKP85b+7kBl7Uv0ab6Q2mYuLyCC64cbWZQkSWpeBtCSJEnKygHARcCHqG/G6d3EooU/IRa3U3sYTvQSLuztPLLg+rx864fNxOz6fK/ojUQIrcY4iHi/foDYJ0ktJBYnvKORRUmSJEmSJEkD2YcItp6ivlYK64GvAy9Nt3yprR1FtM7ZSW3vy8XAu4lFKyVJkiRJkqTMdAFzgZuJmav1hNH3E4sW1jJTU1IsvFjPe3Ep8R4cmnLdkiRJkiRJ0oCOAa4hWinUE0RvBL4JHJtu+VJL6gbeTiwWWet77i/AuUTbFEmSJEmSJKmpTQUuJRaRqyeIzs+K/iDR8kPSXpOIhQGXUvv7aykx49ngWZIkSZIkSS1nOHAW8HvqD6K3Ea0Fzsrdr9SpjgOuBrZQ+/tpGXABsUCkJEmSJEmS1PJeBfwY2EH9YfRa4ErgZGBImhshZWQk8D7gPup77ywC/h6/xJEkSZIkSVKbmgJ8AniC+oPoXmAJ8Hng8DQ3QkrJi4HLgTXU9z65B3grsWioJEmSJEmS1BEa0UqgcDwKXAYcnN4mSA03img1cxvQQ33vibuAM9MtX5IkSZIkSWou+wEXEgFyI4Lo3cDtwPnA9BS3Q6rVEODVwPeBzdT3+t8J/AA4MsX6JUmSJEmSpJYwhwjPttKYMLoH+APwKWBWitshVeNg4FKilUy9r/U1RLuOA9PcAEmSJEmSJKkVTQA+QoTHjQii8+Mx4AvA8biAobIxDfgocCewh/pf0/cRCxSOTHMjJEmSJEmSpHYxA5hH4xYuzI9lRA/qM4FhqW2NOtFk4GzgZmAX9b92dwDXA3PT3AhJkiRJkiSp3R1BtBlYSWPD6E1EOHgeLmKoxpgOXEAsBNiImc69wDPEQpuT09sMSZIkSZIkqfMMBd4IXEv9i7aVG48DXwXOAMaktE1qbUOA2cDFwD1E//FGvBZ3AD8lZup3p7Y1kiRJkiRJkgAYC7wHuIXGtDcoFwD+mljI8FjsHa29xgPvAL4LrKCxr7s/Ah8DJqW2NZIkSZIkSZIq2g84B7gR2Erjw+heYBVwHbGQ3DE4K7XTHAl8EvhfYCeNfW2tBv4DODq1rZEkSZIkSZJUkzHE7NT/BNYxOGF0L/A8cCtwCXAqtuxoJ0OIwPl84kuHRs9yzs+wvwl4GzA8nc2SJEmSJEmS1EjDgNcCVxILuQ1WGN1LtAH5A/AV4O3A1BS2T43RTbRZuRC4AVjD4LxGdgC/AN4HTEhlyyRJkhKw55wkSZJUuy7gBGLG6VuAWSk85nKip2/hWJbC46qyacBxufEyYA6wzyA91g7gV8B/Az8HNgzS40iSJNXNAFqSJElqnP2B04E3AXOBiSk97kbgEeCBgrEI6Enp8TtNYdicHwcM8mPuAG4DfkK02dg4yI8nSZLUEAbQkiRJ0uDoJmZHv44IpU8g3UUGNwOPAouBJ4Anc5cXE4sqamCTgMNyYxbwEmA2gx825z0N3AIsAG4n9qkkSVJLMYCWJEmS0jEReA17A+kZGdaynAii86H0E8ASopVHp4XTo4GDiIA5P/Kh874p17IbuJsInBcAf0r58SVJkhrOAFqSJEnKxkzgZOCVwEmk0z+6GhuIxRWXAytz/z5TcN0KYG1m1SUzCjgQmJ4bBwEvKLg8nfRD5lIrgF8SgfOvsLWGJElqMwbQkiRJUnOYQgTRrySC6WOBYZlW1L9dRFBdOtaXubwN2JK73W5gU+5yD8Vh63pgHDA09/ME4vNKN3sX8xtBzFgeQcwoz499S37Oj1GN2dyGWgrcCfw29++fM61GkiRpkBlAS5IkSc1pNNE3eg4RSJ8IjM+0ItXiCSJozofOy7MtR5IkKV0G0JIkSVLrOISYGX0MsRhemgviaWBrgQcKxj3As5lWJEmSlDEDaEmSJKm1TQSOAI4rGIcBXVkW1QE2Ao9QHDg/BvRmWZQkSVKzMYCWJEmS2s944HBiYcOZBeNQmrMvcjNbT7TRWJT793HgQeCvWRYlSZLUKgygJUmSpM4xBDiIvWH0rNx4ETAdGJldaZnaQfRmXszeoDkfNq/OsC5JkqSWZwAtSZIkKW8U0VP6EGBamcvTgP1prc8RO4HngBXAEmBlmctLgZ6M6pMkSWprrfQfR0mSJEnZGw3sB0woGBP7+Xk8sE/BbScWXB4HDM1dHkXMvt4NbMpdtxPYkru8HdiWu7yVmLG8gWiPUTjWlblufX2bK0mSpHr8P8MCk57nCEJ6AAAAAElFTkSuQmCC" width="85%" style="display: block; margin: auto;" /> --- ## Infer (potentially) causal pathways with graphical model <img src="data:image/png;base64,iVBORw0KGgoAAAANSUhEUgAABaAAAAJUCAYAAADjBIFJAAAABHNCSVQICAgIfAhkiAAAAAlwSFlzAAASdAAAEnQB3mYfeAAAABl0RVh0U29mdHdhcmUAd3d3Lmlua3NjYXBlLm9yZ5vuPBoAACAASURBVHic7N17fFxluff/z7UmSc/0AKUHChba5jRJSkmlFlALVAUBBbEggii6Qd0bRZ/92yCI+5lHQQQRUTZuDm5FEEQqyhZQkDNSsNAATTJN0oZSjj1Rem6TZmbdvz9WYppD0xxmZs0k3/frNS+SdfxOZk1JrnXPdRsiIiIiIiJDUGNFxcHOEof4vpvimzfRHJMcTDbcRIc32XDjMSI4DmjdZSzgAaOAAmAPsBNwwJbWbbYDieB7WwduI7DOGes8529MGuuSfuSd8tra9Zl9tiIiIiLhsLADiIiIiIiIpMuqmTOHJYcPL8JzhWBF4IrwKcIoBMaFGG0rsBJcA+bV4/yV5tvKgp07Gw5fs6YpxFwiIiIiKaUCtIiIiIiIDBp1s4sLvWTePN/cPIN5wGwgP+xcfdACVONYah5LHZGlRdXVKy0YZS0iIiKSc1SAFhERERGRnNUQjc52HicBHyUoOE8IOVI6bAaWGjxjPo/MiseXqyAtIiIiuUIFaBERERERyRnV5eXjC0guBE4COwmYmsbTbQaagV0GvgvaZgBsdZhvOI+gLzQ4DsCIACOBYQTtPdL199Zag0edc39NOns8Go+/n6bziIiIiAyYCtAiIiIiIpLV4tHoBM9zi8D7vOGOBSIDPOQeYDXQ4GC1ma013Hqc24hv77qCxMa1Y6dsOP7ppxP9PcFTCxbkTVq/fmLE8w52EaYYHOz7djDmphh2BLgiYAbBZIYDkQSex+x3bk/ivpL6+k0DPJ6IiIhISqkALSIiIiIiWef16dOHNx8w6lQc5wEn079CbRKI43jR4eojePWJvOTK9eMmvT6Q4nKqPLVgQd7BGzdOz/P8QvCKfShu7VsdpX9F9j3AIzh396gxYx889IUXdqc2sYiIiEjfqQAtIiIiIiJZo668vBz8bxicRVt7i95bC7bUcEt9Z//wnVsWjcd3pCNnOsWj0dGe51Ua/jwHH2otSve11cg2jMWW5KaieHx5OnKKiIiI9IYK0CIiIiIiEioHtqq85ERn3iXOcQq9/zulyWHPGe5xh/d4cU3Ny4N1cr6V5eVHJPEXGrYQ3MnA6F7vbCzBuZ8VFUf/aIsXJ9OXUkRERKQrFaBFRERERCQUyyorR45uaf4izl0CFPVyt7cMewCSj2wvGPn03KqqXenMmI3emj9/xK4dWxc44yScnQ4c1qsdHavM48YmIr+ZXV29M70pRURERAIqQIuIiIiISEY9tWBB3uRNG79s8H/pXWuJrRh/drC4uKj0LxrF286Bt3J29Bjn3CJ8+zzGQb3Y6T2M6yO7m2+c1djYnIGYIiIiMoSpAC0iIiIiIhnhwOorop8xx9Xsf8RzAngY47fDtu186PA1a5oyEDGnrZo5c1hi+PBTMHeewWlAXo87OFaBu7KodsXiwdq6RERERMKnArSIiIiIiKRdQ0Xph31nP26dUK8nWzF3u9/i/qu0ru6NjIQbhBrLyg5NmLsYuAgYt5/Nl+FzaXE8/lQGoomIiMgQowK0iIiIiIikzWuVlWNbWnZ/H2cXA14Pm64GbhsWyb/18Fdf3ZKheINePBod7Xl83uBbQEnPW9ti15L4ekl9/aaMhBMREZEhQQVoERERERFJi/qK6Kdw/AI4pIfNXjPjysLq+H0GfqayDTUOvIay0s+CXYUxq4dN15qzfyuqrf1TxsKJiIjIoKYCtIiIiIiIpFRjRcXBCZe4CeysfW1jsB74wfaC4bfNrapqyWC8IW1ZZWX+6Obmr2DuP4EpPWz6hxZnF5fX1q7PVDYREREZnFSAFhERERGRlFlZUTrPd/YHYNo+Ntlh8OOEzw3ReHxHJrNJu+UVFaOG4X8L5y4Dxuxjs3eds0UltbXPZzKbiIiIDC4qQIuIiIiISEo0VETPd45bgBH72OQZP+JfWPpq3apM5pJ9q59TNJVE/s3gTt/HJgngyuKa+LWZzCUiIiKDhwrQIiIiIiIyIKtmzhyWGDHsJoML97HJFgeXFdfEbzdwGQ0nvVJfXroI7GZgYvdb2N07CoZdNLeqaldGg4mIiEjOUwFaRERERET67fUjjxzXnGx5GDimu/UO/uT7/Gs0Hl+X4WjSR0Hv7uTNwGe73cDxYtJxcjQefz+zyURERCSXqQAtIiIiIiL9Ul1ePr7A+Y9gHN3N6iTw3aKa+HUa9Zxb6sqjFxn8F5DfeZ2DFb7PibqhICIiIr2lArSIiIiIiPRZ62jZx4CKLisd73nmn1NYU/d45pNJKjRUlH7YObsPmNzN6nryEicWv9LwbqZziYiISO5RAVpERERERPpkRUnJFC/PewIo6Wb1sqTPmdF4/M1M59qf+qKiMV5BwT56HHdVWFOzur/nqikrmzTMbFRvtm3x/V3ZOKJ4VUXFtKSfvH8fI9wbzMs7sWj58ncyHkxERERyigrQIiIiIiLSa2/Nnz9i545tzwJzu6x0PEdL4pPFDQ3bM59s/+oqol80xx293d43f35pdd0/+noeB9ZQHm0EjujdHvaX4praU/p6nkxYVlk5ctSe5v813MJuVi9vtsixs6urd2Y8mIiIiOQML+wAIiIiIiKSGxzYju3bfkV3xWd4Juk4OVuLz/3hYV/sz34NFdHj6XXxObvNraralbe76VQcD3azenaBS97lNLBJREREeqACtIiIiIiI9Ep9RdnlZnyum1WPjBp9wMnReHxHxkOlk7PPvTV//og+7+bTr8J1tprV2NicdHwW7IHO6wzOWFke/V4YuURERCQ3qAAtIiIiIiL7VVdWdqo594NuVj0f2d18+qEvvLA746HSb9yunds+3Zcd6ouKxphxZroChSUaj+9J+u5shz3beZ2DWENZ2Rlh5BIREZHspwK0iIiIiIj0aNWcORPN3P/Q9e+Hd8lLLJrV2NgcRq5U+MeOndz3/mbue38zj2/r2j3E+XypTwfMzz8L6DD5YENT0z/Psfj9zQOJG6poPL6HlsRngM6TM5oz9+tVFRXTwsglIiIi2S0v7AAiIiIiIpLdkomW24CDOy3e7fA+VfJKw7thZEqV+zdv5sEtWwHIN+PvJUWMi0TaNzAWNsyefUjR8uXv9O6I7kudl/xk3Xqe3R50J4mYsWjC+AHnDktJff2mldHomb7Hc3QstI9N+slfAJ8KKZqIiIhkKRWgRURERERkn+rKyk4Fd3qXFc4uKqmtqQohUkodNXIkRcOHUzpiuF84fLjXofgciOAnvgD8aH/HWnFkySySHNt5+Y2HHUpjU7Orb2qy2t2536mkMB5/tb6s9MuY/b7DCuO0+vLSTxfXrPjfkKKJiIhIFlILDhERERER6dayysp8M/eTLiuM3xfX1v42hEgpd+q4sfzLxIM4ZvRo76C87sfnOHo3qaCX8L4IWOflIz2PipEj7KwJ4/nWpM4DyXNTce2K+8Du7rrGblg1c+awzCcSERGRbKUCtIiIiIiIdGvMnqYvAYWdFr9v+S0XhxAnTMUrKko+1NMGDjyM8zMVKBu4lsQlON7rtPiI5MjhXw4lkIiIiGQlFaBFRERERKQLt2hRxDn+o8ty52JFVSs7Fx0HPQ/rcRR0fVnZQuDQDMXJCiX19Zvw7D+7rHDuO8sqK/NDiCQiIiJZSAVoERERERHpoqFhxRkYszotXr1z2IhbQgkUNmfnvDV//oh9rfas6+SDQ8HaCQfdjmNVp8WHjWlu/kwogURERCTrqAAtIiIiIiJd+fxL50UOrp1bVdUSRpwsMHbXzm2f7m7Fa5WVYx10u26wO/7ppxNg13Ze7sxdGEYeERERyT4qQIuIiIiISAf1c4qmYnys0+L3h2/feWcogbKEc91PRrhnz+7PASMzHCdrJJ27C9jYafEJK0pKPhBGHhEREckuKkCLiIiIiEgHLhE5jc5/K5j77eFr1jSFkyhrfLyxrKxLn2cP70shZMka0Xh8j5l1vjlhluedFkogERERySoqQIuIiIiISAee2amdlznf+30YWbKMlzR37t4L6mYXFzrcvLACZY2k63J9mKPLdSQiIiJDjwrQIiIiIiLyTw7MOY7ttHhDcW3tP0IJlGUcXODA2r43P/Jl9vp+qCqMx5cBazssNOY7/c0pIiIy5OmXARERERER+aeGiopCYHyHhY6/G/jhJMo6hXUVJfMA3KJFEeDc/Ww/JBg4jGc7LT6grqKkNJRAIiIikjVUgBYRERERkX8y3+9aMDReCiFK1vKwLwKsrK/9BDAt5DhZwxxLOy+L+JFoGFlEREQke6gALSIiIiKSGkcC1wBfBRYCRwB5oSbqBx//iK5LXX3mk2QxZ+e8NX/+CEfkSyEnyTJ+Q5dF5g4PIYiIiIhkkZz7hVhEREREJEu9CkwA7gYmty5rAd4AVgOvtT72/npn5mPuh9khnRd5vr0RRpQQ7QEKelg/dtfObRcAn0rBsQYNz/fWJDsNcfI1QlxERGTIUwFaRERERCR1ngQqgLuATwD5wMzWR3c2ExSku3usIZy+y6M7L0j4/uYQcoTGwcMGZ/S4jePHwLD9HGgVZs3gylKZL1u1+P5mz+tYgfa6uZ5ERERkaFEBWkREREQktTYCnwSuBP4TiPSw7XigsvXR2U66HzW9mmBUdUvqIrcz3AiwDssK8vN3p+Nc2cqMO3A9F6CBkfs9kGe/wbmzU5Mq++W3tOxM5nWsyTsYFVIcERERyRIqQIuIiIiIpJ4PfB94BrgHmNqPY4wiGE1dsY/1m4EVQJyOI6cbga39OF8rrwVchyVJs/z+Hy/3JJOsiHi8BHxwAIfxk0l3V8RjyBSgGTNmGIk9nZd2WSAiIiJDiwrQIiIiIiLp8wxwFEFLjo+l+NjjgWNbH51tpPuR068B7/Z0UMPtdJ2WOeeGXBsFg9+4ARSgDZ6IxuNv1pdHUxkrqyWTyS7XicvGPuciIiKSUSpAi4iIiIik13qCftDfBH5M0Bc63Sa2Pj7Uzbo9wNt033e6Dufewzq24DAS04CGdAbONgmf30U8rgeG92d/H+5IbaLsZySmuU7tWzznNoYUR0RERLKECtAiIiIiIunngJ8BrwC/o38tOVKlADii9dFFNF7XVDR8OIcW5HNoQQEzhg0j6TgeqAK2ZDJomKLx+Pv15aV/BjurH7tv3Vkw/IGUh8pyPnaEdV5otiaEKCIiIpJFVIAWEREREcmcZ4E5BC05Ph5ylm4lnBse372b+O4O8w5+t/WxiX239niHzs2jc5zBHQ76XIB2cN/cqqpd6ciUzTzH3M4XgO9sVShhREREJGt4YQcQERERERliNgAnExR0EyFn6asDgaOBzxHk/xXwNPAY8AWgywDYXFZYHP0bQbuSvnF2R8rD5AAH8zstSlpLS1UoYURERCRrqAAtIiIiIpJ5PvBD4ESCkcO56ingNCAK3MlgGwG9eHESuKdPOzlWFdfWvpCeRNkrHo1OJhjd32FxcUPD9jDyiIiISPZQAVpEREREJDxtLTkeCTtIH7QAdwOVwAnAQwQF9UHJ6+NoZufZHTbICvG94Xnu03T++9LxaDhpREREJJuoAC0iIiIiEq6NwCeBbxEUd7PSCM/zgZ8DM4HzgJfDTZQZhbW1dQ6W9nLzZB7enWkNlKUM+0qXZRH+HEYWERERyS6ahFBEREREJBxTCUYRlxK0sKgEIqEm6sYhBfl8bsIEzp4w3huX591TWL3izbAzZZo5uwNz83qx6ROzqqv73jM6x9WXlc0F98FOi98sXB5/PpRAIiIiklVUgBYRERERSa+RwGyCVhtHAUcSFJyHhxlqf6YWFLx96eRJ0z52wBgiFswt6DuuAD4dbrLMyx827Hcte5puAEb0tJ0z7shMouzizF3RefZJc+5XNohbs4iIiEjvqQAtIiIiIpI6BwAVBKOZ20Y3lwMFYYbqAx94ALjh8RkzVnt53mo6FspPa5gdPbpoefzFcOKFY0ZV1da6suj/mvG5HjbbMnrUAQ9kLFSWqCsvrzT8zjclmonk/zKUQCIiIpJ1VIAWEREREemf8bS3zmh7lACdB4Pmgp3Ar4EbgdcASuvqqC+L/hrj63ttZ/j2MwfHDLWJ9kpq4+cA54SdI/u4G+gyt5DdUbR8+TuhxBEREZGsowK0iIiIiMj+TSdonzGH9lYaU8IMlCLrgVuAm4BNnVcmHT+KGBew1yhoh/vQyvLov1ATvz1zMSUb1ZeXXgDuI50WN2PetaEEEhERkazk7X8TEREREZEh5QDgOOAy4EFgA/A6cD9wJXAKuV98rga+SlBYj9FN8RkgGo+/ac7d0Hm5g+vrjiyensZ8GTPCy+yfRKMjWTfPZL+sqqiYBtbl2gBuLK6ufj3jgURERCRraQS0iIiIiAxl+UAhcCxB0TmX22j0xhLgWuAhetlCI+HsmohxPjBtr8UHWDLv/mWVlR+eW1W1Kw05MybPMvtSF2T4fOnw+vTpw5v95P0Y4zqtejeyu/mHoYQSERGRrKUCtIiIiIgMJVPpWGyupOMke5m2FogDK4C3CUYjj0zxOfYAvweuA2r7unM0Ht+xsrzkAh/vb3QozLujRjc33elg0VDrBz3UNY8ZeTNwdOflhv9vsxobt4UQSURERLKYCtAiIiIiMlhNIiiSHQ3MAz4IXUZsZsoeguLvK3s9aoDtreuHAy+Q2uLze8D/EPR3HtCEcIU1dY/XV5TeirOvdVhhnFlfVvodaldcM5DjS+6oryj7D5z7ctc17ldFNXUPZD6RiIiIZDsVoEVERERkMPCAMoKRzccQjHKeHlKWncBy2gvNLxOMct7Twz43AUem6PyvtR7vdiBl7TEiu/Z8KzliWAXBz/efzOyq+vLSN4trVtydqnNJdmooKzvbOdf1ZoPjxWE7dv1bCJFEREQkB+R+AzIRERERGYpGAkfR3k7jGGBCCDm2EYxkrtrrUQ8k+3CMs4F7U5BlCfAz4I99PH+v1ZSVTco39xJwaKdVSZxdVFxb+6t0nDdV4tFoQdLzRu1vu/Kamq0GfirOuWrmzAN2jxjR48yDEd9vicbjO1JxvnRpqIh+zjnuousgpnURi3xwVnX122HkEhERkeynArSIiIiI5IJJdCw2H0UwgWAmvUvHFhovA2sGeMyZBEXrA/q5vw/8Bbga+McAs/TKqtmlc5K+PUfXdiHOOXdJSe2KmzKRQzKnvqzsK5i7jeCTBntr8swtKKxesTSMXCIiIpIbVIAWERERkWx0BEGxua3oXEJmf3fdQdBGo21U83PA6hSfYzjwPDCnH/tuB34N/AR4M5WheqO+rPQszO4BOo/sdc65/yipXfGTTGeS9KivKLsE535K1/df0hlfKKmO/y6MXCIiIpI7VIAWERERkbDlAXOBD9M+wvmgDJ4/QdCjeWnr40WgjjS1sdjLLcBX+7jPGuDnwC9pn8AwFPXlpYvA7qb7kej37CgYfuHcqqqU9aCWzHp9+vThzaNH3YTxL92sTpqzC4pqa+/KeDARERHJOSpAi4iIiEimRQgm3Gsb4fwxYFwGz7+WjiObXyCYODCTzgJ+34ftXybo73wPQcE8K6wsKzvFN/cHgtHcnS338D5TWFOT6pHjkmaNZWWHJnB/wDi6m9UtzjinpDp+f8aDiYiISE5SAVpERERE0s0jaKFxLLCw9TE+Q+feDlTTXnB+loH3bR6o3vZ9buvv/DPg8XSH6q+G8tKTHfZHui9Cv+85O7+wtvbhTOeS/ml9Pe8CDuxmdbMzFpVUxx/MdC4RERHJXSpAi4iIiEiqeUA5cHzr4yNkboTzaoJRzUtaH3UEhdxsMZxgxPWRPWyzE7gDuBFozECmAWuYHT3O+dwHTOluvcFvEj7/JxqPv5/haNJLrx955Ljm5J6fgF1AN38nGqz38c4uqal5JoR4IiIiksNUgBYRERGRVDiC9tHNJ9D96MlUSwINtBecnwHeyMB5B+K/ga/tY916gr7QNwGbMpYoRRorKg5OkLwXx/Fty57evp1RnscHR43CYD3GpUXV8TvDzCldtbZSuQWYto9NqvyEf2ZpXV22v79EREQkC6kALSIiIiL9MYOgd/OJwEeBiRk453aC0cNLCIrOS8l87+aB2Fff52rgpwT9nfdkNFGKxaPRgojxU2f86y83vsdP161nfF4e9888gsn5wVyFDv7kEv63VcwMX2NZ2aEJ868HO2vfW9mtkd1Nl8xqbGzOXDIREREZTFSAFhEREZHeGA18iGCE82lAaQbO2TZZYNsI56VASwbOmw6d+z474G/ATwj6O7uQcqXD+GkFBU+9vWfP7LYFFxx0IJdNmbz3Ns3Af1tBy9VFVSvfy3jCIa6uuPhAK/C+g7OL6b53N8B2cJcU16z4dSaziYiIyOCjArSIiIiIdCePoOD8sdbH0UAkzeesB54mKDj/HXgzzefLlGHA88BRBCOcfw/8GKgJM1SazAb+SNCSBQPOO/BALpsyiTzr9k+PHcDNkd3NP5zV2LgtczGHpmWVlSNH72n6BvAdeu7L/kjS56vReHywvAdFREQkRCpAi4iIiEibvfs4f4z0Txy4mvZ2Go8weArOnd0MnAv8BrgOeCfcOGlzLnAbMLL1++3AV+rKo/nm+BnGQT3su84593MS/m0l9fU51/8628Wj0QkRjwuBS9jHRJGtNpmzbxfV1t6VoWgiIiIyBKgALSIiIjJ0jQaOB04lKDgfnubzrSUoNj8OPEr2TxiYCkcSjCS/E9gVcpZ0yQOuAi7ba1kDcCYQB1g1Z87EZKL5BrBz6flvkF2Yu9Nc5GdFNTX1aUs8RNTNLi60ZORbGOcDo3rY1Bncu8fZt8tra9dnKp+IiIgMDSpAi4iIiAwdEWAe8EmCgnMl6W2rsYagpcZTrY+30nguCcchwGJg/l7L/hf4IrC188Z15eWVHv41Lrj+euKARzxnN79z0EGPHv/004mUJR7knlqwIG/qpvUfx7yvO8cnAW8/uzxhPpcXxeMvZSKfiIiIDD0qQIuIiIgMbhOAEwnaanwKmNzz5gOy9wjnxwlabMjg9WHgPtqvqSTwXYI2Iz1OqlgfjR5LhGtxHNuL82w2WIzHXYXL40tscE3YmDLx2aXRiG9fICj+9+J9brXgf7+4ZsXidGcTERGRoU0FaBEREZHB50jgZOAUgvYP6Rrl/D7wBPAY8CTwWprOI9nnIuC/gPzW798DziG48dArDqyhvPSz4H0H3FG92ceg0Te723z+WFxbW93n1INMXXl5uYd/hgv6bxf2aifjVYNrC6vjv1cxX0RERDJBBWgRERGR3DcSOAY4DTgDODRN50kCr9I+wvkZoCVN55LsNBr4H+CsvZZVEfR77ndP74bZ0eOcb98EdwZBT+ne2AD2jJl7qNl5D1bU1Gzu7/lzxfKKilEjXMt83+w0nJ0OHNbLXX2HPYm5nxdXxx9S4VlEREQySQVoERERkdx0BEFbjdMI+ukOS9N5VtNecP4b3fT1lSGjELgfKNtr2W3AN4A9qTjBqrKyGUnP/ybOLgDG9GHXhMOe95z/JMaLCd+WRuPx91ORKUzV5eXjC8yfZ46jXTBh6LG0jzrvjR0GdyQj/s9LX61blaaYIiIiIj1SAVpEREQkNxQAHyGYQPAUevtx+757j/a2Gn9DEwdK4DTgTmBc6/dNwMUEo6FTLh6Njs6L8BnnOJegh3l/2sisBPeiw5ZGzL1ku/bUzWps3JbiqClTX1Q0xobnlfg+R5txNI55wCz6/jdb0mFPYe631pz4Y3FDw/Y0xBURERHpNRWgRURERLLXKOAEYBHBBIJj03COBLCcYITzQ8DzgJ+G80huigDfa314rcveBD4LvJSJACtKSqZ4+d7ZOM4DKgd4uHcxGnC20nAN4Oqd5a2O7Nq1NhPF6fqiojEMGzYVkocDxeCKcFYYfM3UgRzbwSse/NblJe4tfqXh3ZQEFhEREUkBFaBFREREsss0gmLz6cBHCUY+p9rrwF9aH88AO9NwDsl9BwH3ELR4afNX4DyCCSgzrr6ioshc8hTgJBd8IiCVrWeagA0O1ppjAx4bcLxnzu12Zk0Ge3zY6Rm+c669FY3zDnDmIh6MclBgzg13ZiNwHAhMMrOJDjcVmAiM2On7rG5u5rCCAsZGBjQ/aDPwnHP2iGf2UFFNTf1ADiYiIiKSLipAi4iIiITvCIIWB4sIJhNM9e9oSeAfwIMEI52rUnx8GXyOIuj3PL31ewdcB1xBloyQf2v+/BG7d2w51sdbSHDTpiTsTPtzy4aN3Lh+AwA/PWwaJ4/t84ca1hj8zeEezy8Y8bcZVVXqyS4iIiJZr7czTIuIiIhI6njAHIKi89kEH79PtY3A0wRtNf4MbEnDOWRwOh+4BRjR+v024EvAn8IK1J1DX3hhN+0TZH6nfk7RVJfMqwSrxHGs4Y4BRoabsqNJ+e3zB65u3u+8jS1ANeaWGFaVMFdVunzFCgtuBoiIiIjkDBWgRURERDJjBLAQOJVgtObkFB/fB15BvZyl/4YBNwEX7rVsOXAm8Fooifqgte/xuwQj/XlqwYK8aZs3lCd8jvawMqDIGYU4DiOkT4IeMay9Y8jq5ua2Lx3wlkGDDyvNWa3vJV9cP2FS9fFPP50II6eIiIhIKqkFh4iIiEj6TKC9n/PHSP1ozC3A34CHgUeADSk+vgwdhwJ/AI7ea9k9wEUMsh7hyyorRx7Q1FTomyvErAjsEHCTCXo0T259pPK9ugtYD6zbkkhsnl/X8EkHjItE3niprOiMrXkjGuZWVe1K4flEREREsooK0CIiIiKpdSBwBvBZ4AQgv+fN+2wFwQjnvwBLAI2QlIE6HrgXOLj1+wRwJXBtaIlCtryiYtRI35+cMBsHFERIjEq6iGfmjwUwbJwP5oFzuC0AzrNtEd9PJsnbCezxfX9rIhJZO7u6unMB/23gEIJJD0cT9GgXERERERERERHZpwkEfXMfBPYQfKQ+VY8ksAyIkQOTrElOMeAygoJz2/X2DsFEmJI+j9H+854RY8nPQAAAIABJREFUchYREREREREREclSB5K+ovNugiLVJcCUTD0hGVLGELTc2Pu6exZdb5nwc9p/5qeGnEVERERERERERLLIQbQXnVtIbdF5E3Bf6/HHZOoJyZBUTNDKZe/r71ZS3y5Guvd12n/u/xFyFhERERERERERCdk0gonY0lF0fp2g8HcaKv5JZnwO2EH7NbgdODvUREPPAtp//r8KN4qIiIiIiIiIiIThEODbBJP8+aSu4OwT9HO+EijP2LMRgTzgR3S8Hlei6zAMk2h/DV4IOYuIiIiIiIiIiGTIBOBC4CmCif9SVXROAE8DFwOHZurJiOxlIvAEHa/LB4FxYYYa4t4jeB22hh1ERERERERERETSZwRB+4v7gGZSV3ROAs+hSQQlfMcC79LxhkgM8ELMJMG/D22vif6NEBEREREREREZRCLAQuBOgv63qRzp3FZ0npyxZyOybxfR8cbKe8DHQ00kbW6n/XU5MeQsIiIiIiIiIiIyQB5wHPAzYAPpKTpPytizEenZcILJ7fa+Vl8GDg8zlHTwf2h/bS4OOYuIiIiIiIiIiPRTlGDitXdIT9H54Mw9FZFemQksp+M1eydBuxnJHifT/vrcHHIWERERERERERHpg1LgB0AjqSs6NxFM2nY+mrhNstcpwPt0vG4vCjWR7Mt02l+nJ8ONIiIiIiIiIiIi+zMR+CbwEqkrOrcAfwG+AIzN3FMR6TMDLiOY/LLt+n0LmBdmKOmRB+wgeK3WhpxFRERERERERES6MQw4DbiPjhOtDfSxDPV0ltwxAfgrHa/hp9D1mwuqaH/NxoecRUREREREREREWlWS+skE40AMOCJzT0NkwI4EXqP9OvYJep5HwgwlvfZb2l+7+SFnEREREREREREZ0qYRtBhoIHVF59cJinXFGXweIqlyHrCT9ut5G3BmqImkr75L++v3lZCziIiIiIiIiIgMOSOARQST/yVITdH5bYLR08cR9M0VyTXDCK7hva/reoLJNyW3fIb21/D6kLOIiIiIiIiIiAwJHkFx+FZgO6kpOr8P3EnQL1qtCSSXHQK8QMfr+0/AAWGGkn4rof11fDjkLCIiIiIiIiIig1ohQTuMd0hN0XkXcDdwCpCfwechki4fAdbSfo23ELSl0Uj+3JUP7CF4PVeHnEVEREREREREZNAZTtBi4zGCydNSUXheBlwCHJjB5yGSTkZwTbcVKh3BBJwnhhlKUqaO4DVNAiNDziIiIiIiIiIiMihUEvSw3URqis5vEIyenpHJJyGSAaOB++h4vb8EHBZmKEmp+2l/bY8MOYuIiIiIiIiISM4aD1wEvEJqis5bCPo6L0QtCGRwKgRq6Xjd3woUhBlKUu4q2l/fc0LOIiIiIiIiIiKSUzyCAvF9QDMDLzonCNp1nI8+qi6D26cIbrK0Xfu7gS+HmkjS5VzaX+fvh5xFRERERERERCQnHEowOdrrpGa0c7z1eJMy+SREQhAhaCezd0/0N4C5YYaStDqK9td6cchZRERERERERESy1gjgPOBJUjOh4DvAdUBZJp+ESIgOIhjhv/f74GGC9jUyeI0kmICw7WabiIiIiIiIiIjspZRgQsHNDLzovIdgQq5TCEaCigwVlcAa2t8LPsFIaC/ETJI5a2j/NzA/3CgiIiIiIiIiIuEbBiwiGK2ZitHO9ajFhgxdF9GxR/pW4PRQE0mm/ZX2178o5CwiIiIiIiIiIqGZSTAqcwMDLzrvIpiccCFgmXwSIlliOPBLOr4vXgGOCDOUhOIG2q8B3XwQERERERERkSElApxG6kY7LyMY8Tk6k09CJMscCrxIx/fGbwn6AcvQcyHt18HlIWcREREREREREcmIQwjaYrzJwIvOawn6RJdn9BmIZKeTgU20vz9aCN5rMnQdS/v1cGfIWURERERERERE0sYjaIlxH5BgYEXnJMGo6UVoUi0RCFrNXEbw3mh7n7wNzA8zlGSFCbRfEy+FnEVEREREREREJOWmAf+PoBg20NHOqwiKbFMy+gxEstsBwB/p+F55BpgcZijJKusJrosdqC++iIiIiIiIiAwSxxGMdm5hYEXnBMFo59NQ4USkswqgkY7vmVvRJwOko6dpvz4OCzeKiIiIiIiIiEj/jSaYBLCagY92fhv4EcGEaiLS1ecJRrS2vWe2A2eFmkiy1X/Tfp18IuQsIiIiIiIiIiJ9NpOgWPw+qevtnJfRZyCSO/II3m97v3cagLIwQ0lW+ybt18q3Qs4iIiIiIiIiItIrbZMKPgj4DKzw/C5BQW16Jp+ASA6aCiyh4/vnz8DYMENJ1vsYHVu0iIiIiIiIiIhkrbHAJcBqUjfaWf1qRfbvOIKbNXv3R48R3AwS6ckhtF83z4acRURERERERESkW3MIRs7tZGCF53UEo52PyGx8kZx2EbCH9vfRRoJRrSK9tYXg2nkv7CAiIiIiIiIiIm0KgHPo+pH//jyeIpggrSCjz0Akt40G7qXje6kKtauRvltK+zU0MeQsIiIiIiIiIjLEHQz8J7CWgRWdtwP/jSZHE+mPWUA1Hd9TdwIjwgwlOesO2q+jj4QbRURERERERESGqkLgZwy8zcYq4DJgQmbjiwwapwKbaX9PNQH/EmoiyXWX0X49fTXkLCIiIiIiIiIyhBiwEHgQ8EnNpIKRjD4DkcHDCAqFSdrfW28CR4cZSgaFT9F+Td0YchYRERERERERGQKGAecDtQxstPMWgskJizMbX2TQORB4lI7vrycJWuKIDNQs2q+rR0POIiIiIiIiIiKD2GQgBrzHwArPdcAlwKiMphcZnOYAq2l/f/nAj9CnCSR1IsBu2kfVi4iIiIiIiIik1FEEE5jtof9F5wRBq46FGc4uMpidD+yi/X22FfhMqIlksGqb1NIHxoScRURERCQl8sIOICIiMsRFgDOAbwHHDuA464BbgNuAtSnIJSJBG5zrgG/utayeoPhcF0oiyWmxWMwDxgHjksnkeDMb53ne+GQyOQ4Y9+qrr+YlEgkikYjNmDHjN+PGjWt2zkUAnHPjWw/jmdnY1q/z6F2huq0dE8AW55wzs+0ENy13mdlu59xW59y21uU7PM/bDmxJJpPbI5HIpt27d2+89tprt6boRyEiIiJDiIUdQEREZIg6ALgQuBiYPoDjvEIwWdW9BCOnZRB6ffr04dvHjBkxIi8vzyUSYwASlhhnLu+fv8tFnNuFWbPnXMsusx0AFTU1m8PKPAhMA/4AzNtr2e8I3rc7Q0kk2chisdgkgtZJ03zfnwIcAkwl6A0+bq/HeIJ/+3NZAtjU9nDObTKzd4D1wLvAOufcu8lkcl1BQcH6WCzmhxlWREREsoMK0CIiIpk1GfgawYjK8fvZdl98gonPfg48RDCyTXLQa5WVYxNNTYW+xywPJjvHZAeTzJiIMQXHwQRFrIF8am0bsBbHRmC9M9Z5sMHHrTezNyDSUFRd/YYF15UEPgr8HpjU+n0CuBK4NrREknEXXXRR/sEHHzzd87zDnHOHAFM9z5vi+/40M5sMHErwb3p+uEkB2AG0dFo2FvBCyNImSVCUfsPM1jjn3nDOvWlmb/i+/2ZeXt7rsVisKcR8IiIikiEqQIuIiGRGOcFo5/OB4f08xg7gHoIRz/r4fw5prKg4OOla5vrOSjyzQgeFQDFB8SobNIOtwrkG82ylj2uIJFn+zsSJtcc//XQi7HAZZAQ3h66nvei/EfgcwU0fGWRisdjIZDI5IxKJzPB9f4aZzXTOzQBmAIeRupaFuwhaYGwhaIGxxcw27/09sGXNmjVjlyxZcr1zDuCZ884779+BRCQS2d56nD1NTU07AYYPH767LwXcWCzmNTU1jW3ddzxAMpkcS9DSYxwweq/HOILWHqPNbLxz7kCg7XEQ/b+BujcHvOWcazSzVc65xkgk0phMJldt2bKl8aabbmpOwTlEREQkC6gALSIikl7HAZcBp9D//++uJejt/HPg/RTlkjRZNXPmsJaR+XM8IvPMuXkuaOFwRNi5+mmnw6rM/Bed772Q53kvzqqufjvsUP0wBtjei21+BXx2r2XPAWehvuo5LRaLDQdKk8lkoZm1FZdntv536gAO3QS8Q3B9vGNma51zbwPrPM97K5FIrPd9f0tBQcHmWCzW2xZJwwluNkaAlUDRAPKlzaJFiyLRaPTAZDI5MRKJTGltPTLZzKY65yYRtCGZQlDEH9aPUySB151ztZ7nrXDO1TrnVmzZsqVehWkREZHcowK0iIhI6nkEBefv0rF/bF9VERSd7yFoASBZyIHXUFZ2FOZOwjgJxweBgrBzpdE7Bo9jPOLvST5WUl+/KexA+zEK+Hfg+z1sUwT8ESjda9ltwDdQb/WcsWjRokhRUdHMvLy8ct/3y4Aygk+fzCAo6PZFC7DGOfea53mrfd9fa2ZvEfQ4fjsSiayNxWLpuiG4iqBAniAYjZzTBdcrrrhiSkFBwQeSyeRhZvYBMzvM9/3pZjYTOJy+FagTwGtANfCK53mvAK/GYrF1aYguIiIiKaICtIiISOqMBj5PUOwq7OcxfOAvwI+AJSnKJSm2as6ciclE88eBk8A+AUxMw2m2AOuAjWCbgBbwmw3b5RxJjG2t27UA+Q4KPBjlgxk2LljlxoAdDG5Sa8ZUF8aThr0E7q94PFK4PL4sC3tJ/5rg53j5PtZ/GriT9snhmoCvA3ekPZn0WywWO8z3/ahzrtzM2orNpfStmLkbaHTOvWZmr7W2gnjN87zXgDdjsVhYN/7+DJzW+nUZEA8pR9q1jqQ+1Pf9WQRF91nOuVlmVkTwyZHe3jhYC7xqZq845172PG9pLBbLxU9riIiIDEoqQIuIiAzcwcC/EvR4PrCfx9hGUPC6AXgjNbEkleqLisa4YXmf8RznOjiBvo+o3JsPvAmsNKwBXIPv7A3PufURs3WuqWnDrMbGlI96rCsuPjAvEpmU8LyJRnKaOWZiVtTak7qQoA3FQLxjZvda0v22MB5/NQWRB+ocgk8QfB24pdO6POAq4FLafyduBD4D1GQqoOyXXXnllYWe5801s7nOuUqggmCCvd7wgdeBajOrc841ep73WktLy2tXX331O2lLPTDXElyXAIuAP4SYJTSxWGx4MpksMbNSIEpwgyFKUJjuzeSK7wJLnXP/AJZGIpGqWCy2I32JRUREZF9UgBYREem/YoLRzl+gfz0uISiM/JSg9+zOFOWSFFlWWZk/qrn5E+DONePTwIh+HGYHsAxYirEM31YO27Fj5eFr1vR68rBMqZ9TNJU9eUXOXJmZN89w81wwKrHPDOK+c3f7zu6OxuNvpjprLxxO8DH90cDJwCN7rZsI/A44ca9lDxO8lzdnKqB0FYvFjkgmk3PNbC4wF6ikfXT6/rzrnKs1sxog7nleDbAiFovtSlfeNPkSwch9gP9Lz+1jhpxYLDYSqPB9f46ZHemcO4pgpPj+JvhNEowmX+Kcey4SiTyrUdIiIiKZoQK0iIhI380HvgOcSu9GYXVnGXA9wci2ZIpySYrUzymaai2Rf3NmF9LX9hqOVZj7u2H/cM6WFpWUxG3x4px9jRsqCw9iT+RoH2+eB/NdMLFmXwrxvsOejDhunFVb+9cMtejII5hAsK0HeylQ1/r1XOB+gsnRABxwHXAF2dc+ZFCLxWKHAXOdc5W+77cVnSf0YtctQK2Z1fq+XwPEI5FITRp7MmfaPOAfrV/fSzCSX3oQi8XykslkCTCn9TqaBxzJ/tsOrQH+Dvzd9/3nrrrqqrr9bC8iIiL9oAK0iIhI7x0HXEZQeO6vJQQfr34wJYkkpVbNLp3j+/Y1B+ez/9F0bXY7bInhHo/4PDgrHl+Rzoxhe3369OEtY0Yc5+MtBBYSjFDtrdcwu6kZ75ezq6vTOeL/h3Ts+Tya4BMGFwE30V6U2gScCzyaxiwCXHTRRfmTJk06ysyOcc4dZ2bHAJN7setW4GVgmXNuWSQSWRaLxVanN23oDiAoshuwnKCQKn3U2sJjjpkdTVCQ/hDBJyN6sgF4yjn3uHPuiauuuur1tAcVEREZAlSAFhER6ZkHnAJ8l/bRlH21B/g9wSjL2hTlkhSqr4h+yjn7d8N9pDfbG6wHu89I/tl2t/w9Hf2ac8XK8vIjnPkn+c7ONtxx9O5TAe8Dt+dZ5IaZ1dUbUhzpo8CTe+XYDEwFbga+vNd2rwBnErTBkRSLxWLjgGN83z+G4ObdB4GR+9ltB8Hrssw5t8w5V3XVVVetJBilPtS8Q3DdNhHcQMnZT1FkkyuuuGJSXl7eccCHWx+z6bmf/2rgCeCJlpaWJ6+55pqNGYgpIiIy6KgALSIi0r18go89fwco6ecxthP08fwxoD6TWWhFRcmHzEWu7WXhuQnsQWfurp35wx+ZW1XVkvaAOWZVRcW0JP6ZOM4Hd1QvdtkJ/Fd+wfBrZlRVbU1BhIMIbvIcRHtRKQ7sJmi90eYu4GtArvUGzlqxWGx6MplsG9l8HMFkcT3djGgGXjazqtZi87L6+vr6xTncribFHiP4hAHADIJCqKTYpZdeOmbkyJHH+r7fVpSex77ndHDAq2b2VzP7azwef0HXq4iISO+oAC0iItLRMOBs4Hv0c/I1YB1wK3AjwceoJcusqCgpi+Bd49z+26k47Flz3J507oFoPL4jE/kGg4ZodLbv8UWDC4Bx+9l8g3PuKt/ZrdF4fE8/T2nAA8BptP+O6xOMHM1v/b6ZoI3Oz/p5Dml15ZVXFpnZ8Wa2gKDgfMh+dtkEPA8s8TxvCbAsFotl3UScWeTnwDdavz6VYJJMSbPWCQ6Pc86d6Jw7EZjDvm+kbDazx3zf/2symfzrD3/4w/WZSyoiIpJbVIAWEREJjCH4eP5lwJR+HqMa+AnwO0CjY7PQqjlzJiYTLdeCO5+eP3a9B9x9jsiNJTU1VZnKNxjFo9HRnrkLPLNvuv3f1Fltzv6/otraP/XjVN8gKNrty7sELTf+0cM2sg/f/e53PxCJRE4AjgdOYP8F50ZgiXPuuUgk8nwsFqtjaLbS6K+vA79o/fo/CCatlQy7/PLLDxw2bNjxrQXphez73zBH0Kv8Ic/zHojFYq9mLqWIiEj2UwFaRESGuknAtwn+2D+gn8d4jKDNxmOpCiWpV19eei7ObsQ4qIfNNplzt7r85M3FrzS8m7FwQ4ADr6G89DSwbwEL9rPx/UnHxdF4fF0vDz8beJFgpHNPv99uA94g6Pu8pvXrtkcj+sTCP8VisYN93/+omS10zh0HlPaweQJYaWbP+b6/JBKJPB2Lxd7MUNTBagHwVOvXvwK+El4UaROLxY5IJpMLzWwhcBLBzevuvGFmjzrnHlq3bt0jt912m25Ki4jIkKYCtIiIDFUfAP4PcCEwoh/7+8BfgKuApSnMJSlWP6doKon8m8Gd3sNmqe5FLD2oKy+fb/jXEvRc3ZctDi4rronfbj2PnB1JMHHdDHoe1b4vLwO3EHxyYci2WLn88ssPzM/PX0Awwvl49l9wXmZmT5rZU8DzsVhM/bRTaxJBOycIRu3PDzGLdOPb3/72iDFjxiwws086504m+DeoO5toHRm9devWR3/605/uzmBMERGRrKACtIiIDDVFwBXA54G8fuzfAtwNXAvUpzCXpEFdeelXDfsx+x6l1oLj9hbs++W1terfmUEOrL6s7AwzdzVQ3MOmfzMv78tFy5e/s4/1txOMDu3N77U+QT/XZoL38S3AS32IPWjEYrE84EO+758EfAI4in33uvWB5Wb2lHPuyaampmevu+667ZnKOoS9BxxIMDJ/fMhZZD9isVhxMpk83cxOBz5I9++nHcCDnufdBzyiPugiIjJUqAAtIiJDRSnwHeAc+ld4bgZ+A1wN6KPlWe6t+fNH7Ni+7RYzzt/XNg7+5PtcGo3HGzOZTTp6asGCvKmbNl7gHD/cV3sUg/WGf1ZhTd2znVadCfyhF6dJEoyObgR+2frYNJDcuSgWi01OJpMfN7NTgYX0XNRcbWaPO+ceb2lpefKaa64Zcj+vLLAEOKb166nA2hCzSB9cfvnlE/Py8k4GFpnZxwgmOO5st3PuCWDx7t2777/++ut3ZjaliIhI5qgALSIig10ZcCnBiOf+fDx/O/Br4Efoj/+c0FhWdmjC3P0EI9C6MFiPcWlRdfzODEeTHlSXl48fhv8jBxftY5MEcGVxTfza1u8PBeLse3R722jnBPAn4DbgCYbQRHitLQI+QjDC+SSgpIfN17QWnJ/0PO+pWCzW2/7bkj6/pL3384nAkyFmkX6KxWIH+L5/MsENs1MI2gZ1ts3M/uycu8fzvMdisVgisylFRETSSwVoEREZrGYD3wU+S//+f7cR+AVwI5qYLGfUlZUtMHO/Bw7ufgtb7FoSXy+pr9dozizVUF7ySYd3C0GBuQvnuPfZXbu/9tXVq58juMHUWdto53cJ2nP8AtiQrrzZpg+TpO12zi1pG+X8gx/8oCqDMaV3/h24vvXri4GbQ8wiKdB6U2ghsAg4AxjdzWbvm9kfzOyuWCy2hCF000xERAYvFaBFRGSwmQNcTv8Lz2sIis63AZooKIfUl5Wdh7k76Gaku8H6pLMvlNbWPpb5ZNJXr1VWjm1paboJxxe6W/+Dd9e+c/em9w/ptLitSPMkcCvwR4Ji9KC2Vy/nU4HTCfrc70tbW42HWkdZqv9sdjuZYLJbCIrPF4eYRVKsl8XoN4B7fd//n6uuumpVRgOKiIikkArQIiIyWBxL0OP51H7uHweuA+4h+Mi+5JD6srKvYO42up/0qcpP+GeW1tW9kelcMjB15dGLDG4CCtqWvbRzJ19cvQa/46YbCEY6/xLY12SFg8YVV1wxKRKJnNLay/ljdF+4AtgMPA486nneo7FY7O2MhZRUmA683vr1kwRtOGQQuvTSS8cMGzbsDDM7DziBrjdSHfA88Oumpqb7NAmoiIjkGhWgRUQk150A/Cfw0X7u/yLwQ+DP6GOuOamuvPSrhv2CborPBrclfL4Rjcf3hBBNUqA+Gj0Wj8XAlK3JJKeubGRjIoEBxcOH726BbzY2Nf2awT3a2b73ve8d2VpwPhWYS/c3W3ygCnjE87xH4vH40sWLFw/mn8tg5wHbgFEEcxBMDTeOZMIVV1wxJS8v73PAecBR3Wyy08wW+77/qx/84AfPod9dREQkB6gALSIiueo44P8RFKD743mCiQUfQn+85az6irJLcO6ndP2dJoGzrxXX1v5PGLn2pz4aPd55zOrNth7+hqKaugf6cx4H1lBWdoEzl9e7PbwXSmpqavpzrnRaUVIyxYt4D1z85ptHv7xrN2eOH8dZE8ZzaEEBBnEvr+D4Wa+8sjHsnKkUi8VGAie0ttY4FejccqTNDuAx4GHP8x7W5IGDzssEraUAJhCMapchIhaLlTrnznXOnQt8oJtNVgG/TiaTd1599dWD/tMfIiKSu1SAFhGRXHMyEAOO7uf+zwDfJ/g4s+Sw+orop3D8ia4jQVuccU5Jdfz+MHL1Rn1F9M599Tfuxh7Xkpzan4kTG2ZHj3M+f+/t9gb/XlQTv6Gv58mEw0aMOPZL48fdumjC+Gi+dfoV1lgS2dV84qzGxuZw0qVGLBY7OJlMfgr4tJmdCIzYx6avm9nDzrmHNm/e/PRNN92U089benQ38PnWr48BXggxi4QkFot5vu8fb2Zfds79/+zdeXxcZfX48c9zJ0v3UkoXqkCBtpl0MqFYoGyyCCiroFIUEeSnsomAaGmhNuEhaS2bLLKoBRThi6IFFGTVskrZCzTJNJPShc2WrdC92eae3x930kym2SaZzGRmzvv16quZ5y5zAulk5txzz/k2MCBulwjwJPCH2trax/XOB6WUUv1NN6thlFJKqbQ7FJhLz1ttLAauBJ5OWkQqbeqCQb+Iew87Jp8bDe53/VU9qxjupwqcAt938XocJ0SEs5MfTnq8v23b4m9OnnxAXmPDwyBHt9koHOIOLPw98P/SE13PWWvHu657soic6LruEcaY9t6fu8BbwKMi8q/Kyso30Ts3ckVtzNfFaAI6J1lrXbz3L0/PmjVreEFBwXeNMWfhzb8Ar2f0CcAJxcXFa8rLy+91HOd2a+376YpZKaWUiqUV0Eoppfq7Q/Eqlo/s4fGLgDLglaRFpNKqKhgcUYj7msCEuE3bxOFbxUtDT6UlsAQkWAEN8Lq/OpRQ1f/S0tLBhRJZCwzt7jH9uQK6xQcHHTRwy6aN/8DwjfhtxnBhUVUo4UR9qllrv+K67inAKUCwg93WA0+JyKPNzc1PzJ8/P+EKeJUVvgM8EP36euCyNMai+hlrbanruj8CzgB2idvcBDziOM7vrbVPoxetlFJKpZEmoJVSSvVXh+D1eD6qh8cvAn6FN2RQZQkBpy4YeAL4evwmY/h+UVXo/nTElageJKARnNJE+jPXlQbOEuHPiTxHJiSgIZpcJ/IiwpS4TU3GyFFFVcu63XYkFaZPn+4rLi4+zBhzioicTPu9XAE+BB52HOefa9aseX7BggVNKQxT9U/FwLLo14/h9QNXqo2LLrqocKeddvqOMeZ84Kvt7LIcWNDY2PjHq6++WvuIK6WUSjlNQCullOpvDgJm07MP2YL3Af0q4I1kBqX6h+jQwZvi140x84uqamanI6aeaC8BXfa/NWyKeG07jxg6lFNG7NT2IOE6f01oZrefIxh4mrghnbd8/AkrG7x2wcUDBnDe6FFtjsmUBDTAsuLiPZw85zVgdNymd2lsLvXX1W1KR1wtLr300oHDhw//hoicLCInASM72HUZ8LCI/KOysvINtEpRtZUPbIn+vQrYO73hqP6urKws4DjO+SJyJjA8bvMWY8y9xphbrLXL2jteKaWU6guagFZKKdVfHIhXsdyTxLMLPI7X4/nNZAal+o9QIDDB57AUGBS7LvCYvzr0TeP9HGSE9hLQh9TWsa65mXxjOHzoEG7dY/f4w9auHTlq9yOfe64JsWaxAAAgAElEQVS5q/PXTvGPNxHfSmJ6ZNe7Lt9ZsYrVDQ24wFeHDuGO8W0LcTMpAQ2wPFh8mIuzCC85t53AbcXVoZ+lOp5LL7104NChQ48GpuO11+io/ckyYKHjOH/XJJDqhlrAj/caNxTYGrd9LPBRqoNS/Zu1doDruicBl9DaKzrWYuDm2trah3RooVJKqb6mCWillFLptj9exfJxPTg2AvwFmAfUJTMo1f+Eg4GHgW/GLa/OLxiw795LlmxIR0w91V4C+tH1G5g4oJC9CwvJM+2/RTO4JxRV1z7e1fnrSiZfKcbY9rbVu67U1TeYT5qbOGbYsLjzZ1YCGjqsio8Yl6lFodDSvn7+GTNmDB44cOAJxphT8YaADWpnt0bgWRH5ZyQSefjXv/712r6OS2WVh4BvRb/eF3g7ZttgvN7QF6Q6KJU5ysrK9jfGXAScBhTGbV4pIrc2Njb+6Zprrsmo36VKKaUyhyaglVJKpUsQbzjgqST++8gFHgTKgXCS41L9UF3p5KNEzKK4ZVfEHFVcU/NcOmLqjfYS0JsiEYb6fF0caRb6q2tO62wPAVMXDKwA9upsv8+bm9k5L6/t2TMwAR3tC/4McHibDYZn/VWhr7V/VO9Ya4e4rnsCXqXzcbSfdN6K1xLoHw0NDY9rYkd1w0BgWzvr8/BaUwF8H/hrzLY78e4A+GHfhqaywezZs8f4fL7zor2id43bvAn4o+M4N1lr3019dEoppbKZJqCVUkqlWilQgVfJ2pPE88Lo8Xrbeg4JBwOLgYPbLAp3+mtC56Qnot7peQKahojLuEAo9HlHO9SWlBxhjDzb1YmyJQENULuPf5JxfdVAQZsNLl/zh0Jd/rfojrj2Gt8ChrSz2zYReRpY6PP5HrLWbk7Gc6ucMQT4DXA5EDso7gfAvdGvK/EuvoJXzfo3YC7eBV2lusVaW+C67sm0357DFZHHfT7fPGvtK2kITymlVBbK63oXpZRSKinGA1cAPwa6zLLFaRkuWA68ldywVH8XDgSOJD75DBubMHPSEU+aFeb5+B5we0c7OEbOzrUpdsVLw8vrSktuEZFfxq6LY2YDPU5AW2uHRSKRE4HpxphjgQHt7LbZGPOoMWbhhg0bnrjxxhvbq2BVqjs2A2vxWmz8APhvdL02Zp/i6N974FU/A7yXkuhU1rDWNuJd0F9YVlY2zRhzCd4dafmAY4w50XXdE8vLy591HOd6a+0T6IBUpZRSvaAV0EoppfrabsAc4Ef07MLnIrxqsCXJDEpljnCw5B8gp8SuCVQUV4euTFdMvdWLCmgQXvPXhKa1tykUCAzxOayl/ercNrKpAhogFAjs7HN4l7jBfxFHSgJLl4W6e55Zs2YNLygoOCna0/kbtJ903mSM+ZeIPLBp06YnNemskmgXvITyALxq57l4lf2b8IaKhoApwAvAQdFjvgH8O+WRqqxird1dRC4RkXPYcYBqCPiN4zj3RZPXSimlVEI0Aa2UUqqv7ALMwLu9s70ETlcW4VVMv5HMoFRmqS4pGZNv5AO8qqwWm6UpMr44HF6Xrrh6q1cJaEBwSourq6t3OG9JyY8wcld3zpFtCWiAcEngWgyXxS3f5K8OXdrZcdbanVzXPQY4CfgOHfR0FpFngIXbtm178Prrr9+SnKiV2sHNwMXRr18GTgeex6t6bgSuxbuw28KPDuJVSTJz5syhAwYM+BHwS7wiglgfAzdu3br1Vn0NVEoplQhNQCullEq2ocBP8QYmDevB8YuAXwGvJTMolZnCJSUXY+Tm2DWBO4qrQ+emK6Zk6G0CGrjWXx2atcN5g4EXgK925wTZmICuneIfbyK+lXiVoi0+KaoO7Wq8HvLbWWt3ikQi38Rrr3EMUNjOKTXprNJhD2AF3l1DLrAFWIlX+Ux0reVnXPDueNia4hhVljv33HPzx44dewpwGbB/3ObPgNsaGxtvvvrqq7/Y8WillFKqLU1AK6WUSpbBwM+AWcCIHhy/GC/x/Hwyg1KZLRwMLAKOil1zjXvQ5KrajB6M1NsEtIGP14wc9eUjn3uuufWcpXsikZV08/1dNiagAeqCgX8LHBO7JmIOKa6peenyyy8fUVBQcIqInGqMOZr4oYWe9caYR4wxC9etW/efW265pSE1kSvVxp/x+kA7tE04E/d4Hd4dR0r1FVNeXv51EbncGHNE3LaNwG2O49xkrf0kDbEppZTKEJqAVkop1VsDgPPx2mWM7sHxz+ANF1yczKBU5ov2M/6ctu03PiyqDu1uMnwYUhIqoDG4JxRV1z6+/ZzBQCVtb8vvVLYmoGuDk88zmN/Hri0NBB+pmTixwBhzFG1/nlp8YYx5WEQecBznP9rjVPUDAaCarj+vvQlM7ftwlAJr7cGu684Gjqftz+Y24M7owML30xOdUkqp/qwnw6CUUkop8H6HfB+wwJ49OP5VvIrnp5MYk8oijjH7gcQlC81jmZ58ThbB/BB43Psapw7OSnNI/UKeyXssIpE2ax+NGnWoMWbnuF2/AB4FFjqO85QmnVU/E8L7+Twe6OjKVARYnbKIVM6z1r4EnFhWVhY0xlyG1588DxgIXOS67vnl5eX3u6571dy5c1emNVillFL9iiaglVJKJcoBvgdcBUzowfFLgTLgX8kMSmUfY+SAHdZEK+VbmVNq/f6RxeHwuneCxV8Ddk93RP3BxKqqD8PBwLvA+Ja1vd97d+C6ESNAk84qs8zDG4zZEQO8l6JYlNqusrKyGjjLWmtF5BIRORfvjrh84EzHcb5XXl5+v+M41lq7Kr3RKqWU6g80Aa2UUioRRwPXAvv24Nh3gfnAXXhVW0p1QYp2uPvcmNfTE0u/VOAU+L4L3O7iO1sLw2OZ10HGtzza/cMPBq7ec68z1o0Y8XdrbXMnByrVn7yK157qQNqvgnYAbXeg0iaaXL7kV7/61fV5eXkzReQcvIGu+cCZrut+t7y8/K5IJDJ/3rx5H6Q3WqWUUunkdL2LUkopxQF4vZr/Q+LJ5w+A84CJwAI0+ay6y5j41i6us22b3m4eQ1x++M6ECcNAvpXuWPoTY1gR+7igqYmvP7fofU0+qww0n45bcIAmoFU/MG/evA+uuuqqixzH2R24Bq8nNHiDXi/w+XwrysvL77HW7pW+KJVSSqWTJqCVUkp1pgj4O/AKcGSCx34GXA5Mwks8a+JHJUSEMXFLH09csaIhLcGkxya6umBjOCAysPAqYFAX51qfrKAygbg7JuVEGJuOWJTqpcfwWld19FqgLThUv2Gt/aSiouJyx3EmAbcBLb+zC/AqopddeeWVN8+ePTv+97tSSqkspwlopZRS7fkS8AegBpjODn0QOvU5Xn/ovfGqYOqTHp3KCQaGxC1tTEsg6bMOYVE39ruk613M33odTSYx7qb4JYHB6QhFqSS4jo6roDUBrfoda+2HFRUVP4tEIhONMb+l9b1goYhcnJeXt7K8vPzqyy+/fEQ641RKKZU6moBWSikVayhggXeAc0lsVsAWvITz3tFz5FqyUCVfQdzjXKp+BkAc/tyN3bq6QCQi3JOMeDKFiLMtfs24ZmA6YlEqCe7Hm6MQ3+i9HliX8miU6qZoa45LIpHIJOB3QMvg18HArIKCgpXl5eVXzJgxQy8QKqVUltMEtFJKKfASfecCK4ErgUQSNY14LTYm4LXcyKlb/VUfMjsknAekJY40GrBxyz/o/b+pF5q9f9s5wxh3h9cwcWSHpLRSGSIC/IYdLzZp/2eVEebNm/dBRUXFTx3HmWiMiZ0HMgL49aBBg94tLy+fZa3Nud/zSimVKzQBrZRSuc0H/AgvOfUHYFQCxzYDd+JVPJ8HfJT06FROE2FL3NKwtASSRnu++25979tnSHeqqLOKMWZ4O8ubUx6IUslzJ/Bp3NqqdASiVE9Za9+/6qqrzhORfYB/0FrVvwtwteu64fLy8h9aazVPoZRSWUZf2JVSKncdDbwJ3AV8OcFjFwFTgXOAD5Mcl1IAGFgbtzRm9fjxOVcd5ZrI3b04fAuNkQeSFUumEGGPHRZdE//zpFQmqQd+G7f2SToCUaq3KisrQxUVFd+OJqIXxmzaA7jbdd2a8vLy6WkKTymlVB9IpLenUkqp7HAoXq/mg3tw7PN4bTZeSWpESrVHWB13w7nZOnzQ3kAoPQGlx+Sq2lfCwUAY8Cd6rIEHiurqNlWXlAzqg9D6LWOYIHHdciUSWZ2eaJRKmtvw2mTlAYzOz+e/kycd6gpjHGGsC6MNZgzIWLw7mgoNDBAYiMGHbL+LZCfApXVWwwbvsdkK0oChXoRPjLDWIJ+I43xiJLJWXOdTnzEfbSgsfH+/JUuaUvy9qyxUWVlZDZxmrT3Ydd1fA4dHNxUDfy8vL3/acZyZ1to30xelUkqpZNAEtFJK5Y5JwFzgVLoeWhZvGd5gwYVd7KdU0ggSNnE/qo5r9ifHEtBRdwNXJ3qQK+bupEeSAUQ4IG5pfXFtrbYJUhknvG/ROCJ5flwzSYxM+uUHH/7v8fUb9gCYMXb0WeJylsHrY+C9Wra98iI7fLGdD6//Lq1/y/a/DN4JBQMiCA44EEEY0ljfFA4GVgNhjCw3mDoj7nKTN6B24ltvxbcJUapL1tqXgCPKy8tPBubjJaABjnJd9/Xy8vK/RiKRX82bN++99EWplFKqNzQBrZRS2W8kcBnwc6AwwWM/wEta30XrwBilUsJxeC2+itUYDsVLxuaWvOZ7ac6bh5c06q53/TU1L/RVSP1V7RT/eCI7tBV6zbSXglOqHwkXFQ2lMG9/wRxoRKYZmCbNjAHACAaYNXYM/96wkWYRxuUXpCvUfLyL2pMQg4CXoG5uJBwMfAi8ImJedRz31U35A5fst2TJ1nQFqjJLRUXFw9baf0UikR8YY64GdsVrG3qGz+ebfuWVV97d2Ng4Z/78+XqhQymlMowmoJVSKnsNAi4CrgDaG8jVmXXAdcDNeH0nlUq5TfkDlwxprG8g9sKJcIKAY7zbx3OG/626NeFg4D/Asd09xojcnWv/nQCciO+E+EyzEXkpLcEo1YmqYHBEAZGjBXOMwRwEMhnBMdFrJe1dMRmTn88pO+3EA198wbiC/NQG3D1fBk41Rk4VMQxprG8OlwZqRFhsDP+ORHgmEArpQFDVIWutC9wzY8aMBwcNGjQDmAEMAQpE5Nz8/Pzp5eXllY7j3GatbUxvtEoppbpLhxAqpVT2cYDpeG0Kriax5PMWvP7Qe0f/1uSzSpto1dyzcctj3wkWH5qOeNJO5E8J7e3k3dNnsfRj4rUZils0j6chFKXaEHBqg8Gp4dKSX4VLAy8W4H4K5u8GzgEpoZufzX48aiT5xjAmr9NaonrgC7w7mVYBtcAS4HXBLBLMIuCN6Fo4us//osc09PR7bEcewhQDFyI87HNYFw4Gng6XllxWGwwGk/g8Kstcf/31WyoqKq5yHGdPvPekLcnmEcANOqhQKaUyi1ZAK6VUdjkar3J5SoLHNQF/wuvzvDbJMSnVc8Y8gkibqt+IOD8Gcq61ROHmrY80DB38Ba19WzvzvL+qKueG7tXu45+Eu32IVYs1k0KhN9ISkMp5AmZ56eRDRZwz6pBvGdzRvW0Gs2dh4acn7DS8yWd4UuBD4FMMax3DxyK+T31bt66duGLFxi5P1ImqYHDEAJGxmMgoV8xYgTHGmNEGdnPBb7wWHN15LYpXAHwNka8Z5Npoy44HEHOfv6ZG/52qHVhrPwMunzNnzp+MMVcbY06JbppI66DCX1hrq9IYplJKqS4kOoRKKaVU/zQFuBY4JsHjBPg7MAdYkeyglOqtuqmTdpHG/A9p2798W5OYPYM1NR+nK67eCpcG7kE4M3ZtUyTCUF+bFs/v+qtDe7Y5riRwO4YLujq/GM4urgr9ueVxdUnJmHwjbYbwfd7czM5xFZQGfllUHbohgW+lX6kLBm4R+Fnc8rX+6tCstASkctY7gcBk18gZ4pgzEPbo0UmEz8TwqiOy1HXMch8Srhff8tLq6i+AnYD1SQ06Qe/su+8o120skogpwsgkA/sKHEDibb9ahIG/ODj3TaquXpXEUFUWsdYe4bruDcC+McsucJ/jODOttTpwViml+iFNQCulVGb7ElAO/JjEhpMBvILXV29xsoNSKpnCpYH7Eb4bt5zRScWeJqCXl06e5op5pYvTb4m4jI3ts5oLCejo97gaGBizLK7PLZr8du076YpL5Y6lpaWDC3HPEpFzTNvkWHc0CrzlYF4T3Fcjrnk1EApl3IVhAeedkpKiiJFpxsg0MAcilJDYnbeC4SUDC5oj3B8IhbTPr2rDWutEBxVeA4yN2bQFuN5xnKuttdpGTiml+hFNQCulVGYagpc8nknbZEt31AFlwMJkB6VUX1hWWnygI87LccvbIi7+QCj0flqC6qWeJqABwiWBRzCM6+jcxph/F1XVzI5dy4UE9B3jd/97ozB930EDt39fAo8VV4dOTHNoKsuFAoGxjs+cb0R+BoxM4NBVBhYJssi3rfGp3rbN6K+WlpYOHihNB7nGnATm5EQqwg18LHA3ec2/9b9Vt6Yv41SZZ+bMmUMHDBhwBXApMCBm0yrgFxUVFQ+nJzKllFLxNAGtlFKZJQ84D69X8y4JHvtx9Lg7geakRqVUH5kzZ85Ex3GOOP2fD51uRI5su9Xc56+u+UF6Iuud3iSgeyLbE9C1wWCw8n//e/svn3/uAOxWUMBXBg1ibVPTNa9t2XIvsAx623lXqbbqAoH9cbg0OvgyvxuHNAGLBB53XZ7MxArnZAjtMzngRJzjjCPHIxxO94YvbhP4vzyXmyaGQsv6OkaVWay1X3Zd99fAD2ib43jacZyLrbX6M6OUUmmmCWillMocRwM3AIlOjd8K3AL8GsjK6iqVPay1o13XPdwYc7SIfB0YD7BPbe3/C4SX/ZG2713EICcUVS97Ih2x9oYmoJNHpk/3hcPLFn/7nZXTaus7vOP6E+A14EW8tkOvAw0pClFlmeUlJcWu4SqQ6d08ZAnG3Nvkcn8m967vC+F9i8YRyZ8OMh3hkG4cImAe8AlXTKypWdnnAaqMYq091HXdG4H9YpYbgRsdx5lrrd3cwaFKKaX6mCaglVKq//MD1wGJ3kbuAg8ClwHvJTsopZJhxowZg4cMGXKQ67pH411k+QrtvD8RkcvOePih/drpBf2hNEWmFIfD61IRb7K0l4AWdvjG+zQB3c7zZWQCurZk8hXGmF//a/0GXt+yhTe3bGVlQ0NX5c5b8ZLQ/wVeiv7Z0OfBqoy2oqRkt2Yv8XwWXc9dWGngHkfMfZoo7Z5lpcUlRpwzDJyJN+OiM40Y+b3PVzh34ltvfZqK+FTGMGVlZWcaY64DRsesrxGRKyorK+9F74hRSqmU0wS0Ukr1XyPxBgz+lMSG9wAswusRvTTZQSnVG9baAuDAaML5KOAAOv75jgBLgKdFZOEP//GPz5qN1ADD4vZ7Zu3IUd848rnnMqa1THsJ6Hb0aQK6PZmWgA6XBo5FeJS4ZOC65uabD6mtexQ4FDgk+veAdk4RaxVedXRLlXQo+RGrTLRy6tThTU3b5iDmZ3T9c/QcyE1F1cv+ZbwLwSpBb0ydmj+4sX66gZ8D+3ex+yYD1w8aMuy63V5+eVsq4lOZwVq7k4hcJSLx76NfcBznImttVbpiU0qpXKQJaKWU6n8KgAuAq4DhCR4bAmYBjyU7KKV6ylq7VyQSOdoYczTwdTr/uV5ljFkkIoscx3naWvt57Ma60sBPRLgj/iADtxZVhy5Kcuh9RhPQvRcuLS1CIq8AO8Vtqhs8ZNi+ccmoPGAfWhPSR9J1H/21eBdAWhLSr+L18FU5ZHlJyQmukd8Bu3WyWyOYhx3j/mZS1bJXUxVbLqgNBqca416CcDqdX4xfZYycW1S17OlUxaYyQ1lZ2T7GmN8Ch8UsNwO/a2hoKLvmmmv07hellEoBTUArpVT/chJwE7BXgsd9CFQCd+FVjSqVNtGE86GO4xwiIicC4zrZ/WPgBRFZ5LruU/Pmzeu0XYyAqQuWPARySvw2YzinqCp0Zy/DTwlNQPfO6ilTdmqINL0CFMVtqjcuhxWFQq934zR70bZCupjO3xtvxrurpCUh/QLatiNref9m3N+COa2T3Zowche+SKX/rbo1KQsuB71TUrJ3xGt/cjodDy0UMAvyCwpn7b1kif7bVG2Ul5efBNwK7B6z/JGIzNK2HEop1fc0Aa2UUv3D/sCN0K0BPLE2AvOBmwG99VSlxRVXXDEqPz//iOjgwGOAzpKmm0XklZYq58rKyjdJ8ENfuKhoKAX5L4GUxG1qEsPpxVWhBxP9HlItHAxUAMd3to8xrC2qCp2UjOcLBQI7+xz+3fWecqO/etl9yXjOvvLOhAnDIoMKH29vYJkx/LCoKnRPD089Bq8lTEtCen+8O1I6EgHqaE1IPwe838PnVv1IXUnJmWLkRrxWWO0RkIWuT+ZMfrv2nVTGluuWBwJTXIf5wLGd7PY/EXN+cU3No6mKS2UGa+0QESkTkZ/T9vX9acdxfmqtXZ6u2JRSKttpAloppdLrS3h9nn9CxxU97XGB+4CZQJdVjUolk7V2EHBwzODAfen457cZr2p0keM4i9asWfP8ggULet3GoHaKf7xp9r2O2aGNQsQYftSLJKTqx1ZPmbJTY6T5CUEO3GGjkev9VcsuS+LTDcb72W5JSB8CjOjimLW0JqRfBN5C+wBnjNXjxw9oHDr4dwJnd7iT4VlcM9NfU/NG6iJT8cKBwJE4XEPHPaIFI79Zu/PoKzJpPoBKjTlz5kw0xtxqjPl6zHKTMeZ3W7ZsmX399ddvSVtwSimVpTQBrZRS6TEIuAiYAwxJ8NingV+iAwZVikyfPt3n9/unRCucjzbGfBUo7GB3Fy/ptghYXF9f/9y11167qS/iqiudfJSIeZId+4JGQM7xVy/7U188r0qPqmBwRIG4T2I4IH6bgf9M8k8+zixc2JctiHyAn9aE9KF0Xu0P3l0qr9GakH4RqO/DGFUPvVNa+uWIG3mwvZ+vqPUCs/zVoTuM3qrfLwiYcDBwjoHrgaHt72NeaBZOC9bUfJzi8FQGKC8vnw7cgncHTItV0SGFj6cpLKWUykqagFZKqdQywJl4bTM664vbnlpgBqBviFWfS+bgwL4UDk7+f2DuZMcKbNfAZZnQz1h1LVxauicSeRgItrP5jUJf/jF7vv32+lTHhfc6Hlsh3dndANB6R0BLQvpZ4LM+jlF1oTYYPNzB/Zu0TUJtZwyPYvLOL1q69H+pjk11rXaKf7yJ+P6A97uqPR+6xp0+uar2lVTGpTKDtXYnEblKRC7Eu9AIgIg86vP5LrTWamslpZRKAk1AK6VU6uyH16v54ASP+xy4Fq9HdGOyg1IKYPbs2bvm5eUdGq1yPgGvPUxHPgGe7+7gwL5WVxr4ngj3smMlNCLc3+j4frJPVZXeTpuhwqWBYxHuA3ZuZ/MbEZdvBEKhlF306MJQYBqtCelDgIFdHLOK1oT0YmAZWmGbMrXBwLkGbicm8RTjE4N7XlF17T9THZdKjDegdvLZYG4ChrWzS72IOaO4puahFIemMkRZWdm+xpjf4b2Gt9gKVNTW1l6/sG/vsFFKqaynCWillOp744ArSbzPcxPwJ+BXaIWcSjJr7S6u6x4ZTTgfCkzuZPctIvJybwYH9rVwcPJ0MPcB+e1tdsR8e1JNTW2q41I95yWUAjOBebSfHPwvjc0n+Ovq+qTFS5LkAfvQmpA+AhjVxTEfAW/QmpB+Db342CfCJSXnY+R22v1MZN50myPfnlxbm9YLbCoxy6YUT/RFnH8IBNrZHDFi/l9RTc29KQ9MZQRrrROJRH5gjLmRthc933Qc5xxr7Zvpik0ppTKdJqCVUqrvFAAXAJV00JuwE48ClwIrkh2Uyk29GRwIvGCt7fcJsHBw8slg/kb7/anXg5zrr162MNVxqcRVl5SMyUfuwHBSe9sFs2hLQeHJ+y1ZsjXVsSXBXrQmpA8Fiun8PfkW4G3aDjf8oo9jzHq1JSUzjJHr2t1ouHfw4GHn7fbyy9tSHJZKglAgMMTn8Cfg1HY2RwR+UlwdujvFYakMMnv27F3z8/NvEpHTYpabjDHXGWMqrbXay18ppRKkCWillOobJ+G12+hqQFW8N4FfAM8nPSKVUxIcHAht+zg/Za3dmKpYk6k2GJxqjPsgwh7tbTeGRx18F0ysqvow1bGp7gkHJ09HzO0Ydmlvu4EFzS4XBUKhfn9RpJtG493y3ZKQ3o/O/61GgDpaE9IvAO/2bYjZJRwMlAEV7WxqRMxP/TU1d6U6pu6oneIf74vkd+tOKl9Bwbq9lyzZ0JPnWT1+/ICmoUO7PadiY0HBB/stWdLUk+fqKwJmeTAwS7w7KHacEWA4r6gqdGc6YlOZw1p7vOu6t0Ob9xQrHcc531q7KF1xKaVUJtIEtFJKJdcU4Cbg8ASPWwNcBdyFl1xQKmFxgwOPAXbqZPftCeempqZn5s+fvy5FYfa5d/bdd1SkufF+4Gsd7LJBYKa/OnSH6WetRHLZsuLiXZ083+0gp3SwSz1iftZfk4NJlA+U0lolfRTt97+OtZa2FdJvAW4fxpixaoOBs43X3ipeA4bT/FWhR1IeVDeFg4HPgRHd2VeE+4trQqf36HlKAz9HuLHbBxif319VVdeT5+prtaWB041wDzvOCHAxfKs///9W/YO1dpDruuV4g8Bb2kEJ8H+O4/zCWqtt8pRSqhs0Aa2UUskxCpgL/Jj2e5V2ZBtwHd6QQR2SphISNzjweODLnez+KfCciCzy+Xz/tta+m5oo0+ONqVPzhzTUX4fhkk52e9q4/LIoFFqassDUDlaPHz+gfsigC40xV9Jxu6J3XeN+Z3JVba7234xv29FZz3aATcCrtB1umPPtJGpLSg42Rp5hxwrzra6YUybX1PwnHXF1VyIJaKC+EWdcaXV1wu1awh3pjloAACAASURBVMHA23i9y7unHyegAcIlk0/DmP9jxxkBm1zjHjy5qrYmHXGpzFJWVjbNGLMA7wJhi49F5OeVlZX3pysupZTKFJqAVkqp3skHfopXvTw8wWMfBS4GVic7KJWdZs6cOXTQoEHTYvo4T+1k934/ODAVoomHW+l48JsL5q8Yp8xfVaX/FlNIpk/31YVDZ4GxwO6d7Pl3aXJ/WhwOZ02VfhLsiteqoyUhfQDtD+Bs0dLXvSUh/RzeRamcEQoEdvc5vI7X8iTWFgzf9FeFnklHXIlIMAENYi7w19T8PpHneGefyftGXJPYhZ5+noAGqC0pOdEYWQgMiNu02hQ0HVC0ZLlWsaouWWvzXNe9EK+1y+CWdRF53OfznWet1fZeSinVAU1AK6VUzx2N126jvUnrnXkL+Dle306lOnTppZcOHD58+CHZPDgwFaqCwRGFuFcLnNvJbk0G/tTscmUgFPooZcHlqOXB4qNdnOvpvMryIwwX+atCD6Qqrgw2BK8FVEtC+qt0fVF0FW0rpJeRpRepPjjooIFbNm96EeQrcZuajZFji6qWPZ2WwBKUaAJa4NXi6tCBCT1H6eSbEXNxQoFlQAIatrfjuI+4z8CCWeT3Fx9rFi7UFmiqW6y1e7mu+3u8dmctNojIzMrKyjvI0tdSpZTqDU1AK6VU4ibhJZ6PS/C4j4E5wB/R3pyqHe0MDjyUHau1Ym3v49zQ0PDva665pkcDp3JFXbD4eMH5PbBbJ7vVY1joi3D1xFBoWapiywWhQKDA58jJgvml8YbudcIslKbmC7Tqucd8gJ/WhPThdFplDni/o16nNSH9OtDQhzGmTEdJVYGLiqtDt6Yjpp6IT0B/3tzMoo2btm8/athQRua1bXXsiJk8qaamtjvnDwUCBT7D/2IHgLrAA5+3dvH4yqBBTBgQ18EkQxLQAHWlJb8WkSva2XS5vzp0TcoDUpnMlJeX/wi4nrYzN55yHOdca+37aYpLKaX6JU1AK6VU9w0CZgKXs2P/yM40Ab8DygFNEKo24gYHHk3n1W2rgMUi8qLP53vUWrsmNVFmj5VTpw5vatpWgZjzgYJOdnWN4XGQmzKlOrK/ig6FPA+4EBjb+d6mxiFy6aTq2kWpiC3HjKM1IX0Ind9RAbAV746dloT0YuDzPo6xkCQnvWuDwcMN7jPEf6+Gu/xVoZ8k87n6WnwCunrbNqavWLV9+8VjRvPT0Tt0G7rGXx26vDvnry0p+bYx8mDs2uLNm/nx6ve2P54zbld+MDJuJmYGJaAFnOWlgYdFODFuU4PP5St64VElylo71nXd24FvxSxvBSocx7nOWqtFJ0ophSaglVKqu04CbqXrCrJ4i4BL8G5tVqrlg8pXo1XOx9F5Ne72wYEi8p+5c+dqj+IkWR4M7uUiFSCn03kSDqBK4I68vIK/TXzrrZzqm9tTAk44GPyqQX4Y/W/cWSU/GN4zUD6pKvR/Ru8QSZVheL2jY4cbdvb/KQLU0ZqQfhHvolgyfQ34JnAlSbhg++wRR+SNXffpUhM3tFHg1bxtDYdPXLEioyq84xPQKxsaePDz9RQNLMQ/YIDsVVho8s0OH+/WFPkn796d9hLhksAjGE6KXXOBDxsapba+3oTr6ykdOJAjh8XNCs2gBDTAOxMmDIsMLHwNKGqzwfCsvyr0tfREpTJdeXn5dOB2aL2DAFjsuu6P586dmzH/PpRSqq9oAloppTq3D3ALXj/NRNQBvwQeS3pEKqNYa4cAB8b0cf4KHf/+3SoiL7W01fD5fG9p5UzfCu0zOeBznStBpndj94hgnnWM3Nsc4aFAKLQ5ZpvBu0PiT8AnfRJsBngnEJjc7DOnGZEzgb26ccg64LrCTVtu3vPdd+v7ODzVuTy833ktCekjaZtIac9aWhPSS4BX8e766Y0H8H7nWuAOvP72PVJbMvkiY8xv45a3ihPZt3hpeHnPQ0yP+AT0xkiEYT5fl8cZ5Pii6mVPdLZPKBAY63P4AO/noEPrmpt3aPORaQlogOWlk6e5YhbjtavZTsR8p7im5qE0haUy3OzZs8fk5eXdAsS+p9gqIr/y+Xy/1fd0SqlcpglopZRq3wi8D78XEvfhpAtfANcANwI6AC4HxQwOPFREDjHGHA7kd7B7BHib6ODAdevW/feWW27JqIq8bLE8WHyYK84MDCfQdUU0wCYM/wQeeW1L/QtnrVx5A3AGXhLuCGBzZwdnCwGzYp/JU5rFOc6IfBco7eahHxq4zdnWcPvEFSs29mWMqlf2om2FdDGdf37YjDcMtSUp/QKJVzJ/CQjjDVZcDswA/pXgOXhj6tRBQxrrVwOjY9dFzGXFNTXXJ3q+/qCnCWgMf/NXhb7X2S61JSUzjJHrujpVtiSgAWqDgRsMXBq/XFQdKtE7MVRvWGtPjA4p/FLM8suO45xtrc24i19KKZUMmoBWSqm2HOAHwHXEfWjtggvch/dBOWerH3ORDg7MLu+UlOwdMXIOcB5thwq1a7PrctF778vLm7e0vKcSH3wvAn/v00DTKBQIDHF8HOkIJwocD3y5+0ebN42RmzflD/jrfkuW9LZSVqXeWGB/WhPS+9N5L/VmvCRyS0L6OaA7g7kuAmIrl58HfgG82d1Aw6WBnyPcGLe80retIZBprTda9DgBDfWNOONKq6u/6GiHcDBQBQS7OlE2JaDDRUVDKchbTlxvejGcWlwVerCDw5TqFmvtTiJyjYicG7O8DbhKe0MrpXKRJqCVUqrVwXjtNr6S4HHP4vV5rk56RKpfSnBw4FrgRRFZ5LruY/PmzftfaqJUvbF6ypSdGtzmcxC5ANizvX0+a27m3HffY9k2r3OEzxjsuF2ZvvOIj/D+n7/iOLy2KX/gkv2WLNmawvCTankwuFfEuNOMyzRjzDRB9iexO0MagUdwuckfCi3uozBVegzGG2YYO9yws9dDaNu240W8pLLE7eMALwP7Rb+ORP9+AJgFdNoPX8CpKw2sQtijzQbDdH9V6IEuv6t+qhcJaBBzgb+m5vftbaoLBPYXh9e6c5psSkADhEtKLsbIzbFrBvNKUXXNQemKSWWXaDX0AmDXlrVom7UfW2u7c0FOKaWygiaglVLKq3y5CvgJ3bv1vsX/gNnAvez44VllkWhPv8OiVc7H0vkwys+AZ0VkEbC4srIylJooVV8QMMv3CRwiImcg5jRgZ4D3Gxv5yer3eL/R67QzyHG4affdOGzokPZO04yhBuQVg1kCUuf4CsP9baBhKBAoKIAJrjFFYqREYJrxhtSN6sHpBMNLYO6Txua/F4fD65Idr+qXfICf1oT0V4HxXRyzEXiN1oT0i0A9XjuXN2l7scPFS0b/Diing/YetfsEvmFcnoxfzvTWCr1JQHeWVA2XBG7D8NPunCfbEtCrx48f0DB08CpikoMAxmVKUSi0NE1hqSzTQTX0RhG5rLKy8g70c4RSKgdoAloplcvygZ8CFcCwBI5rwvvwOwfY1AdxqTTTwYGqPaFAoMDnyHEvbNpy8awPPzzyi+aIAdglL48/jN+dwMCBiZ7yC4HlDoRdkTpjeB9j1kaMfFwoeZ/uXVX1qUnih9IPDjpo4Kb160eTz64OzmhcM06MTASKHCgSL1G4wwCyLa7LYKfb1+bCwH0Y333+qqpOq1RVzhhH2wrpfen8Ym8TUIWXkN4br81L/Ouv4CWfr6admQvhYOA+4PttD5Dzi6uX/aHH30U/0KsKaMARM3lSTU1t7Jr3usYaYGR3zpFtCWiA2mDgKuNd0NhO4Ibi6tAv0xWTyk7W2lNd172dthd2n3Ic5yfW2g/TFZdSSqWCJqCVUrnqaLx2G/4Ej/sX3sCalUmPSKWNtTYvEonsE9PHWQcHqo6cDPwFGASwc55vwz177rlhwoDCzqrie6oZ+BT4RDCfAhhkPRgBNhukSYRtGOpNa1JqqGDyBBkEptAYGYgwCi8J2G55dmceW7+BijVruXPPPQi2n2DfhvACjnlCTPMTxUvDOlxJdWUoMI3WhPQhQMJXb/CS0AZYgXc30kKAN6ZOzR/SWP8pMDxm362+bQ27ZvrAy94moAWuLq4OXRG7VldS8l0xcn93z5GNCehlxcV7OHnOKtpeGFnprw5NSFdMKntZa0dHIpHfGWO+HbO8XkRmVVZWLkhbYEop1cc0Aa2UyjXj8CqmzkzwuBXAz4HHkh6RSjlrrROJRPbtyeBAx3H+Y61dn6pYVb/yI+APtFYJvwacCHxau49/konkHYuRY/H6yQ/v4BwZ45H165n5gdeyfNf8fB6YsBcj8/JcgbCDeRrcJwYNGf7cbi+/vC3NoarMlgfsQ2tC+ggSa/vi4iUOnwd+GS4NDEd4us0ewoP+mtCpSYk2jXqbgAbWFPkn724WLozEnPMJ4NjuniAbE9AA4dLAiwiHxK61VzGuVLKUl5efiTdsNXbg8ULHcc631n6eprCUUqrPaAJaKZUrWtptVOJVX3XXVuA6YD6gVa4ZLG5w4FFEe/l2YPvgQJ/P97jeFpnzDHBl9E+LfwHfw3uNaEPAWREI+JsdDjBGpomYaQaCtNPeoj/b6rqctmJV84qGhjyAnfLylt0wfvwRP1qxol/1rlZZx+DdnXQ2cB7dv5gjAHsVFqy6a8/xe++a33oTizH8sKgqdE+S40y5JCSgwXCcvyr0JEB436JxNOe9TwJDRbM2AR0MzMIrUNjOGM4pqgrdmaaQVA6w1o6NRCJ/MMZ8M2b5Y+DHFRUVWvSilMoqmoBWSuWCw4HbgECCxz0KXAjohOoMFDc48BvAHp3svklEXm2pcq6srFySqjhVv5cH3A6cE7P2J+BcvBYZ3fLG1KmDBjc0TDFQDFKEoQgvybYnHbd7SR3hMxzqEKkDU2dwl7s+ebv47TDA68Au0T1/B90bVqZUD40ALHARXnVzghlWyDOG03femYvHjGKoz0eemN0n1NR8kOQ4Uy4ZCWgR7i+uCZ0OUBcMXC7eBfZuy9YE9LLS4gMdcV5us2i4y18V+kmaQlK5w5SVlZ1jjPkNra2yBLh106ZNs2688Ua9y0gplRU0Aa2Uyma7AtcAPyCx17swcDHwn74ISvWNGTNmDB4yZMhB3RwcuE1EFuvgQNWFwcDf8YagtbgGuDxZT/DG1Kn5g5u37In4ihwYI8KuBjNKcMcIzliDjAJG41Xs9+R921bgc+AjgY8d+BRj1gryiQifIOZdmpvrisPhdZ2c4yjgSVoruM/Ha0WiVDIZvPZYN+IlWXv9OWVEno/zR43aNH/tRzuTwAWj/iopFdBQ34gzrrS6+otwMFBLgrMwsjUB/c6ECYWRgYUbgYKWNYFXi6tDB6YxLJVDrLXjXde9B/hqzHJYRH6ghRFKqWygCWilVDbKw6tcrgCGJXDceryqq9vIgg+q2a6dwYGHEfPBMU6bwYHAi9ba+pQFqzLRSLw2GwdFH0fwXlfSmngNBQJDIo6TX5DXONBpLhjgGpPvRCIDcZyNAPXwBcC6kSM3Hfncc8l8HbsUuCH6dRNwDF7PXaWSoQj4PV7/50QJsAloBNzdCwpGf7kgn0GOw0DHoQlZ++T6jbOBe/H+HWesJCWgQcwFYsxSg/tSoodmawIaIBwM1AGTYpY+8VeHxqQrHpV7pk+f7isuLp6B9xmm5T1tMzCvtra2cmFM/3allMo0moBWSmWbI4FbgckJHOMCdwGzgc/6IijVe9HBgcXAIdE+zsfSeT/v7YMDGxsbF1199dVfpCZSlQX2wqv4nRh9vAWv3/OjaYuof7gT+HH064+B/YGMb2ug0u4E4AK8OQtbon824CWVN8c93hJdi328vQ97KBDY3efwXuzJRbinuCb0w77/NvpeNxPQTXTR1kfgVQeWitdKqDM7nCvLE9BPAV+PXYu4FAZCocY0haRylLV2P9d176XtHQqvuq575ty5c99JV1xKKdUbGTUMRymlOtHTdhtv4VU1vtzVjir1YgcHuq77NWPMyE52/wj4r4gscl33iXnz5mliTPXE/niJ5tHRx58DJwEJVwpmoZ8BJcA0YAzwCHAI7QxiVCoBj0X/9FoBDIkvDzTRuwJyhvAkhhMAp6NdDEwTbzBqF8xjIKckMbr+zfCFN8rSs7apiUJf/hC83wNKpYy19g1r7VQRuU5ELsD7bDPNcZwlZWVlP6+srPxjumNUSqlEaQJaKZXpetpu4wvgKrxqab2drZ+w1o52XffwaFuNr7uuO96YDq8nbBaRV2IGB74JsR8dlUrYMcCDtFbWrwaOAzK+si9J6oFv4Q0l/BIwBbgDOCOdQSnVotlxBhratvMXQ04N8BLD+w48Ld7rWWcGdbbRwMcu7pMGkzsJaJEtLTUMzSIct3wFDa77HrAUeBFYjFewoHfLqT5nrd0KXFheXv5PvOHHXwKGGmPuKisrO8Hn851jrdWLI0qpjKEJaKVUJjscr19zIIFjXOCPwBXoB4i0ix8c6LruVwAj0m4euWVw4GLHcV5cs2bN8wsWLGhKbcQqi50NLKD1dvM3gBPxWk2oVmuBU4HngELg+3j91a9LY0xKAWBEmuLvgTJGcu7zjiB/BtNVArrzcxi517hOMyanrutunyOxbFs99a4LMATvTo9DYvZbhZeMXhL98xpeD3Klkq6iouI/1topruv+HvgOgDHm267r7metPdNa+0KaQ1RKqW7JuTdkSqmssDMwHzgHbbeRUdobHOi6rg4OVOk2C+81peX1ZBHeh7yNaYuof3sFr3fsn6OPrwZCwONpi0gpwAdb2rmlaUjqI0mvwUOGP7Rl88b1wE49PYeI7x6DHJDEsDKAM6TlRqql2zrtLLRX9M+Z0cdb8N6vtCSkn4e2vciV6g1r7WfAqWVlZWcZY27De13b3XXdZ6+88spb165dO0OLMpRS/Z0moJVSmcTgvdn/DbBLAsetByzabiMt4vo4f8MYMwygg9YaOjhQpZIP73Xh/Ji1e4Cf4A3fUh27B9gPuAiv1+xfgQOB2nQGpXKbU1j4WaSx7XVKg/lymsJJm91efnlbbTCw0HgX6nvi9eLq6upwSUmOJaBl+8/K0q0JdW4ZzI5V0mvxktEtrTvewGtjpFSPVVZW3jNnzpz/Oo5zH3AQ4IjIxWPHjj3IWvt9a+2KdMeolFId0QS0UipTTAFux3uz1V0C/B8wA/ikL4JSO4pJOB8KHOW67rhO+jh/DLwgIot8Pt+T1tr3UxepynEDgHvx2km0+C3wc7SXeHf9Am8o4ZF4Pfj/gTegcEM6g1K5a+8lSzaEg4HP8e6UAsAV9kpjSOkj5m6M9CgBbVrvbsg1239WEkxAt2dXvDZOJ0YfNwPLaU1IL8G7c0SphMydO3e1tfYw13XnAGV4F4H3d133zfLy8gsrKiruTXOISinVrkRuXVdKqXQYjPfmagZetWJ3vY3XbuOlvghKtbriiitG5efnHxFtq3EMsGcnu+vgQNUf7Aw8Qmu1WgS4GO8il0rMznj9T/eOPn4SL+Gid5uotAiXBF7FEFu5G4m47BQIhTanLagkiSbXR7Q83hiJMMzX9q2RwG3F1aGfRfevAyYl+DQNEZdxgVDo83BJyY8xcmfsxnXNzYzMi6thMj6/v6oqo4e1Lg8G93JxV4L3PR5Sm5Jv5yO8yuiW1h3/xbtrT6lusdZ+zXXde/AGFLZY6DjOudZa/VlSSvUrWgGtlOrPTsIbMrhbAsdou40+Fj84EOhscGAz3vT4RY7jLNLBgaofGI+XJC2KPm7Aa+2zMF0BZbjPgW/jXewbDBwLXAmUpzMolbvE8LqhTQLa5/NxAPBMumJKFzHmz0ZkXmJHmUcCoZrP+yai/sslclBLbdZHTU3smp/P2qY+f7sylh2rpKvxZpW8Gv2zHL1QrzpgrX3GWjslEoncZYz5ZnR5uuu6U621p1trX0trgEopFUMT0Eqp/mgiXuI5kQnuAtyNN0zs0z6IKWclODjQxRv2uMhxnEUbNmxYfOONN/b6PlalkiQIPEFrpdAXwMl4VWeq56rwkvgP4mVw5uDdWv63dAalcpMj5lUxcmHsmsEcTw4moB3j+7NIcwUJ3EFmiNzddxH1Z85xLXnewMCBPOufxIubN532k9Xv1wNTo38OpReDHbshD9g3+uen0bWNeEnpltYdLwHr+jAGlWGiAwpPjg4ovB3vYvBerusuLi8vn+c4ToW11k1zmEoppS04lFL9SgFwKV4F84AEjqvDe6Oecx8u+0rs4EDg68DwTnbfPjjQcZynrbU5VzmlMsJRwEN4vYoB3gWOA8LpCigLzQV+Ff16G16y5s30haNyUXVJyZh8I2vw+qK2WFlUHZpoMrySNNEWHNFjnsL7Pd4da9eOHLX7kc891wyQKy043pkwoTAysHANMb3Dgc2Fm7aM2vPdd2MHB/oAP14y+hC81zg/bX/WUmEVrX2kl+C1QWpMcQyqHyorKwsYY/6Kd8EdABF5vLm5+ez58+drgY5SKq00Aa2U6i+OxOu/6k/gmK3AdcCv0TfevWKtHee67iHRKucTaNtLLt72wYGu6z41b96891IUplI9dSZwF5AffVyDl3z+MG0RZScH+Cde+ySA94D90btSVIrVBUteFuTA2DUH9/BJ1bUvpCumZOhJArq2NHC6Ef7SrScQrvPXhGZuf74cSUC399/IGB4qqgp9pxuHDwNKaU1IHwSMTH6UndqCN/tkCV6l9At479VUDrLWDhCRa0Tk4pjljx3HOcta+++0BaaUynnagkMplW67AtfgJYgS8SjwM7wEh0qQtXYX13WPbGmr4bruXgAd9HHeIiIv6+BAlaEuAW6gtULtGbyexRvSFlH2coEz8PqXBoA98NpyHI1eJFQpJIb7EdokoCPiXICXmMspQwYP++eWzRvX043WET7h7r6PqP8xwgXxa+JKd1sIbcRL+r6I934WYC+8ZHRL64798e7y6yuD8RLgh+AN1AVYS2tCejHesMP6do9WWcVaWw9cUl5e/hzexfcRwBjXdZ8sLy+/tra29lcLFy7UOTlKqZTTCmilVLr48NpmzKX1lvjuWA1cBDzWF0FlK2vtIODgmMGB+9LxLaNtBgcCL1hrNXmkMo0PuBmI7QX7AN7FLv0Q3rcm4Q3Pakl4/RbvQoBSKVHr9480+b4PadvOKyJOZHLx0vDydMXVWz2pgPaOK/k9yHmdnVvg1eLqUJukfS5UQNcGgwcZ3JfaLAqf+eobvjxxxYqGJD3NYLz3XS2tOw4DxiTp3N3VhNevP7Z1RyjFMagUs9bu7rruX4GDY5afj0QiZ8ybN+9/6YpLKZWbtAJaKZUOpcACYFoCxzQBv8PrL7q5L4LKJtOnT/f5/f4pMRXOXwUKO9jdBcLGmBejfZyfstZuTGG4SiVbIXAPcFrM2m/xeszrIJ6+txzvv/0TeBcCLsYbonVnZwcplSzF4fC6umDgbwI/jFn2GTevHPhBuuJKF9dE7nbE6TQBbYz5c6ri6U+McefF39NlkAVJTD6D1yKjpUr65ujaONr2kp5KYvNPEpVPa0V2i5Yq6ZZK6Zfw2tupLGGtfd9ae7jrunOAMrzik8N9Pt/b5eXlZ1dUVGhBj1IqZbQCWimVSgOBWcAVJHYr4gt41dJaqdGJuMGBx9D57bbbBwc2NTU9M3/+fJ2orrLFCLw+xIdFHwve6851aYsod82k9Zb0JrxBkP9NXzgql9QFg37BDdH2bh8xDocVLQ29mK64eqOnFdAAVcHgiPi1WOtGjtzUMnxw+/NleQV0bUnJt42RB+OWt+YZ354Tqqo+SXE4+XgFGrGtOyanOIYI3mDv2NYdy9C2a1nBWnu867p3A6OiS2KM+c3atWtnL1iwoCmNoSmlcoQmoJVSqXI8cBswPoFjPsZLYNyLvvndwezZs3fNy8s7NFrlfDzw5U52/wR4XkQW+Xy+f1tr301NlEql1Di8qtvS6OMG4Gzg/nQFpLgP+H7064/weqHq8EeVEnXBwF8ETm+zaHh7c/6AA/ZbsiTjEi69SUD36PmyOAH9zoQJwyIDC6uB3WPXjTG/KaqqmZGmsOLtCuxHa6X0wcCgFMewEXiN1tYdLwFatJChZs+ePcbn891jjPl6zPLrjuOcpp8NlFJ9TRPQSqm+Nha4lsSGDLp4SYtL0Te520UHBx6E9yHkaNreRhlPBweqXFMCPA7sFn28HjgFeD5tESnwbil/AS/xDPAmXoXftrRFpHLGsuLiPZw8pxbvDqztBCqKq0NXpimsHotPQDeLkGfafpzr6wR0owgFcc+ZiQno2mBggYFz4pY/LfTlT9rz7bfXpyWorvkAP21bdxST+s/0q2jbS/pVvLtcVAaYPn26r7i4uAyYg/czBfCZ4zhnWmufTGNoSqkspwlopVRfMXhJ5xuAkQkc9zZwAfBKXwSVSXRwoFLddiTwD2B49PEavLsulqYtIhVrN+ANYHT08b3AWekLR+WS2tISa0Tik80Rg5xUVL3sibQE1UPxCej29HUCul0ZloCuLQ380Ah3x68bOLeoOnRHGkLqjWHA/2fvvMOkKq8//nlndgGpglIVFaQvNWABLNh7iYpGjSYmURM1Vko0ohOI2LDFEvvPlqixxWhiwy4iKhaWZQERsNEVpLO7M+/vjzPXnblzZ3Z62T2f59kH7p17557ZvXPve7/vOd+zJ/XWHSNJbdydDTYi91vHuuNtpPJOKWICgcCYUCj0TyTTHsSS4/bly5ePU0sORVFygQrQiqLkgoHAPUR3XG6IDchM/J2IB12Tw9040BiTqHEgRPg4a+NApQlzIvAY9c2b5gFHAF8XLCLFi1HAm9T7/19MfTMuRckZb44ZU9b1+9UfEFs1tCHosyMrPp9XMv0lVIDOnOpBg0YaQm8SO756o29l1cGmcVSL9STaS3pPxGM6nzgNDh0v6Y+BrXmOQWmAQCDQKRgM/iPcP8bhbZ/Pd1ogEFhWsMAURWmUqACtKEo2SbfJ4IvAL9O77gAAIABJREFU+TRBwSidxoGhUGiG3++frgNDReEipMrCqQyYCRyDWvcUK2cBD4b/HwSOBrTcV8k5CysqhoZ8zKR+osphQXN/+d5FbLkQhQrQmbFgyJCdbKjuI+ozPh3WWX9wWP/P5i8tQFj5oDUwlHrrjv2pr0jJF7XAHKKtO0pm8qcxE2HJMYn68dRq4PTJkye/VrjIFEVpbKgArShKttgfuBvxpkuW5YiA9FROIipCAoFAl1AotG84y/kI6v1qvVgNvGWtnW6tfe2vf/3rkjyFqSjFjgGuRSa8HJ4DTke9hYudu4Fzw///AdgLWFS4cJSmwoKBA8+wxj7i8dIbG5u1OGbE7Nmb8x5UiqgAnT7V/frtQLn/NSN2ZpEErY+j+n9e9UpBAisc3Yj2kh5O7ARNrnGypJ1M6feBov8eNlauuuqqg5AePJ3Dq4LAX30+3+RAIBAqXGSKojQWVIBWFCVTOiBC0Nkkf00JAfcD45Hu2o2WCRMmtGnZsuVeET7O2jhQUTKjGfAQcGrEur8hTUv1Aan4KQdeQyYtAaqBvWnk9wKlOFgweOA0a+1lHi+9S03dUf0WLNiQ96BSYMnQodtvCAYTjrVqmjXbli0x/YtevZpv2W67lg1tN6hfv/XmqaeK1j7ti2HDOgbral4DhrhfM3BZ38qqmwsQVrFRDgwm2rpjQJ5jqAMWEm3dMQ8dC+eNQCCwcygUepIIG0Vr7RvBYPC0qVOnrixgaIqiNAJUgFYUJV2cJoM3ATumsN/nSPbbrFwEVWguueSS7dq1azc6LDjvg2T3lcXZXBsHKkpqtAGeBg4NL1tgMhAoVEBKWuwAfAT0CC//GzgBFRmUHGPBN39QxX8MHOXx8sfBEIdVVFX9kPfAlJxROXBg53JjXwMGxbxoeLTfnCptiBqfrsAI6jOlRwENTkhkmfXAh9Rbd8xAqmeUHHHOOeeUd+3a9QZr7UXU60XfWGtPnDJlykeFjE1RlNJGBWhFUdKhF/B3JKM3WTYDNwLXID5wjQKPxoH7kLiE8afGgdu2bXv1+uuv/zFfsSpKidMV+B/iYwlQg3gK/7NgESmZMBQREhwx42pkMkFRcsqSoUO33xasfQcvQRI+KjP+o3vNmbMq33Ep2Wf+4ME9sMGXgT7u1yzmnRYbNh7WY+lSbYyXPGVAX6KtO/qTf01hMfWC9HvAp2gFVNYJBALHhkKhh6i3/dkGTJw8ebI2EFYUJS1UgFYUJRXKgUuBvxDbPTwR/0OaDC7NQUx5x9U48GAS+zEuB96z1k4PhUL/veaaa77LT5SK0qgYALwE7BJe3giMRRvYlTonAf9CxqMWOBnJcFeUnDJn0KD2zWzoZQx7erz8ncU3tn9l5cy8B6ZkjfkVFQfg4wm8m+29HQxxdEVV1cZ8x9UIaQvsSb11xyjEni+fbEQqCh1B+m1AJ5GywJVXXrm7z+d7hgj7GmPM/T/88MMFt99++7YChqYoSgmiArSiKMmyL3APkumQLCuQJmFeTX9KBlfjwMOpF8G8WAO8aa2d7vf7pwcCgcV5ClNRGit7Ay9Qb/WzAjgSyXhSSp/rgQnh/28ERgJzCxeO0lRYMnTo9jXBupcsdm+Pl7dhzMR+c+Zqpl+JYcEsGFQxAam483ts8nKr1m1P6D5zpjaszR09qRekRyONH315jmE59T7SsxHbJxVM0yAQCLSw1t5prf1NxOpPfT7fCYFAYGmh4lIUpfRQAVpRlIZoD1xHak0GLfAYcDEl6NMWCARaA3tHNA78GfE/+2Zr7fuOrYbf7/9UO0UrStY4HrHY2C68vAg4Ivyv0jjwAf+h3pN3CZJNt6ZgESlNhi969WobbNH8vxj28d7CPrix2XZ/zFZTPyW3zBk0qH0zQv8HHBdnk//4t2w7ufeiRSpE5pfWiO2SI0jvj3dmei6pBeYQbd2hSSIpMGnSpHOMMbcjzaABvgdOnTx58msFDEtRlBJCBWhFURIxFriD1AaJlUiTwZIpXQ03DhweCoVGh32c90fsRrwIAp8Rbhz4/fffv6slaIqSEy4AbqM+a2oWcAywumARKbmiLXLPGBBeno5MNNQVLCKlyfD54MGtmtu6B8GcHGeTxcbYc/rOmfd6XgNTUqJ64MATjLF3Al08N7D8fWPzFheNmD270fQhKXG6Ue8jPRxpdpiKvV82WI6I0Y4g/T7Ss0aRfjYx/uiTJk0abYx5CunLAfJc9NfJkyf/BW0krChKA6gArSiKFz2RJoOHprDPFuAGYCrSHKxoyaRxoM/ney0QCKzLV6yK0gQxSEO6qyPWPQ+chj4YNmb6IpMM7cLL04DxhQtHaWpUD6o4x8ike5wJaPOUra37Q//587/Pa2BKQioHDuxc7rM3YjkjziZbseaCfnPnPpDXwJRUKQcGE23d0TPPMdQBC6kXpGcA82iawuqpiO/zVGB95AuXX355x/Ly8ieBA5x11tr/1NTUnKnN1RVFSYQK0IqiRFIOnIf45rVKYb+3gN8DC3IQU1ZwNQ48iMQNUn5qHOj3+/8XCAS+zU+UitLkKQPuBn4bse5BpKpCs2EbP4cDL1Lv2/pb5O+vKHkh3LjuSaBjnE2Wgb24b+W8p03TFKWKBgu+BQMH/hpjbyTemM7wVYjQCQPmVH+S3+iULNENEaMdQXo09ZZc+eJHxD/ase6YQQnaC6aBAV5DROhpwK1EeGgHAoGyUCj0V6TXj8NCa+0JU6ZMqcprpIqilAyG3M4sLgXUC1VRSoNRwL1ARQr7/ABcDtxHkT2IXXHFFZ3Lysr2C2c5HwbsmmDzNdbaD4wx71lrp0+ZMmV2vuJUFOUnWgP/QqwXQK4pk4FAoQJSCsKVwJTw/7ciXqEfFi4cpamxaODA7nXYf8b3hQYLs7DmT/3nzn0rj6EpYaoHVxxjrJkKdmC8bYzhxVBN8Neasd6oKEOqZSKtO/qT/6S6xUR7SX9K49Q8egOfI6L/V0g29P1EfNZJkyadZoy5D2gZXrXB5/P9JhAIPJ3vYBVFKX4MuRWN2gIbcvj+iqJkTlvgeuAcku9QbZGstAkUSRaANg5UlJKmC/Bf5HsLku18HjK5pTQtDPA4cEp4eTniDbqsYBEpTQ4LZv6girONZP61ib+dmW5NcKJm2OaHeYP77+2zvuuQial4rLXwp/6VVffmKy6loLQD9qBekB5F4irHXLAREWodQfotGk+/ij8B10Ysfxxe95Mn/qRJk4YYY56lPrHRGmNuN8ZcFggEtHpNUZSfUAFaUZo2RyLl7t1T2OcL4A9EDDwKQSAQKAsGg0MifJy1caCilCa7Ay8DvcLLm4CTgf8VLCKl0GwHvIuICSANCg8govxXUfLBvP79d/X5ffdgOCzBZiEwz4RM8OYBc6o/yFtwTYiFg/ofHMJ3KfUVMt4YnizDf2GvOXNW5ScypUjpSbSX9DCST7LJFsup95Gejdh4lOI9rAyJfahr/auIEP0pQCAQaBsMBh82xhzvbGCtfcvv958SCAT0+6goCqACtKI0VTohWT3xGrZ4UQvcjDQGy/sAKhAI+ILB4LBww8DRiFdo3KwktHGgopQCeyKev47f6krgKORhTWna7Io89DrnxsPArwsWjdKkqR5c8StjuQEZP8XFYD6wNnTL8h07PXvAW29p5l8GfNGrV/NgixanY+zFwKAGNv8a7IX9Kuc9n4/YlJKjNSKgOoL0GOL7vOeKWmAO9YL0u8CSPMeQLkOR+3GZa70FngauABYB5qqrrpqAWHU4gv83Pp/vpEAgoFZaiqKoAK0oTZCxwF3Ajins8w7SZLA6JxHFIcXGgSuAd62100Oh0EvXXHPNN/mJUlGUNDkWsVpwfAMXIxNLXxQsIqXY2AeptmkWXj4fuX8pSt6Z37dvG9u8/DJj7aUkngAH+Bq4M1QXenRAdfXyPITXaJjXv/+upsz3GyPjzoSCP/C9ganNNmy6q8fSpVvzEZ/SaOhGtJf0CKB5nmNYjojRjnXHDGBLnmNIlmnAZXFeqwHuQfo3rL7qqquOAh4F2odf32atvWDKlCn35z7MnNMauS45n20tkjyxqWARKUoJ4SVAryN7jQnXeby/oiiFYTfEbiNRGambtUh5VV6aDAYCgU6hUGj/JBsHbrDWznKynLVxoKKUFGchTU+dbJqPgKMBLdNU3FwI3Bb+fy1yD3uzcOEoTZ3qfv12MOX+8cBFQIsGNg9hmGktj5iausf7LVigiTkeLBk6dPuaUO2xIWvOMNiDaLip3Gbg9ub+8ut6fPaZVrgp2aAcGEy9IL0P0CPPMdQBC4m27phHcegpLZEM7t0TbLMJuAOYGn6me5aI6gVjzL3GmD8GAoGa3IaaVTojyVsHI1V7XeNstwyYBbwGPAWsyUt0ilJieAnQa8m/cb+iKLnDAGfTQCMdD55Css1y1kRj3LhxrVq3bj0yycaBW6y1M4wx030+3wzgA21soSglyUTguojl14AT0YopJT73IvcxgO+Rh8DFhQtHUWDhoEE9QwT/DOZ0ksuc3GzgeWt4YmN5i+kjZs/enOsYi5kvhw9vV7tty2EYcxri7dysoX2AjRj7YKjWXqeZ5Uoe6IaI0Y51x2ikR0E++RGZpHcE6fcQvaYQHIBUJTU0QbQGmDZmzJh79ttvv1uBX0W8NjsYDJ54zTXXfJWrILNEf+BKpCeJ23qkIWqBfwJ/RaxJShW3d/pK4NsCxaI0ElSAVpTGTQVwP7B3Cvt8CZxLDpoMejQO3I/4DxxRjQOB9wKBgJZXKkrp4kfsE86JWPdQeLm2EAEpJUM5ck/aN7z8OSIEaMmrUnAWDR7cqc4GzwL+COyU5G51GGZhecHim96vsvITUxxZjjll4aBBPUPGHmMtRxtsojGgm+XWmHtDQfu3iqqqH3IZo6IkoAzoS7R1R38aFmSzSRBYQLR1x6dAKE/Hf4hoQTkRi4Arr7zyyp19Pt911Au5K621p0yZMuXtXASYIWXAVUgFcLzm9smyFZgMXE/+/j7ZZBP1NnkgvaDi2bAoSlKoAK0ojZNy4FLgLyTvZxZCxOrLgI3ZCsTl43wY4g0fj58aB9bU1Ey/7rrrCjXDryhKdmkFPIk0GHS4HricJiC6KFmhC5IFtnN4+VngJPT8UYqEiKZ5FyGl/KmwFMwr1tiZPuub1aeyckGpC9J27Fj//PnzBxhr9zTG7mNlDBivfN0TbeqolADtgD2oF6RHU+8PnC82IPYYjnXHB+SugrUDYgvSOYV95o4ZM+ax/fbb72LkXg5QZ629bMqUKX/LeoTp0wF4BmlSmYj1wHeIltaNxM+2AP8FTqX0Kv1UgFayjgrQ8dkZOATJIN0hvO4bpFvti+Tmot4NOBTxvd0RuaH9ED7WPCT7Z30Wj9cFKWMdjJjpt0EyETYiM3ZLwz+fhv9tTGwH7IeUluyG3DjWI3/ja0lultIgXYFHI7+/TuH3XY00xJuJDACC2Q29QYYBD4T/TZZK4HdAxh2KXYLzgdR/f7z4qXGg3+9/ORAIfJ3p8RVFKTp2AP4DjAovB4ELEE96RUmFnyEP2E4J9OVE27koSlGwcPCAvaz1nW6xvwA6pvEW65Ax2SxrzYfl8HmvuXOLtrmyBd+CwYN3NaHQUGvsXsBeSFO31mm83bdYHgfzWL+5c+dkN1JFyTl+oB/R1h1uK4N8sJxoL+mPgG1Zeu9TEYuJlGjfvv275557brtmzZr9NEFnjLl3+fLlF9x7772FroRrj2gtXs/PIeAFxJ7yRcQWJZLtgWMQu46j8M6Ifw+xGspaklceUAFayTqFEqBvBMa51r2BiK+pinV/Rvx1IvkYudh7GdzvSqyYeg7SZA1gCCJAHk78cpo64GXkC7gwxXjdGOQiPg4RMxOV8NQijXcCiLiZLkcClyDiYLI3w1XA/4DHSGzNUEZsKfWfgakpxgjSTTeyVPtboHsS+20juqTvRmBC+P/tET+ns4nvh9wc73PHoTNwBXKT6ZJgO5AJhIeBaxDfylzSEikZGocMfpKhFrmZXEXizxyXyy+/vGN5efmYsK3GoYigH4+N1toPIhoHfkKJZ/goipKQHsj9sk94eTPwC2QgryjpcAbwSPj/IeA45IFQUYqON8eMKev2/cpDa/H9skzO1ZYN7hSfTWAWWGsX4jMLwC6whBbYoG/ZgKqqVSbHJd5vjhlT1nnlyo7+Zr7uBE1fi+2LoS9yfe9Dww0ZE/Ej2GcImcf6VlW9nevPoih5pg2iMTjWHXuR3sRUJmxGksoc6453yCzB7Hng2FR38vv99uSTT/6qd+/eu0Wsfre2tvbEa6+9Nmd9hxqgDHgV8bh2U4lYUyarveyDaBgDPF77D/BzSuf6pgK0knUKJUCXIxc9ty9tALEMSJZ9EeE60hj+RyRDJl5zmkQC9PnALSTv97MFaaZ0e5Lbu+kLPIqU7aSCRUqZzyG1Uo6WwP8hwmm6fILM5sajmAXofYCnabhkqAXeM8Q+RHieSOoZHeuAPwBPpLhfsuyPnMO9U9jnfSTruTqVA6XaOBA5Z97z+XzTly1b9nYRzHAripIfRiDCoHPN/QF5WJlRsIiUxsItwMXh/28ARgJVhQtHURJyNnDuCR06HD915657A4djOZzk/aKTIYQki6w2sMJiV1rMKiMCwiZrTRBj1wMYa9dh5PnPWJ8vZGw7ACNZfAZLGwytwHQE29VAZytiWUey63W7BMvL1sdLLdZveq3H0qXa50NpSnQj2kt6BMnbJmaL5UR7Sc9Ant2SoRtSod0unQMPHz48eOSRRxpjjJMM96W19rgpU6YU4l5+LeL57OZFpEl2qkla24X3PdDjtQmINlEKqACtZJ1CWnDsiszCRXokBRFB660k9t8xvP/OrvUnI+URiY671LXuHER0vjOJ43pxGfKFTIURSEZxJrOfHyNlHquS2LYcmdkbk8HxoHQF6H8D00muc3FLYm++LZCMq7FJ7B8Pi2Qnp3quJGJ7xEf1bJJ/KNiMNESYRhIVB9o4UFGUNDgY8eh1Kk2WIKWHCwoWkdKY8CNZ9EeElxciGWXrChaRonhzETJhYhCB51DCY8wMGvKVKlst5j2DnW7xTe9fWTm70AEpShHREknqcaw79iVxVWkuqEPup5HWHfOIX616Aekn4rH77rtz4okn2hYtWjjPsBuAMyZPnvx8uu+ZBkMRe5Iy1/pXkaSJdG1LWgGvIJMMkWxB7E8Xpfm++UQFaCXrFNoD+jjgOaKFs2WI904iUdUgs0pHutb/HTivgWN6CdB/RwQ858LzI2I18SLwNZJVsHP4eGcS62lrkZLifzVwbIceiEjnZVj/Zvh9qsJxdEIypM9Auuy6+QzJJG/o4jgBESrdcb+EfM6vgJXI378cmc3sDQxCZu9GIL/3UhSgH0CE+ki7jGpkAuArxPu5K/L7/Xl4u80R2/qR39MhHseqRkSWD5Df31bkbzYS+CWS5e6moUmSZDkGOXdTyaB5Cfg9cl7HxeXjfCiJZ7d/ahzo8/leDwQC2p1cURovzZFr5WcJtjkTaWjqVBNVIkLhd7kNTWlidABmAb3Cy68g9/p8911QlHhMJNqj/DHgLETkieLzwYNbtTDBYdaa4WCHW8tw413CXUosN4bZ1vKe8TGjZcu2s7vPnJlsdqWiKJJlHOklPZrkkqmyyY+IQOsI0u8hegFIdfDbSBZ3WnTo0IFf/OIX7Ljjjs4qC0yePHlyIN33TJGXEOvVSL5HxrqZWoLsgmg67srpfwKnZ/je+aBQAnQ58jurI/eNG9uFj7Mpx8eJh/NZof57lQ4tkGvDZrLn9Z4MrZHrQNJ96gotQAPcimQHRPIKIvbG88cZD9zgWvc5IsQ2lG3pJUCHqPdCfhN5eP42zv6dEBsLt/i9ChkoNuTz60cu1O7ZsFXAb5AuqfH2G4d4Cbv9fachv5NEx1wWjt1hJXA8IpwmQ9dwfCOBoxNsV4wCdC31Qshi4I+I+OxFa+SLG3nuBYCrXdv9gFhqPE3887QMObevJ/pvtg6Zbf0qwWdIRFdktvnEFPZZi5QW3ev1Ylhw3sfn84221h6NDHjisRJ4x1o7PRQKvXLNNdek+zkURSk9JiMPIa/EeT0y2w+kZ8AJZLeBrqI49EfGMc6E/rWIVZaiFBq3+Hw3YvWXtPfngiFDdiIY3NMa+zMwfcE6fsuZeC3ngp+8qX3Y+SF8s8t9vg97zZmTTIWmoijJU4YkN0Vad/Qnu9Y4DRFEqtkc645vgH+QwXWpRYsWnHTSSfTs2TNy9ZMbNmw465ZbbsnlpNUwJLnOza+RHk7Z4GJkXBxJCEn0i2cZC6JJnO9a9zKieaXKCUTbdK4EHvLYblckqdJhCtHWtO8SX6tyWIVoZcnSBkl2OxjJ+u9OdJJmLZItXom4JLxC4t9bIrogWe1HAHsijgqOZrQZsaR5G5mU+B/RCYnJ0hv5fUfySPi9HYYgNqgHhbd3kmC3It+pJ5EEykTH9yHJiKciTd4jNbI1iE3xPxDf8WzRBUn0OBz5/XWkfkJsG7ACsVl+CfGI94zfS4B2PJQzZUW8g7pohjzMjnCtvwJ5kHCzN/LBIr8MG8P7J1Pa6yVAO8xETv6G4m6GnJQHudbfjYiSiTgHEVYj+RGxxkiUUebwW8TrN/JGE0JO5Llx9hmJ+P1GcgKSfZ4qXudMJMUoQDvMQ/5mK1KIYQ/kvIgUkL9DLpBLknyP05Gsl0juI/qzJYNBMuFvIbVJoqeQG9hPs7iuxoGHIFn58dDGgYqigEyyforcr9zNWPzIxFjkPfBR5J6lvu9KLjkeeAYZjFvgNHLXb0FRGsIg1m+RWWLTkErEjMdOFnzzh/bbBevva4KmL9iextDJQlcLnYylE4YdG36nVA7JajCrwK4Es8JiV/kMi8Au8Id8C3vNnftNFo+nKEpqtEOeVx1BejTRFqf5oIYMLYSMMRx44IGMHl2fo7d58+YFzZo1O2Dq1KnLE+yaCm2JToi4k9jq+S8RUTBbz7rliIbRybX+GuDKBPt1RRIII3H6lqXKc8hYyeFTvPW+A5HEkUz4DBH2G6I1khQ4jtR0DYtogZchYm0ytEES8S4m+UbA3wGTkImIVJpGHodYv0ayF/Ah8r38G6INNTRp9FV4O6++OUMRkX9oEvG8iiTXrkxi23jsgGiz55O8T/13wFXIREfU768hMTETjiH5ruQ9kdmnyDL/OqQT6XsR69ojX5hdXfufQazAF494AvRW5OE6WVGxCyJ4R87QbEIyRxNlelUCA13rfkV9R/dkeCi8TyT3Ih1avTgNmQFxqEG+fLkoUy1WAXobcqGdl2IMzxA9ixVEJkE+TvF9HkAyyB22IudistkhuyO/D/ekRyKWIReKfwcCgZbAqIjGgcOoz/p3U4fMrk7XxoGKooTxIffjkUAF0dfS5ojYHOmR/zfgEkqn07dS2vwFGeiC+CvuR+r3aUXJFINUdl4Yse56vJtb5YyPhw8vbx3a2NFXU9bJQrnxhdoBhIxve6w1WF8ba6yTcVVjsJus9QX9JrgeIGT964CaUDC4uv/AgavMU0+prY2ilA5+oB/R1h2JnvuKimHDhnHkkUfi90vu1+bNm7dVVlae/corrzyawdu2QqrFq5HsUpDr9TKiLTpBROFrMjiWF9OIta5YgPyd4tGYBeghSGbuLhkc50KS8x/vjWRNJ0q2S8TLwEkkb9ERT4D+DngNb0vdeGxFXBfejFj3c+BxUmtYWon8bdeksI/DGEQPS9ch41+IAP6TLYjbbL1QLEbS0CN9ccsQf5xhiK2FAR4kVnz+P5IXnxNxJ8mLzyBZtNOQcmSHVshMxd/j7LMvseLzh8iDeyr8CfERjvSAOh25sHqJ39u7lrfQ9DwS7yF18Xl35CISyUOk91A7BfnyOd+5FoiFRrxzxaEMEZGvQc6vZLA+n+/+s88++7HOnTuPtNa+FgqF9iX+hSoEfGqMmWGtfW/r1q0v33DDDbn2W1IUpbQ4FxGfQap2HDogZVaO/18QseFIt6mvoqRDABnUj0XGRs8gGWFqAaDkCz+SDBKZbHAVMv7LKyNmz65FhAu3eJE61dUZv4WiKHkliPgOV1Gf4NYGEf0c6469IauVElnj008/Zc2aNZx88sm0atWKli1bNh8+fPgj69evv2DmzJmnk3rzvmOAOxD9I1KHGUqs+AypJQUmyyPECtB9kSaTS3NwvGLmKKRKze2L7bAWsXVZgyQUdkCsp9LRLQcjoq87+xwkCfdLxBrDjwjUXT22OxwR5Q8j+vknFVoi1iVu8Xl1+KcNksTqttltgeijjh/5QYigG/m7qEOym39EMpU7exx/EHAXoh+mwqmI1uqlITlWPN8jWlJn5O/knug6ORzXEYSTVItFgAbx0r2L6DKI7ojgdywyy3G8a595SPfVbPBQGvs8jDz0RP6iDye+qOjVxO5+Us9CX4E88Ed69LRCbioveWzv/rK0Q2a9vDyPGisPprHPCcReCNz2KcmyFJlsGBWxbh8SC9CDkcxptz2NJ+3bt6dPnz6rRo4cWd22bduTkcaaGONZ4aGNAxVFSZZuRDexdSY6d0PuOU4GxzZkoi3ZhryKki0sIvz1Rx4wd0EaBB+IVH0pSi7xIw9pZ4SXLXApkg2tKIpSaDYgVWzvUT+e60a0l/QeZGijkS2++eYb7rvvPk455RS6du1KWVkZhxxyyJ6tWrWa/8Ybb9wdCoUm0/AE8xAkQ3bf8PJJRDeA3cvr0OGfbDMXGTu3da3fi+ISoN8hOtP1O6ITHu8isW0IJE5yHIhkoLuT6uoQreZBpOGlu3pyO0QPOR4RNHduIAbCx3iKWPG5DrEzvYvY3/2eiNWEOwFxLySx5pdJHNeLqcj5CHIe3Ii4E0Qmv+6AjGOvJvr3swOiN05BhHtHv60G/orYAq+L2L53+DP82hXDWGRM/EaSMY9CJk7cevFn4fj/S6zG2BlJKp5A9Ll+EJK0ezkeb1jkIO9hAAAgAElEQVRoLkMyrCLT9o9GMgrOdG27GTiF9MzB3XxJfP/kRHyNiLiRAuGeCbYf6Vq2yANSOjxNtADtvL+XAO1lVv8IIrAuTPP4pcRq0jPs39e1vJzMSnpnEC1Au88HhxZIlvvlJBgItG7dml122YWePXvSu3dv2rRpA3KR9ZrlWwW8ba2d7vf7Xw0EAkvT+QCKojRJbqd+MBRC+i4MQgY9ziBwLTJgezfv0SmKsBHJdPoIyewajXRsz1aigqJ40Qwph3Xs2izia6lVIIqiFDPLEIHOqUBvhWgwjnXHvkiiQUFYv349Dz/8MMcffzz9+kmew6hRo/wdO3Y8/9lnn/31tm3bbgRuQu79kXRAxLpzqU8k+4RYzWUIseTKuisUjmGMRwxPxmxdOOqQ8byDO0lyq+v1VGiN6Fdu8fkrZOxWmWDfLcjzxbtIg9/TkojjRiQjN5L1SCbzB3H2+RARuS8EbnO9djrwAun9vRzNZ174+N96bPM9EvMHwHSiNaBfI4m5TtXCA8DviZ5QcfgCOAvRKa9yvXYuyQnQHZBxTaRWbBEh/WriTzKsRKr2n0E0yd0iXpuAWJPM8hKg15K+x0embEVE5dlIKrrD7zy2vZD0RGMvkmn+F49PiRaguyAP5F4nljuT9SvkZEsHL9P1eJmyVcgsSWTav+Ph+TQyA/MmsRfwxkK6md6jXctVZOaZ7j4ndvLYZggyEOjtfqFly5bstttu7LLLLnTv3p2uXb2qRH5ik7V2pjYOVBQlQ44k2gd/I9Kf4Vnq+zYsQ0qr5uQ3NEWJYSlSMvgS9RZWc5BEBkXJNs2Rio9jw8tBpPHqwwWLSFEUJT02UZ8l7dCNeh/pfZAK6u1id80NNTU1/Otf/2L06NEcdJC0QerduzdnnXVWqyeeeCKwbt268xCx+V5EjPsdIoC57UUmEfsc7GVVkE7CWrJ8TqwA7baWbcych9iORPI1cm59l8L71NGwTUp3wpXgEYQQ/+R44nMkfwM6EpvtPRm556ejqaxEMpAbagb4LjKBfUnEupaISA/i5312EjFMQbKeI/W/Y5EJgIb8rK8g1p97CiI+J8N8JIn4Q+qbPvoQEfrEYsuABlHtz0X8n+PxT0T5zxbzs7xvF2LFxnJivZhT9SSO5CtECIj0z/HKfAU5QScith2Rfgx+RPA/BfFkeR8pvXgfufk0FkE6nc65bYidiKlAfITSpZtruVn4OJF+y8sInyfl5eV0796dHj160LNnT7p06RLPTgNcjQOBdwKBgJYdK4qSCW2QZidB6rNI6pCyqxbh5bmISJ2LkkVFSYfpSBXRtPDyHchY7Z2CRaQ0Rloh2TwHh5drkAyppwsWkaIoSnZxvORfCC+XISKiI0jvSayomHVmzJjB2rVrOe644ygvL6dTp06cffbZPPXUU52WLl16O/BnpCq+p9fuSMWem/Ye63JpSemVdOjWBhorzYGLXetCiMNBKuJzspxLrNPD/SRvPwH1Am7k+d0HOBRpapgqf6Rh8dnhHqIFaIcNSCPKZARwx9bkxoh1LRCL15kJ9mtLrHj/IWIDkgpVwA2u/Y4Ddi1GARok5fsAYj88iED9+ywfb13Dm6S0r9cFzS0+Q/olDCAn3lqiBWiv4zq8gPjRTSPW1xhEIN8//ANy0s5CsnEfp7Qb+Xg1ZmwIryqArnib02fC9kQL0KuBy0488cRH+vXr91MHYDfWWlauXBlavnz5N6tWrXp3/vz5z//4449vkn5GvaIoiptrkGte5MxX5LXxTSSbIN2mHIqSK24CBiB+euVIxsoe6ESJkh3aIYKGY6u2DbHF+3fBIlIURck9ddQ3OFyL2AnkhXnz5vHDDz9wyimn0K5dO7bbbjt+/vOf8+ijj7JmzRqvZoIOf46z3utZP5fjWa/3buOxrjFyHLEayr+Bt3N0vDNcy0EkezkVapDnIHe29a9IXYBeSmq2uwuQZFa31/XDSHPGZJnusW4oiQXoM4n1Kr+a9LK+70SyyB3N2Q8cUqwCNMSfgXqcaMEuG2SS6esVS7sk12WaYew+ttcxIrkVse64CXkQS0QZMrs5GrgOKW35C7mdGcwV29LYJ182NDEKcyAQ+MfSpUvv9vv9LSPXr127liVLlrB48WKWLFnCli1bfEjpzq7Um+IvRmZ6Z4d/ZhHuOKooipICeyHeufHKLpYgFlQXIJN8zs+P4Z/VSGmdohSK85GGN3sipbbPIxlb2egdojRd2iMWL04Dq82IZ2QmFXKKoiilQjeksujn+T7wihUreOCBBxg7dizLli3jjTfeoLY24WPuq8QXOb2SA9NJWksWLwG6hce6xsgYj3V35ehY3Ym1j3iT9DKtn0XijEz4HBVn20S8SOLmjF5UEytAv+C1YQIWIJnmvoh1DSVTHuxaXoO3kJ0MaxA9KrLh5+hiFaAPA8bHeW0i8B+8PZDTpXmW993isW5rlo/rtb/XMdy8izyMjURKBQ8h1qDdTQvEc/s4xDumKfh8el0kviS6W2k2iDlXNmzY8M8uXbq0BOkC/Pnnn7N48WLWrUsqUb9n+MeZ+duIWHPMRmxV3iH58g9FUZom5UipWgjvipkQMri7mOhBDcg17QGiS74UpRBsRYTBj5CeC8OQskZ3ZoyiJEsnRGgeHF7ehIyLUynrVRRFKUUMUp0+jQJm7m7cuJGHHnoIa5NKyPQDvYBFHq956TW5FIS93turiVxjZH/X8mZyZ4u2t8e6dO/RmxDP6EhRdldkEmZZCu/zYRrH9sp0TvV9tiCfIfL7mihh1RDbA+19MjtPK4kWoIcUowDdDUl1dz/UOjQHnkAM8bM1S5XJRdSdog7ethxe67z2TQX3CZSKpcdM6tPvd0I63Y5BvmC7x9lnVyTr42fkR8T09p/ID15WFi/g7ceTVay1T9XV1Z1SV1dHbW0tn3ySbg9FQGbsnEz2C8PrllMvSM9AOv4mM3mhKErTYBySORqPyPuzRQYsm5FSq5uBFbkLTVFSYjni4fcmMn78JdJ4+qZCBqWUJF2QLKCK8PI6pPlqMg2NFEVRSpkKpCfIyEIHAiQrPgMchPTcuhexYIi0FPXSTRqqJs8Er4zrplCRVUZ0IzwQHSJXFdpenuSfZfB+nxGbFdyP1ATodJ6L3I0Ca0jPNtgtQLeOtyGwG7ENPDcgumu6uPXmHYtNgPYD/yC2mZ7bA6UXksVyapaO2z3L+3rZVGxEvmjlEevc5QGp4NUkL117jO8QUf+J8HIPpKnUucAg17bdkE6yF8R5L687QrzJhIbwulDnC69Zpx75OPDtt9++6Mwzz6R9+/Z06dKFnXfemW+/dfe0zIiuSGfSo8PLTgPDSOuOqmweUFGUkqEX9V5fcbueEi0834FkPKsHvVKMzETGMw+Fl29AHkhfKlRASsmxKyI+9wov/wAcjmTXK4qiNFbKkR5SfyHzyu1CUY5Ycv0aGa9ei9hheOkmuRSgvd57dQ6PVyy0J/Z5YmmOj+cmk/4fXnaCifqueZGOfXDItZxu4q37fRI927nFZxDHhNPTPLYXHdIVBnPFJGI9YmYgpW4LXet/gXeTwnQYksV9tyJWDV5Uu5YHEjsrkMpx3X8/9/unyxIkk20o0knezanEz04OEjubl2imJRHZbviXCtuI/TvuReIvbVaora39ZuXKlY86y3vssUc6pu+pUIbMbF2ImNvPRWb1nkMsb/ZHur0ritL4uRsZrMe71jkDmXXIA8nOyH1CxWelmHkYGdeAjJ3+Qb2YqCiJ6IFk0Dvny0rkWUXFZ0VRGjOjkOzP6yhd8TmSVshz7UIkkc4rm7RnDo/vVWGe1QyzIsWrr1Y6mbzJkm1vby/v7lQFaLcInA7ZeI+GyEcPtFbFlAF9ANIlMZIfELFzLSI4v0+0f85tSGbL3AyPPQSZlUq182lzYktRPkNS5L2YSb1vHMhnGUV6HjjuUgDn/bNJCLgeESCPiFjfAegNzI+z349AZAO9RJ1p49EcsfooJG8QfbPoggi1H+f4uD+MGDHi9xs3bjwC2HH33Xdnjz32+OVHH320Y/j4+5D7bOyuiHfm8eHlIGJkH2ndUU1+LoaKomTAx8OHlzerqWntD4VMuc8XNTDbZu0ma0yNNaZmyJw5Y5FyRS+cJhY/IP5/t9M0SgeVxsPFwABkvNkesdXam9x2vVdKm37A60j1H0gm1MHAFwWLSFEUJbe0AaYC55F+FXMx0wmx4fKyEx2Rw+N6vXe2kgeLGS/v63haWTbwmizZlsH7ee3bGCZkvMiLt3uxCNCdkGyUyKxaC5xFfcr8p4gv5R0R22wHPIk01XP7pKRCC6ST60Mp7ncUsbMsMxJs/z5SBhrJ6aQuQBvEx9BNomNnwotEC9AAHYkvQC8nOnt5cJztEnEkhe8MO53YLPuLyEMDo0AgsHncuHF/R6oCzJgxY/b76KOPIs+dbogYPZx6j+ftchiSH3lwH0D959+ANKR0BOmZeFuXKIqSRT4ePry8dW1tT0uwjwmZThjbFWM7gulsrelqsB2xdMTQBmhOTdji3Qch15xRuQGwrKsL0tLnY3Mo+nUD1oJp4/dt3L9Nm9cmdun6zI7l5QtNTY2/34IFefm8ipIl6hA/6A+RLKd+SGb0CehkqhLLUOBVZLwLUjJ8ELC4UAEpiqLkmCOBv5OZTWihmQs8gyQwun+2INXqyxEd5wXXvoMQcTETwdKLHfBOHpud5eMUI15e25n2QUuEV1JBG9LvX+YVa2NNXPD6XH9HqsCyRjEI0D6k6aDbbuE24D+udXcCByIPCw4DkEys32QYx3jgMZLv8ujD257ioQT7PIfEGnkin4mUtixJ8rjOPu4yjvfw7vCaDbwuwokE/0+Izl4egnhlJ+u/40NKZArN80jMkT7fpyLnZq6zoEEmW8YDLay1vxo/fvzVN954o2Nivyz849w4yxDT/dFIhvRwxPA/l5YhbagXvx2WUy9Iz0Ye9HM5y6kojZb5ffu28TXzDQvi6+czpg/YvjZEX2q29gDKDIAJO/RY+aobx4Y/xW/+xG++ixGfAbo1KzfnduzIie23b+s35kTgRAhBszLmD6pYhmEB1iw02IVgq21Z8PN+ny5IpTGHouST75Ex5AykHPc44CogUMCYlOJjOPAKIhqAVIAdTNMol1YUpenRCXm+/UWejhdEbBicn7VIItUQ0rfuBBGXjyQ5zeEdYvtzNUOE6WcziMGL44gdma8nP3oCFDapz8trO5dWD172HqlaZkTiFauXqN4Y8LJTXAE8lc2DFIMAPQE4zLXuY+ILkL9FxM3dItadhdglPJZBHAOAyxDLiWQ4F9jDte59EtuBbECybf4Ysa4FMrNwNMmJ3zvHifEOj3UOnYju+poqo13LFvgqwfazgN9FLPuQ0tfLkjze5YjfcqHZhkwO3Bmxzo/Mqu5Feh1NHdpT35jSk2nTpq0aP378Y9ba3wHNrbW/J/5Dch3SOLAK6fQLYiuzB/WC9Chy7+3TFckwGxte3oTY0jjNDd8ht40HFKUksWPH+qsXzO3vC/n3Ars3hr2AASHwGyI6fudgSmnGxo28vSG6P0av5s05v3NHDmvXLlH9ZTcs3cAeYJ3g6sqYP6jiG2CWtfYDn9/M2lDW4pMRs2erXYdSLHyOTOQ/jXyjrkKaEv6rkEEpRcO+SOWfkywyDxGflxcsIkVRlNwxFrgL7wZkidiKd5axV8axe/1KRIQGSaC6jVi9IR3uJvmEt/XAu0hyYyS/IfsC9G891r1KAh0A7wSudK0fdmh4k5yxEcmsjWzCODSHx/PKdB5A+n0bBnisy0QDKma8PlefbB/EAO4GZ2vJjwE1yIXmLaKF8PWIwByvkR+I+Pcu0TNWGxGRzd2s0M2uxBfAQkiH1EfjvO7wc+RBJTJui1zA3mpg3+6IbYHbuuMxRLRNVPLRHcnI6O9aPxvxMYwnYD+MlJtOQwbVwTjbeXEI8D+iP+t7yAA9Htsj2bmRlhC1yO/tvwn2M8AVwBS8ZZZvic5Gjsc2ZAbT4UZkoiMdmiPnmnuy4RvqfclToScyAfE7ZBIlYeOuCRMm9A+FQlXI72PN5s2be9x1110bUzym+/iOID0aGEb+/b2WUy9IO9nSW/Icg6IUnKohAyp8QQ73GXOYlWt4Xry3ItkaCnHUF4v4rkbGwDs3a8blXbtwYNs22dS66xDRb7q15uVNzZvPGDF7dqJBt6Lkg6nIhDfIGHIUUFm4cJQiYAxSWeZk4H2CJMmovZiiKI2NLsBk5Bl3XYo/2bAgaIfcgy8h+rk9XTYhFeKpWC2cgVTiRxJERMeGNKVkGYG3+Hk0iXWRcmJF6D8jY5dUcPq3RArAn5Jcr61NRPf1ugW4NMXjg3zOI13rupIbIXcUsba0twMXpvl+VUSL0FuR32W8Cu/jgH+71u2FVIWnwj3AORHLq4DOKb4HwHfU97EA+D8SO0csITrRdwWwE1m0qvPKgG5L9soB7gbuj/NaB+CfHjGcQ2LxGSTD9gpEWHRojfhBj0ROjFT4HCn5cOxADkGyjKtc2/VGsnjPIVYgfZCGxWcQ4fKPxIrcv0QuBJMQwTfyM3REvKIDRF88QIS7M2g4e3qf8M8qJIP3ReTvHC8zui/if3whsX+jmxs41jrkbxs521dOvQXJ3UQ3cOkAHIrYTTgXw63IwH9UA8fKNduQct2Pif7Sd0fE0xeRjON38O6w2gY5t0YDJxIrZCfkhhtuqB43btyLwDHAjq1atToPuCHFzxDJ4vCPc7NtjcxCOoL0/kjGfC7pitx0jw4v1yE3+UjrjnnETo4pSknzRa9ebUMtWhyEsYdbOJwQu2CyfqIHkev6agzfW2uCxtotGLMVbK0RoY0Q1Pqg/Mrvlo36rqZ2YN/tWqz5Q8eOXx/Wrt0PBtsOud51Ijtle2WEPeuNsRNb12zdUD2oYjrwcijEyxVVVV9n4RiKkipXIn6PRyP3wv8g92gVG5smRyLjY+eaNwMpxW6sXo+KojRtVhAtcOULg2gXN5CeoBaP20jd5/dpRPOJtIL1I9XpB5P5EN15LzcLgJcb2LcWEYBbRaxzW7AmwzBi9aNk2Uq0AJ2uPco7xArQZxCt5WWLTxD9JjJb/HikGj9VEdXpfxXJbBq3vegbRAvUXZDE07ezdQCvDOhscjUys+Z13H8Dx7rW30tsk754GET8c5/MdwIXJNjPKwP698DJxJZgLEEE4yBifdE7znt+iIjWXgJkPG5GZvy82IRYXKxFLsy74T1ZUIOUkT7ZwLEeDm/nxXIkW3kt8jlbIan28UTIhmZNHHZEBPx47/MDIlS3ob7Bi4PTgHIU0TfGQmRAOwxHPKF3ivN6EOmOvgaZFNg+/LMz8TOMd6SBDGiAyy67bLgx5iOylwXdEN2I9pIeQf67vf6IzBY7gvQMvD2kFKWo+WbkyO02b9hwrMWejuEwMs/y+B5YCGa+wS602G+MYUUQu7IZ5at2nzNndQqa9mBkwD4FGXDEML9v3zY0b97NmGDHkKWzCZmdrLF9fNDHGvpg2YWMjUHMJwb7WLAu9MSA6motc1fySRvgA+ofMF4HDif5fiBK4+BYpLLRGeu8g0xMbIi7h6IoipIqP0NsQ0dm+X1/RKp803lWvABJkHPzeyQLNRMm4G2dejqSrNcQHyPP4g7zgIoUY3Bn0kLyGdBLEe3M4Tmie7ElSz9EF4rURJYiY69cVEG/giQ3RnIk8FKK73MrcJFr3VXIc1M8Sj0D+hhi+/C9jkzIZIVCCdAXIyn8kVQif5xUTsKOyBfILQqeSHzvHi8B+hxkBuwVUsxQRcSxQ0jPjPxy4BrSe3jfAJyE+Ac1RCIBOhXuAc4neQsPt49eMgSR8+MOYr94hRSgQWaA/kVi+5FUSEqABhg3btz/gCMAjDHjb7zxxmlZiiEZyhGhyhGkh+Pth5RLgshscaR1x6dksRxEUbKFHTvW/8X8uQdYfKdbGail0+25FvjMwCxrzacW5lNXt6D//PlJXTOSpC2pTZzG8M3IkdttXr++j/XZPmAGG9jLwp6kl20RNPAGhsfstrrn+i1YoOKPkg/6IJV1jjXarcRPEFAaH6ciVWFOosdLyHOEWoMpiqJkhw4+n+8v1trzrLW5sH68EtFU0qEMmYge7lpfB5xG+g3YzkKcANyf93VEO0pGg7sXqUiPZBjSXykZhiDjG3ciWbIC9IdEa2MfkP7kwb8RcTaSXI23vETgKuR3l6wFYD/EJSFSU6pBtMRE1iGlLkAb5PwY4lp/NvGdLVKiEAL0Hoh4FPnH3BReX53GMfZHvsj+iHXrkBNsqcf28QTo+5Ds32nh5YYujrVI6cgUEvs2N8QeiEC6f5Lbh5CB8pXICZUMhyOf6WDS8xidC4xDBPpUGYacrMnOsp0NTA8vF5sADXJe/ByxSnF/MZPhW+AJ5G+YtNdkAbKgG6IrkhntWHeMIrpEJx9sRG4MjiD9Npk121SUjFg0eHCnYKjuD9aYc4i+2SfDd9byrs/wYciaWS02bvykx9KlqdpJFQUWfIsqKvrV+djTGLsX1oxCrA5SmWzdYuBJE+K2PlVVyQ60FSVdDkXsz5yx5O+ABwoXjpInzkZs4Zwx/4tIQ66SvPYqiqIUGb7y8vJfW2tvq6urS9e+oSHWINnPmSQtDEDEVbdOUotYhN5B8gl4ZYhl61Ri9aQ1yLNzstZzJxErgL+OjFkaSsLaCalu9Goil6wAfR8yHnKoAXYhdasTEBH2faJ/Jxb5XbkTUxvCh1i4xrNM8yP2nj1d6+9D3BYa0j93QHQFd7b5ozSc1FnqAjSIZclzrnU1yIR9ug06OyP2vFMNMtDKFVVIqUAko4jNWF5K+p0pwdu3diEiTrlJJEA79EaaER6CNPxzLpirEf/cfwOPIzYZ2WJPpDTgsHCMHZEL2NbwcechWRkv0rBHdjyaIRe9vRDxsBfQg+jfXS3yZZ6LCHvPktnfBupF2+OA/ZBMYmcm7rvw+z+NXGAjPXWGE33h2IJ8/oY4keiL2wKk8WM2McjkwUFI05q+yMXKOVdCSHbzYmA+4kc0ndjvQ9KMGzfuJWQyAeCyadOmNeTFnU/8yEyhI0jvg3x3stjHLCmWE+0l/RGZTRApSoNUD+nXx1jf+VhzNtHNVxNRh2EWlhcsvun9Kis/yb4ddPHwxbBhHYN128ZgzDFYjiKVZseGGVh7W99+Fc+ap55KpYmuoqTCn4Brw//fitzbZxUsGiUT/DQsFvwBse1zxilPIp6U2iBVURQlQ1q0aHGgMeb/tmzZskuOD3UJkkmbKccgfQDKPV77CEk+e534Fl3liHZ0DdJfyc1WROt5M4WYmiHJa2670keB8wj3dfHgaMR7eufwstsTOVkB+pfE9i17HxGNPyF1L+S/IBYWbh5B+rs1lFzZBtF5xiECrZd1isOBwGvETgL8A+nHFs+9YDDwGJI4E8nq8LqGxPfGIECDTM67rZEtcl5NJblE2HLgAOAXiHhdC7TNtzhUDCQjQLtxGpLkOyOiDfnzn2uNDNbzVXLYFsl8b2xiQgvkQrc522986aWX7u3z+WYCGGNWtmrVqmcgEMj6cbJIW+Qi7gjSIxGRPp/UIpMPjiA9m9jmooqSFgsGDTgihO9Sg03WF2uthaeN4cVgkDcqqqoKWcVQMOzYsf758+fvaQgdieUUTNz+Cq4d+cIYbmu2YdMDpZodrhQ1BnkwOTW8vAKZrI83yG5N/Ic/pbCcS2LvTrcv5z+QxBP1/lYURcmAbt267RIMBp9cuXLl3nk43DIkoS5b+sVJiPgYr/fR90hF+FfU2zB0RfSlw4ifXLEJEU7TqSa/CG+BfTliDTob8cBujyTEHUl0lfZryO8nsvdasgJ0S0QAbx/n9ZXIZ4ukGhHAvfAjCZWHeLy2BbGWfRXR6tYgAnwHYCCSrHko9Yk+F5JYgIb4Pde+Ryad30B+j+VIUuZxyO/P3a/HIsmUzzdwPGg8AnQzJHnSy3q2Bvk7vY0kWv6AaC7bIxaz/REXhFFEfyc2oAL0TzQkQCtKUTBu3LhIU/1Lpk2blo0Z33zSk2gv6T3IvClbqiwn2kv6fXIwYaA0XhYOHrBXyJrrkAzJhthmDK8BT20ob/H0iNmz9VxzUTVkQIU/ZM4AfoVUyDTEtxamrNih44MHvPWWCkZKNtkOaUA3Irz8AfI996qkuQKxYtNzsLjogjwU9Y3z+kTguojle5BMMu0poSiKkiZjxoxpvWrVqgcWLVo0tqamJl8a07mIT3I22RsRJ7OVub0IEba9KvOTwYdkTe+Xxr6ViFPAg4itgkOyAjRI9urjKRzzM0R8jEcL4CHglBTe04tkBGgf8Dekh1m61CKC7WNJbt9YBGiQCYh7kaaZ2WAD0DYXJvCKouSOQMT/J15yySXJlvsXC4uRMpuLECG6AzKzdjFS4pNNW5t4dEVmZq9GZoXXI1nRjyAX+grybx1SrLhLvpo0CwcO7D9/0MDnQtY4olRcLMwyhrOb+8u79J1TdUzfOVWPqPjsTcXn86r6VVb9afkOHbsbQkdheJLEZfA7G7iny/erP58/aIC7oYmiZMIWJEvJ6SewN/EfbvcmOqNIKQ5uwLsnhUH6vESKz9MQKw4VnxVFUdJg3LhxrQ477LA7Kisr186bN+/kPIrPSxAhM9t8gGTc3kJm1e+bEFuvIaQvPoPcn44G3kpxvzeRZ5V4VhPJ8gRi2ZuO77MXW5FKs4lI5na6JGP/EQIuAC4lPVeBJUhGdLLic2NjM2LD8gcy//vXAi9D0xRZNANaKWnGjx//qrXWKV0pNi/obNCNaC/p4dTb4OSL9cisseMn/T5SrtPU2ANptDqRzAZPJc0Xw4Z1DNbWTMVwFtENb93UgX3W4r+1f2XlzHzF1xhZNHBg9zpjz0fuz/FK/wTDDELm4n5z536cl+CUpsBopDTTqdDxyrSZjVhwJNtEWsk9I5F79jfIeN/BII6WwQcAACAASURBVCXMF0asux7x/VYURVFSJBAItP76668v+/DDDyfOnTu3EAlRv0Tsk3JJF2Qc+gvEViAZKhHR9j7ENzhblIVjmUTiasGlyP3tXuonVx9DhFSHOSRXxRlJOXAUkok9GMnGbYNYbkbiZF0nww5Ik8dTgN2S2H4l0jD6QeQZPRU6A38OH8vdO87NAuTvdwep95I6gtjz8mDEMzsVbkEqQx3W4N1QsiHmEX2+/BMR5VOlJXL+nYF4nCeTyPwjMnHyCtLrbQ2oAO2gArRSMlx66aWjfD7fjPDi93V1db1uvfXWdQUNKreUIaW0kYJ0IRocLibaS/pDUm++UIrcjGSsP4MI0UsKG05+mT9owFgwd5I4G3w9xj4UqrU3D6iuzkcWf5NhyW67tahp2+rkkGWikS7l8QgZuH+r8V86ZM4ctx+doqTDeUiTOhCbjcMQUdphFXJdGEL2Gx0rqeNDmkaOQEqeHW95P/IgHllyejUwOa/RKYqiNAImTJjQxlp70ccff3z5e++917K2tiB9W6sQETSf1SvdECuFXkg1bavw+o2IveMiJHt6hefe2cOHJAiNDsfRFskqXhI+/oeUZlWP86zfFRlbbYdkkf+ICMJVwFwyb9juQ7SEPcPH6YiM8VYjdhXvAF9meIzGzg7IRMRu4f/vgOgy65CM+0WIF/hCPGzqVIAWVIBWSorx48c/b609FsBae91NN910eaFjyjNdkJuv4yU9moayJLPPJsTnyvGSfofslScVEy0RcWV3RHB/CJl9X5Vgn5JnXv/+XU2Z704jTSfisRm4vbm//Loen33WmCeBCo4F34JBA04Ecy1yLsZjsY/QuX0qq6fnKzalURPpwfcD8sDyJTIxug15kLmPaJ8+pTCchWRFgTTF6Y+Izw8CZ4bXW6QUt9T6ZyiKohSUQCDQwVp7UXV19WWvvvpqq3XrCjrsPQF4rpABKIqSHipACypAKyXFhAkT+oZCobnIQ/DWUCjU5+abb/6m0HEVED/Qj2hBehjJlYdkE6fBoWPd8TGZ+YcVC4cS3bl5LVLe9Tey13m6aKgeNOBcg7mB2LIyhzowD1BWO7nfpwuW5TO2ps4XvXo1D7Zs/gdC/BnDjnE2swbu02xoJQuUI70CnHLSOUhX7/aIzQPINX4nRKBWCkMbJONmR+S+/zkyWfA4IlSAiM8XIiW1iqIoShJcfvnlHcvLy89ftWrVpa+++mqbxYsXFzqkj5Hre6aZsIqiFAAVoAUVoJWSY/z48fdZa38XXnxg2rRpv0u4Q9OjDVIa7Vh37EX+m+rVIeUnjiA9G/FiKsVB08PUZ5E5fIc0xnwIjxKbUuPj4cNbtq7Zdi/YRN1+n7e+4IT+n89fmLfAlBi+6NWrbbBF8wkYxgHNvbcyc0P+4AkDPqv+Iq/BKY2NzsBHQPfw8nNIo7tIn/fxSEM7pTBMQzKbneea2cj9yWkSGQR+R24aVimKojQ6rrjiis5lZWWXbNu27cJ33nlnu1mzZhEKFYWzw+FEJ8UoilJCqAAtqACtlBwTJkzoFgqFvkAsEoJ+v3/I9ddfX1XouIqcbkR7SY8grniVM1Ygs/eOl/R7ZN6hOB/sgIjnXo0bFiC2HE9TmuI6VRUVu/h9PIOcEzEYWIlhQt85VY/kOTQlAVUVFb38Pu4nfsOTDcaaX/WdO1dLNZVMGIZcq1uGlx9HuriDXPO+Q7zwgnmPTOmF3JvKI9atA7YP/78WOA25PymKoigJuPLKK3f3+/1/staeWVlZ2ey1115j48aNhQ7L4R208a+ilDRNUYDujHQCjeQWpKOmopQU48aNmwo4/s8vTJs27dhE2ysxlCNNLBxBeh+gR55jCCICbqQg/SnF2UDiVKR7bjxmIo0K381PONlhfkXFAfh4krgZ8uYp06zmvL6zF67Ja2BKUlgw8wdVnG0kC7KN9ybc0Ley6gpTnN8rpXjYDZmo/Brx9I/srnQ69eNHS+wY+njg+RzHp8TyX6RBpN/jtRqk4/2/8xqRoihKiTFp0qQKY8xE4NQVK1aUvfzyy3z99deFDsvNGODtQgehKEr6NEUBWlEaDRMnTmwXDAa/RLJT8fl8B95www1vFjisUqcb0V7So5FOvPlkA+I16lh3fIB05y0Gnqe+rDke05GS9M9yH05mzB848DcYey/e4sVya82v+8+d+2q+41JSp3pov91M0P8IsK/X69byxIodO55xwFtvlbxdjJIz/MgkWgDpsfA98C3i9/wdMmE5ymO/IPJQfFBeosyccqB1+P+RDXyDwPrw/3+k+CdsjiC5BJLN1NtE1SCf8RvgL4COmRRFabJMmjRpiDHmMuC0LVu2+N98801mz56NtZby8vKaurq6LdbaTUi/A4tUmIBcV7cRfd/YiEzc1ob/D9AXmaDNlJeRa76iKCWMCtCKUuKMGzfuUuCm8OLMadOmjaZEbRCKlDJk8OQI0vsA/cn/9XM50V7SHyEDv3yzCzAX70zTSELAM8CfgIJ3LPEi3GzwLryaVRpm4K87WZsMlhZvjhlT1vX71X9FRMQYjOFF3+ZtJ/VetKgQ3x2ldBiO2Gz0Di+HkPuq10SVgwUGInYQ+aYF0Aep4OkMdEEqOroitkmdwsstw9umwubwzypkInRZ+N9V4f+vAr4AlhCdMZ5rmgFVyGdO9Hdxsw74K3AnjaNJsKIoSspMmjRpNPAnY8xRhJ9p6urqsNZubNas2YPGmOsDgUCmY+BjgGeRZ6lMsMAeyPOPoigljArQilLiBAKBZps2baq21vYMrzp12rRpTxQ0qMZPO2Qg5Fh3jAI65DmGWiRL2hGk30UEgHxwAXB7kttuQx70pyLZhEXBgsEV51nLHXjcBw3cWxfijxVVVTUFCE3JAtWDK041lvuAVh4vv9R8w6YTeixdquKTkoiWwI3AHxABuiGRMwTcA5yXw5h2QPyo+wD9kMnRPsjEYOxEWn6pRe5BC8I/C4H5iKVULgxExyF/n2SwiNj8N+A66jP4FEVRmhSBQGCfYDA40RhztOul9cDffT7fDYFA4IcsHGov4A3qeydkwjPASVl4H0VRCowK0IrSCBg3btxpwD/Ci99u3ry5/1133VU0HSOaCD2pF6RHA0NJLSsrGywn2kt6BrAlB8fxIeXm+6Swz0ZEiL4GsRgpGPMHVUxERAg3tRbO6V9Z9VCeQ0qK6kGD9jcEvZpAxmDxf9u/snJmOsf5ePjw8tY1W5IulwyGzKyKqqqiMwpcUFExxPr4DyLORWN5pfnGTcerCK0kwSHAI0hmcUPj5q3ATkA2Ht6LpfomU7z6HHxGZg0bOwGLEBuRRL+PUPjnQeBqpAmwoihKkyMQCBwcCoWmAHu7XloN3OXz+W4NBALZmpwbgCTGZCM5J4jYXxWiukhRlCxTaoNYRVG8MePGjXsL2A/AWjv1pptu+nNhQ2rytEZEaEc82B95aM4ndUgWWqR1xzyyY9HSFxERUi3nXo1YxtyCeHHmlerBFScay1PE3v9qDKFT+lZWF22zqvmDKqaTpMesgZXLdui4czp+x/MHDTgOTNK/B2s4rf+cqsdTPU4+qKqo2KXMx+sWesW8aHi035yqMwsQllJ67AjcC/wcETQTZRtfilzfUsUHjAAOBw5F7h2pXl9TYW34X8ezE6A5kq3mQyp9csUGpLfBK8BLpC4sPACcRfznmCDyGZ5GGjV/mV6YiqIopcvYsWP9/fv3Pwm4AhFxI/kWmObz+e4LBAKbs3jYbsD7wK5Zer9HgF9l6b0URSkwKkArSiPh0ksvHebz+T5Csm5rQqHQoJtvvnlhoeNSonAaHDrZbLkWGLz4EfGPdgTpGaSfrRdAssrSYSFwJSIQ5MWzfGFFxdCQjxnElgNus4ax/edUvZCPONIlFQEawGfN0X3mzv1v6scZ+BzYpDOgi1mABqiqqOhS5mO6hQr3awYu61tZdXMh4lJKkjOBu5DrtleFiwW+AnYnuQZ+nRCx+YjwvzumGVcofNyFSCXMcsSbeVXE/1cjXs6pVke1Q66ZkZ7SHZH7WUdgZ2RCcqc0Yycc+8vhn9dJXCXzM+BjvJ9hgsjf5XXEoqPoG+EqiqJkm0Ag0CwYDP7CGHM5YtcUyVLgVp/Pd08gEMh2FVhbpEJyaJberxap/NFJREVpJKgArSiNiPHjx//dWvv78OIL06ZNO7agASkNUY5kJDhi9HCkbC3fLKZekH4P8exMRjxpBnyCh7CXAh8hjQrfyOA9GqRy4MDO5cZ+SKwdwyYMx/abU5XT42eDVAVo4Ol+lVVjUznGF8OGdQzW1XyHnJtJUewCNIgI7feZ18AOdL0UtD6O6v951SsFCUwpRXYDHkWu2xbvsfTRQLzJn+0RL8tfAvuSmndzHdIE9nPqfZadn0I31myD+FH3QQTpvogI0ZfUnjdqkKzofwAvEN0o0CD3HLe44WSlf4A0IH0n9fAVRVFKm0Ag0DoUCv0WmYDb2fXyl9baG/x+/4OBQCDl6rgkaIbc9w5uYLvXgTEkZ1N4N9KHQVGURoIK0IrSiLjkkks6+P3/z96dx0dVnX8c/5w7IWETcAW1LihCIARU3Ks/l7oj7nGpirYqtlpUTAKyJB4TQCBBFFqtWKviLq6AS1utWveFVpaQgBvuCy4IYUsy9/z+OBMzuZlkQpKZO8vzfr3yMvfeM8m3FGZ57jnPCawkNItKKTW8rKzsWZ9jia2zM3YZdn1B+nBswSKeqrEFjvqC9CvYGXSRHIwtXre33/ULwFhs8btDvXTUURk7f7/mJVSTntUuijOyl1Ys6OjfGQttKEDXqMzaXQcsXvV9ax+wMjdnjIGtmhGcDAVogJVDh+5q3Lp3sf/Gwq0NuhyYU1HxoR+5RFIKYD/gT8a+lw5//jPYD9jHhZ3Lws5wzgPOovWbMnn7+r+BncWcTLYBhtKw8ucgWt+O6mdgATAfW5QeiW2/4bUCuyJnfjuzCiFE0hk/fvz2nTp1Gg2Mpmnf5cXATY7jPKm1bs3kkrZwgIexr3EteQ84GviW6K+Dm4F9sK1ChBApQgrQQqSY/Pz8K5VSfwkdfrhly5bBc+bM8XtmlGi7AHb5XHjrjmy2btZcR6gvhNT3k36Phplps7FvetvLEIOenVW5OUVASZMLSk3KXrp8Skf9nljzFqBdQH/51S/XT+zZg8O6d2/0GAV/GrCs4i+0UlVuzvvYYtEv5nz7HWvq7GSZ/bt25fRtG98PSZYCNEBlbu4whfsq0MVz6c0B2YOOUPPnt2djNJF+DgAexH5I9hqIfY4cje1XvG0rft5a7M2454F/Ap93TMyEk01D65EjafrvMZIvse1Awp/kPsPeuHyUOLVyEkKIRDFhwoTeGRkZfwSupWnf/teB6SUlJYuI/fPjrcDVUcZ8iP0c8x229V+018SZ2Bu9QogUkuF3ACFEx/rss8/u2GOPPS7HLlHt17lz52uAGT7HEm0XBCpCX/NC53pgW3fUF6QPoe29Q1trZ+yy8lNCx7XAUuwb3GXY4kB7eoCCvSmaB5wO3A0UY2dJtFnV4MFDwEyK8KvmD1i6fGp7frbfjDE8+qPdR0wBW1zTpABt7MYtrSpAfzB00H5Bt3Hx+fu6Ou774UfWBW1dts6YJgXoZDJw2bLFlUNy/qgM93guHbqqasXVtG3zOJG+3sPesJlG0w/f/8A+J0ZbHfJfGvofv4lts5HqqkJfs7HF5//Dbr44nMjFfGj8+rIl9NiJNGygKIQQaUFrvZcx5hpjzCga7yVjjDHPAFNLS0vfjFOcfKIXn7/HPr/Xr6aMNjGqGvnsKkRKkhnQQqSgwsLCI4wxr2D/jVc7jjNwxowZsoQpte1C480ND8T2Y0tm1dji6VRgXVt+QFVuzsvYGXa/MLCiRgUOGrp06YZ2J4wj7wzooDE88MOP5HbtwoDOnU1Xx4n4mh50zOCcJSsqov78IYNuxaiIHyK+qql1qzZvdoIYjuvRo9G1ZJoBXa8yN+fPCq7ynF4fdOmfU1HxjS+hRLIbDjyObbcRzYfYHscPYvs3iwbDsP2xzwP6tDDOYGeJl2FbngghREorKioaqpTKB35L45ubLvC4MebG0tLSqO/3OtDZwCO0vCpzI/a961th51YDe7TwmFLsJBQhRIqRArQQKaqgoOAB7BsUgKfLy8tP9zOPiLtuwH409JI+Art5VjL6HigHbmErNtqqHJJzljI85jldp1wOG1BR8W5HBoyHSAXozcbQzYnSjUWZ8uylKwpbGlKRk5MZUHyJankm/Q91dWyf0XjxVDIWoN8bNqzrNjWblxjo1+iC4q7spRWX+RRLJK+TgZuwK1Oaswb7Qf0BGn8QF5EFsJtZXQCcT8urNv+N3cw26Z7XhRAiGq314cFgcJxSajiN6zdbgEdd1y2dPHnyB3GOdRDwEi33cq4FTsWu8Am3ErthbSQ/AntjW1IJIVJMvHuICiHiJHSH/KfQ4WmFhYVn+ZlHxN0GbL/mW7EbN/XFLmE+FZiObZ2xybd0W2cH7BL3lcAoWvHaZcDBROr7bG5JxuJzuxh1wUtHHdViyy1HqVOiFZ9TyQGLF280iiuaXDBcUpGT0y/CQ4SI5CBs8fMZmi8+LwGuwM72Go0Un1sriG1jMgrbT/ta7Ky5SI4B3gH+BQyORzghhIgxVVxcPKK4uPgN13VfVUqdQkPxeb1SarbjOHuVlJSM9KH4vBewkJaLzwb7/O0tPgPUtPC4MqT4LETKkhnQQqSw/Pz8UUqpO0KH32RkZAyaNm3aTy0+SKSTDGAAjVt3DCTxXxvew852a3bZdTOzn7/rlNm5/96LF/8c03Qx0uYZ0IBj1Cn9ly9/prnrK4fkLDTml/7ezUqVGdD1KnNznlBwRvg5A3cOXFYxyq9MIinsDtwMnEnk50sXOzNsJ+wmeZE+gIutF8D+ex0DHNbMmDrgb8AEGm7CCyFEUhg1alSn3r17n6+UGgvkeC5/Y4yZFQgE/qq1blNrug6wA3a/gmg36ydgVwZF8h72M4fXGuzs5/VtTieESGgyA1qIFDZz5sw7sbOzAPoEg8FpfuYRCacOu7nhXOws6RzsrtTHATcCi0jMD/AHAC9gZ8ftF2mAMlzT9CQ3JWvxub1cxcXNXVs2eHBvYzghnnkSRcCoidhi4S8UXFSZnb29T5FEYnOAPwHLgbNoWnx2gfuBbGz7iP2RD9IdKQg8hr1pegh2xrNXBvAHYAX2BoEQQiQ8rXX34uLia/r06fOhUupeGhefVwPXrl+/fq/S0tIZPhafOwNPE734/DeaLz5D8+30SpHXTCFSmhSghUhtxnXdPwKbAYwxlxcUFBzjcyaR2H7GFnc1MALYEbuk+WJgNrAYT8HOR8djZ1E8gF0OCEDl0Oz+2Bnd4dZUd+o8N47ZEow5beWw/hFbbGTARUCnOAdKCP2XL68EnvCc7qwyAxf5kUcktH7Y58Y5wDYRrr+AvTl2EVC/HLoO2+5IdLy3sa8Bx2FfB7z6YDeFXIhtPyWEEAlHa71TcXGxdl33U+xeJ7uHXV5ujLnYcZx9SkpKbp01a5afrfMc7A3W5laf1HsO+GOUMZEK0J9hJ8QIIVKYFKCFSHE333zzKmBq6FABt2utO/sYSSSXIHaW9DzgGmyBpRd2U8NrsR/uW70xYAw42M02VwJ3AH2UG7iQpjMT/37A4sUb4x0ugWRS0+ncSBeUMiPjHSaROLhzvOeMIa3/TEQjDjAeO+v56AjX38H2ID4O+F8ccwnrBWwv7vOADyNcPwVYht3MUAghEoLWun9xcfFc13U/A24Atgu7/IrjOMNLSkqGlJaWztNa1/kUM1w5duVPS/4LnIO9+dqSSJ8bSpo5L4RIIVKAFiINdO/efboxpiJ02H/Dhg3jfQ0kkt16GjYxPBLI8jcOYJddjwI+LPnqqz9Uu40naRsn+HdfUiUQQ9M2HCtzcg4Ecn2IkzD2WVb5KvYGxi8U7Pvh4MG7+RRJJI4e2Fm0U2n6PLcOexPuUGy/Z+EfAzwCDMLuD+AtYmyLnbl3B5AZ32hCCNFAa71/cXHxPNd1VwCX0/Da4hpjFhljDispKTlKa/0s9rktEVyB7b3fki+B04DqVvy8zZ7jD4B725BLCJFkpAAtRBrQWtcopS4n1DrBGHP9uHHjvBtbCNFa/bAbAN6BLdAkkm4P/vDTjsetXMWda76nxhiAJQOXVK3yO1gCOLAyN7dRsdkoLvEpS8JQYJQxj3tP1ypzki+BRKLIxrZ5OD3CtWexrYluJXFaEgmoBaZjV+q8HeH6KOzN090jXBNCiFhRWutji4qKFrquuxjbqikQurYFuM9xnJzS0tIRpaWlb/oXM6LhwF+ijFkHnAx80cqf6b1JeAPRZ00LIVKAFKCFSBPl5eVvGmNuDx1mBoPB+0aNGpWWfV9Fm3UCxtH8UvSE8VNdkJnffMsJKz9g6tdff4m83gGgjPtLb+OKnJxMFBHbcqQbVwUWec850fscitR1HravcLbn/JrQteHA5/EOJVptOXajwnzA23rpAOAtmu4TIIQQHUprnVlUVDSyuLh4meu6/1JKnRJ2eZ1SarbjOHuVlJSM1FpX+Ra0eQdgV5cEWhhTC5wNLN2Kn1sT9v3y0O8QQqSBDL8DCCHip6amZkJWVtYpwB7Afj169JiI3WxOiGgOwe5qnVQz57+urWXe9z+eDLyLXZr9L58j+UopRr501FETjn755boMpc4wmO39zpQINmRmvte9ZvMmoEv9OQMH+xhJ+KcAKItwfjG2/+Wn8Y0j2igI3IztEf0EsHfYtZ2xq3jOp+kmpEII0S5jx47dpnPnzr93XTdfKeVt5/U1MHfLli2zpk+f/rMf+VqpL7AI6NbCGINtI7K1763DZ0BPRFYSCZE2ZEaYEGlkzpw565RSF9HwQj9x7NixB/mZSSS8rsA04DWSrPjssT/wT+yb5GE+Z/GNgd67fv/9CQBGmUt8jpMwDli8uBZ433O630tHHSU36tPLOCIXn+diZ8RL8Tn5LMU+/z/lOZ8JPApcGPdEQoiUpLXepbi4eFrnzp0/B24BwovPS40xF33zzTd7lJSU6AQvPm+HbTXVO8q4G2hb7+b6AvS72M3MhRBpQgrQQqSZsrKyV5VSc0KHGa7r3jtmzJguLT5IpKuTgUpsUaal5XfJ5FjsG96Hsb2s046ruLhqvwG7AMf5nSWRGMMHnlMZfdZ+8ytfwgg/aOzNtnBbsLO7rqDxkmGRXNYBZ2JXwYTPtAsA94D0whdCtJ3WesgNN9xwh+u6H2HfM/cMu/w6cGpJScm+paWl98+dO7fWn5StlgnMp2kLKq+7gdI2/o76AvR4EmejRSFEHMjMHiHSULdu3a5fv379sUqpHCA7EAhMJfruxiJ97AjMAi7wO0gMrA19DcBuqlKGXaKdKqqB7i0PMadSm7EaFfWmQit+VupQKsLs1rqMXYDVcQ8j4m0atmgQrhoYAbwc9zQiFgx2g8JvgLtouKkaCB2DLUYLIUSraK0PDwaD41zXHQ6osEuuMebZQCAwRWv9ll/52kBhnw+PiTLuJeAP7fg9W4BXsa2QhBBpRArQQqQhrfXmwsLCkcaYt7Aby109duzYBTNmzHjJ72zCd3nYwuyOfgdpxmbgp7CvTaFz64E1V+6041W9AoFAj0CAnoEAAcX7o1Z/dmFo7Hek+C7bCrXcYHrR8syVLBT50X8Wj5k0mhlojFqnVOOJOKbl3ociNVxM0+Lzz9gVIG/EP46IsXuxrxv3Y9//gF0ROhfbYkXeBwkhmjV69OisXr16na+Uus513VylwuvObFBK3R0MBm+ZPHnyR35lbIebiN6WqAK7oqQ9q4K2AJPa8XghRJKSArQQaaqsrOy/BQUFU7H9uxzXdeddf/31Q6ZNm/aT39mEL/YEbgdOjPHvqS8gr23h6yfPf8O/mi0gG3BW9t7pas/Jb7FvltOCwRhjzDyl1NQoQ6O14KoJGvWgk0Z9oh3HbDSehaCOcrv6k0bEyWHAHZ5za7HPg2/HP06bDQP2aubaT3TcKo+Dgd2bubaG5Jkt/iiwEXgMyAqd64Rddn4wkIyFIyFEDGmte7iu+zvsRrXe9lzfAbfX1tbOuemmm36If7oOcRlNb8Z6fYW9Obu2nb/rKWBJO3+GECIJSQFaiDTWvXv3ydXV1ScDBwK/qqurmwn83udYIr4c7JvOcmCbVj7GOwu5tV8/hh4bEwrcKqilYVYbqF+KC2nDCXSaZ9y6UtrRt1spFnVy3O+Droo+OEW4ruraZAa0cTb5FEfE3q+Ax6HRc8QG4DfAf31J1Hb1faojqcH+b13Tzt/hYAu3zRWg/wMc2c7fEU+LsG2mHqXhhtz2wBPYGxMbfMolhEggkyZN6hsIBK51XfdSmq6KWgXctn79+rmzZs1K5vcLJ2InobRkHbb4/FkH/D4pPguRpqQALUQa01rXjR079mLXdRcDXYDf5efn/2vmzJkP+Z1NxEVP7GyHAdiZX62ZmZzoH8qrgW0bDlUP35L4ZMCSJV9WDc55AcUJbf0ZLtyTdrsUK9PkBowyJtH/vou2yQSeBPqEnTPYljPJVnyOJhM4H5jdzp/zG5ovPierx7GrwMI30hoC/B0415dEQoiEUFRUNEwpdQ1wvjHGWzN5HZheUlKyiOTfRG8QdmPulupCtcA5SOFYCNFOUoAWIs3NmDGjMj8/f6JS6mYApdTtBQUFb5WXl3/idzYRcz8DE/wO0cG+pVEB2uzmWxIfKYd7jGlzAfq7DZ06P9+zbtPgDg2V4BzYw/spMqDUN76EEbFWDBzgOVeKbcmQii6m/QXoSzogRyKaAuQA54WdOwd4GnjQl0RCCF9orZ3QhoLjgF97LtcCTxljykpLS9+Nf7qY6AM8h52Q0pJrgH/EPo4QItWl3QQnIURTM2fOvEUptSB02BN4RGud6WcmIdpGfew5seMn++7by5coPspct+Ep7Iz16h18+QAAIABJREFUrWbg/gMWL67t4EgJzxj29pwK1hjzuS9hRCztR9M+l08DN/qQJV72B4a24/E9gdM7KEuiMdg2VO97zs8Gdop/HCFEvGmtuxcXF492XXcVsIDGxee1wPRgMNi3pKTknBQqPnfB9mKOtrJlMtHbcwghRKtIAVoIAWAyMjIuA74MHR9YXV092c9AQrSNWeU9Uxvc4p3pmPL6rl69GWUeadujnXs6NEwSeOmoozJQ7Bd+TsEnORUV7dnlXSSmW2m8AvBr7N4Hrj9xYsa7kd7Idvysc4HwDTlTbZO+Ddg2JeE9XLencWsOIUSK0VrvUlxcrF3X/RR70yn8RvRq4HrHcfqWlJRcP2XKlC8j/pDk5GBXeBwcZdz92BVDQgjRIaQALYQA4KabblpjjPktEAydKigoKBjhZyYhtpZRvNPknFHeZZRpwcXc24aHLR64bNmyDg+T4HZZs2Y/PJsLGdTbPsURsXM2cITn3CjsBqmpZp7n+ELCN2jdOr/zHLfluSXRVdF0Fvyl2J7QQogUUlRUNKy4uHheqPB8A7Bd2OX/GmMudhxnn5KSkula67U+xYylMqKvankVuzok2XtcCyESiBSghRC/mDlz5n+wS60AFHDX2LFjd/ExkhBbxwm+6T1llDrFjyh+G7S08i2gcmseY+Ce2KRJbG5ADfeeU8q84UcWETOKpjO5FgKLfMgSD88C4T3MdwJObMPP6U/jWXK1wAPtyJXIbgZWhh0HgEk+ZRFCdKBRo0Z1Kioq+m1xcfG7Sqn3gItoWA0TNMY8Zow5vKSkZFhpaek8rXWdj3Fj6VLguihjPgbOArbEPo4QIp1IAVoI0Uj37t1LlFIvhg53dF33wby8vICvoYRopYHvV602sMJzeljlvtl7+pEnAdyzFWNrqA0+FKsgiUwZc6b3nCHwnB9ZRMycBuSGHQeBQp+yxEMdTQvFF7fh5/weW7yvtwhY09ZQCa4WGOs5dxaQ7UMWIUQHGDduXM/i4uJr+vTp85FS6gEab0C7Tik123GcfqWlpXmlpaWv+5UzTk4A/hplzI/ASaTu87wQwkdSgBZCNKK1duvq6i4Gvg+dOnL33Xef6GcmIbbSAs+xUsHA731J4reMuvtpaKvTMsPCgVVVP8Q2UOJZMWTgITQuTAIsy1669BM/8oiYucpz/BCNZ7umors9xyOAHbbi8Q5wgedcKrbfCLcAWBx27AB/8CmLEKKNJk2atM8NN9xwa1ZW1lfALcBuYZc/wvZ33uPGG2+8Rmu92peQ8TUIeJjGeyB41WJbVTXZT0UIITqCFKCFEE3MmjXrS2PMSEJ9v5RSN+Tn55/kcywhWsUoN9Ly8Es/2XPPznEP47Ps/638Cvhna8Ya1D2xTZOYHBP4U4TT98c9iIilvsAxnnNlfgSJswoaF1MzsZvttdYJwK/Cjr/DtvZIdeWe44uAtHv9ECIZaa0PLyoqWug4zkpjzNU03kD1deCcysrKASnc3zmSPtjn7l4tjDHY9hwvxSWRECItSQFaCBHRzJkznwNmhg4dpdR9BQUFff3MJERrDFpaudyAdwO5Xbb06O7dSCstKKOizlhU8O2GrKx/xCNPIlmVm7sXmHM9p+vcOvc+XwKJWDmPxu953waW+pQl3u7xHG9NGw7vc+YD2Blyqe4JGi8/3w5bjBdCJKDRo0dnFRUVjSwuLl7uuu6ryu79Ud86qAa4z3GcoSUlJYeXlJTMnz9/futWhqWGLsCTwB5RxpUA8t5HCBFTUoAWQjSre/fu44GXQ4fbA09prbs2/wghEsbsJmeMmbhkyJBuPmTxVWZ19dPAMuymMs193XbA4sXpUFhqxMUtxbscVfHQoMrKr/1JJGLEuxHpPF9S+ONBGm8kNQwY0orHbQec6jmX6u036tVgl6qHG+FHECFE87TWuxQXF+ttt932S6XUvUBO2OXvgOnBYHCvkpKSkVrrdLnpGM7Brug6JMq4R4AbYx9HCJHuWuoBJIRIc1rruvHjx59dU1PznlJqT2BIdXX1nTTtCSlEQsnOHvTIqqoVNxroF3Z61ywTHAcU+5XLD31Xr95M6wpOaaVy8ODDwHjbEbiBINN8CSRipSeNP3wbmvaJT2U/AguxfT3rjQQKojzufCAr7Pi/wJKOjZbQFgCjw45lBrQQCUJrfbjrule7rnsGTesZ7xtjbq+urr5v1qxZm/zIl0CmA002WfZ4DbsyxsQ+jhAi3UkBWgjRoptuuumH66677kyl1OvYZVy/LSgoeLu8vLzpDFMhEoSaPz9YNXjwjSjjXU44tmLooEdylqyo8CWYSAgf9OuXFVTcQcMSXbAHj+xTUbHCp1giNg6k8Yq/CuALn7L45V4aF6AvAsbTcjuNSyL8jHTyH2AT9n0P2F7YuwJf+pZIiDRWUFDQrWvXrhcCo13XzfFcDhpjFgYCgVu11i/7EC8RXUr0G40fYwvUW6KME0KIDiEtOIQQUd18883/M8ZcEXZqZmFh4ZG+BRKiFQYsX/5AhF7QWQFX3VuRk5PpS6gOFFCKLk58X8Z7BQJx/X2xUtclS4MZ7Dm9MWDUOD/yiJg6wHP8li8p/PU8EN5WZifgxBbG59D4z60GeCgGuRJZDXbWd7gD/QgiRDrTWu9VXFw8rWvXrp8Cf6Vxm42flVKzHcfpV1paeoYUn39xFHBblDE/AifTuN+9EELElBSghRCtMnPmzPuUUnNDhxnAI2PHjv2Vn5mEaIkC4zhcDXg3mxkWCER9Y54U4v0iHlAq+qAEVzl48JkKmhaalZrab/nyz32IJGKrn+c4ndpI1KvD9gEN19JmhN7NBxeSnkWK9z3H3r9LQogY0VofXlxc/Kjruiuxr9nbh11eCVy7cePGXW+88cZrtNarfQmZmAZiNx1saaJFLZCH/XMUQoi4kRYcQohW27x589WdO3ceYow5xBjT2xjz2OjRo4+cM2eOLN0SCWnAkop3KnNzyhRc3+iC4dLKwYOWDFy+Yo5P0YQPVubkDDXKzMPTegPF+9Wdsmb4k0rEWF/P8Se+pPDfPUBh2PEIYAfge8+4DJru85Bu7Tfqef+ueP8uCSE6kNa6ezAY/K1SarTrut5VSq4x5tlQm40XkZ7FkeyAvWHYq4UxBrgM+HdcEgkhRBiZAS2EaLU5c+Zsqa2tPRP4KnTq4KysrKbFHCESSMamLRqUdyk1SqmbVw4Z9BsfIgkfrBzWfwfj8BTQzXNpg4t70QGLF7fUD1ckr+08x19HHJX6VgDvhh1nYjca9DoZ6BN2/B22hUc6+spzvH3EUUKIdpk0adLexcXF01zX/VQpdQcQXnyub7Oxd2lp6Qit9QtI8TmSLsAzwN5Rxk0G5sU+jhBCNCUzoIUQW+WWW275urCw8DxjzAvYD7DnFBQULC8vLy/1O5sQkezz4YdbKnJyzshweMdA77BLGcao+SuH5pw4YEnFO74FbDXzNaiPWx7SpGDSdnVqCw4t/z6rusN+Z4wszc3d1mxxn0Gxp+eSMYqLBy2tXO5HLhEX3hsOCf/3NYbupXEf44sB7yqQSzzH99HyZoWpbIPn2Pt3SQjRRlprBzjZdd0rgRNoOjFuiTHmz9XV1Q/MmjVrU/wTJhUF3AUcFGXco8ANsY8jhBCRyaxFIUSbFBYWXmKMuTt0aIALy8vLH/QzkxAtqRw8+DClzL+BLM+lamPUiIHLl7/sQywRY0tzc7fNxP0HETYQM0rdOHDpch3/VCKOPgL2CjvuFzqXSv4KXOE5tx9NexhvB3wJdA47NxRYGvp++9D1rGau19sGWOc59x8g1TYnPhk7o7De88BJPmURIiVorXu5rnsxcA1N29pIm422mQJMiDLmdeBYYHPs4wghRGTSgkMI0SZlZWX3ANNDhwq4q6Cg4FD/EgnRsoHLl79h4A8RLnVXyiySdhyp58MhQ3bKxH2ZCMVnUE9lL11eEu9MIu68M+e6+pIiMfwILPCcGxn2/YU0Lj6/R9Piczpp0q7HlxRCpICioqJDb7jhhvtc1/0GuIXGxec1SqlpwWBwT2mzsdXOA8ZHGfMJcCZSfBZC+ExmQAsh2kMVFBQ8iH3zA/C9UuqQsrKyVJtdJlJIZW7OTU02JbQ2gvlt9rIVT8c9lOhwHwwevHdQmYXYHeG93gi6nJBTUZHO7RjSxRtA+M3Ro4BX/IkSM62dAQ1NZ/V+B/wK22bjf8C+YddGA3+O8DPSZQb0Fdg/23p3A7/3KYsQSUdr3TkYDJ6jlLoW+5zktdgYM7e6uvo+abPRJocDL9B0ZV+4H4HDgJVxSSSEEC2QHtBCiPYwwWDw94FAoC9wMLCDMWbBuHHjDps+ffrPfocTIpKByyrGV+XmrAWmeS51BfVkVW7OjAHZgyaq+fODfuQT7bcyd+DJQcz9wLYRLr9KTd3wnJUrpficHj6ncQF6T1KvAL01/gF8gS06A+yE7b/6KY2LzzXAw/GNlnC87QE+8yWFEElm0qRJ+ziOc6nrupcppbybd24G5htjZpWWlv7Pj3wpYm/gSVouPtcAZyHFZyFEgpACtBCiXWbNmrXp2muvPSMjI+NtYDdgUDAYfERrfYrWus7vfEJEkr2sYnpVbg40LUIrYNzKlSsO+nDIkPP6LV36XfzTibYyoFbm5ow1MJXIbcZeCbqcIsXntOLdSDPSjPh0EgQeBMaGnbsY2/s53ALg+3iFSlDZnuNPfEkhRBIIbSp4TDAYvEYpNZymK60/AO5yHOdOrfWP8U+YUnoATwE7RBl3NfByzNMIIUQrSQFaCNFut9xyy9eFhYWnGmNeBboDJ1RXV98KXOVzNCGalb2sYnrl4EF1SqkyvB+UDEfXEXxrxZCBvx20tPItfxKKrbFs8ODeVcrcpWB45BHq2az11Wf1Xb1aeiCmlyWe44N8SZFY7qFxAfpUYH2EMenuEM9xpJYmQqS1CRMm7JyRkXGZ67qjgF8p1ejtVB3wNHB7SUnJv5G+zh2hE/A4MDjKuOnAHbGPI4QQrSc9oIUQHaawsPBkY8wCIBA6VVxeXl7qZyYhoqnKHXQaqHnYGSVeroK/1bnkS7/gxFWVOygPo25DRZ4NpGBuncvonIqKmnhnE77bCwjfl2AjdtZYKvUb3Zoe0PXewrbOiuRbbIuO5lYxpUMP6GygMux4A9CL5v9MhEgb9bOdQ0Xn07FF0XDfAPc6jnOb1lpa13Ss24A/RhnzBJAHuLGPI4QQrScFaCFEhyosLBxtjJkddmpUeXn5nb4FEqIVqoYMGYAJPknzy/M/cXBH9V9W+UI8c4mWVe03YBfqMm4DTmtmyGYwV2YvW3F3PHOJhPMZtkVUvVOBhT5liYW2FKD/iC1kRFJG4xnSXulQgC4EZoQd/wM40acsQiSECRMm9M7IyLgEuBzbg9hrsTFm9rfffvvQ3Llza+ObLi3kA+VRxvwX+D/sTTMhhEgoUoAWQnS4/Pz8KUqpCaHDoFLq3LKyssd9DSVEFJ/su2+vLXW181CMaGaIAXUPGbWTsv+38qu4hhONfNCvX5bbJetKAzdii2GRfKxczhxQUeFtwSDSz59p3BLqAeBCn7LEQlsK0D2Br4EuEa4NAZa18Nh0KEC/BwwLOx6N/XskRLpRxcXFx2CfY04DMj3X1wL3OY7zV631irinSx8nY3vzB1oY8xV2ZcsXcUkkhBBbSQrQQohYUAUFBXcCl4aONyulji8rK3vVz1BCRFO/iR2ggc7NDNtoYHbnQKfpfd9/f2380gkDzqohORcaKMGwRwsjH60h8Ichy5b9FL90IoEdA7wYdrwJ2BVIlb8fbSlAAzwEnOc59w7Nt+aol+oF6P2BxWHHQaAv8Lk/cYSIP611r2AweI5S6mogJ8KQxcaYuZs2bXqgvLxcZtvG1r5A/T47zdmEfQ5+Ny6JhBCiDaQALYSIiby8vMAee+zxKHBm6NTPSqmjysrKZBMfkfA+GDx476Bj7sRwdAvDfgRmdOveY/Zub76ZSv1kE9Kq3IHHusopw7BvC8O+MYo/DVxaISsuRDgFfEDjJePjgWn+xOlwbS1A9wS295xbB3wf5XGpXoC+j8Yz5J+l2c1NhUgtRUVFwxzHGWWMuYimKyTWKaUeVkrdrrWW9/PxsTPwNo3bSHm5wFnAU3FJJIQQbSQFaCFEzIwZM6ZLIBD4J3B46NRXGRkZv542bdpqH2MJ0SoGnKrBg65SSk2l5VknXxul/kxN3R0Dq6p+iFe+dPDesGGdutVuPlsZrgMOaGGoAXN3ViAzX2ali2aMB6aGHX+HndW60Z84HaqtBei2SuUC9F7ASiAj7NzpwNP+xBEi9saNG9czMzPzXKXUn4DcCEMWG2PmBgKBB7XWsiFz/HQBXgYOijKuAJgZ8zRCCNFOUoAWQsTUmDFjtgsEAq8Cg0KnqowxR8ycOTPaDCshEsKKgQP3cDICt4A5PcrQTaDmOYZb+y9fXhmXcCmqIidnuwyHUQauqnOcXymlCASDEccqqFC418oGkSKKnsBqoFfYOY3tI57spADdcR4F8sKOK4HB2BmGQqQSpbU+2hhzqTHmTJq2HVsPPOg4zlyt9X99yJfuFLZN0rlRxv2dhpaHQgiR0KQALYSIuTFjxuwaCAReh196ti4JBoPHzJo160c/cwmxNSpzcw9VuNOBI6IMNcA/jOJvnddteKbv6tWb4xAvJVTm5h6qVHAkRl0EdANYsU9/qvbpzz4ff8SAjz4ks7a2fvjnYG4YkJ0zT82fH7k6LURjU4AJYccbgIEkf29fKUB3jCOAV2j8+ehC7KaVQqSECRMm7JyRkTESuJzGbYnqrQDm1dTUzJ02bVqq9MlPRtOBsVHG/Ac4DqiJfRwhhGg/KUALIeIiPz9/kFLqPzT0m3zbcZzjZsyYsd7PXEJsrVb2Iq73M4oFBuZnDxj0rBRKm7IzzJ3zMFyKYp/wa0HHYcEJJ7Kps21DmVlbyz6ffLyp76ery3fGuUl6b4uttA2wCugTdu4F4HjsjaNkJQXo9ssC/kvDai2A/2Fb/8jsZ5HUtNYZrusON8ZcppQ6CQh4hlQrpR52XXduaWmpbGLnv0uAu6OMWQkcSupspiuESANSgBZCxE1hYeG+xpgXge1Cp97YuHHjCbfddpv0kxNJxYBTlZtznoLrgGGtfNjnGB5Wjln41XY7vXn0yy/XxTJjIluZm5sN7snGcC6q+d6Gmzp34c39h/FN797eSxuUUncqpWZqrb+IbVqRYkYBd3jOXQnc7kOWjiIF6Pa7Bbgm7NgA/we85k8cIdpv0qRJ+ziOcylwMY1vvNV7yxhz15YtWx6RCSEJ4wjgX9ibYs35AVt8/iAuiYQQooNIAVoIEVfXXXfdIY7j/BP7ARal1IvdunU7RWstbQpEUqrMzR2mlHsNhvNpvHFVSzYoxUuuYWEno57rt3x5srcAaNF7w4Z17VGz8TAX51jgNCB7Kx6+5OPdd1/wzv4H7O0qdS6NZ27VAg87jjNVa13VkZlFynKAf9O4WFqLXcb8ii+J2k8K0O1zETDPc+6vwB99yCJEu4wePTpr2223PdUYM0op9Ruaft5fq5R6VCl1u9Y6Vs8Rom32At4GdmhhTC1wAvBSXBIJIUQHkgK0ECLuCgsLf22MeR7oHjr1zy1btpw6Z86cLX7mEqI9VuXm7uUa92oUl2A3PNsaSzG8phzexjjv9F+2bKVK4pYAFTk5fQIBDlKuOdg46lAMh9HybB6vILAIl1uzKyp++ZA1adKkvQOBwNXGmCs8P881xjwLlMjyYdEKewFLaHgNAvgWOJDk7Ae9A9DDc+4LYtcX1AH29JzbDHwVo98XSwcDL9N4A7bVwBDsJmxCJIWioqIcpdRFwGU0tLur5wJvGmPmVVdX3zdr1ixpX5V4tgPeBPpHGTcKuDP2cYQQouNJAVoI4YuCgoJjgYWEPvQppZ7q1q1bntY6bdsSiNTwyZ57dt6yTdcRoC4ATgIy2/Bj1mJ4G8U7yqj/oVRlnet+nFNRkVAbzRhQK3JydlNKDQgok2swB6PUwZhfNhzdWu+h1P3BoHkkp6Lim+YGTZw4cY+MjIwCY8ylQJfGkXg+NCNals6LlpwBPE7j98LvA4djNycUqW9n4F1g17Bzm7GtN+RGlkh4WusdXNe9ANszONK+FF8AdzuO83et9ep4ZhNbpRPwPHBMlHFTgEmxjyOEELEhBWghhG8KCgpGYAsAnUKnHvz0009HzpeN2kSKqMjJ2S6g1DlgLkDxa9r3ulunYLULKx2lqlxjVjmYz3HVVyazbs3XPXf+LhZ9pSuzs7fPCAR61znOjsqY3Ywy+yjFAGAAhv5A13b+io+BB1CBB7KXLl25NQ8cP378jp06dboKuBrY1nP5dWB6SUnJIpJ4NrmIqRuBYs+5l4ERgOxNkNp6Y/us5oadM8AFwEO+JBKiFfLy8gI5OTlHu647CtvSynuTO2iMeUkpNddxnCdlYkdSuAM7s7kljwPnIJuiCiGSmBSghRC+KigoOAN4lFDvXGPMI9tss82F8oZZpJqVQ4fuaoLBE1HmROBYoFcMfs13CtYY+BbUDwAY87NSuC5UK6g1Sm1WUIsxtg97qHDrGnoopQJAN6APmN7ATjTcIOootcAbCp7H5bn+FRVL29tuZNy4cT2zsrKuBK7FZg73vjHmpkAg8JjWWj64iXAKeBj7oT7cq8BwpAVDquoDvAgM8pyfCkyMfxwhoisqKsp1HOcSY8wF2BsoXlXA3+vq6uZNnTr12zjHE203DpgWZcxi7MqMjbGPI4QQsSMFaCGE7/Lz8y9QSs3D9pUEmL9u3boL5s6dW+tnLiFi5aWjjsrY5cfvDgXnJGM4AcxQGm+ul2pWY3jBoJ7L2Lz5hX0+/NC7gVmHGD16dFavXr3OVUoVAf08lz8GZjuOc4dseirCdAdeA4Z6zr8OnEzTzfZEctsDuwnlXp7zC7BtWeQmlUgYWutewWDwHKXUSODXEYb8DCxwHGee1vpFZLVPsjkTmE/D559IvsT2qv8yLomEECKGpAAthEgIBQUF5wH3EZoJDTzTvXv3s6VQJNLBkiFDumUac4BS5iCFOcQYDgJ+5XeuNloHvKeMecso3gm66u2W+jnHwqhRozr17t37fKXUOJrOcvwW+KvjODdrraW4KMBu4vcvmvZQfQ9blPwi7olELOwLPA3s7jm/EMgDZCNk4TuttQMc47ruSOBsGu9zAGEbCm7atOmB8vJy6VmfnPYH/oNdddac9cAR2E1zhRAi6UkBWgiRMAoLC/OMMQ/QsOT/+WAweKbs1i3S0cqhQ3c1bu0BoLLB9Lf/ZQBNd7dvt5/qgmwTcMhQW/W2YBOwCsxKUKvAVAUd3h+0ZEWlSpBZhFprx3XdM4DxwDDP5e+B2TU1NX+eNm3aT/FPJxJML+A54BDP+e+B87AtG0Ty+i1wJ0171kvxWSSEoqKiHKXUxcCF2A0yvT4A7g0Gg/OmTJnyeXzTiQ62C/A2LU80cLE3QBfEJZEQQsSBFKCFEAmlsLDwFGPMY0BW6NTLGzduHHHbbbfJhlBCYDcFVIFAtgnQTxn6AH1A7QimD6jeCrOjsX2Qo77Gf15Tw11rfuCptWu5cdedOa3XL22p1wFfY1gDfKsUXwNrXMy3GOcTMupWZb9f9Wl7ezfHk9b68GAwOE4pdYrnUrVS6u91dXUzpkyZIktc01tP4FngMM/5OmASMIMk+jsvALuqajK2z6rXo9hin7T7Er4YP3789hkZGWe10GJjHfC0tNhIKV2xm90eGGXctcCtMU8jhBBxJAVoIUTCyc/PP0kp9QTQOXTqVcdxhs+YMUM2hBKilT4aNqznhpoap4tS3Ywxmcp1O9c5Thej6joFUJ1cE9h41aefHvrvdevmADhKrXht0KAjDh006Gc1f37Q7/yxUlRUtJ9Sajx2aXP4+6Aa4BHHcSZrrVf5k04kgO7YjQmHR7j2CHAl8GNcE4m22guYR+TC3l+Aa4CUfa4TiUlr3dl13RHGmJFKqRNoutHvLy02AoHAg1prmYCROhzgCeC0KOPuAi6LfRwhhIgvKUALIRJSfn7+CUqpJ2nofffali1bhs+ZM0d6tgrRcRSwnIY+yceSJq0GioqKch3Hud4Ycw4NvecBgkqpR1zXnVZaWrrMr3zCVwoYC0yl6eZQ34auzYt3KNFqCrgcmIm9oRBuM/AnbIFHiLjIy8sLDBw48BjgImxbBe/fS4BVwP2hFhufxjWgiJeZwHVRxvwTewO0LvZxhBAivqQALYRIWPn5+f+nlHqGhjfqi4GTy8vLv/MxlhCp5nJgbuj754CTfcwSd1rrPY0xY4wxl+PZ7MkY80IgELhBa/2GT/GEv04CHgC2jXBtEfBHZIPCRNMP+BtwZIRrnwNnAe/GNZFIW6G+znnAxcCeEYb8pJSar5S6T2v9OtJiI5X9nug3viqxLaDWxj6OEELEnxSghRAJLT8///BQEboHgFLqY+D4srKyj/xNJkTKyAJWA32wH373BZb6GcgPWuudXNe9Erssv5fn8uvA9JKSkkVIgSDd7A08hv134fUTUIS9gSN9hP21DXZmeiENe0iEex7b7/mHeIYS6UdrvYvrunnASGD/CEO2GGP+pZSa5zjO01rrmjhHFPF3JHZmc2YLY37AboL7YVwSCSGED6QALYRIeGPHjj3Add1ngR1Dp74GTiwvL0+7IpkQMXIDoEPfp3XvQa11D9d1f4fdtGxnz+WlxpiZVVVVD8xP4T7ZookMIB+4kcjFzU+x7TruQnoKx1sn4HdACdA7wvWfsYXpO5GbRyJGtNa9gsHgqUCeUupEGrd1glBfZ2B+bW3t/TfddJPcCEkf2cAbRF5JU68GOB54JS6JhBDCJ1KAFkIkhfz8/Gyl1D+A3UOnflJKjSgrK3vdz1xCpIjtgM+AbsAWoC/2Rk/aGj16dFavXr0uVkpNpOF5p97HwGzHce7QWm/2IZ7wxz7wMFLkAAAgAElEQVTYQmak9g4AFdgi9fy4JUpfDradxk3YWeqRPAP8AWmTImJgzJgxXbbZZptjsX2dTyPy7NZK4FHHceZprT+Oa0CRCLbH3njYp4UxBjtb/v64JBJCCB9JAVoIkTTGjh27izHmeWNMbujURmPM2TNnznzO12BCpIa/AleEvp+MbS2Q9rTWma7rXoCdET3Ac/kLY8zMTZs23VleXr7Bh3gi/gLAaKCUyBuJgS043AI8gWwk1dG6Yos112BnFkbyLXA18Gi8Qon0EHo9OAE4HzgVe9PW62vgYcdx7tda/zeuAUUi6QT8Azg6yrgbaViBJoQQKU0K0EKIpHL99ddvW1dXtwi7SQdAnVLq8rKysnt8jCVEKuiPna3lAD9iZ/1KUTVEa+24rjscmAgc7Ln8PfAXx3Fma61/jH864YMdgALgWiK35QBbiJoLzMb+mxJt1wc7m/kq7J99JBuAP2NnRf8cp1wixWmtHeAwY0yeMeY8YKcIw9YCC4H5juM8p7WWG0/pTQH3YmfHt2Q+cC7SHkgIkSakAC2ESDpXXnll965duz6O7ZcG4Cqlri0rK5vjZy4hUsACYETo+yuB233MkrC01se7rns9TWc2rVdK3aGUmqW1/sqPbCLu+mFnQ59L8++rq4F5oa+345QrFQSAY4BLgDzsjMJIarArOKYA38UlmUh5RUVFByqlzsf+294lwpCNSqkFSqmHgOdlM0ERZiJ2JVlL3sQ+v0kbLyFE2pACtBAiKWmtM6urq+8Fzgs7Pbt79+5jtNauX7mESHJHAi+Hvv8YOytaNlVrhtb6kGAwOF4pNYLG76m2KKXuDQaD5ZMnT/7Ar3wirvbHFqJPouX31x8AD4S+PoxDrmS0P3Ah9vXduxFouFrgIezy9U9iH0ukOq31/q7rnoO94bFXhCG1xph/KKUedhznaa11dZwjisR3NvAIdjVZc1YDh2DbBQkhRNqQArQQImlprZ3q6uo52Jma9Z4IBoMXzpo1a5NfuYRIcm/R0GLiDOApH7MkBa31YNd1x2L7gmaEXXKNMc8GAoFbtdYv+BRPxNdAbH/ii7D9ilvyJrZP8XPAyhjnSmQKW3Q+GftvaGCU8T9gW5v8GZCVBqJdioqKcpRSedi/e/0jDHGx/1bnO47zkNZaZtmL5hwAvELLz/3rgV8Dy+KSSAghEogUoIUQSS8/P/8apdTNhGYbGGPeqaurO/XWW2+VmQVCbL3zsLMKAV4DjvAxS1KZNGlSX8dxCoHfAZ09l990HKcceEpWaaSFntjWEfnAbq0Y/wnwL+AF7MZV62KWLDFsB/wGOBYYDuzaisd8iC06/w3pTy/aoaioaL9Q0fkcYO9mhr1rjHk4EAg8qrX+Io7xRHLaE3sDv3cLY4LA6cCieAQSQohEIwVoIURKKCwsPMsYcx/QJXTqE8dxhs+YMaPSz1xCJKEAtk1A39DxodgPVaKVtNZ9jDGjjTF/wBbawn1ojJkVCATu0Vpv9COfiKtMbIH1wtB/m9uwMNwW4I3Q1zvYvtHJfkN1D+yS84OxmwgfgH2uiWYd8ARwH/ASslmXaKOwonMetnd7JO8C813XfWzy5MnS1kW01jbA60BulHGjsTfRhBAiLUkBWgiRMq677rpDHMd5moYdyn9yHOesGTNmvORnLiGS0HXAzND3j2I3YRJbqaCgoFvXrl1/D4yhoaBf73vgdsdx/ixLutPGttj+oBdgVxa01CPU61PsjaC3gaXAKuDzjg7YAQLYmYD9sW01DsIWnVuaFehVAzyP7ZO9EJCWWqJNioqKhjmOc6YxJg/Yp5lh7xlj5htj5kvRWbRBBvAMDRujN2cOcHXs4wghROKSArQQIqWMGTNmr0Ag8AyQHTpVo5S6vKysbJ6fuYRIMttgi1s9sUtGBwAf+ZooieXl5QWys7PPUkoVAAd6Lm9WSs0LBoM3T548OZ37AKeb3YEzsZsW/h9NW7a0xgZsIXpl6GsV8AWwBjtj+scOSdqYAnbE3ujdCfu/oz/2OWIAdmZpa2Z5e63Dth95DngS2+dZiK0Seq49LFR0PgM78z4SKTqLjvIXGu9FE8nzwAigLvZxhBAicUkBWgiRcsaPH799bW3tkzT0rjVKqdKysjKNLN8VorXKgILQ97cC1/qYJWVorQ93XfdqbPExvAWBMca8qJSaXVJSsgh5rkonXYGjscXoE2m+J+3WqgG+wxajvwE2hs5twN5Yqu8z/RP272KP0PG2of9ug53dtw2wC7bwvCONN9psjyXYwsxz2HYjtR30c0UaycvLC+Tk5BxqjMkLzXTeuZmhK7AbCT6otV4Vx4gideUD5VHGVGA3Hfw59nGEECKxSQFaCJGSRo8enZWVlfU3bN9NAIwxi2pqai6YM2dOqm/uJERH2BW7MVon7K7tuwNrfU2UQrTW2a7rXgdcRNPZryuNMbMDgcA8rXW1D/GEv/bG9l4/GNvCYl9sL+lktgFYjO1p/RbwJvCVr4lE0tJad3Vd9yTgDOAU7GodLwO8bYx5MhAIPKa1/jiuIUWqOxlYQMu97L/G9r7/LC6JhBAiwUkBWgiRylRBQUEpMIGG57ulwOnl5eWy5FKI6B4Afhv6fhwww8csKWnChAm9MzIyrgKuoKF/fb2flVJ3KaX+IsWTtJYF7IctSO+HbTE1AOjlZ6gWfAdUYduCvIftW12BLD8X7XD99ddvm5mZeYox5kyl1Ak0bDodrg54BXgyGAw+NWXKlC/jm1Kkif2A/wDdWxizCbuy5e24JBJCiCQgBWghRMorKCg4B7gbu8wZ4AfXdc+9+eabX/QxlhDJYH/srEWAL4G9sEv4RQfTWme6rnsadgPIQzyXXWPMv6U9h/DYiYZidH/szOneofM7A91i9Hs3YHtNf4ed4fcRtthche1D/VOMfq9IM5MmTeqrlDoOGKGUOp7IKwE2G2NeU0otqqure3jq1KnfxjmmSC+7YFdx7NbCGBe74eyTcUkkhBBJQgrQQoi0cN111w1VSj2llNozdKoOmFReXj7dv1RCJIWXgKNC318E3O9flPRQVFQ0TCl1DXA+TfvtrgJu27hx49/Ky8s3xD+dSCJdgT40FKUzsUXpTOys6q409HgG22LHANXYfsybgM2h/36HbXdwdehxJ2N7NwvRYbTWTjAY3E8pNQLbWmNYM0M3GmP+DcwPBAJPaa2ltZqIh67AyzTdTNhrLHYfDSGEEGGkAC2ESBv5+fk7KKXm01BMA7ize/fuf9Jay6xOISI7BVgY+n4pth+tzMCNA6317saYq4wxlwHbeS7/CNzrOM5crXWVD/FE+vkd8PfQ938B/uRjFpEitNbdgeONMSOMMcOxG11G8i2wCHjyp59+emHOnDlb4hZSCHCAJ4DTooz7O3Bp7OMIIUTykQK0ECKtaK0zqqurZ9H4g/PrSqmzy8rKvvErlxAJTAHLgUGh42MBaV8TR1rrrsFg8EKl1NVAjueyMca8Atyxdu3aJ6UoI2JoJ2zLDQe7qdaeyM0o0QYTJ07cLRAInGKMOVUpdTR2Rn4ky4BFjuMsAN7RWrvxSylEI7OAa6OMeQU4HmlVJoQQEUkBWgiRlvLz80cppf4MdAqd+tJ13XNuvvnmN/zMJUSCuhyYG/r+OezyexF/qri4+BhgNHZmesBzfY1S6m6l1J1a6w/jH0+kgTdp6FE+FLsqQogWaa0zgsHgr5VSJwEnAUOaGVobuqG20BizcPLkybJhtEgElwF3RhlTBRyG9MAXQohmSQFaCJG2CgsLjzDGPIad1QXSF1qI5mQBq7H9ZA2QC1T4GSjdTZgwYeeMjIyRwB+BPTyXjTHmRaXU3G+++eapuXPn1voQUaSmSUBp6PuJwFQfs4gENn78+B07dep0FDAi9NWrmaE/AS8YYxYFAoEFWuu18cooRCucgG394t2PIdwP2BtzcuNXCCFaIAVoIURaKygo6As8Buxff84Y88imTZsuu+2226r9SyZEwrkB0KHv78LOCBI+y8vLC+Tk5AwPBoNXKKVOxLZHCPc18PdgMHjnlClTPvUhokgt+wL/C33/BvBrH7OIBJKXlxfIzs4+xHGck40xJ2H/rjT3WbMKeMYYszAQCLyuta6LX1IhWm0Q8DrN3zwBu1Hrb7DPh0IIIVogBWghRNobPXp0VlZW1gzg6rDTK4Gzy8vLl/sUS4hEsyPwKdAF2AL0xRY3RYLQWu/puu5lwO+BnT2XXWPMy8C9gUDgCa213GATbbUaO+veBXbBbg4n0lBoJcbxSqkTjTHH03Sz1HobgZeMMc8aY56T1hoiCewAvAXs3cIYA4wE7o9LIiGESHJSgBZCiJCCgoILgTuArqFT1caYUTNnznzIx1hCJJK/AleEvp8MFPmYRTQjNCv6aNd1RwFn0rRX9CbskuL7Kisrn50/f34w7iFFMrsN2/oF4BLgXv+iiHgaM2ZMl549e/7add1jsRvS7k/znyc/Vkq9YIx5wXGc5+Sml0giXYF/AwdHGXcDUBL7OEIIkRqkAC2EEGHGjh070HXdx4GB9eeUUnO7des2Wmstu1qLdNcfqMS2efgR2B3Y4Gsi0aJJkybt7TjO5cBF2NmqXp8B9zmOM09rvSq+6USSGo69gQEwHzjHxywihrTWDrBvMBg8HjhOKfVr7J4AkWw2xvxHKfWs67rPTp48+YP4JRWiwzjY57Uzo4x7BDgfOwtaCCFEK0gBWgghPMaNG9fTdd17jDGnh51+Gzi/vLxclo2KdLcAu6EUwJXA7T5mEa0UmhV9nOu6I4HTsa1UvN40xsyrra19ZNq0aT/FOaJIHl2A77GzBNdh2/PIDdoUMXHixN0cxznOcZzjjDG/wf7/25wq4F+O4/wT+LfWemN8UgoRMzOB66KMeQ27AmBL7OMIIUTqkAK0EEJEpvLz869WSpUBnULn1hljrpw5c+YDfgYTwmdHAi+Hvv8AyMb2ghVJYty4cT0zMzPzlFIXYzeR874f3AIscBznQeB5rfXmuIcUiS78RtSxwIs+ZhHtoLXeyXXdo4H6r/4tDP8eeNEY80/Xdf81ZcqUz+MSUoj4uByYG2XMJ9jWHGtiH0cIIVKLFKCFEKIFhYWFRxpjHqTx0vW7gGvKy8ul9YBIV28DB4W+PwN4yscsoh0mTpy4WyAQ+C1wGdAvwpBNxpgXgfmyeaEIMwq7ZwLALcAYH7OIrTB27NhtunbtenAr+zjXAUuAFxzHWQS8obWWG44iFQ0HnqbpngnhfgQOw25ULoQQYitJAVoIIaLIz8/fAbhbKXVK2OmVSqnzysrK3vcrlxA+Oh94MPT9a8ARPmYRHUNprY8wxlxsjDkb6BFhzHpgkeM4j/3888/PzZo1a1OcM4rEsQvwBfazxMfA3v7GEc3RWvcA/s8Yc7Qx5mhgKLbPbSQGWA68gG2t8Yq01RBpYD/gP0D3FsbUAichqz2EEP/P3p2HR1Xdfxx/nzuTEDZ3xLWiAkmYJGpxo9alrfvSqpVad6wtbW2t0ATB1vobWzdC0Lp0o4v7irtWbdVqtYoLiCQZEpYqbqi4gYQlkLnn98cZSjJZJoHM3JnJ5/U8PNzlzOQjTpKZ7z33e2STqQAtItI9G1pyTGXjAjzN1trJ06dPvx4tQiJ9SxhYDOyW2B8DvBxcHOlN0Wh0gO/7J1hrTzHGHIvr9ZusyRjzmDFmporRfdYc3OxZcAv3NgaYRRISLTUOMsYcYq09CFdcC3fxkAZjzLPW2ufWr1//3FVXXaXWAtKX7IR7/7JrF2MsMA64NROBRETylQrQIiI9MHny5Eg8Hr8bKGt1+J/r168/+7rrrvsoqFwiAfg5brEegHuBUwPMImlSVVU1sH///scZY07B3aLcYTEa+Dsw0/O8f6hNR59xGXBpYnsSUBNglj7rkksu2dPzvK/i7kQ5CNeXvyv/NcY86/v+s/F4/Nkrr7zyg/SnFMlKg4EXcHcFdOVS4DfpjyMikt9UgBYR6aHzzz9/UP/+/W8wxoxrdfh94Oyampp/BRRLJNMGA+8CWwJx3MJVbwaaSNIqMTP6OGAscCwwsINhzcDz1trHQ6HQ49FodGFGQ0om7Qe8mtj+N3BYcFH6hrFjx4ZKSkrKjDGH4IrNB9N2jYqOvI1bOPZZz/OejUaj76Q5pkguCOHWrzg+xbi7gDPQnY4iIptNBWgRkU1UWVl5ijFmBrB14pA1xvzZWvtzLVAofUQNUJnY1kJkfUg0Gi3yff8IXDH6RNwFiY68aYx52lr7mOd5T0Wj0bWZSylpZnB9oHfCXYQaCnwaaKI8M3ny5C379++/n+/7X7XWjjbGHMTG9xydeRN40Vr7n1Ao9HQ0GtWFQZH2fgecn2LM88CRuAurIiKymVSAFhHZDJMmTdrT9/07jTH7tzrcYK09a/r06XMCCyaSGbvgih0FuAXqvgQsDzSRZFxiZvQxxpgTrbVHA9t1MnSVtfYZY8zj8Xj88SuuuOLdTOaUtPgLcF5i+3TcbEHZBNFoNByPx/cyxhxojDnQWnsgMDzFw9YDs3GLwb7ged6L0Wj0s7SHFcltk4GrU4xpwN1l8Hn644iI9A0qQIuIbKZoNBpuamqqBH4NFCYOtwDTv/jii1/NmDFjfXDpRNLuDlzhCdyHuuoAs0jAotGoB+xvrT3OWnssbgG0zt5v1gJPAP/wPG+WZkfnpBOBBxPbdwBnBpglp1xyySV7GmNGG2P2BQ4ERtNxj/XWVgAvW2tfCoVCzwOvRqPR1enOKpJHxgJ3A14XYz7BLa68OCOJRET6CBWgRUR6SWVl5X7GmFtptQCQtfbVUCh0dnV19YIAo4mk05eBDbP93wf2ANYFF0eyycUXXzykoKDgMOCExJ+tOhnaAswDnvY872ng+Wg0qtdR9huIK9YUAZ/h2nC0BJooC0Wj0Z183x8NjE600jgAGNKNh74JvAjMSbTUmBuNRv20hhXJX/vh+qF3daFnDfB14OVMBBIR6UtUgBYR6UXRaLSoqakpCkxi4+yKNcBlgwYNmqYPjpKnnmXjAmRnAbcHF0Wy1fjx4wt22mmng33fPxY4jlYX6zqw0lr7b2PM84nC2xwVpLPWE8DRie1DcX1T+yoTjUZ3B/a21o621o4G9gW27cZjP7HWvgy8YoyZtXbt2lerq6tXpjWtSN+xO66ovH0XY3zcDOkHMpJIRKSPUQFaRCQNqqqqDgf+Buy64Zgx5ql4PH7eNddco76nkm+OBx5NbNcCe6MV4yWFaDS6RzweP9oY83XcBYyuinRrgFeNMS8YY14EXopGo19kIqek9FPghsR2Na4VT96rqqoa2L9//zJgL8/z9rLWVgAVwBbdePhKYK4xZrbv+3Osta9dfvnli9IaWKTv2gZ4CShOMe7nwLXpjyMi0jepAC0ikiaTJ0/e0vf9amvt+FaHVwO/1mxoyTMGiAGlif3DgWeCiyO5JhqNevF4fC/ga4mC9CHA4C4eEgfmA68CryTaHcWi0ajaP2Tel4C3E9vzgUiAWXpdoq/5MN/3I7gC816JP8Ppuo/sBk3AGxuKzaFQaDawUO8BRDKiEHeXxtdTjJsB/DD9cURE+i4VoEVE0qyysvIUY8wfgO1aHX4uHo//4Nprr9UCJ5IvxgN/Smw/ARwbYBbJcdFoNByPx0cbY74KHAwcRNufoR1ZBcwxxryamFU6NxwOL1KhLyPqgLLE9ghycPGuaDQaBvbwfT9irS01xozCXVQrBfp382k+B94Aaq21rwNzGhsbG2fOnBlPT2oR6YIBbsG1BuvKE8A3Uf96EZG0UgFaRCQDqqqqtgdqaPsmeA1w2dtvv12jD6eSB/oBS4AdcO03ynGzokV6g7nkkktKPM/bUJAeg5uBmkoTri3MXGvtG8Dc5cuX199www3NaczaF10JXJzYngBcF2CWLk2ZMmXroqKiEfF4fLgxZqQxptRaW4q7Pb+wm0/j44rs84B5nufNA2qj0eg7aYotIj13OfDLFGNex/Wub0p/HBGRvk0FaBGRDJo0adLx1to/Aju3OjzX9/3zrrnmmrlB5RLpJf8HRBPbfwW+H1wUyXfRaHQ7YH/f9/cH9gcOwPX6TCUOvGmtjRljGqy19UBDKBRqiEaja9MYOZ8dBPwnsf0UcGSAWYhGo1vE4/ERxpjh1toRnueNsNaOxF20SDWTPtmH1tr5iddKLTBvzZo19TU1Nat6P7mI9JIf4NpqdGUp7vfGe+mPIyIiKkCLiGTYhAkTtiooKJhqrf0BG38OrweuGTRo0KXRaHRdgPFENscQXC/Y/kAzMAz4MMhA0rdccsklI4wx+xlj9rbW7mOM2YeuFzdsLQ4ssdYuMsYsAhZ5nreopaVlUTgcflv9pbvkAR8A2+N+nw0BVqTriyUKzLsZY4ZZa3f3PG833/eHGWN2w/3c6e7/89betdY2eJ4331o731o7P3FR4rPeTS8iaXYc8BAQ7mLMStzdNPMykkhERFSAFhEJSmVl5THGmD/iFnDaYJ7neeOrq6tfDSqXyGb6E64fNMBvgEsDzCLCL3/5y10LCgr2icfjextj9sb1Kt4DCPXgadYDb1lr3/Q87x1r7TvAO9baJb7vv1NQUPC+CtTcysY2U6cA92/Kk0yZMmXrgoKCnYwxu1hrdzTG7ArsCOyS+LMb3Zvp3pEvgA0XGBb7vr8oFAo1Ao3RaPSLTXxOEcke+wLPAQO7GNOCK1L/MxOBRETEUQFaRCRA0Wh0QFNT06XAJNwMMnD9c28vKCiYeNVVV30aXDqRTTISaMC9nj/DXWDRreqSVS644IJ+2267banv+6XGmLJED+AyYHe6njXXmTjudu73gGXAe9baZcDSUCj0YTwe/yAUCn3w6aeffprH/ae/A9yT2L4ZOBcgGo16wHbxeHwIsF0oFBri+/5QYDtjzHbA9tbaHXD943el+wv+deZzXD/6xYk/izzPWwQsjEajyzbzuUUke+0BvAQMTTHufOAP6Y8jIiKtqQAtIpIFqqqqDgP+TNtFtT4Afl5TU3N3IKFENt0jwAmJbX3Qk5wRjUYLgWG+74/Y0DvY9/3hxpgRuIspPZk13Zkm3MWZz4BPgU+Az4wxn1prm4Dl1tpVoVCoKTF2eUtLyyprbVM8Hm8qKiqy0Wh0eS/k6FI0Gh2wdu3afuFweKtwOFwQj8cHAwNCoVA/YKt4PF4EbAVsZYzZKh6Pb7dw4cKz+vXr5w0cOLBl6NChbwNb4tph9OZnjs9wBea3cS1TloRCoSXAkjVr1rw9derUtLX+EJGsNQRXfE61OO2VpF6YUERE0kAFaBGRLBGNRouampqmAFOAfq1OPWet/fH06dMbA4om0lOH4m6BBVgElAB+YGlEekE0Gi1saWnZ1fO8L+GK0cOMMV/yff9LxpgNx4oyHGs1rt96HNdeoiOfd3BsEFCQ2A4BW7Q6NxAo7K2APdACfAS8i7sA+x7wgbX2fWPM+77vLw2Hw+9Go9GmALKJSPbqDzwDjEkx7i7gDNydhiIikmEqQIuIZJmJEycOD4fDv7fWHtHq8FpganNz81V5fPu25JdXgP0T2ycCD3cw5jjg7xlLJJJm0Wh0G2AH3/d3tNbuCOxgjNnJGDPU9/2djDHb4PoXb8vmt5rIatZa3xjzGW6G98fAJ8aYj6y1H1trPzHGfOx53kfxePzjeDz+SWFh4UfRaFQXqkSkJ0LAfbj3GV15FjgGd8FOREQCoAK0iEiWmjRp0lhr7Y3A9q0OL7bWnj99+vSngsol0k2nAXcmtl8ADkk6vx/wY+B7mQwlki0mTpzYf8CAAdsUFBRsiytKb+P7/pbW2oHAIGPMVsaYwRv2rbVbGmOK2Fi43jrxd3/czGuDa4nREytxM4+h7SzqlsS51dbaZmPMCmCdMeZ/x4AVifHLcW1Dls+ZM2fwCy+88ERzczPr1q2bB+zdwzwiIj1xI/CTFGPqcO9B0t66SEREOqcCtIhIFps4ceI24XB4qrX2PDb+zLbGmDuNMRdVV1cvDTKfSBfCuAXAdkvsjwFebnX+90AF8NUM5xKR9GoEihPbw3C9mkVEetuvgF+nGPMe7v3He+mPIyIiXVEBWkQkB0yaNOkg4A/W2vJWh1cD0wYNGnR1NBpdG1A0ka78HJie2L4XODWxXYTr9boe2C6AXCKSPjVAZWJbi5CKSDqcDtxO1/WMFcDBuBnQIiISMBWgRURyxPjx4wu23HLLidbaX+EWkNpgkTFmwrRp0x4PKptIJwbjFhTbEnd7/0jgTeC7uMWAwLUMWBFIOhFJh8Nw/VYBHsf1ehcR6S1fB56g68VS1+F6Pv8rI4lERCQlFaBFRHLMhAkTdgyHw1OBM2n7c/xpa+2F06dPnx9QNOnbCnEf+JK1ng35W2Ai8E9gwyKb+wGz055ORDIlDCzD9ahei7vLYVWgiUQkX5Tj1pXYsosxPu5C98yMJBIRkW5RAVpEJEdVVlbuZ4y5ATig1eH1wB+am5t/dcMNN3zRyUNF0mEfIIq7JfZhNhajd8HNei7ALWp2IO52WC9x/nQ2zoYWkfxwF64ABPAt4JEAs4hIftgFmJX4uysTcRe8RUQki4SCDiAiIptm1qxZS3feeeebttxyyw+MMWOAAbif6weEw+Gzx4wZ89GsWbPqA44pfceHuFlH9+D6vu4ALMUtRFiCm7XUDxgOjGj1uFrg3xlNKiLp1h84ObG9EngswCwikvu2AZ4D9kgxbjqpFyYUEZEAaAa0iEgemDJlytYtLS1RXOEv3OrUbGNM1bRp01Tgk0xJXpX+deApYHJivwU3+9lLbN8NnJXJgCKSdtvgFhoNAx8AOwM20EQikquKcO8jvppi3L3AabiL4SIikmVUgBYRySOVlZUlxpjrgCOTTj0dCoUmTJ06NRZELulTDHAbcEZi38cVmy3t33dYXP/n/TOWTkQy5Xng4MT2aNzFKBGRnvBwd1adkmLc88BRuL7zIiKShVSAFjCHZzsAACAASURBVBHJP2bSpEmnAFdba1vfqrjOGPOHcDj8m6uuuurToMJJn1AEPIsrLHspxn5B14sJiUhumgxcndiOApcFF0VEctT1wAUpxsRwF7s+T38cERHZVCpAi4jkqfHjxxcMHjz4XGPM5cCQVqc+B6YOGjToumg0qpkiki7bA6/hbr1PtebEEOCTtCcSkUyKABvWIXgN3ekgIj1zCfCbFGOWAl8B3k5/HBER2RwqQIuI5LmJEyduEwqFLgIm4BaB2+AdY8yvpk2bdhvqzSnpUQq8Agyk65nQY4CXM5JIRDJpMbAn7nfMLrhikYhIKuOBP6UY8wVwKPBG+uOIiMjmUgFaRKSPmDRp0ghr7dXAya2PG2NeNsb8orq6+tmAokl+Owr4O64A3dn7jnOAWzOWSEQypfXt898H/hpgFhHJDd8E7qftotrJ1gPH4RYnFBGRHKACtIhIH1NVVXUAMB04KOnU057nXVxdXT07gFiS3y7AFaI6YoErgF9lLo6IZMiRwD8S2w8BJwWYRUSy32HAE7i1JDpjgXOBWzIRSEREeocK0CIifZOpqqo6Dbgc2L3VcQvc53ner6qrqxcEE03y1I3ATzo4HgfuA76b2TgikgGFuP7ug4FVwHaA1h4QkY7sAzwHbJFi3GSgOu1pRESkV6kALSLSh7VaqDAK7NjqlI+7/XFyTU3NW4GEk3wTAh4GjqX9+483cB88RST/3M/G1k/HAE8GmEVEstOewH+AHVKMu5GNbX1ERCSHqAAtIiJUVVUNBH4KTAG2anVqnTHm5nXr1l163XXXfRRMOskjg4FZuMUJWy9KuAa3UKEWwxTJP+cCf0ts/w73u0ZEZIPtgReAkSnG3QGcjZskISIiOUYFaBER+Z+LL7542/Xr10/GFQj6tzr1BXBNKBT67dSpU1cEk07yxDBgNrANbd+H7Ah8GEQgEUmr7YEPcBed3gF2CzaOiGSRLXBtN1LdBfV34ESgJd2BREQkPVSAFhGRdiZOnLhzOBy+1Fr7PdquQr4S+H1LS8vVv/3tb5cHFE9y30G4D5ytX1tfSxwTkfzzMnBAYrsCqAswi4hkh/64ljyHpBj3CvANXB95ERHJUaGgA4iISPZ5+eWXV7700kuPHXjggfcYY4biWiYYoB/wVc/zxo8ZM6Zw3333fePVV19tDjat5KB3gbfY2BcW3AfM2cHEEZE02xF3kQnc9/9/AswiIsELAXfi+sJ3JQYcjrsTT0REcphmQIuISEqTJ0+OxOPxycAZtO3d+xlwQygUulatOWQT3Aj8JLE9FdeDvFOLhg/vZ4qKtrchf7sWP+R5Jj7AWNvPYgosDAKwmK2NtcsxWINdZWAdsC5OeFXYi/sm7n0Samr6ePclS9am9z9NRFrZG5ib2H4JdxeEiPRNBpgBfD/FuPdwPyveSXsiERFJOxWgRUSk26qqqsqAS4FTaPs75FPgRhWipYcMcA8wNgQP1+416pde3Cv2jB0J7GwxQ8HuAAwBdqDtApmbawWu5/Qy4CMDH/rWLsUzC7HewvCaNQtHLF6s2f0ivWcJrv+zj5sRvSzQNCISlKtIccEZ+AQ4GGhMfxwREckEFaBFRKTHfv7zn+/jed7/Ad+k7e+ST4Brmpubf3fDDTfodklpx44dG2psbBzlGf8Aa4ms8v1Rp/33zUMtFD46Yng2vS/xgbeBBVgWGEMMn1dHjhpVb2bOjAcdTiQH/QH4UWL7HODWALOISDB+grv7qSurcW03ZqU/joiIZEo2fdATEZEcM3HixPJQKPQr2s+IXgnclFis8INg0kk2iEUiO3gh9gMz2sOOtpaDgK1bj/mkpYUz33yLx0eOaNPfJUutwvAG2DkGM8cSeqGktvatoEOJ5IDjgUcT2/cCpwaYRUQy73TgNujyV/063OSGf2QkkYiIZIwK0CIistmqqqoqgEtoX4huNsbcEo/HL7/mmmveDSadZNJbw4YVrR/c/6s+3uG4GUyju/O4xrVrGRouYOtwTq6P/KaBpy32adbFnyxZsGBl0IFEslB/3F0yA3ALig3BFZtEJP8dDvwdKOxijA+chrtAJSIieUYFaBER6TWJQnQVbpZL60riOuAez/OuqK6uXhBIOEmbRWVle8bhOIw9BjgUV2jqLS3AMgzLsHxgLR8bzy7Dmrg15gtjbRxr1lhj1xprWjD+SmCQxRRgbT9jzABrTMhYuwXGhow1QyxmiMXuaGAorghW0It51wLPG3gyHvIfG/VGw6JefG6RXPcIcEJi+3DgmQCziEhm7I/7Xh+UYtwE4Lr0xxERkSCoAC0iIr1u0qRJe1prfwb8EOjX6pRvrX0ciE6fPn1OMOmkNyyuqNh+vd9yKsacYeCAzXy6j4FGDAvBLMD6C+Meiwu9fstGzJ37cW/k7cqC0SO389YWbB8PMdz6ZiTGjgRTbLAlwPab+fSzMdwRj3N3JBb7sDfyiuSwHwJ/TGxfC/w8wCwikn4VwLPANinGXQZE055GREQCowK0iIikTWVl5W7AJGPMeUBRq1MWeMwYM23atGkvBJNOempeRcXAQhv/lsGcAfZIILwJT/OWgZet4VXrm1fXG9NQUVf3eW9n7S1v7b33Vmvj8VJj7X4YewCu2L7nJjxVHMvTBnNHi7UPRmKxpl6OKpILdgLew30G+S8wPNg4IpJGI4DngR1SjPsj8OP0xxERkSCpAC0iIml38cUXD1m/fv1PgAuBrZJOv26MuW7gwIF3RqPRlgDiSQoNe5cMIx66wMD3gS168NC4wbxmrX3GerxSQOiV4bW1y9KVM1MW7bPPkJb16w8wxu6P4etYDqRty5lUVmLsTSHfu35Eff1/05VTJEu9DuyT2C4B1JZJJP/sCrwA7JZi3EO49UPiaU8kIiKBUgFaREQyZsqUKVu3tLT8FPgZsF3S6f8aY64dOHDgTdFodHUA8SRJQ3n5aGP8C7GcRvdnO38M5jmLfdr3eaQvtJ2IRSKDvBBf8yzHWzgG98G7O3xjeNxY/7qRdQ1PpzOjSBa5DLg0sV0FTA8wi4j0vqHAv4HiFOOexf3ObE57IhERCZwK0CIiknEXXHBBv6KiolOttb+g/QeUFcAtnudNra6uXhpAvD7NgmksKzvJGDsF2K+bD3sLuCPk2QeGz5v/hnEtVvokC2ZhJFLhh8xJnrVn2G63GDCvg391cd38+/ryv5/0CfsDryS2nwO+FlwUEellQ3Df16NSjJuD+95fme5AIiKSHVSAFhGRwESjUa+pqek4YArwlaTT64B7rLVXT58+fX7m0/U9DeXlYwz+VODgbgz/HMNjnvVvHVHX8IyKph2L7TUqEvLNWQbGWTcrLJXZxtgpxbXzn0l7OJFgeMD7uL6wLbiFPpP7wB+Mu31fRHLHlsAzwOgU4xYChwAfpT2RiIhkDRWgRUQkK1RWVh7ieV6ltfZ4XIFiAx/4O3B9TU3NM6jQ2esaysvLDfZqsMemGGqBpzD8Lh7nyUgsti4T+fLB7NGjCwY2Nx9ljD0fOJrU78H+6Rv/4lG1Da9nIJ5Ipv0V+F5i+zTg7lbnPFzx+aBMhxKRTTYAeJLUF7D/iys+6w43EZE+RgVoERHJKpMmTdrTWvsz4AdA/6TTC621vzfG/KWmpmZVAPHySiwS2Sbk2WlgxtG26J9sLZbb4yH728i8+bEMxctbC8rLSyx2AtizcB/aO2PB3hEK9/v5iLlzP85UPpEMOAl4ILF9B3Bmq3Nn4QrUhZkOJSKbpBB4GHdxtSvv4YrPb6U9kYiIZB0VoEVEJCtdeOGFQwsKCi4AfgRsm3T6M+DPoVDo91OnTn0n8+lyX2P5qLEGc0OKthDLgBtC4cI/qQDa+xpKSrYl7I03xvwM146gY5ZPMHZCSd38OzKXTiStBgKfAEW4n+dDce04+gGLgJ2BUGDpRKS7CoD7gRNSjPsYOBRoSHsiERHJSipAi4hIVmu1YGElUJF02rfWPm6Mua6mpubpIPLlmlgkskPIcCOGb3cxbBVwY0Fh0VV7zpmzIlPZ+qrZo0cPGLRu7QW4XuhbdTH0ibjPjyKxmC66SC45BogBya/bJ4GjEtuH4NpuTASuSRzzUMslkWwWAm4Hvpti3HLg68DctCcSEZGspQK0iIjkjIsuuuhr1tqfWWu/SfuWEa9ba68fPHjwPdFodG0Q+bJdY/moM8DcSOdFznUGZoRM6DfDa2uXZTKbwILRI7ez6wp+CfwYNxO0I19YuLC0LnZz5pKJbJYhuMLT3cCVuBnPABcA1ye2pwJXAUvY+POpADcrWkSyjwH+hGuX1pUvgCOAV9OeSEREspoK0CIiknOqqqp2B36I++CzTdLp5caYe40xv62urtatnrgF8AatX1ODNT/rdJDlUc94E0bW1b2ZwWjSgfmlpbuFCrxrrOXkTgcZ+8d43FyohSAlR5wN3AKsBK7AFZ63xxWcAebjeshOYePnk36AXt8i2akGqEwxZg1wLPBc2tOIiEjWUwFaRERyVjQaLVq1atV3fN+/yBgTSTrtA/8yxswYOHDgg9FotE/OpFu0zz5D4i3r7sbd/tqRzy1MKa2LzchkLkmtoSJygrH8AdcPtyNz/Bb/26MaGt7OZC6RTfRP3ExIgA+B/wN+Bmz42d1M25n//QHdzSKSfa4ELk4xZh1wMvD39McREZFcoAK0iIjkA1NZWXmkMeYnuNk2yYtXvQPMMMb8ddq0aR9mPl4w5leUHuhZ7z46LWDa201hy8TiOQs/yWgw6bba8vKt++FfY+EcOnjfZuAjg/+dkXUNzwcQT6Qn9sD1gi7CXSD0gE/ZuMispe1rfDDQlMmAIpLSL4HLU4yJA6cD96Y/joiI5AoVoEVEJK9cdNFFO/m+fxbwU2CXpNPrcLd5z6ipqXmGPF7gqqGs7Ehj7IPAgA5OL8dwTklt7JFM55JNs6B81DEWcxsbi3WtNVtrTimtr38s07lEeqgKmNZqf0MhuiNbAVoEVSR7/BS4IcUYi2uP9tf0xxERkVyiArSIiOSlaDRa2NTUdDJuQbdDOhgy31o7o7Cw8Parrrrq0wzHS6tEsfIB3EzDZI0G76TiurrGTOeSzbO4rGzXFmPvB/br4PQ6azi9tDZ2f6ZzifRAGHgNKKf9nSrJtmXjgoUiEqxzgL/R+QUjcMXn84E/ZiSRiIjkFBWgRUQk71100UXFvu+fC4wHtk463Qw8Atz29ttvPz5z5sx4xgP2okTf4Jm07aUKgLXcvc4LfX+v2tpVAUSTXvDWsGFFzYMH/A7M9zo4HTfWnFtcX39bxoOJdN8+uCJ0qgL09sDH6Y8jIimcDtxK6u/ZKmB6+uOIiEguUgFaRET6jPPPP39Q//79TzfG/BjYu4Mh7wF3WGv/MH369Jxb2K2hrOxkY+w9uFmGrVkDvyiui10dRK5U5leUftmz3ne6Oz7uc3kkFtuk3rCNZWXnYeyIbg22ZmFJff3fNuXrpNuC8shE6z7oJ7+XixvDmcW1sbuDyCXSTdXApBRjdgA+ykAWEencKcBdtH9fkSwKXJb2NCIikrNUgBYRkT6psrJytOd54621p+EWu2rNB/5ljJmxYsWKh2bMmLE+gIg9sjAS2dv3+A8wMOmUxZoJJfX11weRqzsWVETOtpZbujveGsaV1sa6PX6DRcOHbxHv3+8DOu6L3ZEnS+pix/T062TKgorI963lT7S/JXqtZ+xhI2vnvxJELpFu6A/MB3al81mVOwEfZCyRiCT7NnA3qYvP1wET0h9HRERyWVc9nERERPLW9OnT50ybNu2HnuftbK39AdC6WOcBh1tr791iiy2WVFZWXjFx4sThAUVNqaGkZFvf4wE6KD4buCCbi8+bwljGbcrj/P79TqX7xeesV1wb+4s1nAm0JJ0q8q15aFFFRfIinCLZYg3wPbr+LJLqdn8RSZ+T6N7M55uAiemPIyIiuU4zoEVERBIqKytLjDHjcIWRIR0MmWOtvS2bFi6MRSKFIY+ngYOTTvnGcG5xbezWIHL1RE9nQAMWE9qzpLb2rZ58ncbyyIvAV3rwkKyeAb3BgrKyU62xd5BUsLPwStHKVYftvmTJ2oCiiaRyC3AWHX8mGQbkXCskkTxwDPAgHawlkeR23OKEftoTiYhIzlMBWkREJMkFF1zQr6io6JvW2vHAN2j/+3It8Chw26BBg56IRqPJM1AzZkF55AYLP+3g1KUldbHfZDzQJtiEAjTG2mhx/fxu95ts2KtkpPFDjfTsvU9OFKABGivKJmFtdbsThr+W1Ma+H0Akke7YFliAWxw2eTb0nsCbGU8k0rcdD9wPFKYYNxO3OGFg739ERCS3qAAtIiLShUmTJu0JjLPWng18qYMh71lrbwdumT59emMmsy2oGPUNa81TJP0+N4YHRtbGTjFgM5lnU3VUgL7uo2U8v9KtMzi0IMzvd2v3T/9mcV1seHf/GxdUlF1prb249bG7Pv2M+z5fDrjK18zheyQ/LGcK0ACN5aP+CuZ7yccN/knFdQ0PBZFJpBtOB+7o4PhIYFGGs4j0ZUcBDwFFKcY9AHwXyPr1MUREJHuoAC0iItIN0WjUa2pqOgwYB5xM+37LGGNe9n3/loKCgnuuvvrqz9OZJxaJDAp5xGhfFJ/XbEIH7VVbuyqdX783dVSAvujd93lk+XKGFhRQXNSPqbvswtbhti1hLd5hpXV1/071/Hbs2NCCxvlvAzu3Pn7Nhx/xry9W8ta6dVhrmV8eSX5oThWg3xo2rKh58MDngf2STi0NrWkuHbF48RdB5BLphkeB42j72eQQYAVudnQBrijWH9dqZovEmP64ftIWWJ449gUQTxz/HPgIyIqWSSJZ7EjgYVIXnx/HvQdqTnsiERHJKypAi4iI9NDEiRP7h8Ph47to0dFsrX3K87yZ1tr7a2pqer0Y3Fge+Q1wSdLhJg9vr5F1dTl123pHBeiGtWvZsaCArUKdr0Nm4Obiuti5qZ6/Ya/IUcbnyc7ON1vL3FWrOXBQu2sKOVWABlhcVrZri7F1wJZJp6pL6mKTg8gk0oGhQAlulvNIYDjwTdK3QPo64GNcMfpDYBnwDq79xwJgIbAyTV9bJNsdhev53D/FuCeBE1HxWURENoEK0CIiIpvhoosu2sX3/TOA7+OKKMlWAI8AM3urX3SiyLiA5A+L1vy4pL7+j5v7/JnWUQF6ZTzO4C6KzwlNcZ8dI7FYU1eDGisid2M5tasxn7a0sG04nHw45wrQAI3lo84F87ekw82YUGlPF24U2UxbAPsCBwCjgGJcwTn5Akk2WMrGYnQt8Erib7UZkHx2NK74nGrm8z9wxWctaisiIptEBWgREZHeYSorKw8GzjHGfJuOCyxLgbuNMXdMmzbt9U39Qg3lkRsN/KTNF8e8PLKu/iu50ve5tc0oQGMN40prY50uYFhbXr51If5SUny4zqcCNEBjeeQ54NDWxwzMKK6L/TCYRNIHhIAIrth8YOLvUtI3qzkT1gCv44rRrwAv42ZOi+SDY3D9nFMVn/8JfAsVn0VEZDOoAC0iItLLLrjggn79+vU7EhgLfBsY0MGwRuCeRDG62wttxSKRHUIeb9H2A6P1jf+VUbUNL29W8IBsTgEaw7MltbGvd3a6saLsx1j7+1RPk28F6PkVpV/2rPcabYt/zcYL71k8b977QeWSvDMUN4PyaOAIYNtefO7luIt2n+MKwS1sbJOxnLYX2wpxffkNsFXi2GBgG2BHYFAv5lqMa0XwBPAcsLoXn1skU47FFZ/7pRj3FK74vCbtiUREJK+pAC0iIpJGEyZM2CoUCp1ijDkDt6hW8mxAC7wK3Ov7/sxrrrnm3a6er6Gi7BfG2iuSnuHRkvrYN3szdyZtVgEarIc3vLO+141lkVcw7J/qSfKtAA3QWBa5D8O3Wx+zxlxWWlsfDSiS5D4DjMEtGHg0sA+b9nmiCdfqYiGu7cVbuL7MHwLnAhfgZlG/svmRAXcRcHtcMXpI4u89cS1BioE9cAsd9tRa4HlcMfoRIKf670ufdTxwH6mLzy/gZknnzKLGIiKSvVSAFhERyZCJEyfu7HneKcaYscBBnQybD9waj8dvv/baa9vMVLVgFpZHFtqkXtMW7yuldXWz0hQ77TazAI2xNlpcP/+y5OOLIpFRcY9Yd54jHwvQiVnQc5IOv1tcMmp3M3NmPJBQkqtKgTMSf4b14HE+7m6PV4DX2Nhj+b0uHtMPeAP4HpCpn2sFwO64YvQoXPuQ/YGde/AcFpf3TuAe4JNezijSG07BvUZTXXB5Hnehqcs1FkRERLpLBWgREZEATJ48OdLS0nK6MeY0XOEjmQ/8xxhzL3D/tGnTPlxYMeoA35rkNhu1JXWxvdIeOI02twANLCmui+2R3P+6sSxSjWFSd54gHwvQ0PEMcA//0JF1Dc8HlUlyxg7AacCZwJe7+ZjPgRfZ2DP5VdxCrD11KK7lxoub8NjetAtte1ofgGv3kcp63KJtdwAPo/YFkh3G4l6TqYrP/8QtOKjXrYiI9BoVoEVERAI2efLkSDweH4ubXTi8gyE+MKv4nbdX7/vmf4/o39y88YwxE0pq66/LTNL06IUCdLui6rOHHRbe8dOP38Hdap9SvhagG8oj4w38qc1BY2tKaud3qzAvfdI+wI+As0m9OBm4uzYeBZ4G/o0rvvaGQmBdLz1XbxkAfAU4HPgmbmZ4KiuAW4AaoMsWSyJpdBpwK9DuF12SJ4GT0IKDIiLSy1SAFhERyR6mqqrqQOA7uNtkd0kesNuyjzhi7usbdm3cZ1gkFnsngxl7XW8UoMH+raRu/nkb9hrKyo43xj7a3UfnawG6rqxsaIGxS2nbe7yupC5WEVQmyUoh3IzHCcBXU4xdgys4P4KbKflxeqNltWJcL+yTgYPp+rPVOmAmcC2Q3BpHJJ3OBG5CxWcREQlQ8kJIIiIiEhxbU1Mzq6amZuKgQYN2s9YeDFwPLN0wYPePPmw9vj7Xi8+9x3wnFokM+t8edlyAYbJGeX39R7QvdkUWDR++RRB5JOsUAD8GFuEWJeus+BzHzXA+F9ea41Tcrfx9ufgMrqf1dbiWIbsDv4BO+84X4u5ymY2bKf6NTASUPu9HuBn4qYrPT6Dis4iIpJEK0CIiIlkoGo3606dP/09NTc2FgwYN2nXku+/+ZNS77/ClZcv+N8ZkboGuXDAoHOJkgFgksg2G44MOlDWMTX6dePF+/UYHkkWyhcHdaREDfk/HfejBLSBYCXwJOAK4GfgiA/ly0dvAVUAZro3JNcBnnYw9BFfQ/wfd768t0lMXAX8g9Wf+x1HxWURE0kwFaBERkSwXjUb9Q2OxlV+ZH6OwpeV/x33dxt2GtZwD4Bl7BtAv4DhZw/je7ORj1mNEEFkkKxwEvADcA52+Dl7E9TgehSukLu1knHTsDVzhfifgHDqfFX0kbkb0vXT+/0JkU0wGpnZj3GO4FjLNqQaKiIhsDhWgRUREcoGx7WYohvDfDCJKFvvawvLyPYzxxgUdJJtYazt6nXQ241Xy15dwxab/4IrQydYCfwHKca04HgVsxtLlp2bcwm/lwHG4Wc/J/6YGGIsrUlcD/TMZUPKOwV00urobY+9DxWcREckQFaBFRERygtku+Ygl/F4QSQL0QYrzxrd+NdhUt7THgWUpxuQN3/fbvU6MYUgQWSQQBhgP1OOKoMlagBnAHsAPEuOkd1lcm4MjgAOAf3UwpgCYhPv3V39o2RQG15N8YjfG3gWcBqxPayIREZEEFaBFRERygMUfmHzMM6YpiCyBsdyecozh2914pqeAD1OOyhPG2navE+vT7vUkeWk48AzwJ2BwB+cfAyLAD0l9gUd6x2u4AvMRwOsdnN8D9zPqVmCbDOaS3BYC/gZc0I2xfwbOxF18EhERyQgVoEVERHKBMQXJh9b6fp+auWSNdw+wenOfx1hz8+anyR1+KNTu9mrPozCILJJRE3Czab/Wwbnngf2BE4CFmQwl//M0sB+uEPhu0jkDnAXMAw7OcC7JPf1wfcTHdWPsb3EXnPx0BhIREUmmArSIiEgOMJY1ycfC4ZY+1SvUhlq+APPgZj7NigGDBz/SK4FyhG9Mu9nOFlYFkUUyogi4CbiW9otxfoErTH8NNxNXguUDdwAluAXjkouCu+DadUzOcC7JHQOBR3C9nFOZimvPod7uIiKScSpAi4iI5AAf2rVRCK0PbRVEliD5lls26wksd+46a1a7Yn4+KzRm6/ZHrQrQ+WlP4BU6ngn5OFCG6xGr2Y/ZZTUwBTfbuSHpXBi3oNydwIAM55LstjWuXcuR3Rgbxb3GREREAqECtIiISA7w4P3kY9bzhwUQJVCl9fXPAO9s6uM9z25eATsHWdP+dWLxkm/5l9x3FG5Wc0XS8eW4xcaOo32rB8kuLwFfxs1UTZ6lehrwIrBbpkNJVtoR+DcwJsU4i5v1fFnaE4mIiHRBBWgREZFcYM1b7Q7hFQcRJUgGfCy3beLDG0bWzn+lVwPlBNP+dWLtksznkDQ6AXgYNyOytUZcgerujCeSTbUWN1P1BNzFg9b2Bl4ARmQ6lGSV3XGvg/IU4+LAD3B9n0VERAKlArSIiEgO8EMtseRjxrB/EFmCFrfczKb1sLy5d5PkBmPtAcnHPJ92ryfJWacCD9C+3/M9wL64IrTknr/jFimsSzq+K24RybKMJ5JssDcwC9dupyvrcQtZ/jXtiURERLpBBWgREZEcUDKvcRHwWetj1nKwHTs2FFCkwERiscVYXuzhw+IhE7ozLYGymAVj4ZCkw00jR42qDySQ9LZxuEXswq2O+UAV8F202GSuWwx8Bbgv6fgOwNOkngEr+eUA4BlgaIpxq4ETgbvSnkhERKSbVIAWERHJAQasMbyUdHjIgvnzDwwkUODMzT18wD9H1Na+l44k2ayhonQfYOfWxwzMMjNnxgOKJL3nTNzsxtYXoeK4ovT0IAJJWjThZrn/Len4UOBf/Ax2jQAAIABJREFUwMiMJ5IgHA88C2yTYtxyXD/4x9OeSEREpAdUgBYREckRvrWPJR8znjk1iCxBi1t7D64w0z3W3py2MFnMs16714e1pt3rSHLOl4EZtH0vv6H4vKk90iV7+cD3gRuSjm8HPAJslfFEkknjcG12+qcY9xHwNeA/6Q4kIiLSU+HUQ0RERCQbmHD8UVrCv6dV0cliz3hr2LCLdl+yZG2A0TIuEos1NVZEHsRyVjeGf9avafUjaQ+VZWKRSKGBc5KaZVs879FgEkkv2QG34GDrYtQ64DRckSpX7AhEuzj/O6C2F75OCTCxi/NXAO/0wtdJNwtciLvQMKHV8WJcv+9jE+ckv0wGrgJMinFvA0cAi9KeSEREZBOoAC0iIpIjSuYuWNpYFnkKw1GtDm+zdvDAs3GzIfuam6EbBWjL3X2tQA/geZxu2/cK/XdJbe1bgQSS3tAPeBDYpdUxC5xLbhWfwc3aHd/FeQv8qBe+zo9SfJ0Z5EYBGty/yURga+CcVsePBK7EFSslP4SAG+ne90AD7jXQ59pMiYhI7lALDhERkVzi8ZfkQwYmzx49uiCIOEEqro09C6QsphrLzelPk13s2LEhA1PaHYc/B5FHes01QHLf92ogHxfY/C4wYDOfowA4vReyZJsfAS8nHbsIt/Cc5L4i4G66V3yejVtoVsVnERHJapoBLSIikkOKi0c9uKBh/iIMI1od3mNQc/OPgeuDyhUEA7YR+xvwjuli2GfFsfrXMhYqSyxoaPgBhuKkw+/6PvcFEkh6wzeAHycd+wfwywCyZMKWwLeAuzbjOY4HhvROnKyyFjgJeI22s+FnAC8By4IIJb1iW+BRYEw3xj4FnExP1kMQEREJiArQIiIiOcTMnBlfUB6ZZpNbbhj7fwtGj7yzeM7CTwKKFoiSuvk3ATcFnSOb1JaXbw3+ZcnHLVRHYrF1QWSSzdYf+Att+8AuAk4lv/r+Wtr+N45j8wrQ41I8fy77EPg2bsG5DXfADMHNiB8XUCbZPMOAJ3B9y1O5D9eCqs+1lxIRkdykFhwiIiI5ZmVh0c3AgqTD2/jNBTcEECctjMlsjcjLm5oUFFj/t8D2SYffLFq5ql37FskZVbji1AY+cB6wIpA06fMCrki8weHArpv4XNsDyXdHvLSJz5WtXsUtotja2cD+AWSRzbMP7vXZneJzDfAdVHwWEZEcogK0iIhIjtl3zpz11pqq5OPG8N2G8khe9Dsd5GX2LcrW4VBGv166NJSVnWwMZycft9ZM6osLMeaJ7XH9fVu7HleszTdvAc+32veAMzfxuc5i48xgcDOFGzfxubLZlUB9q32DK1BK7jgSeA7YMcU4i1tochJtL9SIiIhkPRWgRUREclBpff1jYB5KPm7gz/MrSr8cRCYJ1vyK0jJj7M0dnHqitL7+gUznkV4zERjUav9joF2LlTxyS9L+ODatbUbyhZjk580X64ELk44dDBwaQBbpuR8Afwe2SDFuHW5Bzeq0JxIREUkDFaBFRERyVChcMJ72i00N8Kz3aOM+xTsFkUmCEYtEtvGs9xAwOOnUirA1Pwwik/SKLYDzk45dASwPIEum3EvbRdVGAgf28Dn2BSpa7a+BvF6A81/Ak0nHpgQRRLrNAFHceg6p1mVqAk4A7k5zJhERkbRRAVpERCRHjZg792Nr+D6uH2xrOxEP3xuLRAqDyCWZNXv06IJQiPuAPZNOWbA/GF5f/24QuaRXnEbbmZEfAX8OKEumrALuTzp2Tg+fY1zS/v3kd9Ee4NdJ+0fR/meCZIeBwIPA/3Vj7Ae4Ge3/TGsiERGRNEt1tVVERESyWGlt7NHGirJLsfbyNicsB4U8Hnp3zJhv7zpr1pqA4nWLsf4SS2hmyoHNbWZFbu4X/SfWS17IsQ1reKPXvl6axCKRwvD6tXdZy9c6OH15Sd381P+uks2+l7T/R2B1EEEy7GbaFp2/C/yc7v23FwKndvB8+W4W8ApwQGLf4P4NLw0skXRkB+ARYL9ujG0AjgbeSWsiERGRDMifJd9FRET6KAumsSxypzF8t4PTz8V9TojEYr1XvJWsEItECkMe9wLf6uD0w8V1sZNN+9nxkju+BCxh4/t1H9gDeDuoQL2sFJifdOwWNvZ8XkTbGbynA3d143nH4tp4bPAeMAyIA38Bzksavy8wp5uZs933aTtDfj4QCSiLtBfB9XverRtjXwROBD5JayIREZEMUQsOERGRHGfADhq8xfeA2R2cPizk8XhjcXFyb2DJYbNHjx4Q8niUjovPDQWFReeo+JzzTqDtZJFZ5E/xORUL3J50rLttOMYl7d+EKz73BTNxixJuMAoYHlAWaetIXFG5O8XnW4Gvo+KziIjkERWgRURE8sCus2atIdzyLaCxg9MHUxB+elFFxS6ZziW9b35p6Y6D1q19ClfQaMPA4rjP0XvOmbMigGjSu5LbqjwcSIrg3EzbiyhHALumeMxQ2n5fWFwxr69YATyfdKyj9jySWeOBx4AtU4yzwGW4iyjr0pxJREQko1SAFhERyRMlcxcsDZvQoUBtu5OG/eN+fO7C8tLDM59MektjJHKQF/bmAF/p4PQCvPBhkVhM/ULzwwFJ+/8KJEVwltC2mOoBZ6Z4zDm0XePmBWBx78bKes8l7Se/jiRzwsANwJ+AghRj1+Fev1FcIVpERCSvqAAtIiKSR4bX1i5bh3cY8Fq7k4btfLwnG8sjk63Wgcg5DeWR8Xj8C9ixo9N+i/+14nnz3s90LkmLbYHWdyysBeoCyhKkm5P2x9H1z65xSfu39GKWXPFy0v7egaSQbYF/Aj/txtjPgKOA29KaSEREJEAqQIuIiOSZirq6z9fhHWXhlQ5Oh4CrF1ZE7ltcUbF9prNJzy0YPXK7xorI3cbNoitsP8K8bgrXHzKqoeGDjIeTdNkjaX8RffOW/PuAla32RwIHdjL2ANzChhuswvVE7muSF3ZMfi1J+pXjLgJ3p/3Jm8BBtJ+5LiIikldUgBYREclDFXV1n4fXNB+K5S8dnbeWk1tsfEFDeWR8prNJ9zWWjxpr1xXEsJza0XkDdzUb75DiOQu1WFV+SV6obEkQIbLAKlwRurXOFiMcl7R/P22L133FB7gZ8xtsDWwRUJa+6FjgP8Du3Rj7MjCGjtduEBERySsqQIuIiOSpEYsXN5fUx35g4Yd0PHtyKwN/aiyPPDG/tDS54CUBikUiOyyoiNwP5l6go5nqLcCU4rrY6XvV1q7KcDxJv+SCYV++wJDcRuO7wICkY0XQ7iLNzekKlOUsrqVDaypAp58BfgE8Svf+vW/DzZBels5QIiIi2UIFaBERkTxXWhebYfEOAzrrD3y0F/bqGirKfjGvomJgxoJJO++OGdO/saJsUsijwVpO7mTYh8bYr5fUxaZmNJxkUnKBdU0gKbLD88B/W+1vCXwracxJuJm+G7wN/DvNubJZU9L+oEBS9B0DgbuAK0j9+doHLgLOpu1MdRERkbymArSIiEgfUFpXNytsQl8G01lP1MHG2iv62fiixrKyH80ePbogowH7uGcPOyzcWFZ23qqmLxZibTWwVccjzUNxn32Ka+e/kNGAkmk2ab8vLxpqgVuTjiW34RiXtH8zrtDXV4WS9uOBpOgbhgOzaD8DvyNNwMnAtLQmEhERyUIqQIuIiPQRw2trl5XU1X/HWHMyrk9oR3bE2D8Mal4baywb9R2r9wppZcE0lJWdPO/NxW/F8f8C7NLJ0GVYe2pJXf1JkVjsw0xmlEAkt1Xp63cm3ELbgvIRwK6J7Z2Bb7Q6Z3HtDfqy5Bn0atOTHsfhFhss78bY/+IWynw4rYlERESylD5UioiI9DHF9fUPFhQWlRqYQfuZlo5hBMbcs6A8srCxouxCteboXYuGD++3oCJy9sLySN0LTSvvn/DOe7t8e/GbzFvdUacFM9MUro+U1M+/N+NBJSjJPXyHBpIie7wNPNdq3wPOTGyfQ9sZv8ktO/qaMDCk1b4FlgeUJV8ZYDLwCJ3erdLGC7jFBuenM5SIiEg2UwFaRESkD9pzzpwVxXWxH2I4HMzrXQ3F2t/2s/F3G8sjUxeXle3axVhJoXGf4p0WVJRdGe/f731ruWW170d+vdRNRm9cu5bT/vsml76/lCbfB5hnrTmqpK7+O8VzFvblRej6oiVJ+7sHESLLJC9GOA5XCDwr6fjNmQiTxXbBFaE3WAasDihLPtoCuB+4mu59lp6Bm6H/cTpDiYiIZLu+3E9OREREcG0gFlZETsXyG+v6WXalxcKjnjW3eWvXPj5i8eLmjITMYbFIpDAU4misORPsicD/+mtfvvQDbv80ebIrDPC8z5p9/7w4PJTJrJI1BgMr2PhePY6baZm8uFwuK6X9jNBbaN/PeYOBuNZBg1sdm0TbfrqrgB2BlZ08x1+A85KO7QvMSR03Z3yLtj83XgIOCihLvikHHiD170mA9cDPgD+mNZGIiEiO0AxoERGRPs6ALa6N3b2ysGiUhR/SeX9ogLCBk6yxD8T79/uwsSJy68Ly0sPVK7q9hvLy0Y0Vo64LebyL5WGwY2lVfJ67ejV3dlB8Bljt+9vE4UHcn876Qkv+WgksaLUfwhVK+7JVQPIiqlcm7d9H58XnvuLApP3ZgaTIP9/FLTbYneLzh8DXUfFZRETkf8Kph4iIiEhfsO+cOeuBGfMqKu4osvFx1nIhhhFdPGQrLGf5eGctKI+802h50IZ4omjFqn/vvmTJ2kzlzhaLhg/vt76o6BBj7NEGTgZ/GLbjm83WW8uv3l/aZlW1TpwIHAn8GqjBzYSVvuFVoKTV/pG07YPcF90MfK/VfkHS+ZsyFyVrHZG0/3IgKfJHP6AaN5u5O14CxgJL05ZIREQkB6kFh4iIiHTIgreovPTr1ngXWstxdP99wxqLedFgnw75PDoiFsvbhZcaKyp2tzZ+hMEcDvYoXH/QlOrWrOGcN5fEV/t+KPXo/5mNm6HeVc9uyR9nALe32o8BZQFlSYeetuDYoBEo7uD4EmAPOltY1cn3Fhy7AO+w8We1j2tJsiywRLltOHAP8OVujv89MBFYl7ZEIiIiOUozoEVERKRDBnzqGp4Gnl4QiexlPfszMKeQusja32APBw6Pe1zdWB55F8zL1vqveB6vrizoP2ffOXNyblGsd8eM6b/qiy++bDwOsIb9gQOx8d1cpaermlcbTQbujwzod/1q31+Km1mXvIhaZ/YFXsMV0apQq4F89wTQwsb36xFgH2BuYImywx24OwKS3UwPvhHz1Fm0vVD4Cio+b6oTcTPqt+rG2LXAT4G/pjWRiIhIDvv/9u49yM66vuP4+zm5LwkhQAhLgFEC2cRACUQgQWW8BAEtXqr1Ak5qR00dUZBqdWynilpbFG9UbQttRWmRitZCsaCCLXLRSQSBIORGABMSchGTkNsmm92nf3zPTo7Zs9lzfZ5zdt+vmd/k7Mm5fJ/znOeZ5PP89vtzBrQkSarYugULJuzeseMNKemlJFzIwF+Br8T+FB5LSJaSsBzSlTBqZdeyZb9JqKQrRXOlUFgxd9aJpKO6kt6ki0LfbNLC2ZCeRo3bC8lPUtKbdo0df2uZ8P11wDeAF1XxmhuAK4ietxq+7gAuKvn5H4EP5FRLo9U6A/pE4P/K3L8QeHqI5w7nGdAFYBUwo+S+K4Gv5lNO26q25cY64K1EyxxJkjQIA2hJklSTlfNmHk3PmLelcAkp84mF0uqxF1hFyioS1pDwHGm6hSR5rjdJN41LR2+ZsWzZlqTOWY6rzzhj6r6+vceM2p8ckxboTJJkKmnaCclJkHYBM4HxdW5LH7AkTdObxxRGf/fkZcuGmoV4GHAV8GGq+w2124iQaajgTe3pLfz+RYZuos3EoRYKbRe1BtD1GM4B9B8Dt5T8vJdoyfHbfMppSzOIz7DSlht3A+/Ez1iSpCEZQEuSpLotnzXrqMLYUef39XFRIeGCFKY16a32A9uLgxS2F2dN70xJegqkaR9JkpCOBiZBkkDa/yvUk4ujWS3ItkD645TkzsLYnp90PbSqllDidOB64OwqntMNXANcDbRdaxMd0hhgDXBCyX3XAB/Lp5yGMoBunALRG/70kvu+BfxpLtW0p7cS34/JFTw2Jc63f40Lw0qSVBEDaEmS1FApJCtOO+3MJEkvTFNemZCeRWX/qW83O4iezPckffxo5uOPP9SgFiIFos3C56hwUcOitcBHsC3HcHM5cG3Jz/uIftBP5lNOwxhAN857iG3r1wecxsDPVwN1AF8BFlf4+N8Ci4ge7ZIkqUIG0JIkqalSKDw5Z86s/QXOTpL0HEjmk3Iq7bUYci/w6wSWpGmypHdU35KXPPrE8ib3rO4EPk/lixT2u4cILR9rdEHKxQRgNTC95L7biEXS2pkBdGNMIT7HY0vuuxm4JJ9y2sqpwHeIsL4SS4G3A880qyBJkoYrA2hJkpS5dQsWTOjevX1WX29hJvTNIkm6UpIuSGcCE3MsbVcafahXUkhWFkhX9NK3aveYjhVlFg/MyuuBr1PdIoX7gW8Cfwk834SalK1FRDhb6lIiPGtXBtCN8W3i+9FvD/HZ/iafctpCQlyk+zyx6OBQUuBrwEeBnibWJUnSsGUALUmSWsrK00+f3pvuOyGhMLUA09KUYxOSY1L6pqUUjk1ID0tgchqtKiYSfXLHEzNF+3UTQUwPsJMIELYBuyHZmMDGlHRLAhvThE1pWtgyOknWnbJs2bMZb26lOoi+v58AxlbxvOeBzxIBtr1K21cBeACYX3LfVuAsokd0OzKArt87iNnOpT5NLGiq8qYBNwAXVfj454mA/46mVSRJkiRJktRC5gL3EYF6NWMJEVaqfXURi0yW7tfltG9/9RlEeF46rmnye15d5j1PbfJ7NstcYBe//314lOouUI00byJ6OFd63rwfOD6XSiVJkiRJknKUEO0X1lNdCN1LzAA9JvuS1SBXMnC/3kbMkNbIMY1YdLT0e7AHODPPolpYB3Ad1Z0rr6a91imQJEmSJElquIlED9O9VBdEbyXaeYzPvmQ1wE0M3Kd/m2tFytIEoh3Lwd+BRYd60gj2MmAVlZ8f1wMLc6lUkiRJkiSpRZ0M/JDq23KsAxYDo7IvWXUYT7RUOXh/Xp1nUcpEB3AXA/f9F/MsqkWNJ46J/VR+Tvwv4Og8ipUkSZIkSWoHC4mewNUG0Q8Cr8qhXtWuE3iWgfvyC3kWpaY6DPhfBu7zn2CriIOdQ3Xnwt3AFblUKkmSJEmS1GbGEEHKC1QfRN8F/EH2JatG8ym/n79M9AnX8HEE8HMG7utHgSk51tVqxgBXUd2s58eA03KoVZIkSZIkqa1NB24E+qguhO4tPq8z+5JVg/lET++D9+N3iBmzan+nECFpud9cOCrHulrN6cDDVH6u6wOuBcblUawkSZIkSdJw8SrKh1dDjR3AJzHEbAdnAFsYuA+XA7NzrEv1ez3lLzA8CByZY12tZBzwN0APlZ/fngVek0exkiRJkiRJw9Fo4DJgM9UH0euB9+JCha1uNrCBgftvO/DmHOtSbRLg48RvJBy8T+8HDs+vtJZyHtX3vb8R25ZIkiRJkiQ1xWTg74gFt6oNoh8DLs6+ZFXhZGAZ5duqfAnoyK80VeHFwE8pfxzehPsRIkC+nupaDD0HvCGPYiVJkiRJkkaa44HrqG6hrv6xBIPoVjYeuIHy++4pYGF+pWkICbCYaH9z8L7rIWZEK84/66juvHULcHQexUqSJEmSJI1kc4D/ofoQOgUeAF6dfcmq0GJgHwP3Wx/RgsD+wa3lZOAeyh9rm/FYAzgO+E+qO09tAt6SR7GSJEmSJEk6YCHwCLUF0fcTfVjVes4Dnqb8ftsA/BkwJrfqBHAU0R6lm/L76W4ieB3JCsQFlReoftbz1BzqlSRJkiRJUhmjgPdRfiG7SsZ/A3Mzr1pD6QCuZvB2K08T4V4hrwJHqA6ipcZWyu+XbcR+SfIqsEXMBx6kunPRZuDteRQrSZIkSZKkofUHY9upPoTuA27HILoVnQs8weD7bilwfm7VjRzjgA8QC+INti9uxVnPRwHXEgtoOutZkiRJkiRpGOoEricWP6s2iO4F/o3oa6vWMQ74NLCLwffdI8C7i49V4xwDfArYyOCf/W+At+VVYIsYA1xJzACv5pyzGvtkS5IkSZIktaVTgH+n+pmIKRFe3wB0ZV61DmUq0ZZjL4Pvu03Fx4z0mbj1mknM5N3N4J/188RvHUzIqcZW8Rrgcao7x+wDPoefnSRJkiRJUtubDdxIbUF0L9Ga4+zMq9ahzAS+R7ROGWzf7QFuBv4QFyys1BSin/q9HPq42EmEp5PzKbNlTCfOLdWeVx4CzsyhXkmSJEmSJDXRXGLBwWrDov5xJ3Be5lXrUM4Cvs/gCxX2jy3A14EF+ZTZ0sYBfwT8AOjm0J/jNuCLRJubkWwi8BkO3RKm3NgOXIYLZ0qSJEmSJA1r5wA/pvYg+n7gdUCSdeEa1IuBL1PZApRrgK8AFwDj8yi2BUwFLiVa1Gylss/sCmBSHsW2kNHA+zl0P+xyow/4DjFjWpIkSZIkSSPEucBPqT2IXgYsIkIptYbDgQ8DT1LZPtwF3AF8iOgZPlyNAuYTCzkupfJ2NPcAb8YZuwALiWO+2vPEcuD8HOqVJEmSJElSi3g5EbTVGkQ/RcwOHamzaVvVPGIRvU1Uvi+fI3p+XwVcTPREbkedRP1XEdvzOyr/DJ4hFnEczoF8Nc6itvPDVuK84AUqSZIkSZIkAbFY3S+pPYheB3wEOCLrwnVIY4E3ArcQCxNWs0/3E7Nevwl8DHgTMKv4mq1gEhG0X0L0JP4esJbqv7sbibDexTYPOAn4Dw690GW50QtcT7Q5kSRJw4B99yRJktRorwU+AbyyxufvBL4N/D2wqkE1qTEmAa8BLgQuAk6s8XX2A08DK4t/biZmT28mFjzsv72njloPJ2YyTwWmldzuBGYAXdTeUzgFHiYW1vwR8AsiOBUcTxz/76X6Cw1LgA8CDza6KEmSJEmSJA0/C4j2BdXOgCydCflDonesEyda0xzgo8DdwG5qn/0+2NhNtL/YSCzkt5oIJx8E7i3++avi360Bni8+vqcJtWwmZvS+Gzi2AZ/dcHMccdGom+o/22eJz9XjXJIkSZIkSVU7FbiR+kLBlUQ/2I6Ma1flRhOB9CKiHcWDVL5YX6uNHuBx4Lri9szBcHQwU4me17VcgNhZfO6kzKuWJEmSJEnSsPNiIpisto9w6dhWfI0TMq5dtZlMzGC/HPgGMVO6lh7LzRr7iIsbtwFfAN5D9HFulR7VrexoIjzeRW2f+3U4k1ySpBHBq/iSJEnK2nRiscHFwGE1vkYPsWDctcDSBtWl7EwEZhZHZ3EcUxz9t6cCY+p4jz3AJqJ9x+aDbq8FVhD9p3vqeI+RaBrw58Bl1Hb8/oDoEW1/d0mSRggDaEmSJOXlKGJm7AeBI+t4nV8B1wM3Ay80oC61jklEa48OYBwxM/kwYFTx77YVH7ed6DW+kwiUu6lvAUMNdDLR7/tPgPE1PP8XwF8ADzSyKEmSJEmSJGkoHcD7geXU105hB/AvRAsFSY3xUuAWau/n/WvgLTj5SZIkSZIkSS3g5cDtxGzWesLoJ4CPE7OsJVWv/1is5xhcRMxWlyRJkiRJklrKbOCfqG2Bs9KxC7gBWJBt+VJbGgNcAjxM7cfcSuBSoJBx7ZIkSZIkSVLVjiRmMq+lviA6BR4DPkR9/aal4agT+BSwntqPr9U441mSJEmSJEltqgBcDNxF/UH0XqK1wCJiQTtppJoH3Ajso/bj6RlgMbFYpCRJkiRJktT2ziFCsz3UH0ZvA/4VeBW2DNDI0AG8D3iE+o6dR4F3YfAsSZIkSZKkYWoKcDnRWqPeIDoF1gHXAKdnuRFSRmYDXwK2Ut9xcg9wEZBkWr0kSZIkSZKUo3nAdcBOGhNGPw5cBczIcBukRptMtJq5C+ij9uOhl2hb42KekiRJkiRJGtEmAx8AHqYxQXQvcB9wJfCi7DZDqlkBOB+4CdhNfd//buCfga5Mt0CSJEmSJElqA2cT4dkOGhNGp8CvgE8Cp2W4HVIlTgY+C6yl/u/5c8BngM5Mt0CSJEmSJElqQ5OAxcD91NeG4OCxGvgCcC4uYKh8nEDMzv85jfluPwC8Exib5UZIkiRJkiRJw8UJwBU0rkVH/9gC3AhcjOGdmqv/O9yoCyrdwC3Y31mSJEmSJElqqDnA1cAGGhtG7yQWffs48JLMtkbDWaND5xRYQ3xHj85wOyRJkiRJkqQRZxSxaNu3gBdobBidAk8C/wC8kWgHIg2lAJwFfApYSuNC527gu8CF2DZGkiRJkiRJytwE4B3A7cA+Gh9G7wN+BvwVETAaAqrfFODtwLeBTTT2e7cUuKz4HpIkSZIkSZJawBHAu4DvE201Gh1Gp0Tv6O8R7RVeCozOZMvUChJgLvAJ4D5gP439bm0EvgicmtUGSZIkSZIkSarNBKKFxreA39KcMDoFdhD9o68CFgITm79pykiBCJwvJy5qNHqWc3+LjR8Qi2F6MUOSJEmSJElqQ6OBVwNfA9bSvDA6BXqAXwJfBd4KdGawfWqM0cA5wEeJli5bac53ZA9wKzFb//BMtkySJKkKSd4FSJIkSW0sIVpnvBl4AzAng/dcDzx80Hgmg/fVoZ0AzCuOc4AFNG8GezfwY6J9y+3E4pmSJEktyQBakiRJapxpwHlE+4yLyW7G8nbg18BDJWMF0JvR+480x3EgbJ5HXIQ4tsnv2Q3cTYTOt2LoLEmS2oQBtCRJktQco4hw8rXABcB8su3Luwt4AlhNhNGrirdXEYsqamhTgVnF0QW8BDiD5ofN/dYCdwJ3AD8l9qkkSVJbMYCWJEmSsnE40Tv6AiKUPinHWtYTQXR/KL0SWAOsY+SF0x3AiUTA3D9mF/88MuNaeoD7idACxZptAAAB80lEQVT5TmJWuyRJUlszgJYkSZLycRLwCuDlwMuIWbat8O/zHUQQvR7YQMzC3VD8+dni7U25VVedCUS4fDwwveT28cXb08k+ZD7Ys0Q/5zuBu7C1hiRJGmZa4R+4kiRJkuBoIoh+BXAu0Vd4TK4VDa4X2FYythbHtjL37+VAqJoW7++3rXgfxcdMBArFMbl4/1jgsOLtCcD44phSMo486Of+Mb4xm9tQa4D7gJ8B9wJP5VuOJElScxlAS5IkSa1pAnA2EUi/DFjAgVBW7WM5ETTfS4TO6/MtR5IkKVsG0JIkSVL7OAk4E5hLLIZ3BtCZa0UqtRl4qGT8vHifJEnSiGUALUmSJLW3KcAcYF7JmEW0sVDzbAMe5/cD5yc40FJEkiRJGEBLkiRJw9FkYDbQBcwsGacQrT1Uud8BK4lWGiuBFcAjxOKMkiRJGoIBtCRJkjRyJMCJHAiju4pjBnA8rbloXxb2AuuAVRwImvtD5y051iVJktT2DKAlSZIk9ZtA9JQ+CTiuzO3jgGNpr/9H7AOeBzYATwHPlbn9DNCXU32SJEnDWjv9w1GSJElS/jqAo4AjiP7TR5SMKWVuTyw+bxRweMnrHMGB/49MBMYQ/ZO3Fe/rBV4o3u4BdhZvdwN7gO3A1oPG78rctxX7MkuSJOXm/wEIrfJvEMPewgAAAABJRU5ErkJggg==" width="85%" style="display: block; margin: auto;" /> --- class: center, middle # OUR WORK --- ## Current state of NetCoupler - Series of R scripts -- - Only uses specific methods ??? - Greats well on Clemens' computer - Not readily re-usable by others - PC-algorithm for network - Cox for modelling - Not easily extensible to other models --- ## Converting to a usable R package .center[  ] --- ## Converting to a usable R package .center[  ] ??? We've got to improve on the underlying implementation, optimize the code for better speed and performance, fix up the interface so that other methods can be used like regression or mixed models, and reduce some software dependencies since right now it relies on software that is really hard to install and get working. --- ## Goals: - Submit to CRAN[1] -- - Create simple interface to use -- - Create website with documentation -- - Develop tutorials .footnote[[1] CRAN distributes R packages.] --- class: center, middle # Interest? Feedback? Comments?